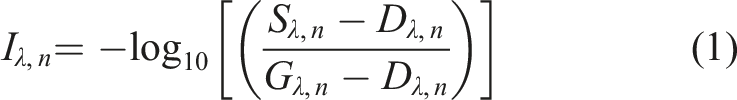Abstract
Near infrared (NIR) hyperspectral imaging and multivariate data analysis was evaluated for its potential to detect and classify Listeria species. Three Listeria species, namely L. monocytogenes (ATCC 23074), L. innocua (ATCC 33090) and L. ivanovii (ATCC 19119) were grown for single colonies on Brain Heart Infusion agar and imaged in the NIR range of 950–2500 nm. Principal component analysis (PCA) was used for data exploration and to establish pattern recognition. Images were pre-processed with standard normal variate correction and the Savitzky-Golay smoothing technique (third order polynomial with 15 points). Two approaches to data analysis, that is object-wise and pixel-wise analysis, were investigated for discriminant analysis. The PCA score plot showed slight separation between the three groups with L. monocytogenes and L. ivanovii grouping close together. It was possible to visualise separation along PC3 (5.64% sum of squares (SS)) and PC4 (3.44% SS). Based on the loadings, differences in bacteria were attributed to teichoic acids, protein, and carbohydrate composition in the bacterial cell wall within the wavelength range 1000–1900 nm. Using extracted spectral data from the hypercubes, partial least squares discriminant analysis was employed for further classification. Classification accuracies above 90% were achieved for L. monocytogenes, L. innocua and L. ivanovii. This was true for data analysed using both pixel-wise analysis and object-wise analysis. The results demonstrated that hyperspectral imaging has notable potential to classify bacteria within the Listeria genus. Nonetheless, in order to improve model efficiency, model optimisation and incorporation of more bacterial strains need to be investigated in further research.
Keywords
Key points
• A new method for detecting and differentiating Listeria species on agar media is described. • NIR hyperspectral image data was used for classification model development. • Classification accuracies of above 90% were achieved.
Introduction
Microbial food safety remains a priority globally due to the increase in reported cases of foodborne illnesses. The World Health Organization estimates that of all the foodborne disease reports between 2007 and 2015, 54% were caused by pathogenic bacteria.1,2 Pathogenic bacteria such as Listeria monocytogenes (L. monocytogenes), Salmonella, Campylobacter and Escherichia coli (E. coli) are among the main microorganisms that cause foodborne diseases. 3 Despite the various criteria for guidance on acceptable microbial levels worldwide, many countries agree that the absence of L. monocytogenes in 25 g of infant food and ready-to-eat foods is the acceptable food safety requirement.4,5 Other examples of pathogens with strict regulations include the Shiga toxin-producing E. coli in ground beef and Salmonella spp. in ready-to-eat foods and powdered infant foods. 6
Hence, the need arises for rapid, accurate and sensitive methods for early detection of pathogens at every stage of the food production process. 7 The analytical methods currently being used often include plating and culturing. These methods are often applied in microbiology analysis due to their high accuracy levels and ease of interpretation. 8 However, these methods are usually laborious, time consuming and often give retrospective information, making them unsuitable for real time analysis.
Hyperspectral imaging (HSI) is an imaging technique that aims to mitigate those limitations by combining both spatial and spectral information to analyse and detect a specimen. 9 The method allows one to visualise the sample and collect three-dimensional data with two spatial axes (x,y) and one spectral axis (k), resulting in a dataset (x, y, k), which is often referred to as a hypercube. 10 Each pixel contains a spectrum representative of chemical components of the given sample due to light absorbing and scattering properties. 11 With the use of multivariate statistics and machine learning algorithms on the experimental data, organism identification and prediction model development for different organisms becomes achievable. In bacterial cells, the membrane macromolecule structures such as lipids, proteins, nucleic acids, and polysaccharides are known to be correlated with the genetic information. 12 For instance the presence of an outer lipid membrane in Gram-negative bacteria has been proved to show considerable peaks at wavebands 1650 and 1900 nm suggesting higher concentrations of liposaccharides.13,14 Similarly, prominent peaks in the waveband range 1000–1600 nm are often attributed to proteins and wall teichoic acids present in the thick peptidoglycan layer of Gram-positive bacteria.15,16 Therefore, differences in structure equates to different absorption signatures of each of the functional groups due to the different vibrational frequencies.
Hyperspectral imaging has been used for the expeditious detection and differentiation of different bacterial pathogens both at a macroscopic and microscopic scale. Dubois et al., 17 pioneered early research studies on the application of near infrared hyperspectral imaging (NIR-HSI) for microbial detection and identification at a microscopic scale by identifying bacteria on food cards. The study showed that Gram-negative bacteria (E. coli and Salmonella spp.) could be identified and distinguished from Gram-positive bacteria (L. innocua, L. monocytogenes, Bacillus cereus, and B. subtilis). Bacterial biofilms isolated from various food sources were deposited in 2 mm deep wells on an aluminium plate and allowed to dry before image acquisition. Differences in second derivative transformed spectra were used to differentiate between Gram-positive and Gram-negative bacteria. Partial least squares discriminant analysis (PLS-DA) was employed using the wavelength range 1200–2350 nm. Classification results for the model were promising, with clear separated peaks observed on the histogram plot of classification.
A study by Yoon et al., 18 showed that it is possible to classify bacterial colonies on solid agar using wavelengths in the visible region (400–1000 nm). Principal component analysis (PCA) was applied together with Bhattacharyya distance, which measures the similarity of two probability distributions, for differentiation of Campylobacter from non-Campylobacter colonies. This resulted in detection accuracies of between 80 and 99%. Another macroscopic study carried out by Kammies et al., 14 investigated the prospect of using PCA and PLS-DA to differentiate between pathogenic and non-pathogenic bacteria in the NIR wavelength range (1000–2300 nm). The authors successfully differentiated between B. cereus, E. coli, Salmonella, Staphylococcus aureus and S. epidermidis grown on solid media with prediction accuracies of between 82 and 99%. Gu et al., 19 provided a unified way of classifying bacteria on different types of agars. The team was able to classify E. coli, Staphylococcus and Salmonella using PLS-DA and support vector machines (SVM) with classification accuracies of 75% and 95%, respectively. Taking a step further, Michael et al., 7 investigated whether it was possible to distinguish strains within the same species and genus using the spectral data of individual bacterial cells obtained from the hypercubes. The K-nearest neighbours (KNN) model produced classification accuracies of 100% for all Salmonella spp. strains investigated.
