Abstract
Currently, there is a significant shortage of transplantable organs for patients in need. Interspecies chimerism and blastocyst complementation are alternatives for generating transplantable human organs in host animals such as pigs to meet this shortage. While successful interspecies chimerism and organ generation have been observed between evolutionarily close species such as rat and mouse, barriers still exist for more distant species pairs such as human–mouse, marmoset–mouse, human–pig, and others. One of the proposed barriers to chimerism is the difference in developmental stages between the donor cells and the host embryo at the time the cells are introduced into the host embryo. Hence, there is a logical effort to stage-match the donor cells with the host embryos for enhancing interspecies chimerism. In this study, we used an in silico approach to simultaneously stage-match the early developing embryos of four species, including human, marmoset, mouse, and pig based on transcriptome similarities. We used an unsupervised clustering algorithm to simultaneously stage-match all four species as well as Spearman’s correlation analyses to stage-match pairs of donor–host species. From our stage-matching analyses, we found that the four stages that best matched with each other are the human blastocyst (E6/E7), the gastrulating mouse embryo (E6–E6.75), the marmoset late inner cell mass, and the pig late blastocyst. We further demonstrated that human pluripotent stem cells best matched with the mouse post-implantation stages. We also performed ontology analysis of the genes upregulated and commonly expressed between donor–host species pairs at their best matched stages. The stage-matching results predicted by this study will inform in vivo and in vitro interspecies chimerism and blastocyst complementation studies and can be used to match donor cells with host embryos between multiple species pairs to enhance chimerism for organogenesis.
Keywords
Introduction
Organ transplantation is the ultimate standard of care to replace liver, kidney, pancreas, heart, and lung. Many factors influence the success of transplantation, including intensive immunosuppressive agents against rejection, surgical complications, and a source of available organs1–3. Currently, a major challenge that affects the field of organ transplantation is the overwhelming demand of organs compared with the available supply 1 . The total number of candidates in the United States on the waiting list is 106,105 as of April 2022 4 . Due to increasing organ demand, there is a need for alternative approaches to generate organs, tissues, and cells for transplantation. Studies in regenerative medicine studies are now exploring the potential prospect of reprogramming technologies and the development of an inexhaustible source of cells and organs through genetic engineering 5 . The development of organs via scaffolding and in vitro conditions faces the major challenge of constructing thick, complex tissue with vascularization that can be maintained in vitro until it is needed and in vivo upon implantation 6 . That together with the limited advance in clinical trials for the transplantation of engineered organs has further complicated the field. However, an in vivo method like blastocyst complementation provides a novel approach to generate exogenic organs of one species in another via the creation of chimeric animals.
Intraspecies and interspecies blastocyst complementation has produced various exogenic organs. Some of the successful intraspecies exogenic organs include mouse–mouse lung, thyroid, liver, and kidney, and pig–pig eye and liver7–11. Intraspecies derived organs appear to be much easier to generate than interspecies chimeras. Two of the very few successful interspecies exogenic organs generated via blastocyst complementation are the rat–mouse pancreas 12 and rat–mouse kidney 13 . The ability to produce interspecies chimeras has proven to be much more challenging than anticipated and predicted. With the generation of these exogenic organs, we are learning that several interspecies chimeric barriers determine and even limit the success of organ transplantation. Scientific barriers that impede chimera formation include (1) evolutionary distance between species, (2) need for stage-matching between species for temporal and spatial synchronization of developmental stages, (3) cell competition and the survival of pluripotent stem cells (PSCs) and embryos, and (4) compatibility of ligand–receptor signaling between species14–19. Of these barriers, evolutionary distance and stage-matching provide promising avenues to better understand the relationships between transcriptomes and genome regulation of donor cells and host embryos used to produce interspecies chimeras. For example, rodents and humans are approximately 90 million years divergent from one another, while rats and mice are only 20.9 million years in divergence 15 . The greater evolutionary divergence between two species, the less cellular and molecular information is conserved. However, key transcripts and genes are required for early embryogenesis and contain stage-specific time points with highly conserved networks that are necessary to maintain and express the necessary transcriptional machinery. Despite many highly conserved transcripts and genes, there remain species-specific differences that contribute to a species’ stage-specificity and variations in temporal development, which is likely influenced by different gestational periods. The combination of species’ stage-specificity, temporal development, and highly conserved developmental networks in early embryogenesis suggest that the greater the transcriptomic similarity between any two developmental stages, the greater probability of generating an efficient chimera 20 .
Numerous studies have focused on in silico stage-matching in interspecies chimerism and blastocyst complementation in human, non-human primate, mouse, and pig early stage embryos. Analysis of single-cell RNA-sequencing (scRNA-seq) data identified transcripts, epigenetic modifiers, and pluripotency factors that were stage-specific to the human, marmoset, and mouse 17 . Another study compared the transcriptomes of pig and mouse preimplantation embryos and found that regulatory networks were conserved during the first lineage segregation and primitive endoderm differentiation, but not during that of the ectoderm development 21 . An additional in silico study demonstrated that the transcriptome of human embryonic stem cells (hESC) was comparable with the pig epiblast cells in late-stage pig blastocysts and indicated that the transcriptome of the donor cells plays a role in forming efficient chimeras 22 . The authors suggested that stage-matching based on transcriptome similarities will allow synchronization of development between donor and host cells in the developing chimeric embryo.
However, there is a paucity of in silico studies that compare all four species simultaneously, that is, human, marmoset, mouse, and pig. Many current studies perform pairwise comparisons between two species, limiting the ability to stage-match the different developmental stages of multiple species with one another.
