Abstract
Human adipose-derived mesenchymal stem cells (hAMSCs) are capable of immunomodulation and regeneration after neural injury. For these reasons, hAMSCs have been investigated as a promising stem cell candidate for stroke treatment. However, noninvasive experiments studying the effects of grafted stem cells in the host brain have not yet been reported. Cerebrospinal fluid (CSF), which can be collected without sacrificing the subject, is involved in physiological control of the brain and reflects the pathophysiology of various neurological disorders of the central nervous system (CNS). Following stem cell transplantation in a stroke model, quantitative analysis of CSF proteome changes can potentially reveal the therapeutic effect of stem cells on the host CNS. We examined hAMSC-secreted proteins obtained from serum-free culture medium by liquid chromatography-tandem mass spectrometry (LC-MS/MS), which identified several extracellular matrix proteins, supporting the well-known active paracrine function of hAMSCs. Subsequently, we performed label-free quantitative proteomic analysis on CSF samples from rat stroke models intravenously injected with hAMSC (experimental) or phosphate buffered saline (control). In total, 524 proteins were identified; among them, 125 and 91 proteins were increased and decreased with hAMSC treatment, respectively. Furthermore, gene set enrichment analysis revealed three proteins, 14-3-3 theta, MAG, and neurocan, that showed significant increases in the hAMSC-treated model; these proteins are core members of neurotrophin signaling, nerve growth factor (NGF) signaling, and glycosaminoglycan metabolism, respectively. Subsequent histological and neurologic function experiments validated proliferative neurogenesis in the hAMSC-treated stroke model. We conclude that (i) intravenous injection of hAMSCs can induce neurologic recovery in a rat stroke model and (ii) CSF may reflect the therapeutic effect of hAMSCs. Additionally, proteins as 14-3-3 theta, MAG, and neurocan could be considered as potential CSF biomarkers of neuroregeneration. These CSF proteome profiling results would be utilized as valuable resource in further stroke studies.
Introduction
Stroke is defined as cerebral tissue damage that occurs due to the lack of oxygen and glucose in ischemic or hemorrhagic vascular events. This condition impairs various brain functions and is considered one of the most common causes of disability and death. Although stroke induces differentiation of endogenous neural stem cells into the phenotype of the most destroyed neuron, the neuro-functional recovery in clinical aspects remains unsatisfactory, and there are few treatment modalities for neuroregeneration 1,2 . Cell-based therapy is a promising treatment for neuronal damage and has potential regenerative and protective effects. In several animal models of stroke, transplantation of mesenchymal stromal cells (MSCs) reportedly improves neural function and reduces the infarct area 3 . Human adipose-derived mesenchymal stem cells (hAMSCs) display therapeutic capabilities that include regeneration of injured cells, immune modulation, angiogenesis, and neurogenesis 4 –9 . hAMSCs secrete inflammatory cytokines and growth factors that enhance tissue regeneration 10 . Furthermore, the clinical safety of hAMSCs has been proven in several human trials 11 –13 . These properties of hAMSCs have attracted intense research interest concerning their use in cell therapy for irreversible cell/tissue damage in diseases such as stroke. Preclinical studies based on mouse and rat stroke models have demonstrated a reduced lesion volume and enhanced functional behavior following hAMSC transplantation 14 –16 . In animal models of stroke, most studies have evaluated both behavioral and histological changes. Histological results provide evidence that is more objective; however, the subject must be sacrificed, which is impossible in a human trial. Therefore, most human trials have assessed functional recovery without laboratory assessments, such as histology. This lack of assessment makes it difficult to elucidate the biochemical effects of grafted stem cells in the host brain.
Cerebrospinal fluid (CSF) is an ideal biospecimen to study various neurological disorders as it is easily obtained from animals or patients and most closely reflects the pathophysiology of the central nervous system (CNS). Transplanted hAMSCs likely alter CSF proteins with their highly active paracrine function and the subsequent host response. However, no CSF proteomics studies have systematically assessed protein changes in the CSF following hAMSC treatment in a stroke animal model. In this study, we first investigated the secretome of hAMSCs from in vitro cell culture condition. Subsequently, we performed mass spectrometry-based label-free quantitation to measure proteome changes in the CSF of a rat stroke model after intravenous hAMSC administration. Our quantitative proteomics data revealed that 14-3-3 protein theta (Ywhaq), myelin-associated glycoprotein (MAG), and neurocan core protein (Ncan), which were previously associated with neuroregeneration 17 –20 , were highly upregulated following hAMSC injection. Subsequent neurologic function and histologic experiments validated the efficacy of hAMSC injection in a stroke model. This study produced precise and abundant proteomic data including the three proteins, which is expected to be used as a valuable resource for further stroke studies.
Materials and Methods
Overall Scheme of the Proteomics Experiment
We designed the experiment in two ways: characterization of the hAMSC secretome and quantitative proteome profiling of CSF from rats treated with hAMSCs (Fig. 1). By comparing these results, we expect to determine the paracrine and treatment effects of hAMSC in stroke model.
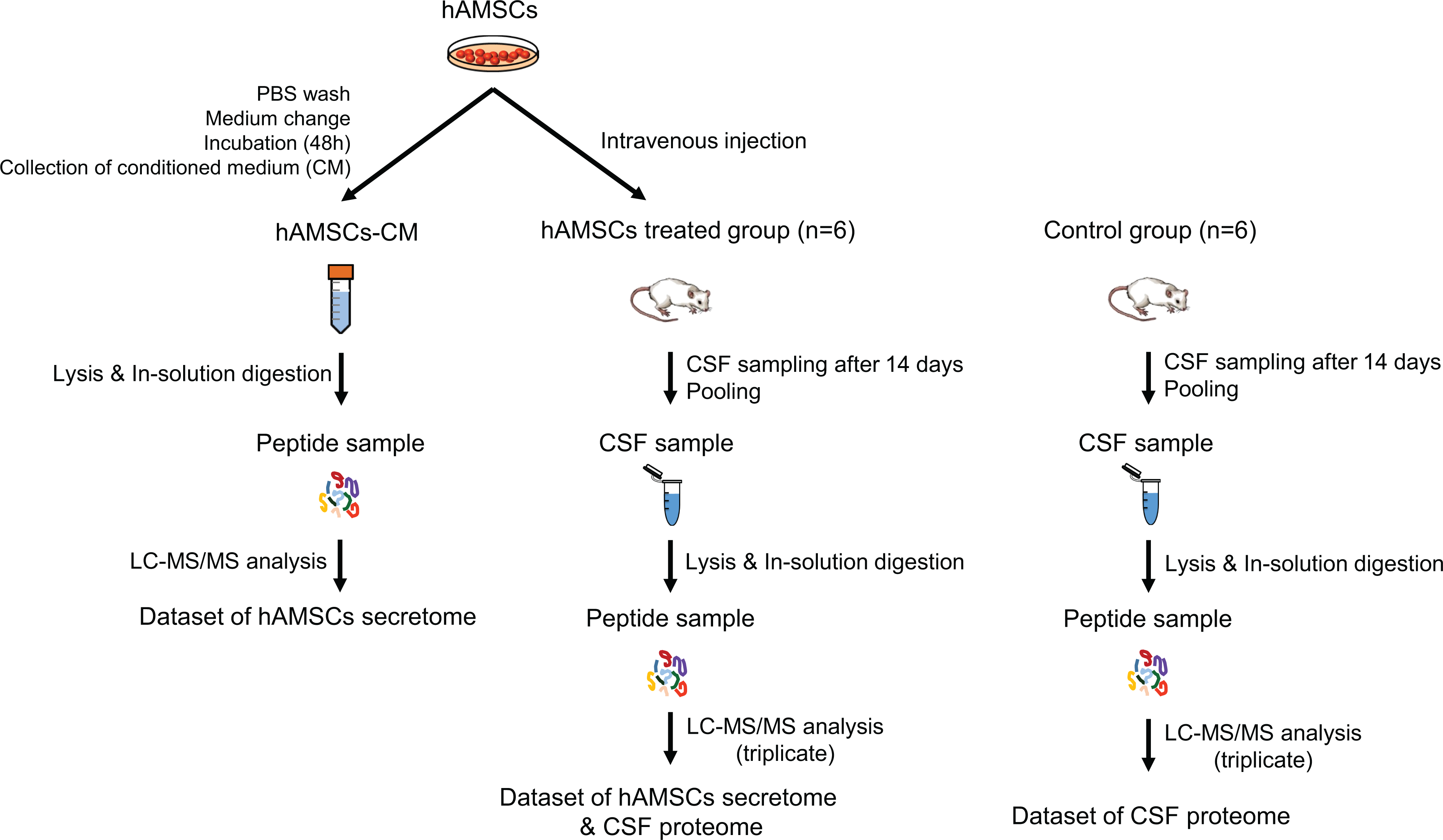
Schematic illustration of the secretome profiling procedure for hAMSCs (hAMSCs-CM analysis) and quantitative label-free CSF proteome analysis (hAMSCs treated vs control group study). Among control group and hAMSC-treated group (n = 19 in each group), 6 were included for CSF proteome study for each group.
