Abstract
This study aimed to explore the function of CLPTM1L in oral squamous cell carcinoma and mechanism of tumorigenesis. The expression of CLPTM1L was detected by immunohistochemistry. The localization in cells was detected by immunofluorescence. Cell invasion, proliferation, and migration were detected by transwell, CCK-8 and scratch-wound test. The possible characteristics of CLPTM1L were analysed in TCGA, GO, KEGG and String databases. IHC revealed that the expression of CLPTM1L in 92 cases of OSCC tissues was significantly higher (P < 0.01) than 29 cases of normal oral epithelium tissues. The expression of CLPTM1L was significantly higher in oral squamous cell carcinoma in TCGA database. CLPTM1L expression was not significantly correlated with the patients’ clinical parameters. High expression of CLPTM1L was associated with worse prognosis. Cox regression analysis demonstrated that the CLPTM1L expression was the significant risk factor. CLPTM1L was mainly localized in the perinuclear cytoplasm. The vitro studies revealed that the knockdown of CLPTM1L suppressed invasion, proliferation and migration. CLPTM1L related genes were enriched in protein processing in the endoplasmic reticulum, protein folding, endoplasmic reticulum formation, N-glycan biosynthesis, and protein hydroxylation. Highly expressed CLPTM1L may contribute to a poor prognosis and increase invasion, proliferation and migration of oral squamous cell carcinoma. CLPTM1L may play an important role in tumorgenesis and would be a valuable target gene for the treatment of oral squamous cell carcinoma.
Introduction
Oral cancer refers to the malignant tumor occurring in the oral cavity and its adjacent anatomical structure, including lips, tongue, cheeks, floor of the mouth, hard and soft palate, sinuses, and pharynx. It is a relatively common malignant tumor of the head and neck, accounting for approximately 3% of all malignant tumors in the whole body. The most common type of oral cancers are squamous cell carcinomas 1 . Although the incidence of oral cancer is slowly decreasing in some developed countries, the incidence of oral cancer is increasing at an earlier age globally. According to statistics from American cancer society, there were ∼350,000 new cases of oral cancer and ∼170,000 deaths worldwide in 2018 2 . Early detection and treatment can significantly improve the prognosis of patients. However, a large number of patients are already diagnosed at an advanced stage. Oral cancer is a complex disease caused by multiple factors, including (1) familial and genetic susceptibility; (2) solar radiation, environmental pollution, and so on; (3) immunosuppression such as HIV infection; (4) bad habits, including smoking, chewing betel nut, smoking marijuana and drinking alcohol; (5) virus infection, containing EPstein-Barr virus (EBV), human papillomavirus (HPV16, 18) and human herpes simplex virus (HSV6, 8); (6) candida albicans infection; (7) poor oral hygiene, lack of teeth, poor repair and sharp tooth tip; (8) iron deficiency; (9) potential malignant lesions and states in the oral cavity, including leukoplakia, lichen planus, erythema, and oral submucosal fibrosis. Faced with the same external risk factors, only a portion of the population will eventually develop to oral cancer, suggesting that there exist differences between individuals that result from their genetic predisposition.
Multiple genome-wide association studies (GWAS) have demonstrated that the TERT-CLPTM1L genetic variation at 5p15.33 is correlated with the risk of various malignant tumors, such as lung cancer, cervical cancer, pancreatic cancer, bladder cancer, glioma, prostate cancer, basal cell carcinoma and melanoma, and oral cancer 3 –5 . Currently, there is a lot of human gene researches about TERT (telomerase reverse transcriptase). Its encoding telomerase may participate in the development of tumor through various channels. One of the important approaches is normal somatic TERT gene mutation as well as the activation of telomere and telomerase, which results in cell aging problems. Eventually, malignant proliferation will appear 6 . Some studies have also shown that the genetic variation of CLPTM1L is associated with the incidence of some malignant tumors 7 –17 . In the previous studies of oral squamous cell carcinoma, we also found multiple genetic variations located in CLPTM1L. Nevertheless, there are few reports of what role CLPTM1L plays in the generation and progression of oral squamous cell carcinoma.
CLPTM1L is the homologous protein gene of CLPTM1. CLPTM1 was reported for the first time in a cleft lip and palate family with a translocation breakpoint that cosegregates in two of three generations. CLPTM1L was extensively expressed in both adult and embryonic tissues, and identified as a transmememal rupture protein 18 . CLPTM1L, also known as CRR9, was found in screening cisplatin-resistant genes, which were up-regulated in cisplatin-resistant ovarian cancer cells and could induce apoptosis of cisplatin-sensitive cells 19 . The present study aimed to explore the significance of CLPTM1L in oral squamous cell carcinoma and the possible mechanism in the development of tumor.
Materials and Methods
Cell Culture and Reagents
OSCC cell lines (Cal 27 and Hsc-3) were obtained from ATCC (American type culture collection, Manassas, VA, USA). Both cells were grown in DMEM/F12 media with 10% FBS.
Cell Transfection
siCLPTM1L and siNC transfection were prepared according to Lipofectamine®2000 instructions and added to each well. After 24 hours, the cell status was observed and the medium was changed. After 48 to 72 hours of transfection, cell extracts were analyzed for other experiments.