In this study, we examined early embryos from humans, marmosets, mice, and pigs. We collected scRNA-seq datasets from these four species’ early embryos and analyzed them using unbiased algorithms. The methods used in the analysis generated two sets of interspecies stage-matching results that were compared with one another and to published studies. Further ontology analysis provided additional insights into important developmental pathways that were shared between the best matched stages for each donor–host pair.
Methods
Data Sources
The scRNA-seq datasets were obtained from previous studies that focused on transcriptomes from different stages of preimplantation development in human, marmoset, mouse, and pig embryos17,22–27. Human scRNA-seq datasets were downloaded from the Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) under accession numbers GSE36552 and GSE66507; mouse datasets from GSE45719 and GSE100597; and pig datasets from GSE112380. EMBL (European Molecular Biology Laboratory) scRNA-seq datasets were also obtained for human datasets under accession number E-MTAB-3929 and marmoset datasets under accession number E-MTAB-7078.
Sequencing Data Processing
Raw sequencing reads were analyzed using a customized pipeline (CHURP; https://github.com/msi-ris/CHURP) developed and maintained by the University of Minnesota Supercomputing Institute (MSI). Briefly, FastQC v0.11.7 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) was used to examine the sequencing quality of the FASTQ files. Adapters and low-quality reads were trimmed using Trimmomatic v0.33 (http://www.usadellab.org/cms/index.php?page=trimmomatic). For additional quality check, FastQC was used on the post-trimming sequences to ensure successful adapter and quality trimming. The remaining sequences were then mapped to the corresponding reference genome (Human: hg19, Mouse: mm10, Pig: susScr3 and Marmoset: C_jacchus3.2.1) using HISAT2 v2.1.0 (https://ccb.jhu.edu/software/hisat2/index.shtml), and transcript abundance was counted using subread v1.6.2 (http://subread.sourceforge.net/)
Integrating Datasets Across Species for Single-Cell Data Analysis
The raw-count matrices were converted into a Seurat object and analyzed with R package SEURAT v4.1.0 (https://satijalab.org/seurat/). They were filtered for mitochondrial contamination and normalized using the negative binomial regression method implemented in the SCTransform wrapper in Seurat. For integration, the 6,000 most conserved genes in all samples were identified with SelectIntegrationFeatures function. Subsequently, the PrepSCTIntegration function was used to identify conserved features for integrating the datasets. Integration anchors were identified using FindIntegrationAnchors function with a scTransform normalization method. Seurat’s IntegrateData function was implemented to integrate the datasets.
Clustering Algorithm for Stage-Matching Species Across Four Species
Principal components analysis (PCA) was used to reduce the dimensionality of the data prior to performing clustering. To calculate the principal components (PCs), ElbowPlot function was performed to select suitable dimensionality, and PCA was conducted using RunPCA function with 50 dimensions. The Uniform Manifold Approximation and Projection (UMAP) algorithm in the Seurat RunUMAP function was used, and the top 20 PCs were embedded into two dimensions. To define cell clusters, an unsupervised Shared Nearest Neighbor (SNN) cell graph was created using the Seurat FindNeighbors function based on the top 20 PCs. The Seurat FindClusters function was used to detect communities using the Louvain clustering algorithm. The FindClusters function also has a resolution parameter, which allowed to control the granularity of the clusters. A lower value of resolution results in a higher number of clusters, and a higher value of resolution results in a fewer number of clusters. Data were clustered using FindClusters with a resolution = 0.7, algorithm = 1 (Louvain algorithm) parameters. The Seurat Dimplot function was used to plot PCA and UMAP, and the parameters split.by = “species” and group.by = “orig.ident” were used to visualize each single-species cluster in the cells.
Spearman’s Correlation Analysis for Pairwise Donor–Host Stage-Matching
After each species dataset was normalized using Seurat function SCTransform, the subset of commonly expressed genes between two species was created using Seurat SubsetData function. The top 2,000 feature genes among commonly expressed genes were identified using SelectIntegrationFeatures function in Seurat. Integration anchors were identified using Seurat PrepSCTIntegration and PrepSCTIntegration functions, and subsequent integration of two species datasets was performed using Seurat IntegrateData function. For the correlation analysis, average expression was calculated using integrated matrix and AverageExpression function in Seurat. Subsequently, a matrix of Spearman correlation coefficients was calculated using rcorr function from the Hmisc package in R (https://cran.rproject.org/web/packages/Hmisc/index.html), and the matrix was visualized using corrplot function from the corrplot package in R (github.com/taiyun/corrplot).
KEGG (Kyoto Encyclopedia of Genes and Genomes) Analysis and GO (Gene Ontology) Analysis for Pairwise Donor–Host Best Matched Stages
The differential expression analysis across all stages of each species was performed using the wilcoxauc function from R package presto (https://github.com/immunogenomics/presto), and the top 3,000 genes of the results were selected by P value < 0.1 and area under the receiver operator curve (AUC) >0.5 using presto top_markers function. Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment pathway analysis were performed using enricher functions (enrichGO, enrichKEGG) in the clusterProfiler package (https://bioconductor.org/packages/clusterProfiler) to obtain the biological function of intersecting genes between the best matched stages of two different species. GO terms with a P value < 0.1 were considered significantly enriched. Bar plots were used to visualize enrichment data using R package enrichplot (https://github.com/YuLab-SMU/enrichplot).