Although there has been considerable work done on the detection of different bacteria using HSI, limited work has been done on the Listeria genus. From a public health perspective, L. monocytogenes is the pathogen of most concern. However, food industries prefer the detection of all Listeria species to ensure absence of L. monocytogenes in their processing plants and products. 20 Orsi and Wiedmann 21 report that L. monocytogenes often occurs with other non-pathogenic species of the Listeria genus. In addition, the presence of any Listeria species above 100 CFU per gram of a food sample is considered a health hazard and a cause for further investigation. 5 Hence, the aim of the study was to investigate the use of NIR-HSI in detection and differentiation of three Listeria species from pure cultures on Brain Heart Infusion agar (BHI), namely L. monocytogenes, L. innocua and L. ivanovii.
Materials and methods
Bacterial culture and sample preparation
Three bacterial isolates were investigated in this study. The isolates included L. monocytogenes (ATCC 23074), L. innocua (ATCC 33090) and L. ivanovii (ATCC 19119). All bacterial strains were obtained in lyophilized form from Davies Diagnostics, South Africa, and resuspended as per the manufacturer’s instructions. To minimize spectral variation from the growth media, BHI, a general-purpose growth media, was used throughout this study. The streaking method was chosen as the mode of microbial inoculation based on a study by Kammies et al., 14 This resulted in single colonies and large areas of bacterial growth where the initial streak was made, allowing for more spectral data collection.
A loopful (<10 µl) (obtained with a sterile inoculating plastic loop (Lasec SA)) of Listeria species stored in glycerol (skim milk tryptone glucose glycerin (STGG), NHLS, Greenpoint) at −80°C was streaked onto BHI agar under aseptic conditions in a class II biosafety cabinet. The petri dishes/plates were thereafter incubated for 24 h at 37°C for bacterial growth. This process was repeated to ensure bacterial viability and purity, whereby a single colony of the bacteria from the incubated plates was steaked onto BHI and incubated at 37°C for 22 ±1 h for Listeria growth. The plates were prepared in duplicate and allowed to cool down to ambient temperature (approximately 21°C) for 15 min prior to hyperspectral image acquisition. This was considered as the first experiment (Rep 1). Following the above protocol, the experiment was repeated once more (Rep 2) resulting in a total of twelve petri dishes. All samples were prepared and scanned in the biosafety level 2 microbiology laboratory and the vibrational spectroscopy unit, respectively, located at the Department of Food Science, Stellenbosch University.
NIR-HSI imaging system and image acquisition
The images were acquired using a pushbroom HySpex SWIR 384 (short wave infrared) camera on the dual VNIR/SWIR imaging system (Norsk Elektro Optikk (NEO), Oslo, Norway). The system is composed of a spectrograph, translation stage, a mercury-cadmium-telluride (HgCdTe) detector, two 150 W halogen lamps (Ushio Lighting Inc., Tokyo, Japan), together with a computer equipped with Breeze software version 2021.1.5 (Prediktera AB, Umeå, Sweden). Petri dishes were placed onto the translation stage which moved at a speed of 50 mm/s under a camera equipped with a 30 cm focal length lens. The SWIR camera had a field of view of 95 mm and with a spatial resolution of 247 × 195 µm. The spectral resolution between the 288 spectral bands was 5.45 nm, and the images were recorded in the 950–2500 nm wavelength range. The plates were imaged with the lid opened on a black hard covered paper material with minimum absorption.
Hyperspectral image analysis
The images were analysed using Evince v.2.7.0 (Prediktera AB, Umeå, Sweden). As stated by Gowen et al., and Feng et al.,3,9 in order to account for apparatus variations, image correction and calibration is necessary in order to obtain meaningful chemical data interpretations. Calibration and image correction was done automatically in the Evince software according to Equation (1). A dark (0%) and 50% grey Zenith Allucore diffuse reflectance standard (SphereOptics GmbH, Uhldingen-Mühlhofen, Germany) was captured prior to sample image collection.
𝑛 = Pixel index variable (𝑛 = 1…𝑁) of the reorganised hypercube
I λ , = Standardised absorbance intensity, pixel 𝑛, at wavelength λ
S λ , = Sample image, pixel 𝑛, at wavelength λ D λ , = Dark reference image, pixel 𝑛, at wavelength λ G λ , = Grey reference image, pixel 𝑛, at wavelength λ
Data analysis
Image exploration and pre-processing
Before model development was possible, it was necessary to isolate the bacterial growth by removing unwanted pixels such as the agar, background information and shadows from each individual image. Exclusion of the wavelength range 2300–2500 nm removed the regions of spectral noise. Principal component analysis was performed on mean centred data with five principal components to visualise the clustering of the different information such as background, agar and bacteria on the score plots. 22 Clusters which corresponded to unwanted information in the score images were removed, thereafter PCA was recalculated. The brushing technique was used iteratively on the score plots and score images to remove the unwanted pixels, 14 until all unwanted information was removed and relevant information maintained. After acquiring the region of interest, each image comprised at least 9000 pixels per bacterial isolate.
Spectral analysis and principal component analysis
A total of twelve plates were used for the analysis. Once the images had been cleaned, all twelve images were merged to form a single image. The hyperspectral data cubes were arranged as shown in Figure 1. The data set comprised of two duplicate plates from the first day (Rep 1) and two duplicate plates from the second day (Rep 2) per bacteria. Bacterial species were arranged in rows and data collection dates in columns. To investigate the chemical properties of the bacterial species, mean raw spectra from each of the twelve plates were computed in the wavelength range 950–2400 nm. Mean spectra of the pure agar were also computed. To remove spectral noise and enhance the spectral information of the bacteria, standard normal variate (SNV) transformation was applied.
23
This attempts to reduce the light scattering making the spectra comparable in terms of absorbance values. This was followed by a Savitzky Golay (SG) smoothing filter (third order polynomial and 15-point smoothing) to increase precision by removing the ‘noise’. The SG algorithm smooths data by optimally fitting data points using least squares polynomial approximation.