Patients and Tissue Specimens
A retrospective cohort of patients with primary OSCC treated at our institution between Jan. 2013 and Jun. 2016 was enrolled. Written informed consent was obtained from these patients. The patients were required to have not received preoperative radiotherapy, chemotherapy, or any other medical intervention, and all clinical data were complete, including the follow-up prognostic information records. The tumor stage is based on the American Joint Committee on Cancer’s seventh Edition of the Cancer Manual. Finally, 92 qualified primary oral squamous cell carcinoma tissue specimens were selected, and 29 normal oral mucosal tissues were obtained from the margins of surgical resection in some cases. All patients were M0 at the start of the study. The study was approved by the Ethics Committee of The Stomatological Hospital of Jiangsu Province.
Immunohistochemistry (IHC)
IHC staining was performed in routine procedure on 4-µm formalin-fixed, paraffinembedded specimens. The antigen retrieval was done in boiling pressure cooker for 2 minutes. The slides were cooled until they reached room temperature. Slides were rinsed twice with wash buffer PBS. Slides were incubated with primary antibody CLPTM1L (rabbit polyclonal, HPA014791, Sigma-Aldrich, Belgium) diluted to 1:200 overnight at 4°C. Slides were rinsed with wash buffer. Slides were incubated with secondary antibody (ProteinTech Group, Chicago, IL, USA) for 15 minutes in room temperature and rinsed twice with wash buffer. Slides were incubated with DAB substrate and stained with hematoxylin. Slides were dehydrated and coverslipped for viewing. 92 oral squamous cell carcinoma samples and 29 normal oral mucosal tissues were selected for IHC. Double-blind evaluation was conducted by two senior oral pathologists. Tumors were scored as negative – 0, weak – 1, intermediate – 2, or strong – 3. Independent scores were averaged. The score of immunostaining was calculated according to “immunohistochemistry score = staining intensity score × positive proportion score.” Samples were splitted in low (IHC score<4) and high (IHC score >=4) expression after quantification of CLPTM1L immunostaining.
qRT-PCR Assay
Total RNA in tissues and cells were extracted by TRIzol reagent (Invitrogen, USA) and reagent Kit (TAKARA, Japan) according to the manufacturer’s protocol. cDNA was reversed by PrimeScript TM RT reagent Kit (TAKARA, Japan). Quantitative real-time PCR (qRT-PCR) was performed on a 7300HT system 5 using SYBR Premix Ex Taq II kit (TAKARA, Japan). The internal control was set as GAPDH. The relative expression level was calculated by 2−ΔΔCt method. Forward primer of CLPTM1Lwas 5′-TAA GAG CTG GTA CTC CTG GT-3′ and reverse primer was 5′-GTC ATC AAT GAA GGT GTT GA-3′.
Western Blot
Cells were harvested on ice in RIPA buffer (50 mmol/L Tris, pH 7.4, 150 mmol/L NaCl, 0.1% SDS, 0.5% sodium deoxycholate, and 1.0% NP-40) with protease inhibitor cocktail (Roche, Swiss). The membrane was blocked with 5% BSA for 2 hours and performed incubation with specific primary antibodies overnight at 4°C, including CLPTM1L (1:1500, ab155119, abcam, USA); GAPDH (1:1000, ab181602, abcam, USA). The membrane was then incubated with HRP Goat-anti-Rabbit (1:2000) or Goat-anti-Mouse (1:2000) for 2 hours at room temperature.
Invasion Assays
Transwell filter (8 mm aperture; Millipore, Bedford, USA) was used to analyze the invasive ability of cells. The matrix and DMEM were prepared in the ratio of 1:6. The 8 µm MilporeTranswell chamber was placed on a 24-well plate and then we added 50 µL solution into each chamber. Matrigel was used in the invasion assay to cover the insert membrane. Cells in a density of 1 × 105 were inoculated in the serum-free medium of 200 µL in the upper chamber, and the medium containing 500 µL 10% FBS were placed in the lower chamber. After being incubated in the incubator at 37°C for 24 hours, the cells remaining in the upper room were gently removed with cotton swabs. Cells in the lower chamber were immobilized with methanol for 30 minutes, and then we stained them with crystal violet for 20 minutes.
Wound-Healing Assay
The cells were cultured in a 6-well plate in the density of 90%. Gently and slowly scratch the monolayer with a new 1 mL pipette tip across the center of the well. While scratching across the surface of the well, we always keep the long-axial of the tip perpendicular to the bottom of the well. Photographs of the same region were taken with the same microscope, and scratch changes at different time points were recorded.
Cell Proliferation
Cell Counting Kit 8 (CCK8) (Beyotime, Jiangsu, China) was used to detect the cell proliferation capacity. Cells were inoculated into 96-well plates till a density of 1 × 103 cells per well. 10 µL of Cell Counting Kit 8 reagent was added into each well at 0 h, 24 h, 48 h, and 72 h. The plates were placed for a subsequent 2 hour incubation at 37°C. The absorbance was measured at the wavelength of 450 nm according to the manufacturer’s instructions.
OSCC Xenograft Model
Six-week-old female nu/nu mice were obtained and maintained in a specific pathologic-free environment. Cancer cells suspended in total 100 µL PBS were inoculated subcutaneously on the single or both flanks (4 animals per experimental group). The tumor volumes were calculated as (length × width × height)/2. Finally, mice were sacrificed and final tumor volumes were measured upon tumor samples were harvested.