Results
Although significant progress has been made in the field of interspecies chimerism, it has become apparent that as the evolutionary distance between the donor and host species increases, the efficiency of chimerism decreases. A greater efficiency of chimerism is observed between closely related species such as rat and mouse when compared with more distant species such as human–mouse, human–pig, and others. While multiple factors play a role in successful chimerism, the difference in developmental stages between the donor and host embryo cells, especially between distant and diverse species, has been proposed as one of the major barriers to efficient chimerism16,22,28. It has been proposed that stage-matching the donor and host cells, at the time of generating chimeras, would overcome this barrier16,22. In this study, we performed in silico transcriptome-based stage-matching between species relevant to interspecies chimerism. We hypothesized that for the in silico stage-matching, the developmental stages with the highest similarities in transcriptome programs are the ones that best match with each other and are most likely to give rise to successful chimeras in vivo. We surmised that this stage-matching approach will help us overcome the interspecies barrier and generate efficient chimeras between evolutionarily distant species in vivo. In this study, we performed scRNA-seq analyses of early embryos across four species relevant to interspecies chimerism studies involving human, marmoset, mouse, and pig. Human and marmoset datasets were chosen, as they were two primate donor species evolutionarily closer to each other and are clinically relevant for future human transplantation studies. The mouse was chosen because it is the most common studied rodent host for interspecies chimerism. Last, the pig was selected, as the end goal in the field is to produce human organs in large host animals with similar organ sizes to humans, such as pigs. The scRNA-seq datasets derived from the embryos of each of the species were compiled together from previous studies17,22–27. The stages analyzed for human23–25, marmoset 17 , mouse26,27, and pig 22 are shown in Fig. 1A. The predicted stage-matching results from the scRNA-seq analysis were then validated by comparison with donor–host developmental stages from previously published in silico as well as in vitro and in vivo stage-matching experiments reporting successful interspecies chimerism.

Single-cell RNA analysis of early embryos across species: (A) Datasets of human, marmoset, mouse, and pig that were used for the single-cell stage-matching analysis. The datasets were collected from publicly available scRNA-seq databases. In some species such as the human and mouse, a single time point consists of more than one dataset. Created with Biorender.com. (B) Principal components analysis (PCA) of the embryonic datasets for each of the four species. (C) Uniform Manifold Approximation and Projection (UMAP) of the embryonic datasets for each of the four species. (D) Embryonic datasets in each species were divided into 20 clusters based on similarities in gene expression across all the species.
Early Embryogenesis Pathways Are Conserved Across Species
The scRNA-seq datasets for the embryos from the four species—human, marmoset, mouse, and pig—were analyzed using PCA and UMAP. For each of the species, PCA plots and UMAPs were generated across all the embryonic developmental stages as detailed in the “Methods” section. The PCA identified specific developmental clusters that are conserved across species (Fig. 1B). For the pig analysis, slight differences in the pattern of PCA clusters were observed due to limited data availability that were restricted to morula through late blastocyst pig stages. The scRNA-seq data for the earlier pig embryo stages from zygote to four-cell stage were not publicly available for the final analysis. The UMAP for each species was performed to identify developmental pseudopaths and depicted in Fig. 1C where cells in each cluster and their respective developmental stages can be observed. Similar embryonic stages clustered closely together and the developmental pseudopath was conserved across species (Fig. 1C). This analysis enabled visualization of cell fate transitions from the earliest embryonic stages and up to late blastocyst and gastrulation stages based on single-cell gene expression programs. These findings validate the potential of using single-cell transcriptomes across species to compare developmental stages and generate better stage-matching for interspecies chimerism.
In Silico Prediction of Best Matched Developmental Stages Across Species
To simultaneously stage-match the four species with each other, we compared the similarities in genome-wide gene expression programs during their embryonic development across all the species. The datasets consisting of all developmental stages, as shown in Fig. 1C, were divided into 20 clusters (0–19) for each of the species based on similarities in gene expression as detailed in the “Methods” section.
The 20 clusters are depicted as UMAP for each species in Fig. 1D, and each of the 20 clusters contained cell types that corresponded to matching developmental stages across species. For example, cluster 1 in human contained cells with the most similar transcriptome to cells in cluster 1 of marmoset, mouse, and pig. The clusters were numbered from 0 to 19 as shown in Fig. 1D. For each of the 20 clusters, the datasets corresponding to embryonic stages of each species were analyzed. Previous in vitro studies demonstrated that hESCs contribute better to mouse gastrulating embryos when compared with mouse blastocysts 29 . Therefore, in this study, we focused on extracting clusters containing the mouse gastrulating embryo stages E6–E6.5 datasets. The human, marmoset, and pig embryonic stages present in these clusters and hence similar to the E6 mouse stages were then identified. According to previously successful in vitro stage-matching chimerism studies, the mouse E6 gastrulating datasets should closely resemble those from hESCs, which are extracted from human E6–E7 blastocyst stages (mid to late blastocyst) 29 . From the clusters containing the mouse E6 datasets, that is, clusters 1, 4, 8, and 17, it was observed that human cells specific to stages E6, E7, and late blastocyst (E7) were largely present. Human datasets specific to epiblast, trophectoderm (TE), and primitive endoderm of E6–E7 blastocysts were also present in these clusters (Fig. 2). For the marmoset and pig embryo stages, the marmoset late inner cell mass (ICM), pig late epiblast, and pig late hypoblast were present in these clusters, which demonstrated similarities between these datasets and the human E6–E7 blastocysts. Cluster 10 (Fig. 2E) further highlighted the similarities between the human E6, E7, and late blastocyst stages with the marmoset late ICM and the pig late epiblast stage. The mouse embryonic stage in cluster 10 is E5 (E4.5–E5.5), which represents the stage of development just prior to the onset of gastrulation. The results of the clustering analysis were summarized in Fig. 2F. The stages between the four species that best matched with each other were the human E6–E7 blastocyst, the mouse E6 (E6–E6.5) gastrulating embryo, the marmoset late ICM, and the pig late epiblast. Clustering analysis of scRNA-seq datasets across the four species was used to predict embryonic stages based on transcriptomic similarities. These predicted results can be further tested by performing in vitro and in vivo chimerism stage-matching experiments to overcome the interspecies chimerism barrier of differential developmental stages, generating successful interspecies chimeras. Future in silico studies would involve analyzing the remaining clusters (Supplementary Fig. 3) and predicting other combinations of developmental stages that match across the four species.