24
To investigate potential differences between the bacteria, score plots, score images and loading line plots of the mosaics were evaluated interactively. Arrangement of twelve cleaned Listeria species imaged plates, consisting of duplicate plates from two different repeats stitched together to form a single mosaic.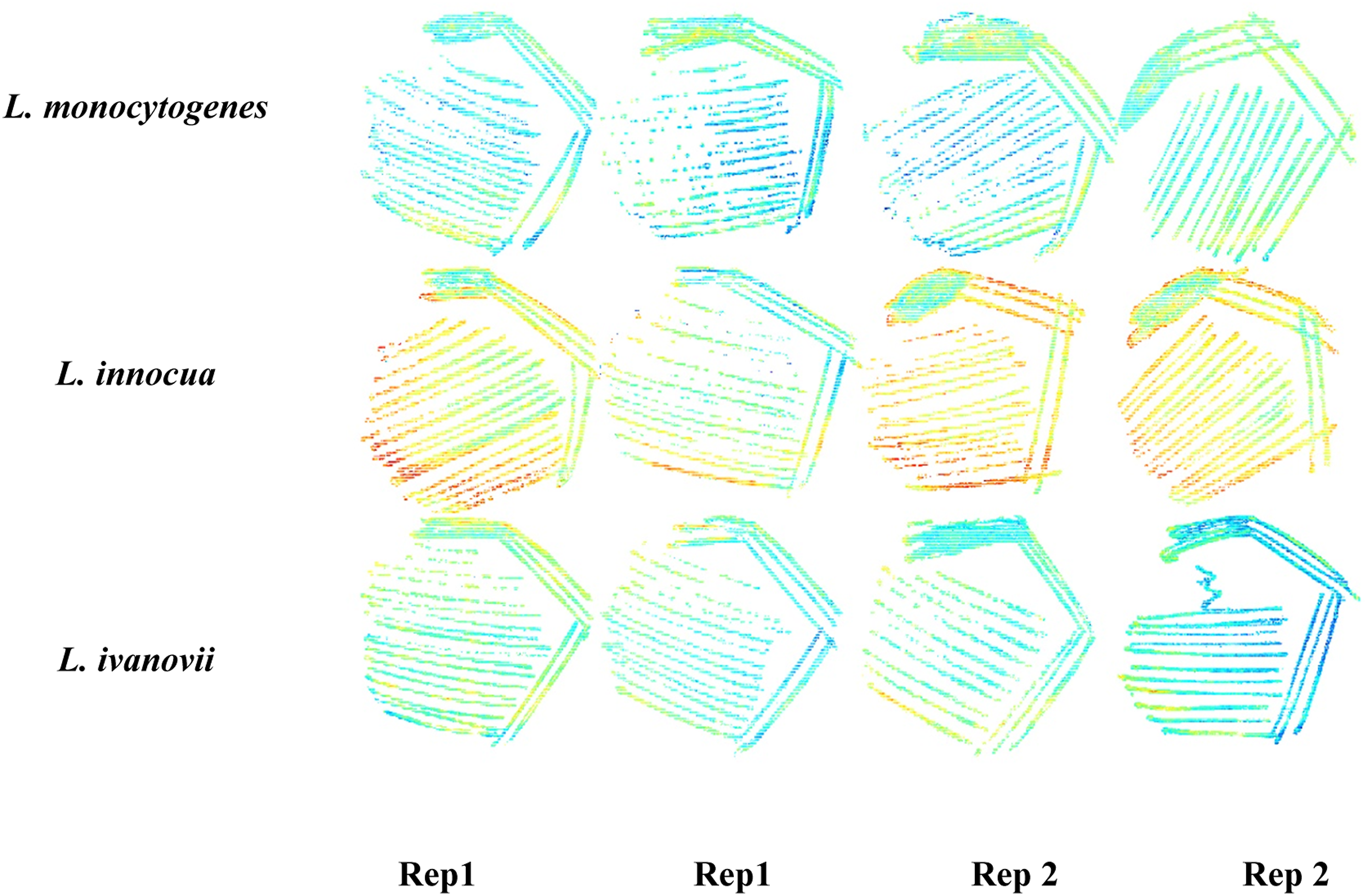
Discriminant analysis
PLS-DA was used to investigate the possibility of classifying the three bacterial species. This is a linear supervised classification model that is based on the PLS approach. 25 The multivariate calibration method seeks to find factors (latent variables) which capture maximum variance and explain variation according to classes. 26 This can therefore be used to predict class labels (Y-data) from the absorbance values (X-data). For this study, both object-wise and pixel-wise approaches were evaluated. For the pixel-wise method, each pixel in the image was used in the model calculations. 27 The dataset was split into a calibration (training) set and the validation (test) set. The calibration set comprised of 83 866 pixels which contributed 70% and the test set had 35 881 pixels which was 30% of the data set.
Sample size distribution for calibration and validation sets for object-wise analysis.
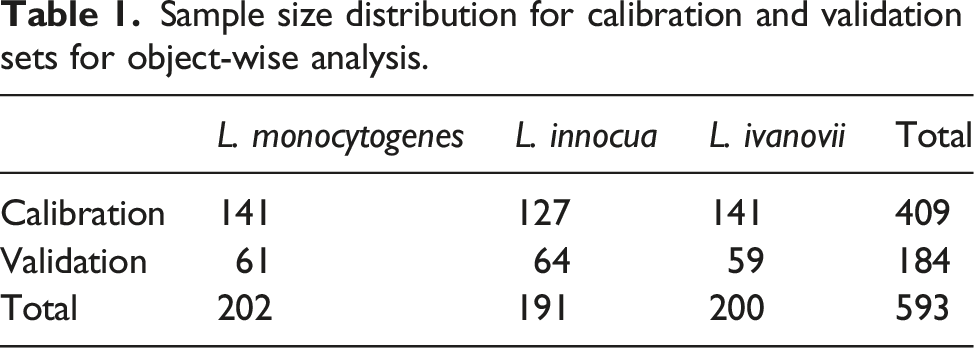
In this study the venetian blinds cross validation approach was used to determine the best number of latent variables. Cross validation enables a calculation of the best number of components to use in the model as well as allows an approximation of the performance of a model when applied to external data.
25
The venetian blinds method works by selecting every nth object into a training set. It follows a 123123123 pattern where 1, 2, 3 are data splits starting at objects numbered 1 through n.29,30 A confusion matrix was obtained, which was then used to calculate the classification accuracy of the PLS-DA models on the validation set. The classification accuracy is a metric that shows the overall performance of a model. It shows the number of correctly classified objects divided by the total number of predictions (Equation (2)).