Statistical Analyses
The associations between CLPTM1L expression and various clinicopathological parameters of patients were evaluated using Chi square test. The survival rates of patients were estimated using Kaplan–Meier curve and compared with Log-rank test. The prognostic analyses were performed by univariate and multivariate Cox regression models to determine the individual clinicopathological variables with patients’ overall or disease-free survival. T test was used to assess vitro findings. P-values <0.05 (two-sided) were considered as statistically significant. All statistical analyses were performed using Graphpad Prism 5 (La Jolla, CA, USA) or SPSS 18.0 (Armonk, NY, USA).
Results
The expression of CLPTM1L was evaluated by immunohistochemical staining score, and the expression of CLPTM1L was significantly different in 92 oral squamous cell carcinoma samples and 29 normal oral mucosal tissues. Representative immunohistochemical staining results were shown in Fig. 1. The statistical results were shown in Table 1. With low expression in 38 cases and high expression in 49 cases, CLPTM1L was negatively expressed in 5 cases of oral squamous cell carcinoma. However, in normal tissues, there were 10 cases of negative expression, 18 cases of low expression and 1 case of high expression. IHC results showed that CLPTM1L was significantly highly expressed in oral squamous cell carcinoma (P<0.01) than normal samples.
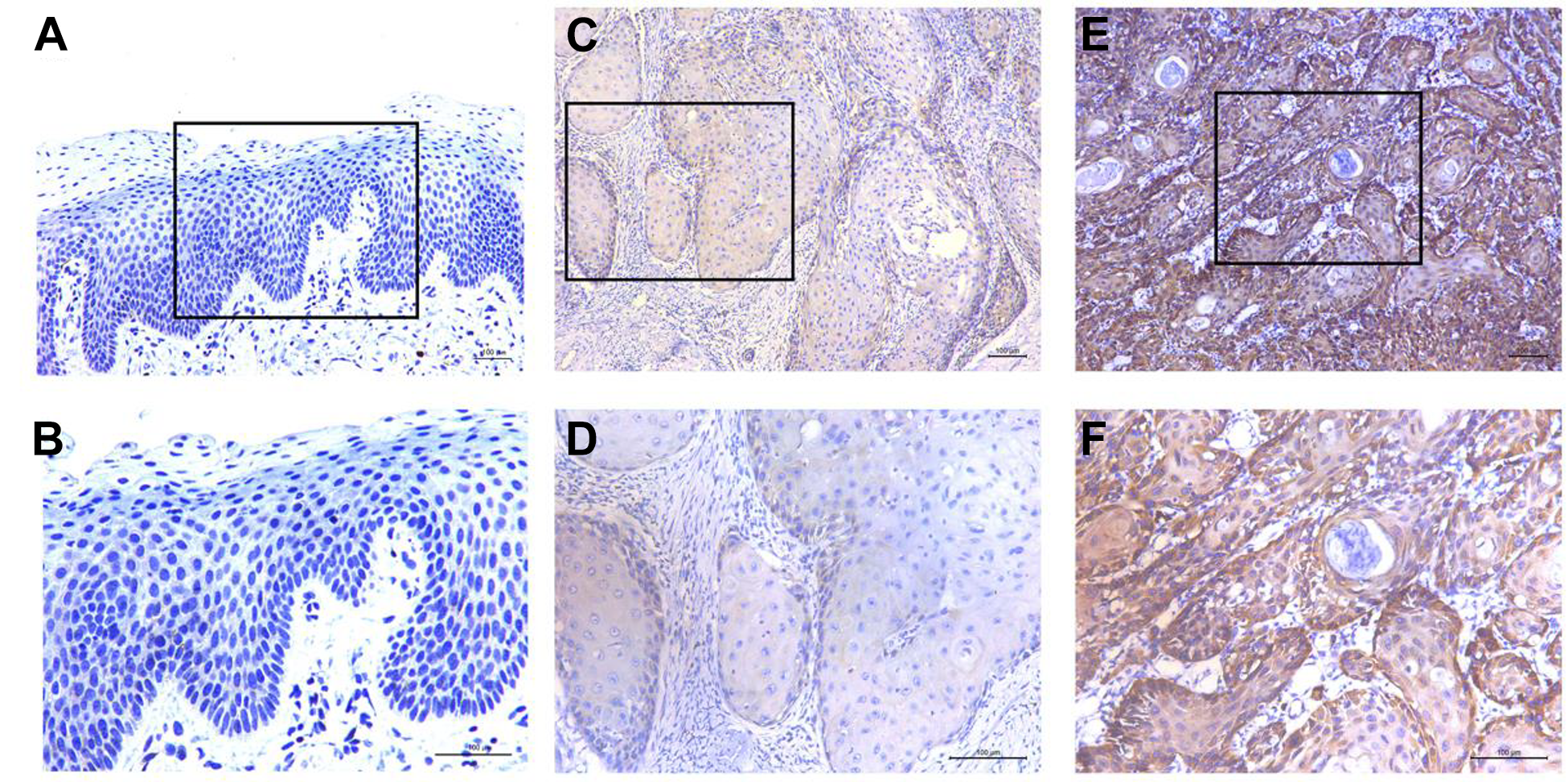
Immunohistochemistry was used to detect the typical expression of CLPTM1L. (A, B) Normal oral epithelium tissue showed negative expression of CLPTM1L. (C, D) primary oral squamous cell carcinoma showed low and high expression of CLPTM1L. (E, F) primary oral squamous cell carcinoma showed low and high expression of CLPTM1L. (Scale: 100 μm).