Cell Culture and Cell Medium Preparation
Human AMSCs were kindly provided by RNL Bio Co., Ltd. (Seoul, Korea). These hAMSCs were expanded and characterized as previously described 12 . The hAMSCs used in these experiments were harvested from passages 5–6. Live hAMSCs were allowed to grow to 90% confluency on 100-mm diameter culture dishes in Keratinocyte-SFM (Invitrogen, Carlsbad, CA, USA)-based media containing 0.2 mM ascorbic acid, 0.09 mM calcium, 5 ng/mL rEGF, and 5% fetal bovine serum at 37°C in an atmosphere of 5% CO2 and 95% air. After repeated rinses with phosphate buffered saline (PBS), the cell medium was changed to serum-free Dulbecco’s Modified Eagle’s medium (DMEM; Gibco, Grand Island, N.Y., USA) and incubated for 48 h. Aliquots of the hAMSCs were then used for cell injection, and the conditioned medium (hAMSC-CM) was collected for proteomic analysis. hAMSC-CM was centrifuged at 3,000 × g for 4 min to remove debris, and the supernatants were frozen at -20°C [ 4 ]. A 500 µL aliquot of the hAMSC-CM sample was loaded on a 0.45 µm tube filter (Spin-x® Centrifuge Tube Filter, Corning, Inc., Corning, NY, USA). Subsequently, the filter was centrifuged at 9,000 × g for 1 min at 4°C to remove cell debris. The filtered hAMSC-CM samples were collected and concentrated in an Amicon Ultra-0.5 centrifugal filter with a molecular weight cutoff of 3 kDa (Millipore, Billerica, MA, USA), followed by the addition of 500 µL 50 mM ammonium bicarbonate and centrifugation at 14,000 × g for 30 min at 4°C. The flow-through was transferred to a 1.5-mL tube and stored at −80°C. Buffer exchange was performed by loading 450 µL 8 M urea in 50 mM ammonium bicarbonate on the filter and centrifuged at 14,000 × g for approximately 30 min at 20°C. Each sample was concentrated by reverse filtration and centrifuged at 1,000 × g for 2 min at 4°C. The protein concentration was determined by a BCA protein assay.
Animal Modeling and Cell Transplantation
All animal experiments were approved by the appropriate Institutional Review Boards of Korea University (Seoul, Korea; KUIACUC-20150324-3) and conducted in accordance with the National Institutes of Health Guide for the care and use of laboratory animals (NIH publication No. 80-23, revised in 1996) and the ARRIVE guideline. Transient middle cerebral artery occlusion (tMCAo) was induced using the intraluminal vascular occlusion method [5]. Using a face mask, male Sprague-Dawley rats weighing 250–300 g (Orient Bio Inc., Korea) were randomly assigned to the treatments and anesthetized with 4% isoflurane and maintained with 1.5% isoflurane in an atmosphere of 70% N2O and 30% O2. A 4-0 silicon rubber-coated monofilament (Doccol Corporation, Sharon, MA, USA) was advanced from the right common carotid artery into the lumen of the internal carotid artery until it blocked the origin of the MCA. Ninety minutes after tMCAo, reperfusion was performed by withdrawing the suture until the tip cleared the lumen of the common carotid artery [6]. One day after tMCAo, the rats were randomly divided into the PBS control group and hAMSC-treated group (n = 19 in each group; 19 for neurological function test/cerebral infarction volume measurement/immunohistochemistry and 6 for CSF proteome study in each group). For the hAMSC-treated groups, 2 × 106 hAMSCs in 1 mL PBS were injected intravenously via the tail vein 24 h after tMCAo, while only 1 mL PBS was injected in the control group. To label proliferating cells, we injected all rats with 5-bromo-2-deoxyuridine (BrdU, 50 mg/kg, in saline, i.p.) daily for 14 days after transplantation. After animal modeling, further investigations as neurological function test, CSF proteomics, and brain tissue evaluation were conducted and analyzed by independent investigators who were blinded to the treatment conditions.
Neurological Function Tests
Neurobehavioral functions were evaluated on days 1, 5, 10, and 14 using the modified Neurological Severity Score (mNSS) 21 . The mNSS is a composite of motor (muscle status and abnormal movement), sensory (visual, tactile, and proprioceptive), and balance tests. Neurological function was graded on a scale of 0–18 (normal–maximum deficit).
CSF Sampling and Protein Digestion
CSF samples were obtained on day 14 after treatment. The rats were anesthetized with 4% isoflurane, maintained with 1.5% isoflurane in an atmosphere of 70% N2O and 30% O2, and placed on a stereotactic head frame in the prone position. A longitudinal median skin incision was performed on the posterior neck, and the occipital bone and atlas were exposed. Approximately 100 µL CSF was aspirated through the cisterna magna using a flexible polyethylene tube. All CSF samples were centrifuged to remove erythrocytes, and only supernatants were collected for analysis. The protein concentration was determined using the BCA protein assay.