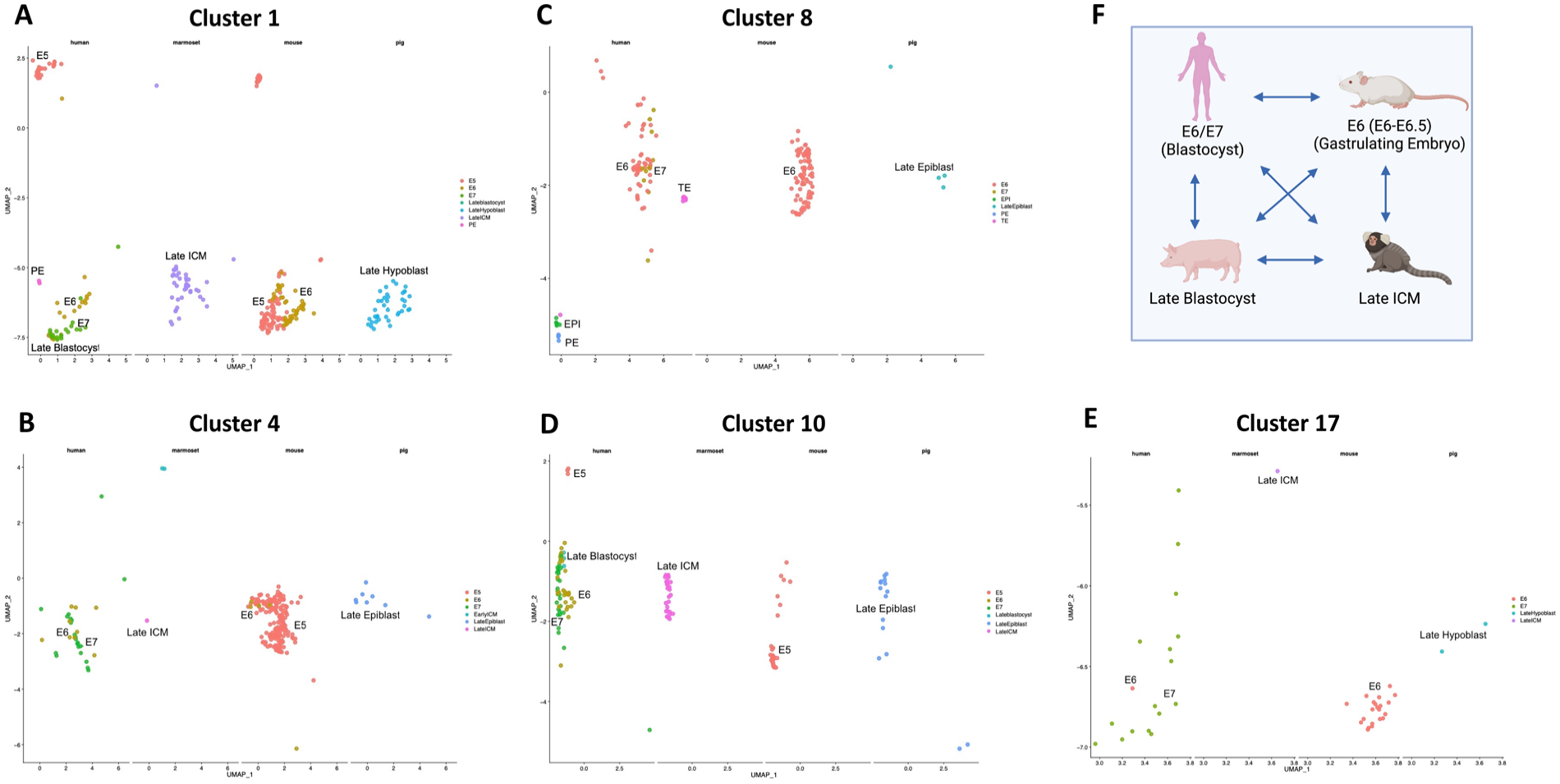
In silico stage-matching across species. Clusters containing the mouse E6 gastrulating stages (E6–E6.5) were selected for analysis (see Fig. 1). (A–D) depict clusters 1, 4, 8, and 17, respectively. These clusters represent the mouse E6 gastrulating embryos. Human stage-matching corresponds to the blastocyst stages E6, E7, late blastocyst (E7), and components of the blastocyst such as the epiblast and primitive endoderm. For marmoset stage-matching within these clusters, late inner cell mass (ICM) was found. For pig stage-matching, these clusters correspond to the cell types in the late blastocyst, that is, late epiblast and late hypoblast. (E) Cluster 10 where the human, marmoset, and pig stages match with those in clusters 1, 4, 8, and 17. The mouse stage in cluster 10 is E5, which includes data from the late blastocyst E4.5 to the early gastrulating E5.5 embryo. (F) Summary of the stage-matching results from the clustering algorithm analysis for the four species. Created with Biorender.com.