CLPTM1L Expression Patterns in OSCC and Normal Oral Epithelium.

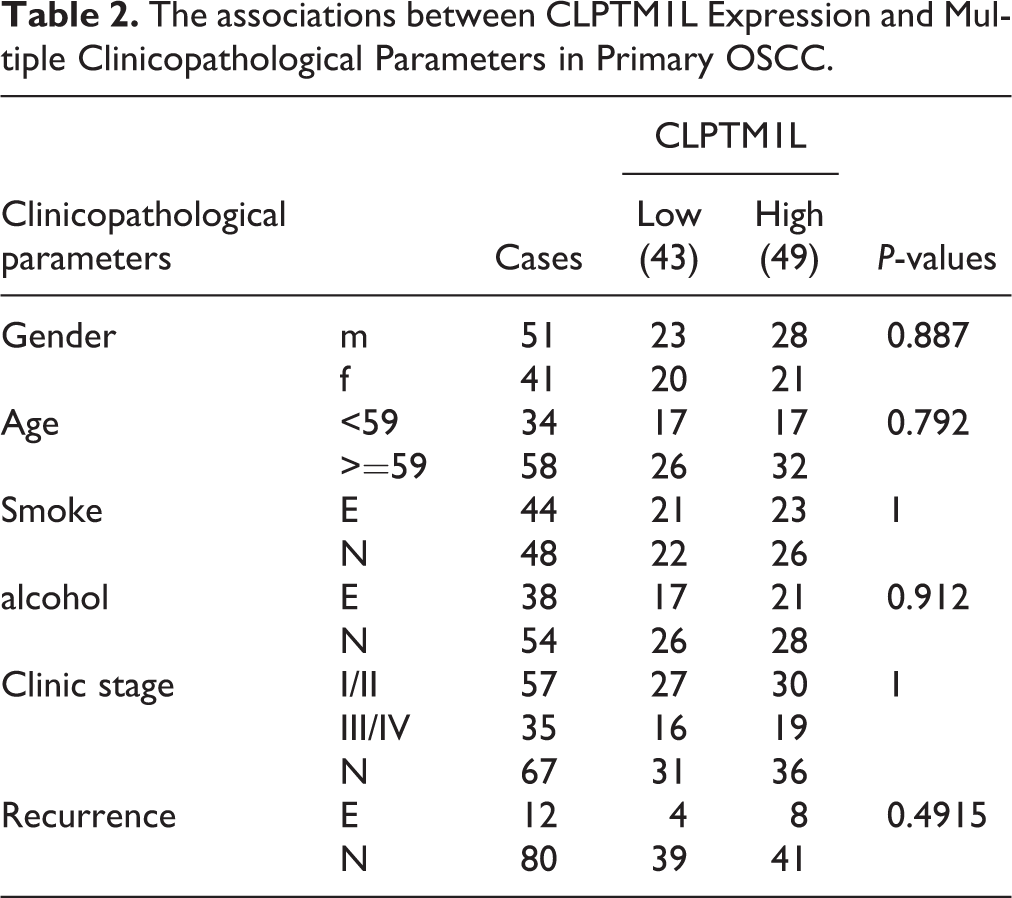
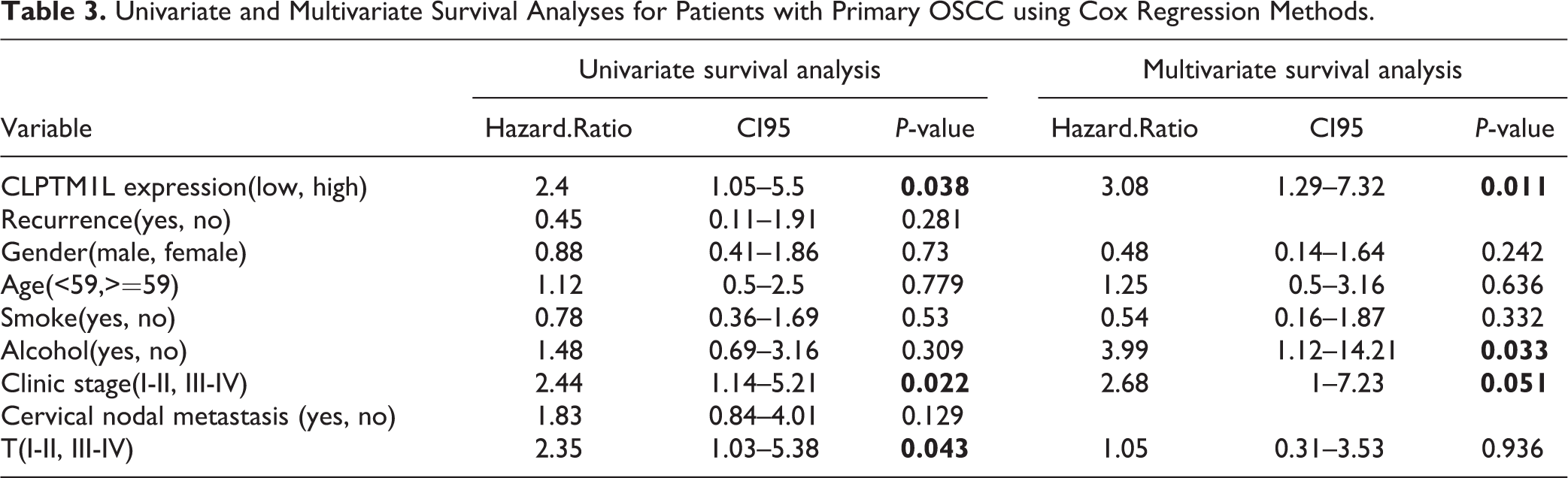
In order to explore whether CLPTM1L expression is correlated with various conventional clinical indicators in patients of oral squamous cell carcinoma, chi-square test was used for statistical calculation between the expression of CLPTM1L and the gender, age, smoking, alcohol consumption, clinical stage, tumor size, metastasis or recurrence of the patients. However, the difference was not statistically significant (Table 2 and supplemental Table S1). In order to explore whether the CLPTM1L is high or low in the oral squamous cell carcinomas as well as its relationship between the prognosis survival rate of patients, this research combined the follow-up data, makes statistics analysis of the patient’s overall survival and disease-free survival time, and conducts Kaplan Meier survival curve analysis (Fig. 2). CLPTM1L high expression is associated with a significantly low survival rate and poor prognosis. To understand the relationship between the prognosis of oral primary squamous cell carcinoma and patients’ gender, age, smoking, alcohol consumption, clinical stage, tumor size, presence or absence of metastasis, and recurrence, univariate and multivariate Cox regression analyses were performed (Table 3). Cox regression analysis of single factor, the