31
Sensitivity, which is also referred as Recall, illustrates the probability of a model to correctly assign the object in the right class (Equation (3)). Specificity, on the other hand is indicative of a model’s ability to detect a true negative and is calculated as shown in Equation (4).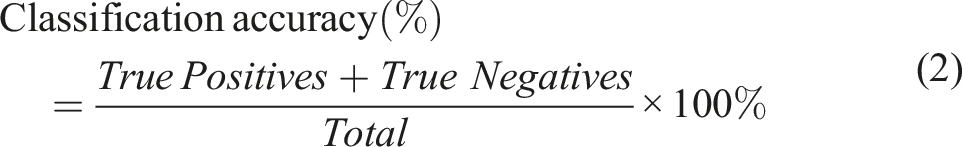
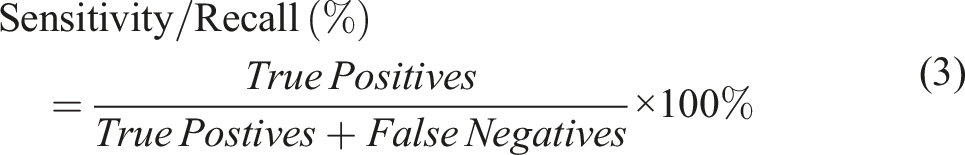
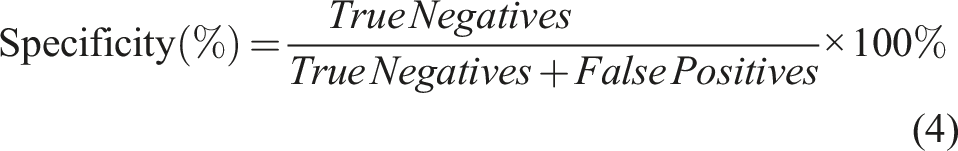
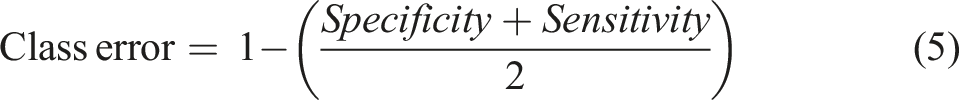
(Using a model to distinguish L. monocytogenes from L. innocua and L. ivanovii as an example)
True Positives (TP): L. monocytogenes classified correctly True Negatives (TN): L. innocua and L. ivanovii classified correctly
False Positives (FP): L. innocua and L. ivanovii incorrectly classified as L. monocytogenes
False Negatives (FN): L. monocytogenes incorrectly classified as L. innocua or L. ivanovii not classified at all
Total: TP+TN+FP+FN (The sum of all organisms used in the model)
Results and discussion
Spectral analysis of bacterial colonies
The mean absorbance spectra of the Listeria species prior to any pre-processing are displayed in Figure 2(a). Note that L. ivanovii differed considerably from the other Listeria species, presenting higher absorbance values. The spectra showed differences for all three species at 1420 nm most probably relating from O-H stretching (second overtone band O-H stretch). Water is a dominant component of the bacterial cell wall. According to Workman and Weyer
32
the wavelengths 1400 and 1900 nm are normally associated with O-H groups attributed to water molecules. Nonetheless past studies have used the interaction of water with the cell wall to explain NIR results. Dubois et al.,
17
used the O-H stretch and O-H deformation at peak 1940 nm to distinguish between Gram-positive and Gram-negative bacteria. The absorbance band at 1420 nm is suggested to be from multiple O-H groups within the peptidoglycan layer hydrogen bonded to water molecules. The absorbance band 1460 nm (Figure 2(a)) indicates differences within the three Listeria species. However, it is worth noting that L. innocua and L. monocytogenes have quite similar absorption values, with a noticeable overlap at this wavelength. The peak is likely associated with a primary amide due to the N-H stretch second overtone. This suggests the presence of similar proteins in the cell wall of the two bacterial species.33,34 Spectra of bacterial colonies (a) Original mean spectra of bacterial colonies (b) original mean spectra of bacteria and of BHI agar and (c) pre-processed mean spectra using SNV and Savitzky Golay second derivative with third order polynomial 15 points smoothing.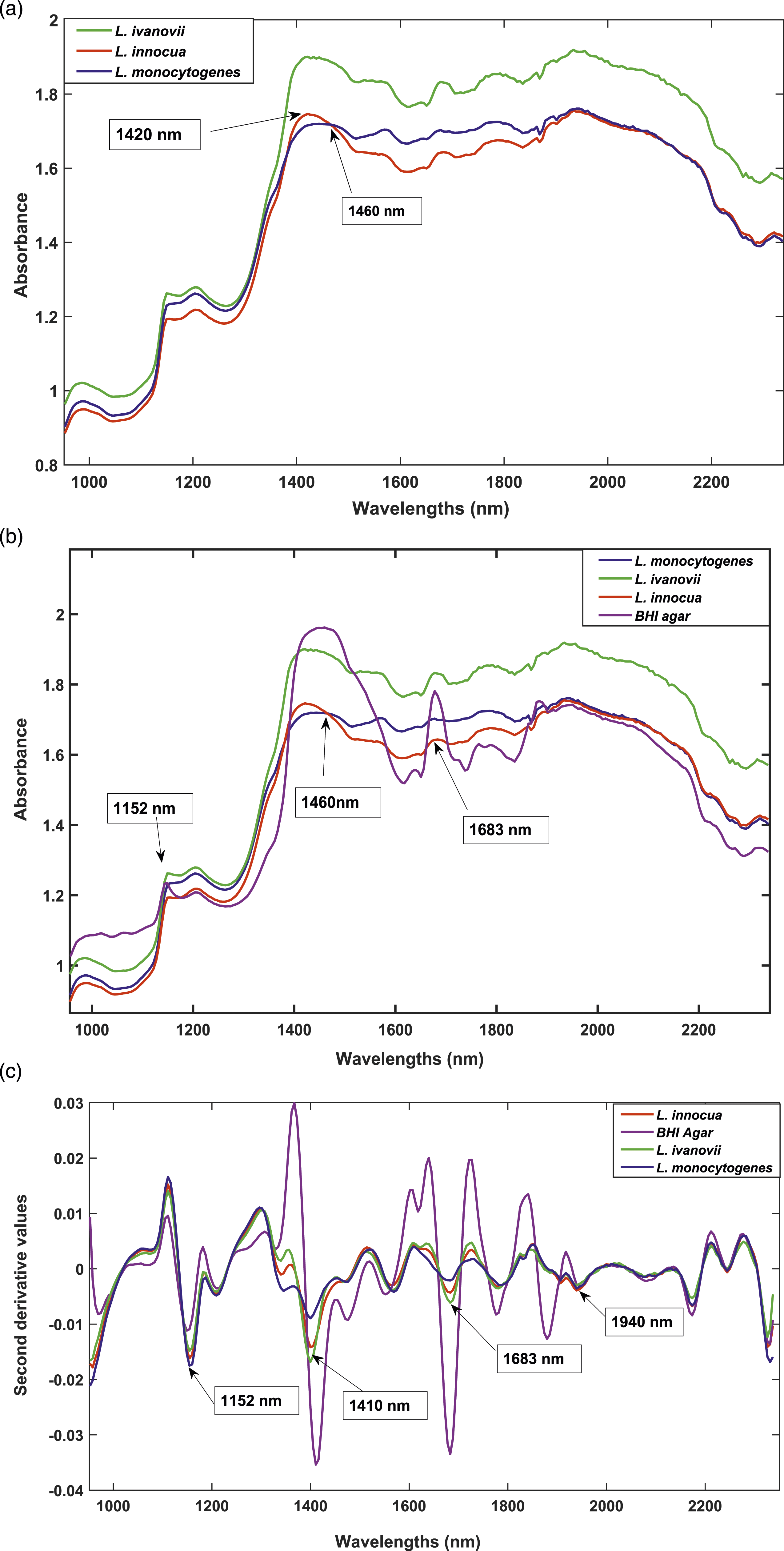
Figure 2(b) shows the raw mean spectra of the different bacterial species together with the agar. There are subtle peaks at wavelength 1683 nm resulting from all the different bacteria, despite also a dormant peak from the agar. This could suggest that the similar peaks in the original spectra of the bacterial species might also include traces of agar. However, according to Gu et al., 19 even though culture media might influence the spectrum of bacterial colonies, correct classification is still possible. Barth, 35 states that protein is measured in the NIR region as associated functional groups, such as N-H and C-H groups. In addition, Workman 36 postulates that the spectra of proteins appear as β-sheets at 1691–1680 nm. Thus, the band at 1683 nm (Figure 2(c)) validate the presence of peptide structures from the CONH2 (amides). Since BHI agar consists of protein from the brain and heart solids, this might account for the distinct peak at that