Human PSCs Cluster With Mouse Post-Implantation Embryonic Stages
In addition to stage-matching embryos across the four species, we used a clustering algorithm to stage-match hPSCs (hESCs and hiPSCs) with the mouse embryonic developmental stages and predict the mouse embryo stages that were most similar to the hPSCs. The analysis contained two human induced pluripotent stem cells (hiPSC) datasets from (1) a feeder layer-free hiPSCs line and (2) hiPSCs grown on SNL fibroblasts. The hESC datasets were obtained from a study that compared hESCs at day 0 and day 10 in culture 23 . The same mouse datasets used in the previous analysis were used for this analysis26,27. The hiPSCs and hESCs clustered with mouse early embryonic stages as shown in Supplementary Fig. 1. The hiPSCs clustered closely with the post-implantation E6.75 mouse embryo stages (Supplementary Fig. 1). The hESCs at day 0 clustered closely with the post-implantation E6.5 mouse embryo stages. The hESCs at day 10 clustered with multiple post-implantation stages of the mouse embryo from E5.5–E6.75 (Supplementary Fig. 1). In all cases, the hPSCs clustered with mouse post-implantation embryonic stages, which is in agreement with previously reported results16,23. No clustering of the hPSCs with any of the mouse preimplantation stages was observed. This may be because the hESCs are more “primed” compared with mouse “naïve” blastocyst cells and match better with the “primed” epiblast cells of the mouse gastrulating embryo 30 . Some variation in clustering was observed between the hiPSCs and the hESCs with the mouse embryos. This variability might have been due to differences in their culture conditions and highlights the impact of culture conditions on the transcriptome of stem cells.
Stage-Matching of Developmental Stages Using Spearman’s Correlation Analysis
In addition to the clustering algorithm for stage-matching, we used Spearman’s correlation analyses for pairwise comparisons between the four species as described in the methods. By performing Spearman’s correlation between the embryonic stages, the stage-matching between species pairs can be determined and ranked. We performed the correlation analysis between donor–host species combinations of human–mouse, marmoset–mouse, human–pig, and marmoset–pig. Subsequently, we determined which developmental stages had the highest correlation coefficients between the species pairs as highlighted in Fig. 3. For the human–mouse analysis, the human E6 and E7 blastocyst stages were best matched with the mouse E6.5 gastrulating embryo. The hESCs were best matched with the mouse E6.75 gastrulating embryo (Fig. 3A). For the marmoset–mouse analysis, the marmoset late ICM was best matched with the mouse E6.75 gastrulating embryo (Fig. 3B). For the human–pig correlation analysis, the human E6 and E7 blastocysts as well as the hESCs were best matched with the pig spherical epiblast (~E10–E11 early post-implantation embryo or later blastocyst stage) (Fig. 3C). Our analyses agree with previously published results where the authors proposed that hESCs introduced into late pig blastocysts rather than early pig blastocysts will give rise to successful interspecies chimeras 22 . For the marmoset–pig analysis, the marmoset late ICM was best matched with the pig spherical epiblast (Fig. 3D). The best matched developmental stages obtained by Spearman’s correlation analysis (Fig. 3E) are in agreement with the results from our clustering algorithm approach (Fig. 2F). This agreement between the clustering algorithm and Spearman’s correlation analysis underscores the strength of our in silico approach for stage-matching developmental stages across diverse species.
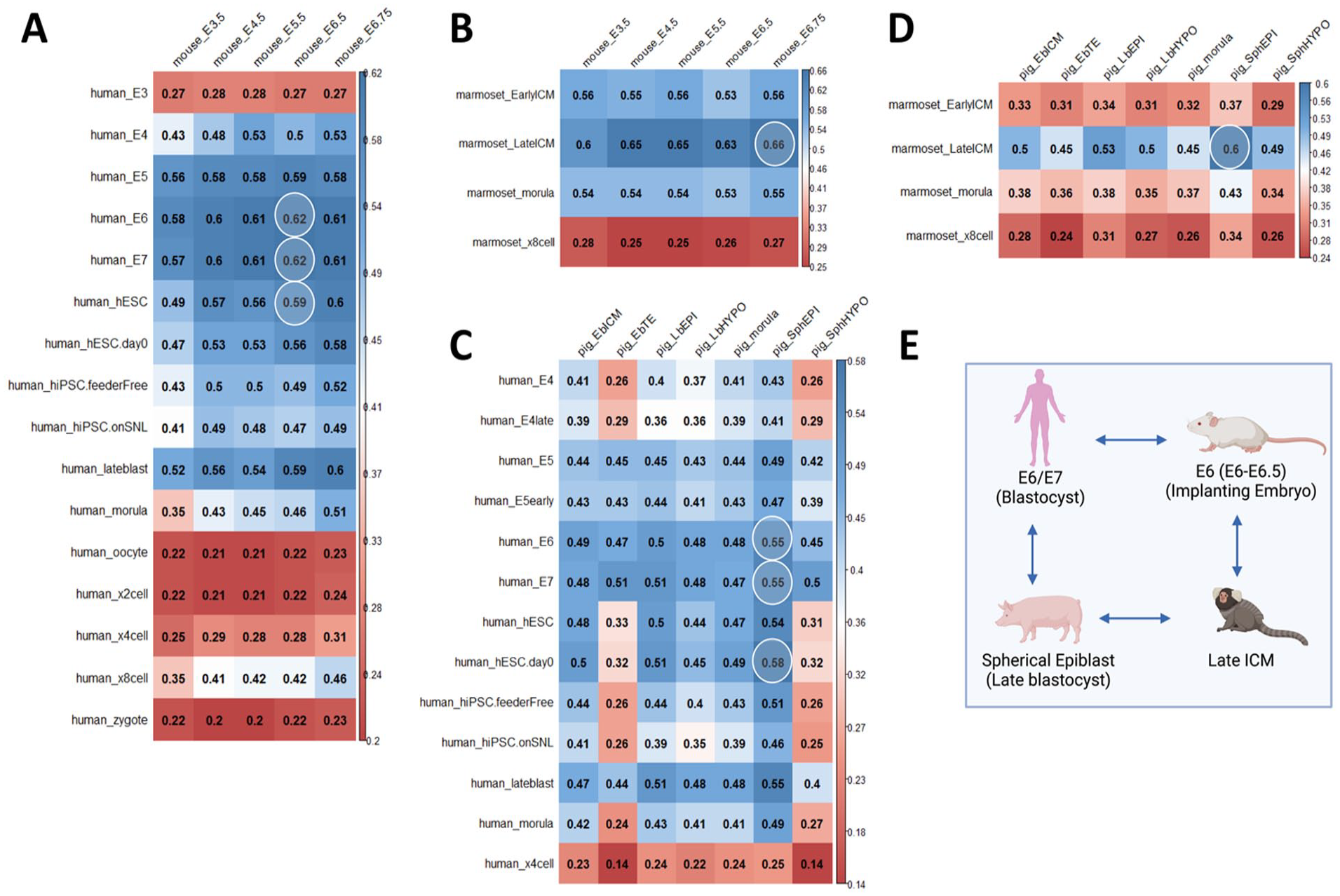