observed indicators, found that patients with gender, age, smoking, drinking, lymph node metastasis, recurrence, and postoperative survival rate has no obvious correlation. The tumor size, clinical stage, CLPTM1L expression was significantly related with postoperative survival rate (P < 0.05). Four commonly used confounding factors including gender, age, smoking and alcohol consumption were added in combination with relevant factors. Then, the multivariate regression analysis was carried out to determine the independent prognostic factors influencing the overall survival of the patients. Analysis results demonstrated that the expression level of CLPTM1L (P = 0.011) was significantly correlated with the overall survival rate of patients, and high CLPTM1L expression would lead to poor prognosis.
The associations between CLPTM1L Expression and Multiple Clinicopathological Parameters in Primary OSCC.
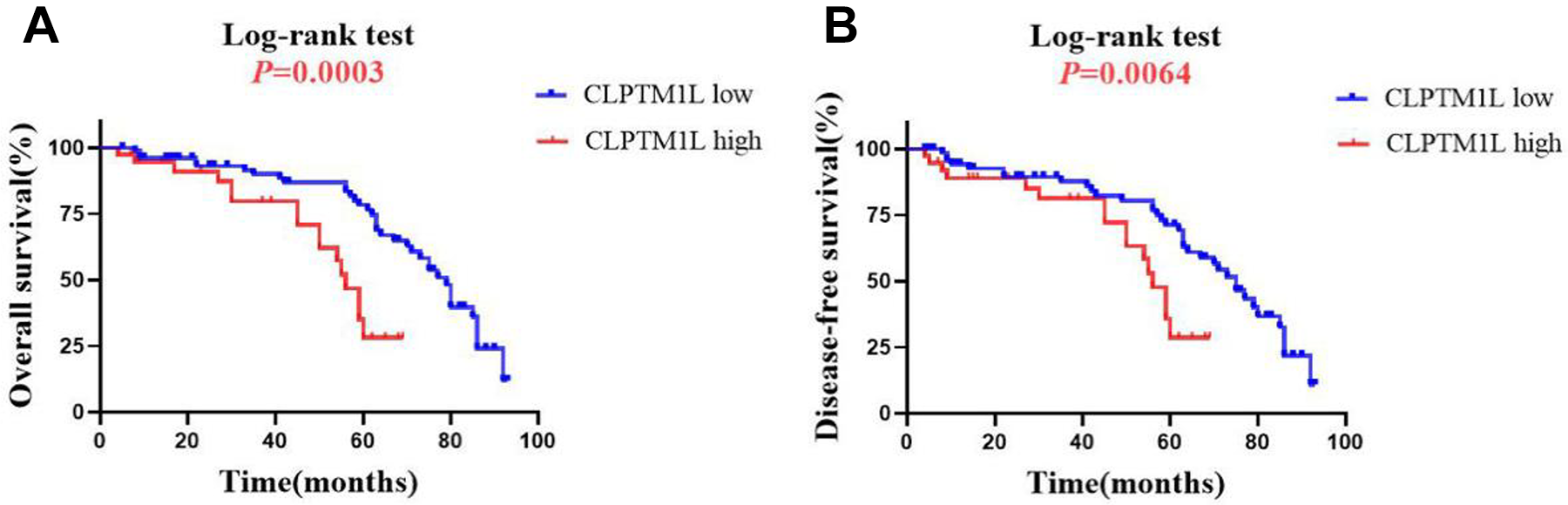
Kaplan–Meier graphs representing the probability of OSCC patients’ survival based on CLPTM1L expression status. (A) High CLPTM1L expression is significantly associated with reduced overall survival in OSCC patients. (B) High CLPTM1L expression is significantly associated with reduced disease-free survival in OSCC patients. P-value from Log-rank test is shown.
Univariate and Multivariate Survival Analyses for Patients with Primary OSCC using Cox Regression Methods.