wavelength. However, though the media (agar) shows the largest absorbance, it is also clear that there are differences in absorbance for the other Listeria species. Gram-positive bacteria are made up of a peptidoglycan layer embedded with teichoic acids and lipoteichoic acids. The peptidoglycan layer constitutes 40–80% by weight of a cell wall.37,38 The structure has cross linkages with four amino acids namely: L-alanine, L-lysine, D-alanine, and D-glutamine 39 Hence differences in absorbance could be as a result of variances in protein structures in the cell wall. 40
To investigate further, mean spectral data was pre-processed with SNV and Savitzky Golay 2nd derivative 3rd order polynomial with 15 points smoothing (Figure 2(c)). The correction reduced the noise and light scattering effects and clearly resolved peaks. The band at 1152 nm, was mainly attributed to the bacteria. The peak is most likely a result of the C-H second overtone stretch associated with carbohydrates in the cell wall.32,41 The spectra also showed differences for all three species at 1410 nm (second overtone band O-H stretch), most probably relating from O-H stretching of teichoic acid12,32 The differences in teichoic acids between the Listeria species could explain the differences in absorbance values, with L. monocytogenes having more of them. According to Vazquez-Boland et al., 42 Listeria have carbohydrate-containing structures on their surface, which can contribute to differences in pathogenicity within this genus. Such differences could be a result of differences in carbohydrate levels in the cell wall, especially in pathogenic bacteria. Another contributing factor might be the differences in length and structure of glycan units found in the peptidoglycan layer.14,15 There was a subtle peak at 1940 nm. This arose from a C=O stretch, second overtone CONH, related to amino acids bound with water molecules.34,43
Principal component analysis
PCA was initially employed as an unsupervised multivariate method to explore whether microbial discrimination could be achieved. Analysis of the complete dataset was done in the wavelength range 953–2300 nm. Pixel-wise analysis was done on the pre-processed data which means that each pixel’s spectrum from the cleaned mosaic image was considered during the model calculation. The score plot distinguished three clusters as shown in Figure 3(a). However, for improved visualisation, pixels were coloured manually where the blue, red, and green markers represent L. monocytogenes, L. innocua and L. ivanovii respectively (Figure 3(b)). PCA score plots of PC3 vs PC4 resulted in a cumulative variance of 9.17% which explained variation attributed to differences in carbohydrates and proteins. Pixelwise PCA analysis for L. monocytogenes (blue), L. innocua (red) and L ivanovii (green). Minimal separation of classes was observed, (a) and (b) PCA score plot of PC3 (5.63% SS) vs. PC4 (3.44% SS).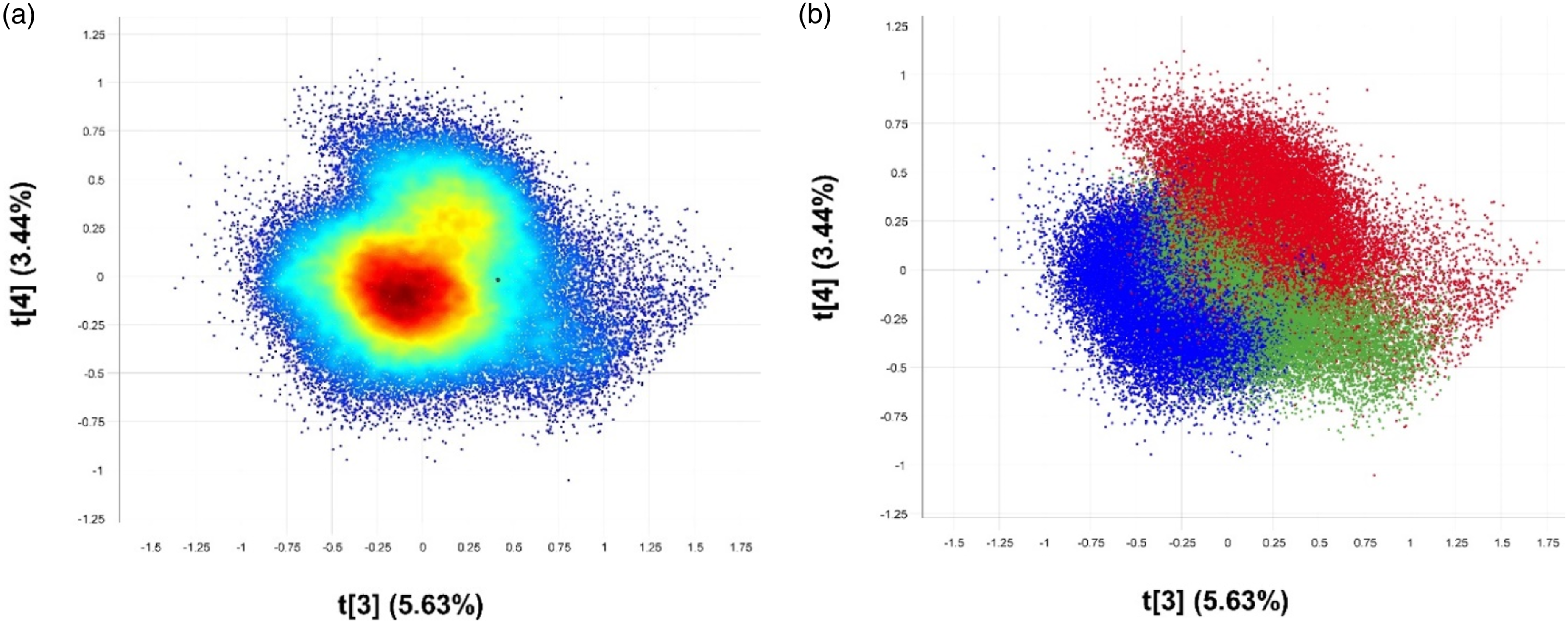
The lack of clear distinct clusters is indicative of similar chemical components since the bacterial species are from the same genus. However, there is a trend in separation along PC3 with L. monocytogenes (blue) being depicted as slightly different from L. innocua (red) and L ivanovii (green). Figure 4 shows the loading plot of PC3 indicating the variables responsible for the separation of the bacteria. Since pixels of L. monocytogenes were located in the negative region along PC3 in the score plot, they were represented by the negatively loaded peaks. Although minor, there were peaks at 1623 nm and 1740 nm from the CONH2 specifically due to C=O hydrogen bonded to the peptide link.