Spearman’s correlation analyses for pairwise species comparison. Correlation coefficients for comparisons between (A) human and mouse embryonic stages, (B) marmoset and mouse embryonic stages, (C) human and pig embryonic stages, and (D) marmoset and pig embryonic stages. The highest correlation coefficients for each pair are highlighted in white circles and depicts the developmental stages that best match with each other. (E) Summary of the stage-matching results from Spearman’s correlation analysis. Created with Biorender.com.
KEGG Analysis Between Best Matched Species Pairs Reveals Commonly Expressed Signaling and Metabolic Pathways
After stage-matching species based on their similarities in their transcriptome, we performed KEGG analysis between donor–host species pairs. We selected the commonly expressed genes from the topmost 3,000 upregulated genes per species as described in the “Methods” section. For each of the species pairs, KEGG analysis provided insight into the commonly expressed and upregulated pathways between the species at their best matched stages and the pathways that might contribute to successful interspecies chimerism at those stages. The KEGG analysis showed that the human–mouse best matched stages both expressed genes specific to Hippo signaling (eg, TEAD1, NFAT, FRMD4A) (Fig. 4A), an evolutionarily conserved signaling pathway across multiple species. This pathway is important for the first lineage specification into ICM and TE in early mammalian embryos. Hippo signaling also plays a role in organ specification and growth. Interestingly, among the common top gene functions in human–mouse and marmoset–mouse best matched stages, multiple genes specific to cell adhesion (eg, PVRL1, PRTG, CD47) and cell junctions (eg, LIMS1, SGCE, CELSR2) were also identified. One of the proposed barriers to interspecies chimerism, in addition to the differences in developmental stages, is the variances in cell–cell adhesion genes between more distant species. It is posited that the differences in the cell adhesion genes between the donor and host species do not allow the donor cells to interact with, synchronize, and develop in the chimeric embryos. Through our stage-matching technique, we showed that the best matched stages between the human–mouse and marmoset–mouse species expressed common genes involved in focal adhesion, tight junction, as well as regulation of actin cytoskeleton (eg, ACTN1, EZR, LCP1) and adherens junctions (eg, PAK2, DLG5, CTNNB1) (Fig. 4A, Supplementary Fig. 2A). Hence, stage-matching donor and host species could serve as a technique to overcome the interspecies chimeric barrier in differential expression of cell adhesion genes between donor and host species. For the human–pig and marmoset–pig best matched stages, multiple genes involved in metabolic pathways were upregulated and commonly expressed between the donor and host species pairs (Fig. 4B, Supplementary Fig. 2B). For the human–pig best matched stages, the topmost commonly expressed genes were involved in fatty acid and fatty acid ester metabolism (eg, ARG2, FADS1, FADS2), carbon metabolism, and sugar metabolism (eg, ENO1, PFKP, PGK1) (Fig. 4B). For the marmoset–pig best matched stages, the top commonly expressed genes were involved in fatty acid and fatty acid ester metabolism (eg, GPX4, SMPD2, HSD17B12) (Supplementary Fig. 2B). These results are interesting because the pig early embryos have a uniquely higher lipid content compared with the embryos of other species and hence upregulate lipid metabolism-specific genes21,31.

KEGG analysis (A and B) and GO analysis (C and D) of the most commonly expressed genes between best matched human–mouse and human–pig embryonic stages. The top 3,000 upregulated genes per species were used for the pairwise analysis. Based on KEGG analysis, the pathways most commonly expressed between (A) human E7 and mouse E6.5 and (B) human E7 and pig late (spherical) epiblast were listed. GO analysis of the top shared molecular functions between (C) human E7 and mouse E6.5 and (D) human E7 and pig late (spherical) epiblast were also obtained. KEGG: Kyoto Encyclopedia of Genes and Genomes. GO: Gene Ontology.