Transfection of si-CLPTM1L (siRNA-CLPTM1L) could effectively inhibit the expression of CLPTM1L in Cal27 and HSC-3 cells. The result of qRT-PCR showed that compared with the negative control group (siNC), the expression of CLPTM1L gene was significantly decreased in the si-CLPTM1L group (Fig 3A, B). The result of western blot showed that compared with the negative control group (siNC), the expression of CLPTM1L protein was significantly decreased in the si-CLPTM1L group (Fig 3C). The localization and distribution of CLPTM1L in oral squamous cell carcinoma cells (Cal27 and HSC-3) were detected by immunofluorescence (Fig 3D). The internal reference is the cytoskeleton green fluorescent probe Actin, a network system that runs through the nucleus and cytoplasm, showing cell morphology. Endogenous CLPTM1L is a red fluorescent probe, and fusion localization shows that CLPTM1L is mainly distributed in the cytoplasm, especially dense in perinuclear cytoplasm. The results of CCK-8 experiment (Fig 3E), Transwell (Fig 3F) and scratch experiment (Fig. 3G) showed that, compared with the NC group, the invasion, proliferation, and the migration ability in si-CLPTM1L group was reduced. It can be seen that the expression of CLPTM1L is directly proportional to the malignant degree of oral squamous cell carcinoma cells.
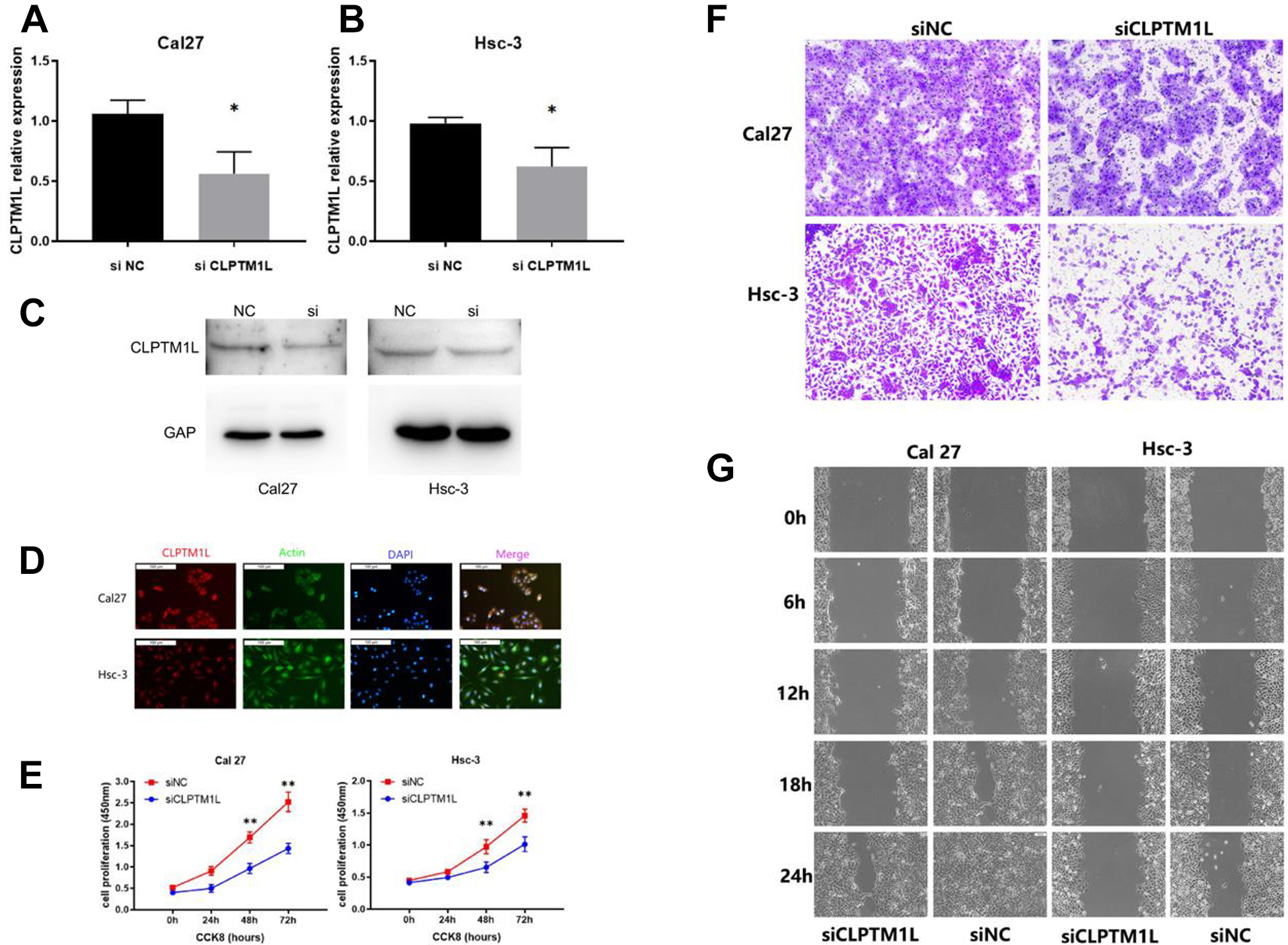
(A, B) siRNA-mediated knockdown of CLPTM1L in OSCC cells(Cal27 and Hsc-3). The CLPTM1L mRNA level after siRNA treatment was measured using quantitative real-time PCR in different groups. *: P < 0.01 vs NC control group. (C) Result of western blot. Expression of CLPTM1L was lower in si group. (D) Cellular localization of CLPTM1L in OSCC cells.Cal 27 and Hsc-3 OSCC cells were stained with Actin (green), DAPI(blue) or anti-CLPTM1L antibody (red). We examined CLPTM1L immunoreactivity using methanol-fixed cells. E: CCK-8: After CLPTM1L was silenced, the proliferation ability of OSCC cells was reduced. (** represents P < 0.01). (F) Transwell: After CLPTM1L was silenced, the invasion ability of OSCC cells was reduced. (G) Scratch wound assay: After CLPTM1L was silenced, the migration ability of OSCC cells was reduced.