36
There is a notable band at 1188 nm (C-H stretch second overtone) owing to CH3 groups from carbohydrates.
34
Loading plot for PC3 showing notable peaks at 1188, 1268, 1340, 1430, 1623, 1683 and 1740 nm.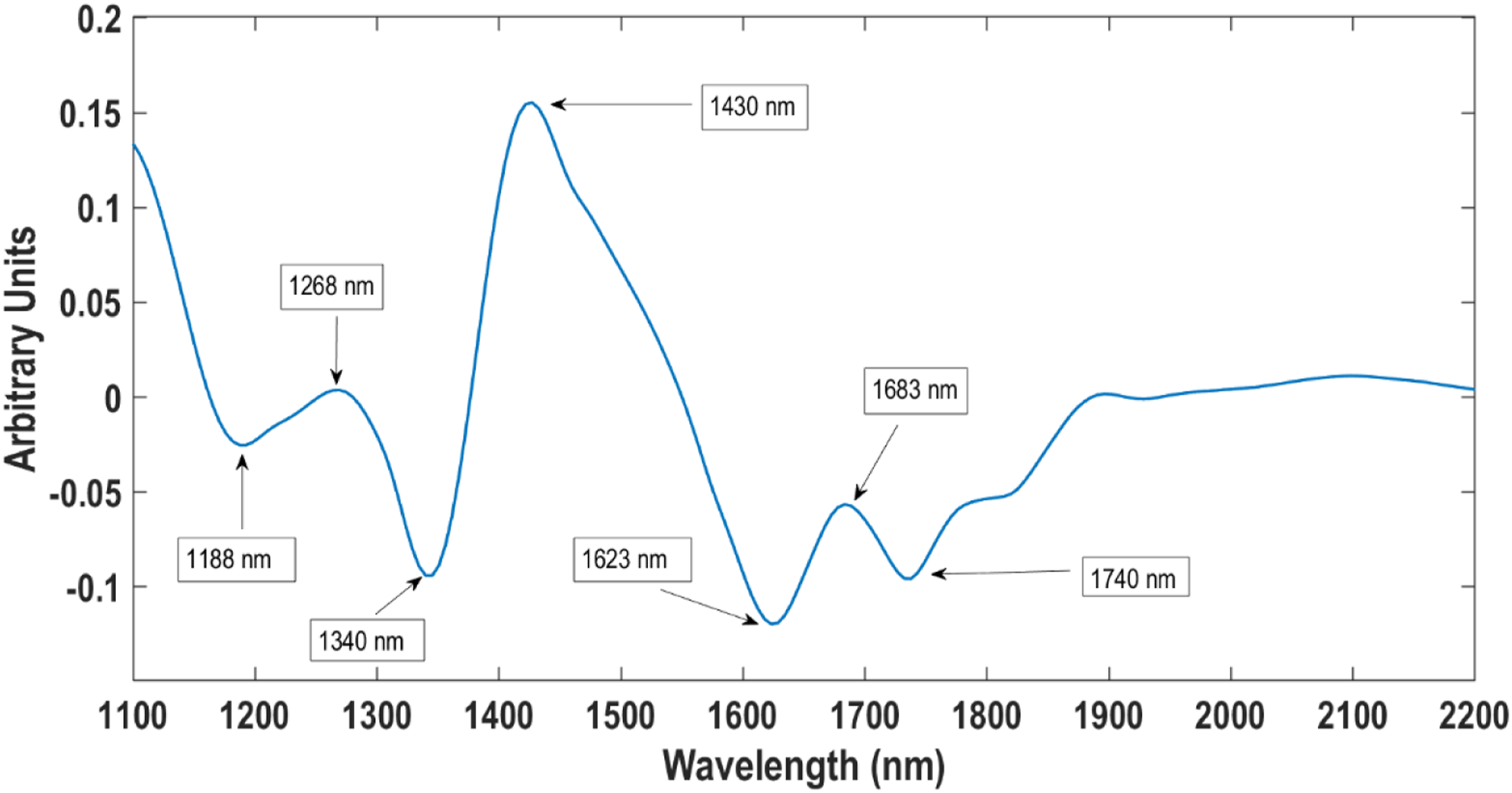
The pixels for L. innocua and L ivanovii were located in the positive region of the score plot and therefore was associated with the positive band at 1268 nm associated with carbohydrates. According to Workman, 36 the peak at 1340 can be attributed to an O-H stretch first overtone suggestive to be from hydroxyl groups in the teichoic acid structures. Also noticed was a prominent peak at 1430 nm (first overtone band N-H stretch) most probably relating to a primary amide (proteins) associated with all three bacterial species. The PCA results showed that it was possible to distinguish the three Listeria species. Nonetheless, for further classification a supervised pattern recognition method was needed.
The slight separation of pixels in the score plot of PC3 and PC4 (Figure 3(b)) could likely be due to differences in teichoic acid and protein composition. Although there is limited biochemical information on the cell wall structure of other Listeria species besides L. monocytogenes, Navarre and Schneewind 15 suggest that pathogenic bacteria might have more surface proteins than non-pathogenic bacteria. L. monocytogenes invades host cells using internalin (surface proteins) via nonphagocytic cells. 44 L innocua has been proven to lack genes that aid in coding for internalin.45,46 Hence the absence in these proteins in L. innocua might be the reason why L. monocytogenes (pathogenic) and L. innocua (non-pathogenic) cluster on opposite sides of the score plot. The loading line plot for PC3 (Figure 4) also showed a prominent positive peak at 1430 nm. The N-H stretch second overtone could be attributed to a primary amide R-C=O-NH2 group likely associated with peptidoglycan layer within the cell wall.
Partial least squares discriminant analysis model
Pixel-wise analysis
Confusion matrix prediction results for pixel-wise analysis for Listeria species using PLS-DA.