Proteins in each CSF sample were reduced with 10 mM dithiothreitol (GE Healthcare Life Sciences, Uppsala, Sweden) and incubated at 37°C for 1 h with gentle shaking on a thermomixer (Thermomixer Comfort, Eppendorf, Hamburg, Germany). Proteins were subsequently alkylated with 40 mM iodoacetamide and incubated at room temperature for 1 h with gentle shaking on the thermomixer in the dark. Samples were diluted 10-fold with 50 mM ammonium bicarbonate, and 1 M CaCl2 was added until reaching a final concentration of 1 mM CaCl2. The resulting sample was digested with trypsin at 37°C overnight (enzyme: protein ratio, 1:50). After the first digestion, additional trypsin (enzyme: protein ratio, 1:100) was added at 37°C for 6 h. Then, the peptide sample was acidified immediately after digestion with trifluoroacetic acid (TFA) to a pH of 2–2.5 22 .
A spin column was washed with 500 µL solution A (Sol A, 0.1% TFA) by centrifugation for 2 min at 1000 × g and 500 µL solution B (Sol B, 0.1% TFA + 80% acetonitrile) by centrifugation for 2 min at 1000 × g. The spin column was equilibrated with 500 µL Sol A followed by centrifugation for 2 min at 1000 × g. The equilibration step was repeated four times. The peptide sample was loaded and centrifuged for 2 min at 400 × g. The flow-through was loaded again and centrifuged for 2 min at 400 × g. This step was repeated twice. The spin column was washed with 500 µL Sol A by centrifugation for 2 min at 600 × g. Bound peptides were eluted using 200 µL Sol B by centrifugation for 2 min at 400 × g. This step was repeated twice using new tubes each time.
LC-MS/MS Experiments
The peptide sample was mass-analyzed using a linear quadrupolar ion trap–Fourier transform ion cyclotron resonance mass spectrometer (LTQ-FT Ultra, Thermo Electron, San Jose, CA, USA) coupled to a modified ultra-performance LC device (nanoACQUITY, Waters, Milford, MA, USA) as previously described 23 . Briefly, the sample was loaded onto an in-house solid phase extraction trap column (150 µm inner diameter, 3 cm in length) and transferred online to an in-house packed analytical column (75 µm × 100 cm). The LC flow rate was 300 nL/min. Solvent A was 0.1% formic acid in water, and solvent B was 0.1% formic acid in acetonitrile. The gradient was 300 min long and comprised 0–40% of solvent B in 270 min, 40–80% of solvent B in 20 min, and 1% of solvent B in 5 min. The electrospray voltage was 3 kV, and the capillary temperature was 200°C. MS scans (500–2000 m/z) were acquired at a resolution of 1.0 × 105 with an automatic gain control target value of 1.0 × 106. Data-dependent MS/MS was performed on the seven most intense peaks with a normalized collision energy of 35%. The isolation window for the MS/MS events was 3 Th.
Data Analysis
The precursor masses of the MS/MS data were corrected using post-experiment monoisotopic mass refinement (PE-MMR) software before a database search. The PE-MMR method was previously demonstrated to accurately assign precursor masses to MS/MS data before a protein database search 24 . The resultant MS/MS data from the hAMSC secretome sample were searched against the composite database of the UniProt human reference database (released May 2013; 90,398 entries) and common contaminants (179 entries) using the MS-GF+ search engine (v9387) (PMID: 25358478). The MS/MS data obtained from the CSF of hAMSC-treated rats were searched against the UniProt rat reference database (UP000002494; 21,668 entries) along with common contaminants using the MaxQuant platform (version 1.6.2.6) (PMID: 27809316). Fully tryptic peptides were allowed with up to two missed cleavages. The mass tolerance was 10 ppm for precursor ions. Cysteine carbamidomethylation was used as a static modification, and methionine oxidation and N-termini carbamylation were used for variable modification options. Peptides and proteins were identified based on 1% false discovery rates at both the protein and peptide levels using target-decoy analysis. An MS intensity-based label-free quantitative analysis was conducted using six data sets obtained from triplicate LC-MS/MS experiments of control and hAMSC groups as previously described using the iBAQ method (PMID: 21593866) within the MaxQuant platform with a match-to-match option.
The P-value was calculated based on normalized iBAQ values using the t-test function in an Excel program, and the fold-change was estimated based on the sum of all replicate intensities. To avoid infinite values during calculating fold changes, we added minimal noise values (500) to all data points. Gene set enrichment analysis (PMID: 16199517) was carried out using GSEA 3.0, which was downloaded from http://software.broadinstitute.org/gsea/. A list of genes identified from CSF samples was loaded onto the software to run GSEA. Curated gene sets related to canonical pathways in the Molecular Signatures Database were used. Gene set size filters (min = 6, max = 100) were used to calculate enriched pathways with 1,000 permutations.
Brain Tissue Preparation and Evaluation
Cerebral Infarction Volume Measurement
On day 14 after treatment, subsequent to CSF sampling as described above, the rats were anesthetized with a mixture of Zoletil (30 mg/kg) and xylazine (5 mg/kg) before perfusion with saline. The brain was then removed and sliced into 2 mm coronal sections before incubation with cold 2% 2,3,5-triphenyl tetrazolium chloride (TTC; Sigma, St. Louis, MO) for 30 min. Both sides of each slice were scanned by a digital camera, and the infarction area was determined using ImageJ (NIH, Bethesda, MD, USA). The infarction areas were measured as the area relative to the area of the contralateral hemisphere. For each animal, the average of the infarction areas from five coronal slices and 10 sides was obtained and evaluated. After correcting for edema, the infarction volume was calculated as follows: corrected infarct volume % = [contralateral hemisphere volume - (ipsilateral hemisphere volume - infarct volume)]/contralateral hemisphere volume × 100.
Immunohistochemistry
The animals were anesthetized with a mixture of Zoletil (30 mg/kg) and xylazine (5 mg/kg) and perfused transcardially with saline followed by 4% paraformaldehyde in 0.1 mol/L phosphate buffer. After perfusion, brains were removed and kept in a 4% paraformaldehyde solution overnight at 4°C and then placed in a 30% sucrose solution for cryoprotection. Frozen brains were sliced into 20-µm thick coronal sections at using a cryostat vibratome (CM3050 S, Leica Microsystems) and stored at −80°C until further processing. Sections were collected every sixth section between bregma levels +1.0 and −0.2 mm to quantify BrdU and doublecortin (DCX) labeling.
For BrdU staining to evaluate the degree of neurogenesis, DNA was first denatured by incubation in 2 N HCl for 40 min at 37°C followed by neutralization with immersion in 0.1 mol/L borate buffer (pH 8.5). The sections were then rinsed with PBS and blocked with 10% normal horse serum, 0.3% Triton X-100, and 1.0% bovine albumin serum in PBS for 1 h at room temperature and then incubated with a mouse monoclonal antibody against BrdU (1:100; Roche) overnight at 4°C. For DCX immunohistochemistry, an immature neural cell marker, the same protocol was used, excluding the denaturing steps. Sections were incubated with anti-DCX antibody (1:25; Santa Cruz Biotechnology, CA) in PBS (0.3% Triton X-100/1.0% bovine albumin serum). The following secondary antibodies were used: Alexa Fluor 488 anti-mouse IgG (1:400; Invitrogen) and Alexa Fluor 594 anti-goat IgG (1:800; Invitrogen) in PBST. Fluorescence colocalization was detected using a Zeiss LSM 700 confocal laser microscope (Carl Zeiss). Analyses of the total number of BrdU and DCX-positive cells in the ipsilateral hemisphere were performed in a similar manner to those described above. We used MetaMorph imaging software (version 7.8.1; Molecular Devices) to quantify BrdU and DCX double staining in the ipsilateral hemisphere.