In the TCGA database, the expression of CLPTM1L is positively correlated with oral squamous cell carcinoma, and the expression of CLPTM1L in 331 oral squamous cell carcinoma tissues is significantly higher than that in 32 normal oral mucosal tissues. Meanwhile, there is a statistical significance (Fig. 4A). The expression difference between 32 paired oral squamous cell carcinoma and normal mucosal tissues is also statistically significant (Fig 4B). We collected CLPTM1L related genes in the TCGA database and conducted pathway enrichment analysis by R language in the KEGG and GO (Fig. 4C, D). CLPTM1L related genes were enriched in protein processing in the endoplasmic reticulum, protein folding, endoplasmic reticulum formation, N-glycan biosynthesis, and protein hydroxylation. Besides, different numbers of genes in positively related pathways are discussed. Proteins and their functional interactions form the backbone of cellular mechanisms. The goal of the String database is to collect, evaluate, and integrate all the publicly available sources of protein-protein interactions and supplement the information with computational predictions 20 . The protein network that may interact with CLPTM1L protein is analyzed in the String database. The circle in the figure is the node and each node represents a protein. The helix in the protein represents the known structure of the protein. The line used is called ‘confidence’. The thicker and darker the lines between the circles, the stronger the interaction between the two proteins. As shown by the results, CLPTM1L may interact with TERT, SLC6A18, LPCAT1, and ATF7IP (Fig. 4E).
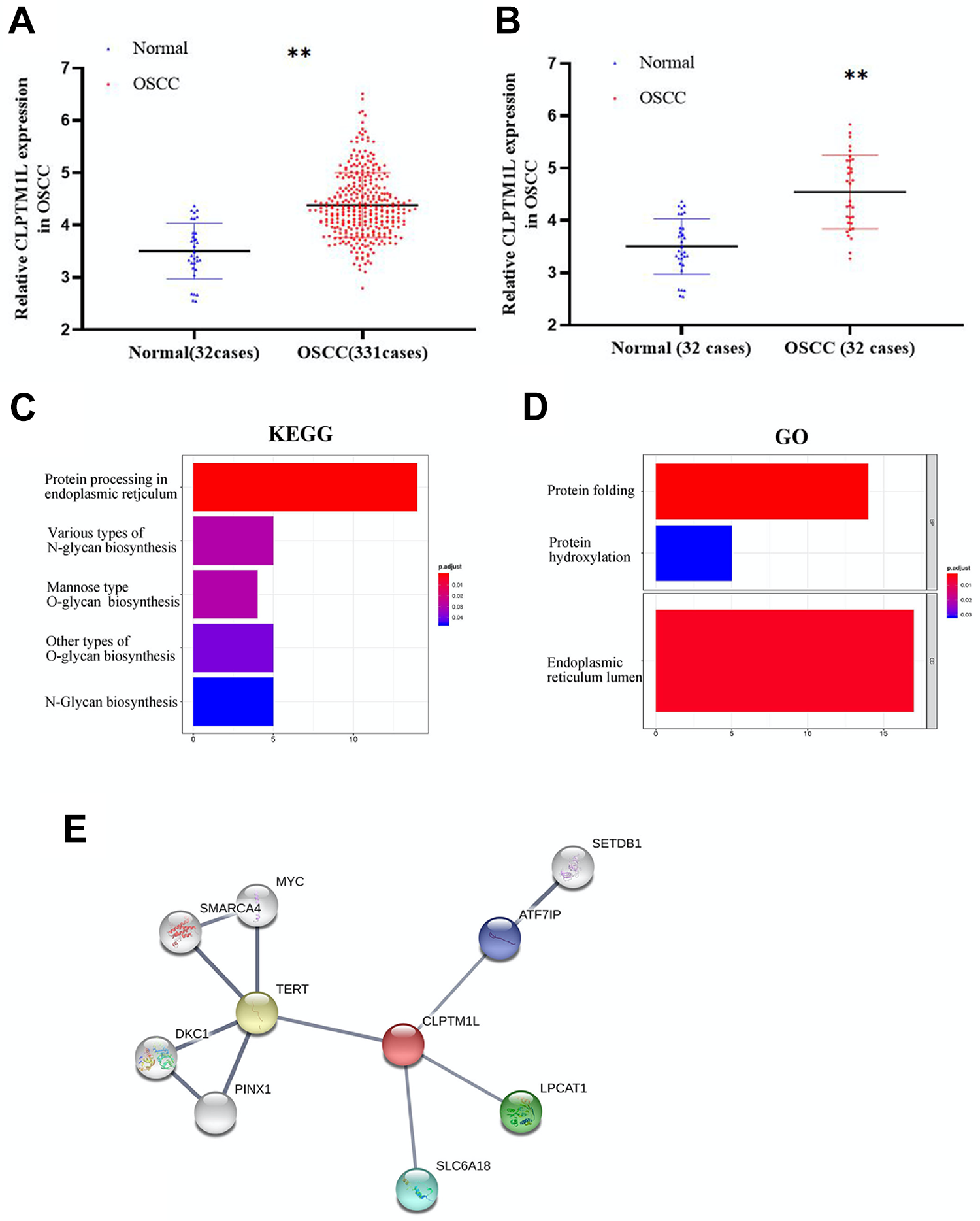
Data analysis of CLPTM1L in OSCC. (A, B) In the TCGA database, the expression of CLPTM1L is positively correlated with oral squamous cell carcinoma. (C, D) CLPTM1L possible relative genes in the KEGG and GO platform analysis. Red is positively related to the corresponding channel, blue is negatively related to the corresponding channel, and the horizontal axis represents the path difference gene number. Furthermore, Y-axis is enriched to describe information access. (E) CLPTM1L possible relative protein in the String database analysis.