Previous research has also indicated an increase in fatty acids in human and mouse PSCs on conversion from a naïve to a primed state 32 . The cells in the human E6/E7 blastocyst are in a more “primed” state, which agrees with the increased fatty acid metabolism we observed in the human E6/E7 blastocyst and its best matched pig spherical epiblast stage. In the pig spherical blastocyst, upregulation of genes specific to glycolysis was also previously observed 22 . A similar upregulation of glycolysis-specific genes was previously reported in primed human PSCs and mouse epiblasts, all the stages that best matched with each other using our clustering algorithm33,34. We observed this upregulation in sugar metabolism in the best matched human–pig stages as well (Fig. 4B). The significance of some of the genes involved in other processes and pathways from the KEGG analysis, such as infections, neurological diseases including Parkinson and Huntington, as well as protein complexes and other metabolism pathways, requires further discussion. These pathways could provide greater insight into understanding genes involved in these processes, some of which are common between the donor and host stages (Fig. 4A–D). Further studies will establish whether they are relevant for our purposes of interspecies chimerism–specific stage-matching.
GO Analysis of Upregulated Genes in Best Matched Embryo Stages Provide Information Into Molecular Functions Shared Among Species
While KEGG analysis provides insight into different pathways shared among the species at their best matched stages, with GO analysis, we gain more insight into the molecular function of specific genes commonly upregulated by the two species. Thus, we performed GO analyses on the human–mouse, human–pig (Fig. 4C, D), marmoset–mouse, and marmoset–pig (Supplementary Fig. 2C, D) best matched stages. Consistent with our pathway analysis, the GO analysis for the human–mouse best matched stages resulted in multiple terms related to cell adhesion (Fig. 4C). From the human–pig matched stages, multiple terms related to establishment of cell and tissue polarity (eg, calcium binding genes ANXA1, ANXA2, NRP1) were observed (Fig. 4D). These results suggest the possibility of successful cell–cell adhesion and establishment of polarity during development between stage-matched donor and host cells in a chimeric embryo. Terms related to regulation of cell shape and cell morphogenesis (eg, genes DYNLL1, APOA1, FMNL2) were also shared between the human–mouse species (Fig. 4C). This suggested similarities in the regulation of shape and structure of the human and mouse cells at the best matched stages which is favorable to chimerism. From the human–pig GO analysis, we identified multiple terms involved in regulation of cell cycle G2/M phase transition (Fig. 4D). Hence, multiple genes involved in this G2/M transition were commonly upregulated between the human and pig stages (eg, TUBA1B, PRCC, RALA) and may lead to synchronization in their cell cycles at their best matched stages. In addition to this, terms related to canonical and non-canonical Wnt signaling pathways were observed in the human–pig (eg, genes TCF7L2, LGR4) and marmoset–pig (eg, genes G3BP1, SFRP2, TSKU) stage-matched GO analysis (Fig. 4D, Supplementary Fig. 2D). Wnt signaling plays an integral role in multiple functions such as cell proliferation, migration, and stem cell maintenance during early embryonic development and is important for preimplantation development of the embryo35,36. As the stages that best matched with each other for the human–pig and marmoset–pig pairs are all preimplantation stages—human blastocyst, marmoset late ICM, and pig late blastocyst—the upregulation of Wnt signaling pathways was expected. Similar to results from the KEGG analysis, the majority of terms in the marmoset–pig GO analysis were related to fatty acid (cholesterol, sterol) biosynthesis and metabolism (Supplementary Fig. 2D). The marmoset late ICM which consists of PSCs in the “primed state” have higher fatty acid content compared with “naïve” cells 32 , and pig embryos have uniquely high levels of fatty acid21,31. Thus, these embryonic stages best match with each other and demonstrated similar properties to each other which can enhance chimerism through stage-matching.
Discussion
One of the proposed barriers to successful chimerism is the difference in developmental stages between the donor and host cells at the time of introducing the donor cells into the host embryos 16 . Studies have attempted to stage-match donor cells and host embryos using both in vivo and ex vivo strategies16,28,37. However, a more efficient and cost-effective approach is in silico stage-matching cells and embryos across different and distinct species 17 . In this study, we analyzed the scRNA-seq datasets of early embryos across four species, including human, marmoset, mouse, and pig. We used a clustering algorithm that simultaneously stage-matched all four species with each other based on similarities in the transcriptomes across different developmental stages within each species. We also performed a pairwise Spearman’s correlation analysis between different donor–host species to verify the results of our clustering algorithm. Based on our in silico approaches, we observed that the stages which best matched with each other are the human blastocyst (E6/E7), the gastrulating mouse embryo (E6–E6.75), the marmoset late ICM, and the pig late blastocyst. Through our clustering algorithm, we also observed that hESCs and hiPSCs best matched with the mouse gastrulating embryo at E6–E6.75. This in silico approach of stage-matching multiple species based on their transcriptome is comparatively inexpensive and more efficient than in vitro and in vivo stage-matching. Furthermore, the predictions made can be used to optimize and enhance interspecies chimerism between distant species when performing both in vitro and in vivo chimerism studies. From our study predictions, we hypothesize that hESCs injected into the (1) gastrulating mouse embryo and (2) pig late blastocyst will lead to enhanced interspecies chimerism. Similarly, we hypothesize that marmoset ESCs extracted from the late ICM will successfully contribute to chimerism when injected into the gastrulating mouse embryo and pig late blastocyst. The predictions of stage-matching made in this study are yet to be tested in vivo but some of the reported in vitro and in vivo chimerism studies to date are in agreement with our predictions.