Quantification and Statistics
Staining experiments and quantification analyses were performed in six sections from each animal. To count the total number of BrdU and DCX-positive cells, we obtained six sections every 280 μm, beginning at a section 1.2 mm rostral to the bregma. The sum of the values from all six sections was used as the final value. Data are expressed as the mean ± SEM. The significant differences between the two groups were evaluated by Student’s t-test. A P-value < .05 was considered statistically significant. IBM SPSS Statistics version 25 software (IBM, New York, NY) was used for the analyses.
Results
Characterization of the hAMSC Secretome and Quantitative Profiling of CSF Proteome
Secreted proteins from hAMSCs were collected and analyzed using LC-MS/MS. We identified 625 secretory protein groups with two or more sibling peptides in hAMSC-CM. The hAMSC secretome was classified according to biological processes using the Web-based Gene Set Analysis Toolkit (WebGestalt, http://bioinfo.vanderbilt.edu/webgestalt) 25 . Gene ontology of the 625 secretory proteins was also analyzed based on biological processes. Several clusters pertaining to extracellular structure organization, regulation of peptidase activity, and posttranscriptional regulation of gene expression were identified (Fig 2). The list of proteins is available in Supplemental Table S1.
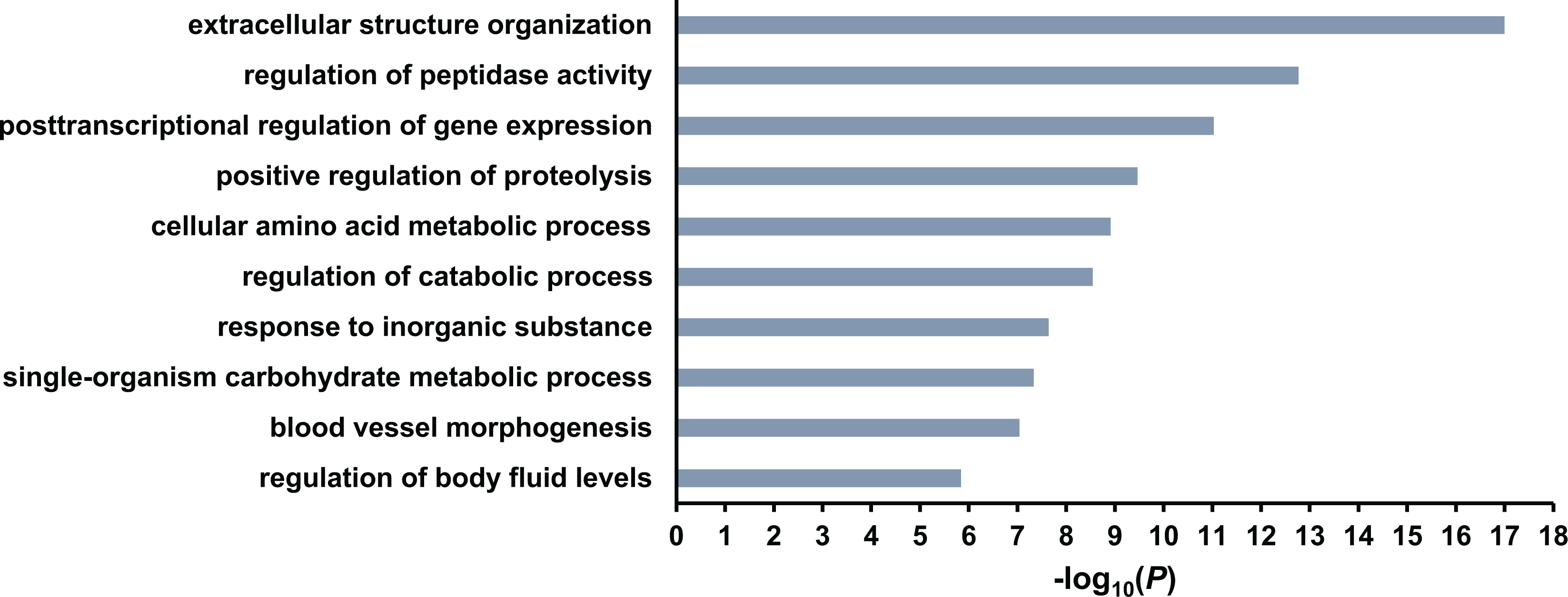
Identification of the top biological processes of hAMSC proteins. The P-value was computed by WebGestalt.
We then compared the CSF proteome between control and hAMSC-treated (hAMSC) groups by label-free quantitative proteomics. The samples were analyzed by LC-MS/MS 26 in triplicate in each group. As shown in Supplemental Fig. 1, six LC-MS/MS runs were performed, and the base peak chromatograms from the triplicate experiments demonstrated high reproducibility among the triplicate LC-MS/MS experiments. Tandem mass spectral data were searched using MaxQuant, and the protein abundance was estimated using the iBAQ method for label-free quantitation. The distribution of the most abundant proteins revealed albumin as an abundant CSF protein (Fig. 3). In total, 524 proteins were identified from six experiments. As shown in Fig. 4A, several proteins were altered by more than two-fold: 125 proteins were upregulated, and 91 proteins were downregulated. A full list of proteins is available in Supplemental Table S2. Since we carried out triplicate experiments, we were able to measure the significance of these changes, as shown in Fig. 4A. Thus, we conducted Student’s t-test using the iBAQ values per protein with triplicate data points. The result is depicted as a volcano plot and shown in Fig. 4B. More than 200 proteins were significantly altered, and some are highlighted in red (Fig. 4B).