We next explored xenograft mouse models to validate CLPTM1L effects in vivo. Cal27 cells were inoculated in immunodeficient nude mice. Knockdown of CLPTM1L inhibited tumor growth (Supplemental Fig. S1).
Discussion
Malignant tumors are complex diseases caused by the combined action of multiple factors. Different individuals have different morbidity and prognosis under the same risk factors, which is due to the differences in genetic susceptibility among individuals. Chromosome 5p15.33 has been reported as a pleiotropic locus associated with a variety of malignant tumors such as lung cancer, breast cancer, glioma, ovarian cancer, and pancreatic cancer 3,9,21,22 .
CLPTM1L’s single nucleotide polymorphisms (SNPs) are in association with many human malignant tumors, especially in some highly invasive and metastatic malignant tumors 3 . In GWAS studies related to lung cancer, CLPTM1L-rs402710 and rs401681 have been reported to be associated with the risk of lung cancer 7,16,23,24 . The high expression of CLPTM1L is in consistence with the hypomethylation in the downstream region, and methylation of CLPTM1L has an active regulatory function in lung cancer 14 . In pancreatic cancer, CLPTM1L is overexpressed and significantly correlated with poor prognosis 25,26 . CLPTM1L might interact with MYH9, a protein involved in regulating cell migration and adhesion, as protein partners for CLPTM1L 27 . Studies about gene expression of cervical cancer have revealed that CLPTM1L is overexpressed in tumors 28 and its increase in copy number is related to the occurrence of tumors, which may play a role by silencing the expression of other genes 29 . In breast cancer, it has been reported that high expression of CLPTM1L is associated with poor prognosis of breast cancer and is connected to the estrogen receptor (ER) negative breast cancer subtype 30 .
In our study, 92 primary OSCC tissue samples were included, and it was found that the expression of CLPTM1L was not related to clinical-pathological factors, but the expression level of CLPTM1L was independently related to the overall survival rate of patients. The high expression of CLPTM1L would lead to poor prognosis, which is consistent with published studies 31,32 . In our further study, CLPTM1L gene silencing could inhibit invasion, proliferation and migration of OSCC cells. In another study of CLPTM1L gene in OSCC 33 , CLPTM1L expression is significantly related to the depth of invasion and poor prognosis of patients.
IF revealed that CLPTM1L was mainly localized in the cytoplasm, especially in the perinuclear cytoplasm. In a study of pancreatic cancer, CLPTM1L was found to be primarily localized in the endoplasmic reticulum, possibly with nuclear staining or nuclear membrane staining 27 . The gene pathways were analyzed by TCGA database. The endoplasmic reticulum, protein folding, and endoplasmic reticulum formation, were all corresponding to the cytoplasmic localization of CLPTM1L found in IF experiments.
In the String database analysis, proteins that may interact with CLPTM1L include TERT, SLC6A18, LPCAT1, and ATF7IP. TERT is the reverse transcriptase component of telomerase, encodes the catalytic subunit in the telomerase complex, is the telomerase rate-limiting enzyme, and is the key to maintain telomere DNA length, chromosome stability and cell immortality. It may be involved in the development of tumors through a variety of ways
The research on the function of CLPTM1L in malignant tumors remains in the early stage. This gene has great value in the diagnosis and treatment of malignant tumors.
Supplemental Material
Supplemental Material, sj-rar-1-cll-10.1177_09636897211045970 - CLPTM1L Is a Novel Putative Oncogene Promoting Tumorigenesis in Oral Squamous Cell Carcinoma
Supplemental Material, sj-rar-1-cll-10.1177_09636897211045970 for CLPTM1L Is a Novel Putative Oncogene Promoting Tumorigenesis in Oral Squamous Cell Carcinoma by Yunwen Hou, Feifei Xue, Yu Fu, Guanying Feng, Ruixia Wang and Hua Yuan in Cell Transplantation
Footnotes
Author Contribution
Yunwen Hou, Feifei Xue and Yu Fu contribute equally to this work
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The study was approved by the research ethics committee of Stomatological Hospital of Jiangsu Province.
Statement of Human and Animal Rights
All procedures in this study were conducted in accordance with the research ethics committee of Stomatological Hospital of Jiangsu Province approved protocols (2016-137).
Statement of Informed Consent
Every patient signed an informed consent.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National Natural Science Foundation of China (81672678), A Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD, 2018-87), The Project of Invigorating Health Care through Science, Technology and Education (Jiangsu Provincial Medical Youth Talent QNRC2016852), Health research project of Jiangsu Provincial Health Committee (ZDB202001) and sponsored by Qing Lan Project.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