A previous human–mouse chimerism study demonstrated that hESCs injected into early and late gastrulating mouse embryos gave rise to successful human–mouse chimeras with an efficiency of 70% when cultured in vitro 28 . An in vivo stage-matching study was done in which hESCs were injected into the late-stage E8.5 gastrulating mouse embryo and the early stage E3.5 mouse blastocyst. The results showed that hESCs contributed to chimerism when injected into E8.5 embryo but did not show chimerism when injected into the E3.5 blastocyst 16,20. In agreement with these in vitro and in vivo studies, our in silico results successfully predicted the best matched human–mouse developmental stages using their respective transcriptomes. In another in silico study comparing transcripts of human, marmoset, and mouse early embryos, the authors generated developmental stage–specific gene expression modules across the three species 17 . The transcriptome analyses suggested that human ICM was most similar to marmoset ICM, which was also in agreement with our results. In a previous study analyzing the single-cell-level transcriptome of early and late pig embryos, the authors observed alignment of the hESCs with the pig epiblast along PC2 and PC3 in a pairwise PCA analysis 22 . Hence, they proposed that hESCs introduced into pig late blastocysts may give rise to successful chimeras. These are also the predictions from our current in silico approaches, which further supports the feasibility of our approaches.
For each of our best matched donor–host species pairs, we performed a KEGG analysis and GO analysis on the most significantly common upregulated genes. The KEGG analysis allowed us to observe the most conserved pathways among the best matched stages. From the GO analysis, we were able to observe the molecular functions of the genes that are shared between the donor and host species. Together, the KEGG and GO analyses provided us an integrated insight into the gene functions and pathways and how they might play a role in enhancing chimerism in vivo. In the future, more detailed studies identifying specific genes that are commonly expressed and significantly different between the best matched donor–host stages will be required.
The host embryo developmental stages for the mouse (E0.5–E6.5) and pig embryos (E4–E11) analyzed in this interspecies stage-matching study were selected to overcome any concerns associated with immune rejection of the donor cells. As these preimplantation host embryo stages occur well before thymus development, the donor cells introduced into the host embryo are able to contribute to thymus development of the chimeric embryo38,39. The developing chimeric embryo’s immune system is trained to recognize donor cells as “self” and hence eliminates concerns with regard to immune rejection or inflammation due to donor cells 40 . Another step at which immune rejection concerns occur is during the transplantation of the organ of interest from the chimeric animal into the organ recipient. The organ that develops in the chimeric embryo is required to contain cells and express antigens exclusive to the donor species. This applies not only to the cells forming the organ but also the vasculature of the organ of interest. Blastocyst complementation studies have been working on developing chimeric animals with donor-derived vasculature 41 . Furthermore, immune rejection can also occur due to host animal antigens being expressed in the donor cell–derived organ used for transplantation. The field of xenotransplantation has been making significant progress in overcoming these hurdles 42 . Genetically modified (GM) pigs have been developed where xenoantigens are knocked out and human antigens that are conducive for integration of the transplanted organ in the recipient are expressed in the GM pigs43–46. Once the inherent hurdles of the interspecies chimerism and blastocyst complementation field have been tackled, the GM pigs would ideally be used as host organisms for development of our organs of interest and subsequent transplantation of those organs in the clinic.
The in silico approaches used in this study helped us to simultaneously stage-match four species relevant to interspecies chimerism studies. In this study, we decided to only focus on the clusters containing the mouse gastrulating embryo and stage-match species within those clusters. However, from the clustering information that we generated, we can analyze the remaining clusters and stage-match different human, marmoset, mouse, and pig stages based on those of interest (Supplementary Fig. 3). Furthermore, the techniques used for analysis in this study can be applied to other species’ combinations to understand developmental similarities across species and cell types in the context of chimerism as well as early development.
Supplemental Material
sj-docx-1-cll-10.1177_09636897231158728 – Supplemental material for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism
Supplemental material, sj-docx-1-cll-10.1177_09636897231158728 for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism by Anala Shetty, Seunghyun Lim, Phoebe Strell, Clifford J. Steer, Juan Carlos Rivera-Mulia and Walter C. Low in Cell Transplantation
Supplemental Material
sj-tiff-2-cll-10.1177_09636897231158728 – Supplemental material for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism
Supplemental material, sj-tiff-2-cll-10.1177_09636897231158728 for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism by Anala Shetty, Seunghyun Lim, Phoebe Strell, Clifford J. Steer, Juan Carlos Rivera-Mulia and Walter C. Low in Cell Transplantation
Supplemental Material
sj-tiff-3-cll-10.1177_09636897231158728 – Supplemental material for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism
Supplemental material, sj-tiff-3-cll-10.1177_09636897231158728 for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism by Anala Shetty, Seunghyun Lim, Phoebe Strell, Clifford J. Steer, Juan Carlos Rivera-Mulia and Walter C. Low in Cell Transplantation
Supplemental Material
sj-tiff-4-cll-10.1177_09636897231158728 – Supplemental material for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism
Supplemental material, sj-tiff-4-cll-10.1177_09636897231158728 for In Silico Stage-Matching of Human, Marmoset, Mouse, and Pig Embryos to Enhance Organ Development Through Interspecies Chimerism by Anala Shetty, Seunghyun Lim, Phoebe Strell, Clifford J. Steer, Juan Carlos Rivera-Mulia and Walter C. Low in Cell Transplantation
Footnotes
Ethical Approval
Ethical approval is not applicable for this article.
Statement of Human and Animal Rights
This article does not contain any studies with human or animal subjects.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is supported in part by NIH grants R01 DK117286 (CJS), R01 DK117286-03S1 (CJS and WCL), and Regenerative Medicine Minnesota Grant 3016-11702-00100620 (JCR-M).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