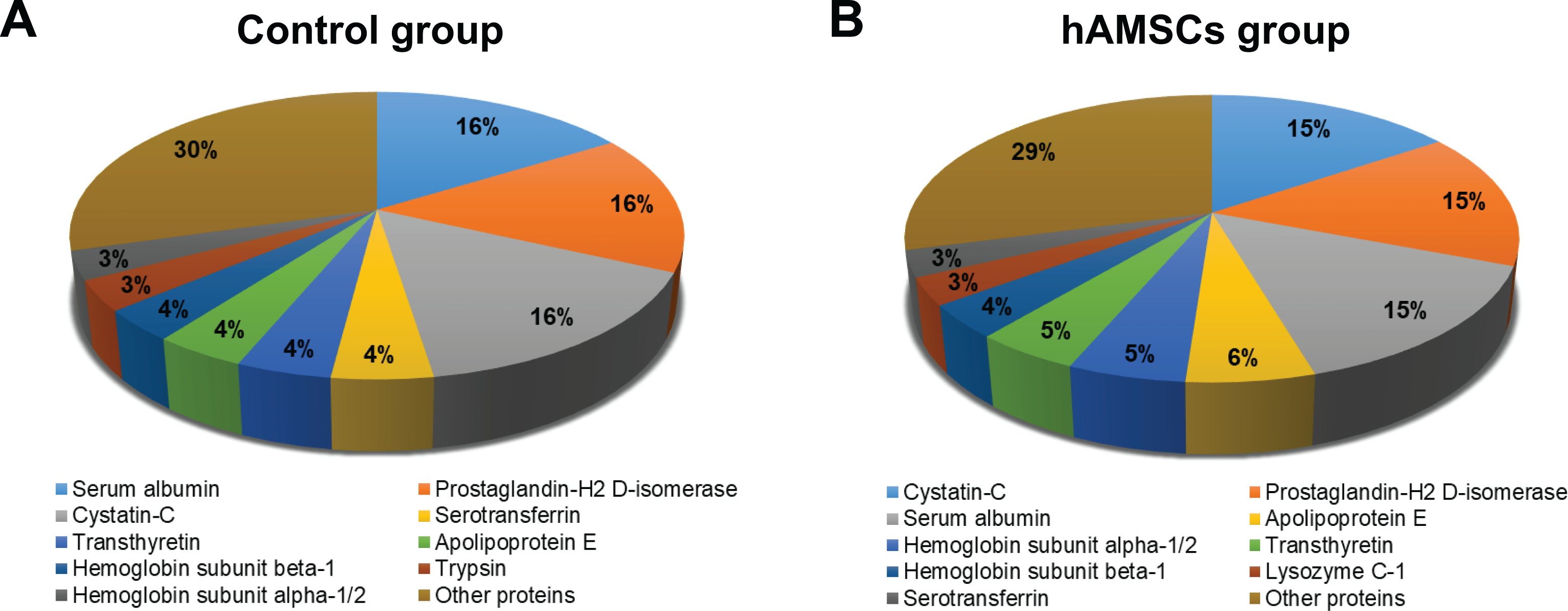
Distribution of abundant proteins identified in the control (a) and hAMSC groups (b).
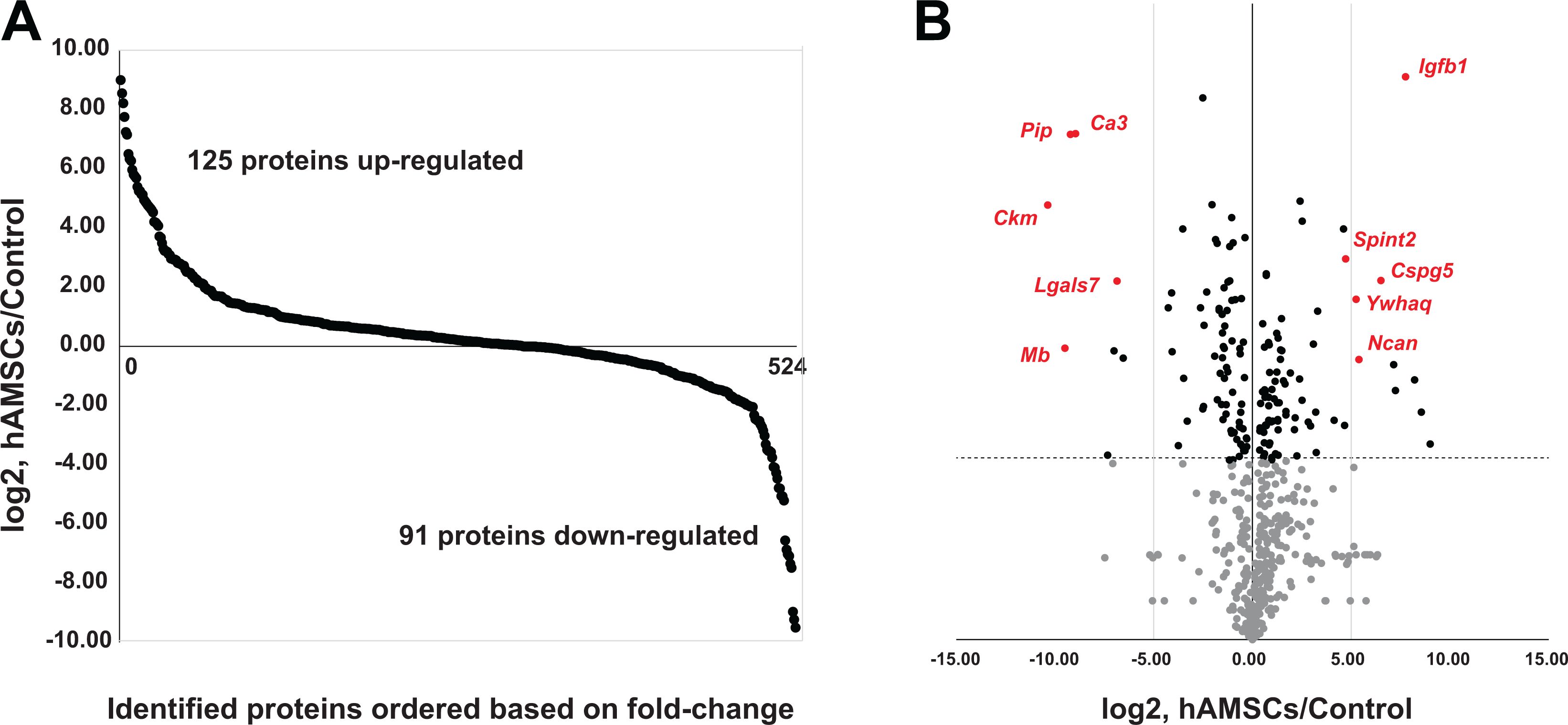
Identified proteins from rat CSF samples. In total, the 524 proteins identified were ordered based on their fold-change (log2, hAMSCs/Control) (a). Volcano plot depiction based on fold-change and P-value as estimated by Student’s t-test. (b)
Enriched Gene Sets in CSF of the Stroke Model
We performed GSEA to identify enriched pathways (Figs. 5, 6). Of 1329 gene sets loaded from “c2.cp.v6.2.symbols.gmt,” 1,224 sets were filtered out, leaving 105 sets. With FDR <25%, 33 and 19 gene sets were enriched in the hAMSC group as up- and downregulated, respectively. At nominal P-values <5%, 23 and 16 gene sets were enriched in the hAMSC group as up- and downregulated, respectively. Of these gene sets, representative enriched gene sets are shown in Fig. 5. Upregulated gene sets (Fig. 5A–D) included pathways such as neurotrophin signaling, glycosaminoglycan metabolism, nerve growth factor (NGF) signaling, and insulin receptor pathways in cardiac myocytes. Conversely, downregulated gene sets (Fig. 5E–H) contained complement and coagulation cascades, fibrinolysis pathway, hemostasis, and the Acute Myocardial Infarction (AMI) pathway.