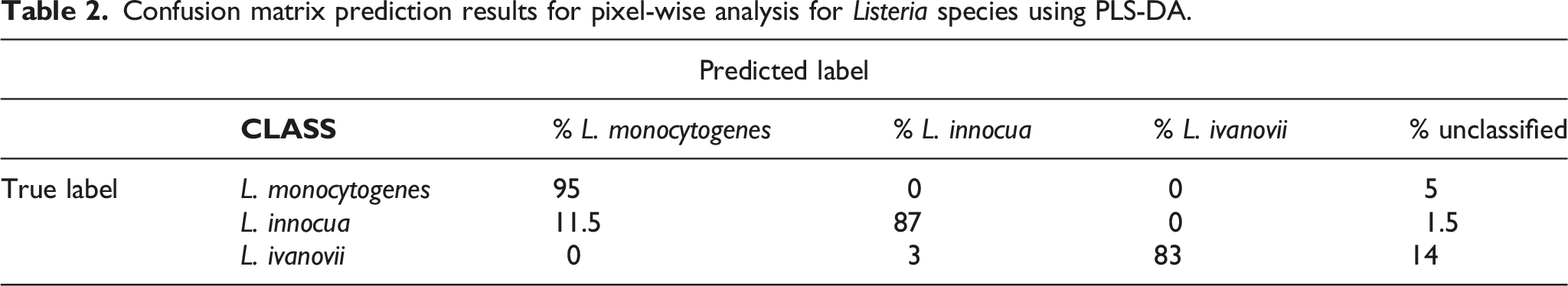

The PLS-DA results also showed pixels which had insufficient information to use for classification which were referred to as unclassified pixels. These pixels also fell below −0.5 or above 1.5 of the Y calculated critical value. L. ivanovii had the highest number of unclassified pixels, that is 1892 pixels (14%). L. monocytogenes and L. innocua also had 485 (5%) and 158 (1.5%) unclassified pixels respectively. L. ivanovii’s unclassified pixels could be as a result of similar chemical structures to those in other species, such as proteins and teichoic acids embedded in the cell walls, 50 making it difficult for the model to classify.
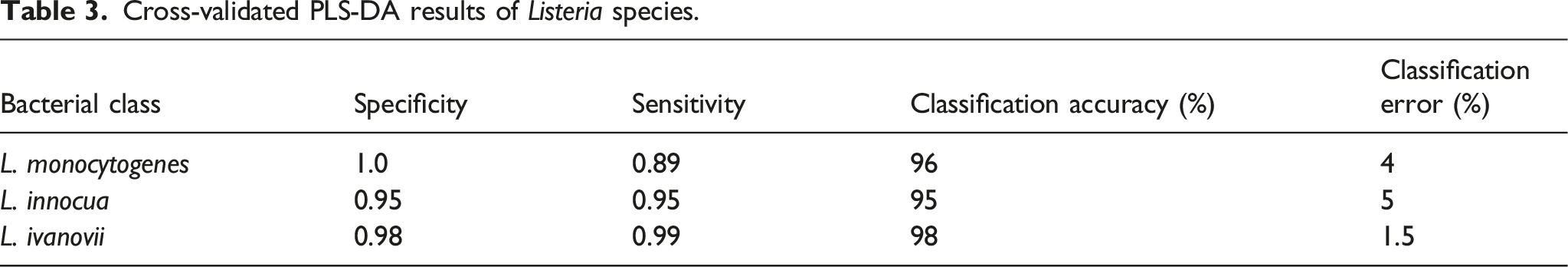
Cross-validated PLS-DA results of Listeria species.
Object-wise analysis
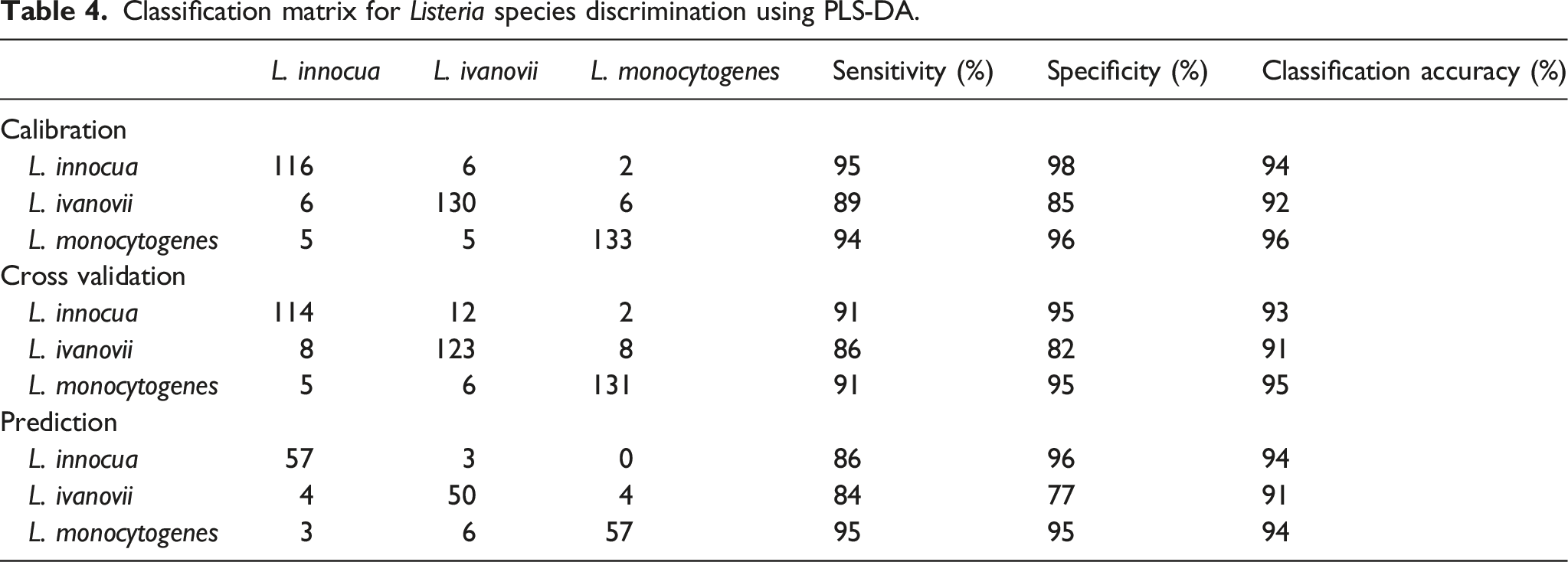
Classification matrix for Listeria species discrimination using PLS-DA.
To attain more spectral information regarding the results of the model, variable importance in projection (VIP) score plots were studied. VIP scores approximate the importance of each variable on the model and are usually used for variable selection.