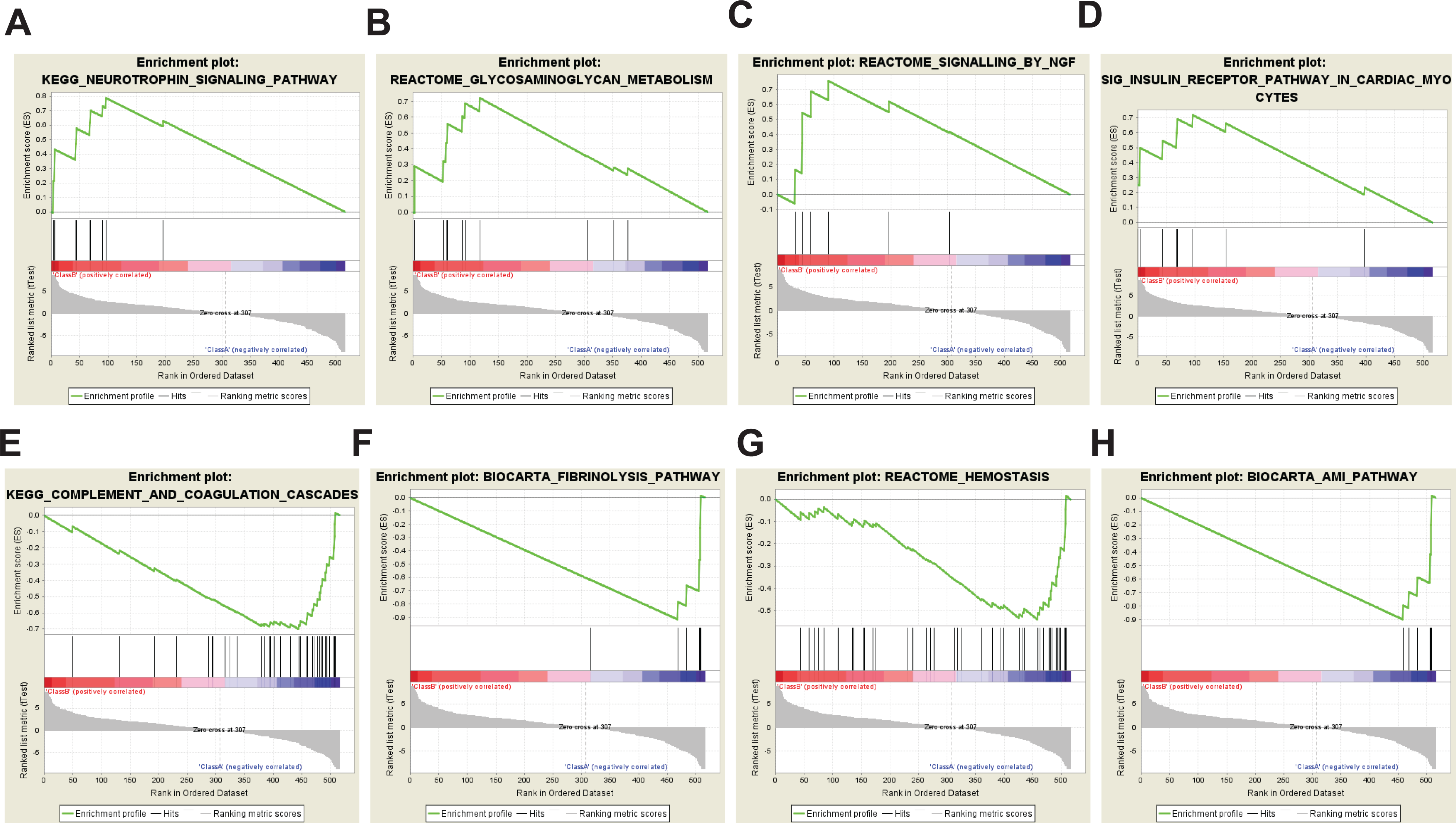
Representative altered pathways in the hAMSC group. Pathways such as the neurotrophin signaling pathway (a), glycosaminoglycan metabolism (b), signaling by ngf (c), and insulin receptor pathway in cardiac myocytes (d) were activated in the rat stroke model
Comparative Analysis of Genes Found in hAMSCs and CSF of the Stroke Model
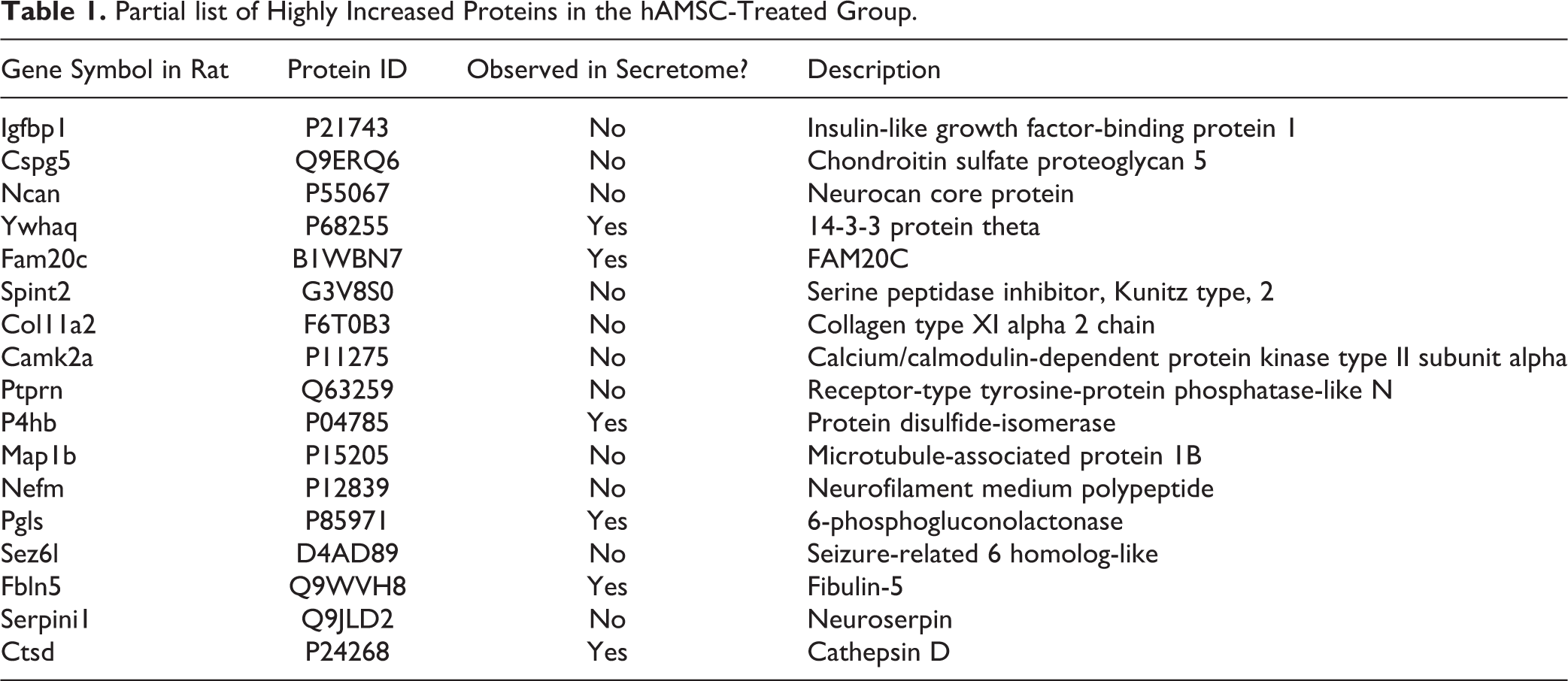
To determine potential key molecules in these enriched pathways, we performed comparative analysis of secreted proteins from the hAMSC and CSF proteome of the stroke model. A full list of genes is available in Supplemental Table S3. Of 524 rat genes observed at the protein level in CSF samples, interestingly, protein products of 141 human genes were detected from the hAMSC secretome. Although we were not able to ascertain whether all 141 proteins were human analogs, some could circulate even after day 9. Table 1 shows a partial list of highly upregulated proteins observed in CSF samples of the stroke model. Some proteins, such as Ywhaq, Fam20c, P4hb, Pgls, Fbln5, and Ctsd, were also found in the human secreted proteins; however, all proteins, except Ywhaq and Fam2c, were rat versions of these proteins. This finding indicates that the expression of these highly upregulated hAMSC-secreted proteins was induced during treatment.
Partial list of Highly Increased Proteins in the hAMSC-Treated Group.

Decrease in Cerebral Infarction by hAMSC Treatment
We carried out tMCAO to evaluate the effect of hAMSC transplantation on neurological function recovery using mNSS. Compared with rats in the PBS-treated control group, rats in the hAMSC-treated group showed significantly lower mNSS at days 10 and 14 after tMCAO (Fig. 6A). Furthermore, the hAMSC transplantation group showed a significant decrease in cerebral infarction volume compared to that in the control group (control 58.4 ± 2.4 vs hAMSC 42.7 ± 3.5, P < .01). Representative histology data are shown in Fig. 6B.
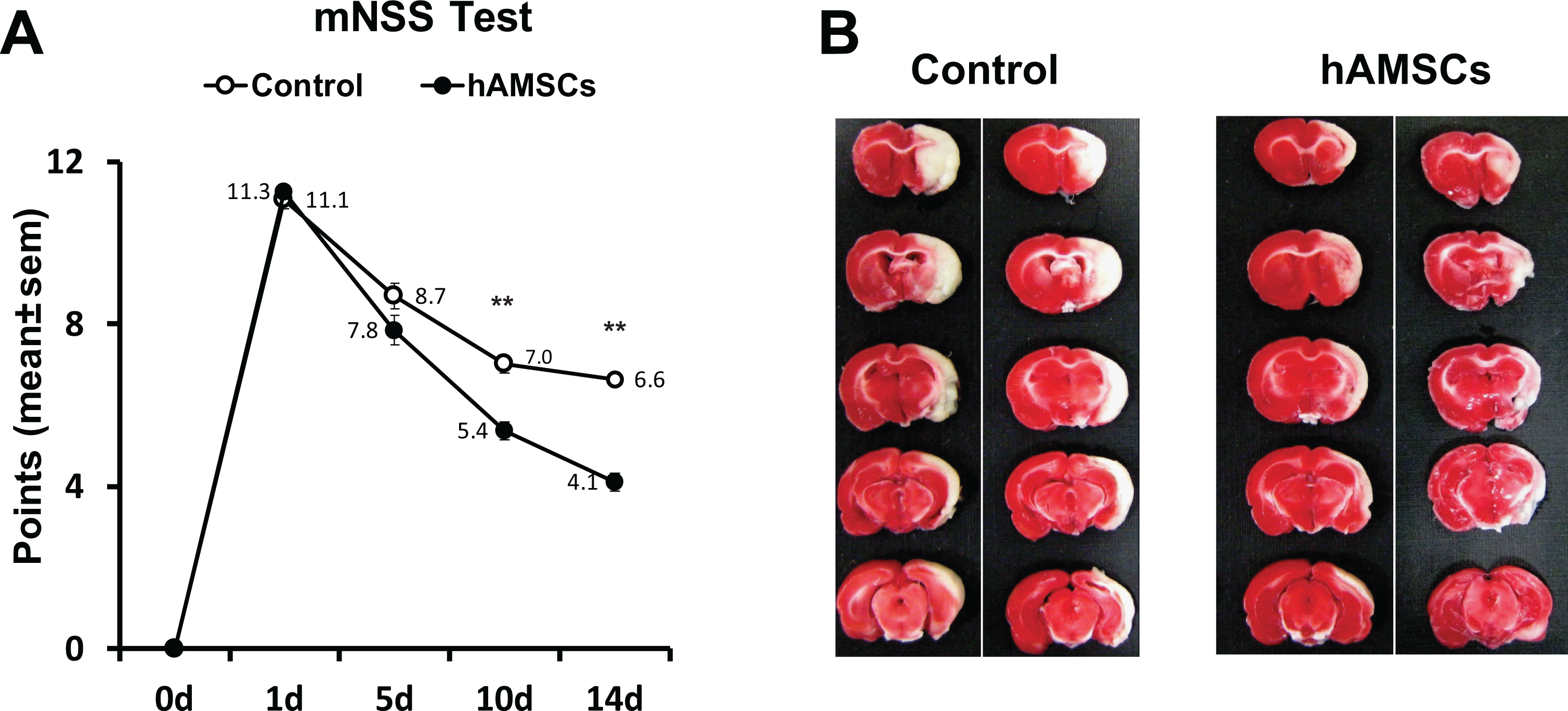
Human AMSC transplantation improves brain damage. Lower scores are shown in the hAMSC-treated group than in the control group after day 10 in the mNSS test. Mean and SEM values are indicated as numbers and bars. ** indicates P-value < .01 (a). Cerebral infarction volume decreased with hAMSC administration (b).
Increase in Proliferative Neural Progenitor Cells with hAMSC Treatment
Proteomic results of the CSF from the hAMSC-treated group suggested neuroregeneration through activated pathways related to neurogenesis. Thus, we conducted immunohistochemical analysis using a proliferative marker (BrdU) and neural progenitor cell marker (DCX). To evaluate the degree of neurogenesis, we counted the number of BrdU+/DCX+ colocalized cells, signifying proliferative neural progenitor cells. The number of colocalized cells was significantly higher in the hAMSC group, indicative of greater neuronal proliferation and regeneration (Fig. 7A, B).