53
Variables with VIP scores greater than one are regarded as important and those less than one are less important for a given model. Figure 5 shows the VIP scores for the PLS-DA model. The VIP scores plot for the Listeria species indicating the wavelengths with the most contributory variables for class discrimination.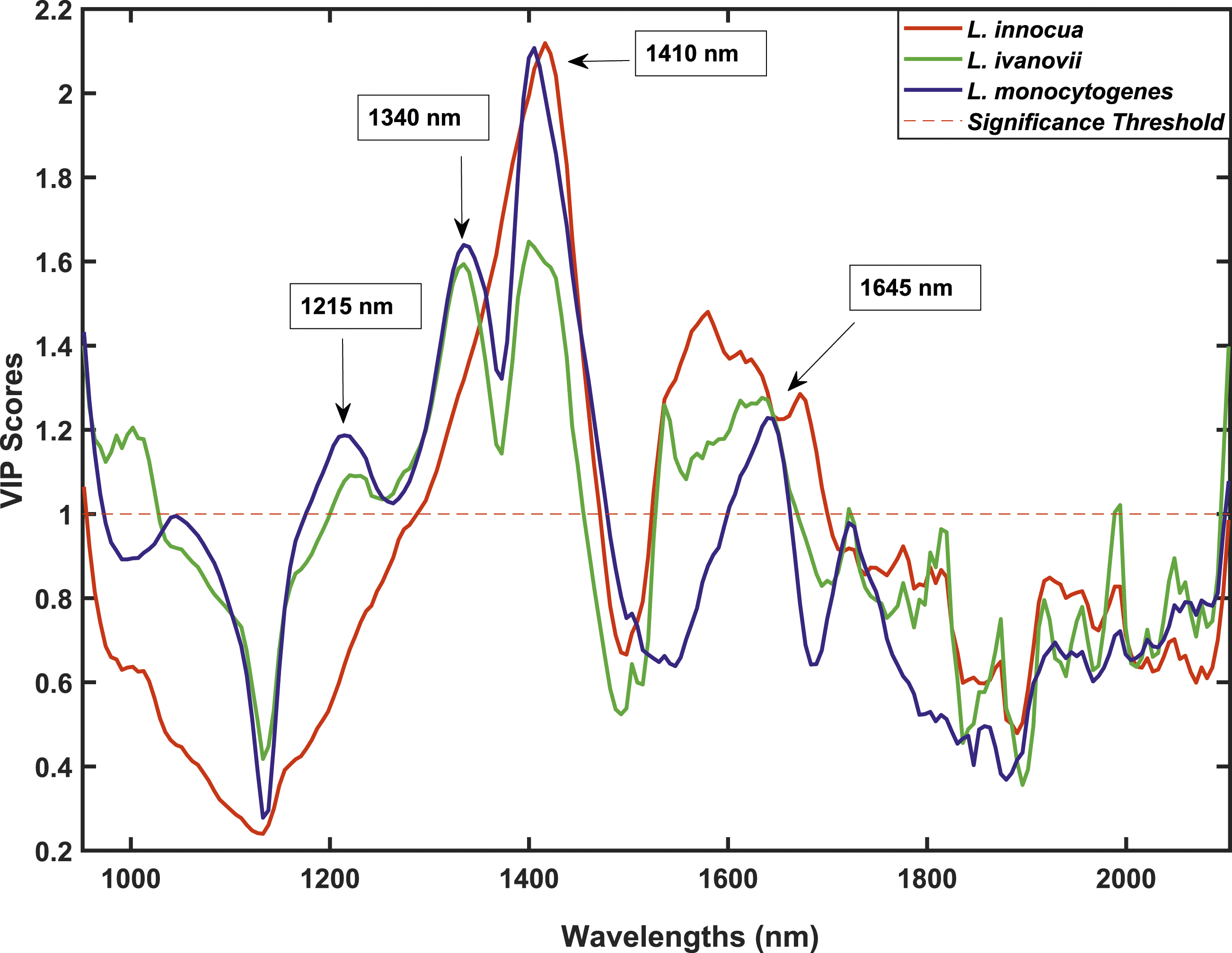
The graph shows wavelengths in the spectral zone 1200 and 1800 nm being more distinct and hence more important. The bands at 1215, 1340, 1410 and 1645 nm were identified as the most contributing bands for class differentiation. Absorbances at 1215 and 1340 nm might be from CH stretch second overtone 54 and O-H stretch first overtone from isolated hydroxyl groups 55 respectively. It is quite interesting to note that at bands 1215 and 1340 nm, L. ivanovii and L. monocytogenes are comparable as opposed to L. innocua. This can be attributed to differences in amounts of lipoteichoic acid in the cell wall of different Listeria spp. Previous work done by Ruhland and Fiedler 56 showed that different Listeria strains have different lipoteichoic acids in some cases being absent all together. This might also be the case within the Listeria species investigated. In addition, hydroxyl-polycyclic hydrocarbons most likely from teichoic acids and lipoteichoic acids can be used to explain the band at 1410 nm. 36 Research by Pucciarelli et al., 57 points out that the wall teichoic acids of certain strains of L. innocua and L. monocytogenes have more N-acetylglucosamine than other Listeria species. These differences in teichoic acids can be considered as a source of discrimination. The band 1645 nm is associated with a C-H stretch first overtone probably arising from glucose and lipid ethers.40,58 The band is most likely a result of sugars in the peptidoglycan layer. Peptidoglycan is a polysaccharide made up of amino acids and sugars. 40 Since all Gram-positive bacteria (Listeria species included) have a pronounced peptidoglycan layer, it explains why theses wavelengths might be of most interest.37,59
Conclusion
The main aim of the study was to investigate the use of NIR hyperspectral imaging for detection and classification of Listeria species. Although there were not many differences in the spectra of the Listeria species, there was still some useful information in the spectral range of 1000–1800 nm. Spectra obtained showed a similar trend due to likenesses of morphological, biochemical, and serological properties of the Listeria spp. However, results from the spectral profiles also showed that differences in protein structures, teichoic acid and carbohydrates could be possible sources of variation between the Listeria species. The PCA model was used to explore the data trends and results showed partial separation of the classes on the score plots. With such promising results, data was further investigated using a supervised classification model, PLS-DA. The results obtained were relatively comparable to each other, with sensitivity values and classification accuracies above 85 and 90% respectively. The VIP scores indicated notable bands at 1215 nm (carbohydrate related), 1340 nm (lipoteichoic acid related), 1410 nm (wall teichoic acid related), and 1645 nm (carbohydrate related) which contributed to class differentiation. Since the PLS-DA produced good results there was no reason to explore non-linear methods. However, future work should focus on ways of optimizing the model such as selecting wavelengths of most importance. By doing so, the model may become more efficient thus improving processing time. The method can then be used in real time in conjunction with other conventional methods of analysis. In addition, since L. monocytogenes was classified excellently and is the pathogen of most concern, future work should also focus on differentiating it from other Listeria sub-species and serotypes.
Footnotes
Acknowledgements
The authors would also like to acknowledge Stellenbosch University Postgraduate Scholarship Programme for their financial support to the student Rumbidzai Matenda.
Author contribution
All authors conceived and designed the research. Microbial laboratory work was done mainly by R.T.M with assistance from D.R. Data analysis was done primarily by R.T.M with the assistance of P.J.W. Writing of the manuscript (first draft) was done by R.T.M. The manuscript was read, reviewed, edited, and approved by all authors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is based on the research supported wholly/in part by the National Research Foundation of South Africa (Grant Numbers: 137998). The NRF’s national equipment program is acknowledged for funding the NIR hyperspectral imaging instruments through the grant UID 105652. The hyperspectral imaging work performed in this article was conducted at the Vibrational Spectroscopy Unit which belongs to Central Analytical Facility at Stellenbosch University.