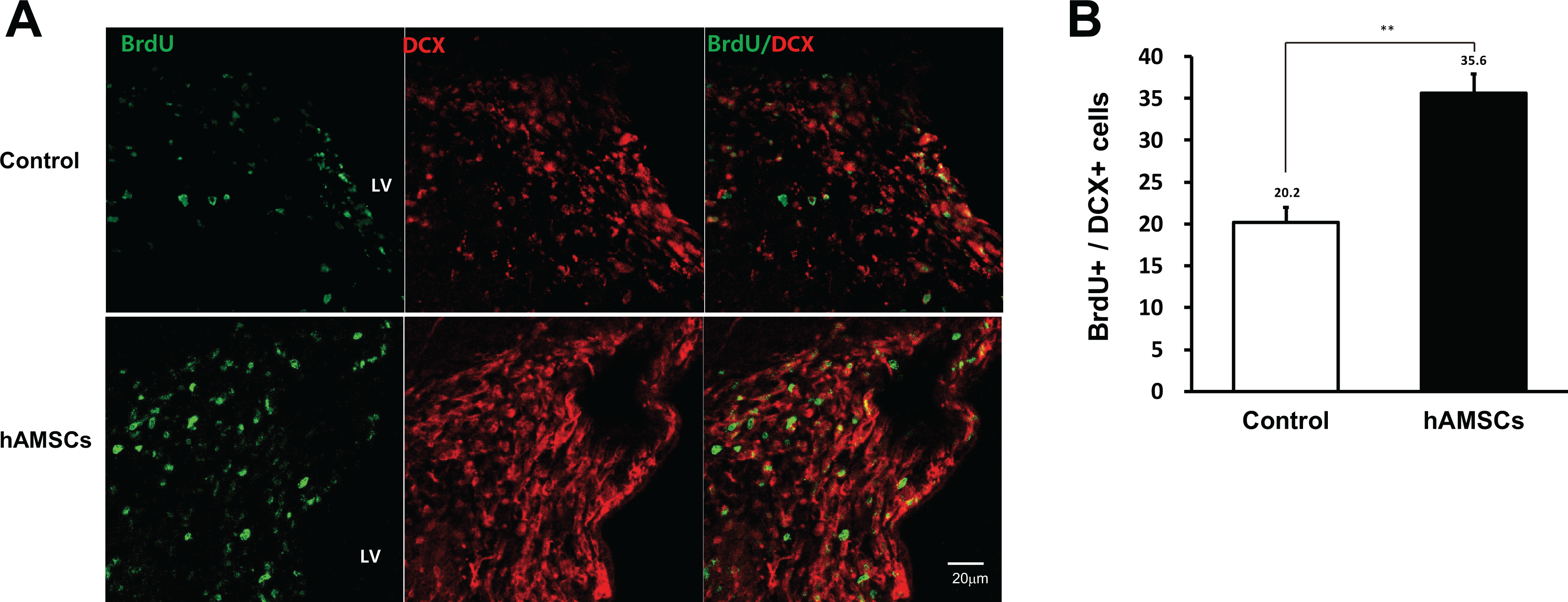
High colocalization of BrdU/DCX-positive cells in the hAMSC group. Representative immunohistochemical images (a) and their quantitative counts (b). Mean and SEM values are indicated as numbers and bars. ** indicates P-value < .01
Discussion
Human AMSCs secrete numerous proteins related to the regulation of cell death, apoptosis, and the wound response. These secreted proteins are expected to show a therapeutic effect following intravenous injection. However, the function of the secretome is unknown. The effects of grafted stem cells in the brain following stem cell transplantation are also unclear. Presently, we investigated in serial, the secretome of hAMSC-CM and CSF from hAMSC treated stroke rat model. Analysis of CSF can clarify the state of the CNS. CSF is a fluid that surrounds the brain as a protective buffer and is a reservoir for the circulation of hormones and biological substances as well as for the excretion of body waste. Since CSF is closely related to the brain, the CSF state can reflect the pathophysiology of various disease states and can shed light on various neurodegenerative disorders in the CNS. Therefore, CSF may be an attractive source of biomarkers following stem cell transplantation in experimental animal models of stroke. Furthermore, the use of these biomarkers and animal models might reveal the effect of stem cells on the host CNS following stroke.
We identified 625 proteins from the hAMSC secretome and performed bioinformatics analysis. The profile of the identified hAMSC secretome featured biological processes related to cell growth, cell regulation, and neurogenesis. The hAMSC secretome displayed active paracrine and neurodegenerative functions. In addition, we investigated the CSF proteomes of a stroke model using label-free quantitative proteomics to determine the altered production of CSF proteins in control and hAMSC-injected groups. From the label-free proteomic experiments, we identified 125 upregulated and 91 downregulated proteins in the CSF after hAMSC transplantation, respectively. Gene set enrichment analysis using differentially expressed proteins (DEPs) showed the activation of several neurogenesis-related pathways. For example, upregulated DEPs included proteins involved in neuroregeneration, such as those encoded by the Ywhaq gene, which is a member of the 14-3-3 family, a highly conserved protein family in mammals 27 . As shown in Supplemental Fig. S2, this protein is 99% identical among the mouse, rat, and human orthologs, thereby making it difficult to distinguish the rat version from the hAMSC-secreted version. In the hAMSC-treated stroke model, 14-3-3 protein theta was identified as a member of the activated neurotrophin signaling pathway. MAG encoded by the Mag gene is a type 1 transmembrane protein glycoprotein and has important positive functions in glia–axon interactions in the CNS and myelination during nerve regeneration 28 . Mag was identified as a member of the activated NGF signaling pathway with hAMSC treatment. Neurocan modulates neuronal adhesion and neurite growth during development by binding to neural cell adhesion molecules 19 . Transplantation of hAMSCs for stroke promotes the production of neurocan core protein (Ncan), which is a member of activated glycosaminoglycan metabolism. Neurocan can reduce expression in astrocytes, which suppresses axonal sprouting, and in turn enables MSC-mediated permissive synaptogenesis following stroke 29,30 . Therefore, these proteins might be considered efficient therapeutic biomarkers in the treatment of stroke.
The neurotrophin signaling pathway showed upregulation in the hAMSC group, and this pathway is related to nervous system development and functions, such as cell fate control, axon growth and guidance, dendrite structure and pruning, synapse formation, and synaptic plasticity 31 . The complement and coagulation cascades, which were downregulated in the hAMSC group, are activated after cerebral injury (including stroke) to induce inflammatory reactions and are associated with unfavorable clinical outcomes 32 –34 . These representative up- and downregulated pathways in the CSF could be strong evidence of the hAMSC treatment effect after stroke, explaining the behavioral and pathological recovery.
Several studies have used hAMSCs to treat damaged cells by directly placing them on the pathologic site. In this experiment, we administered hAMSCs intravenously to subjects with induced cerebral infarction. Indirect intravenous administration of hAMSCs resulted in improvement in neural damage by reducing the infarction volume and significantly increasing neuronal proliferation and neural progenitor cells. This improvement in stroke-related damage may be induced by the paracrine effect of administered hAMSCs, as indicated by the CSF proteome alteration. This result suggests that clinical, intravenous administration of adipose-derived stem cells may have a good impact on stroke treatment.
The limitation of this paper is that although proteomics results and literature reviews have introduced specific proteins that are believed to be related to stroke recovery, direct validation experiments have not been performed on these proteins and follow-up studies are needed in the future.
Conclusions
In summary, we used a rat stroke model to observe the influence of hAMSCs on CSF, which could reflect conditions in the CNS. The proteomic analysis revealed proteome changes after intravenous injection of hAMSCs. Furthermore, pathway analyses identified three proteins (14-3-3 theta, MAG, and neurocan) as core members of neurotrophin signaling, NGF signaling, and glycosaminoglycan metabolism, respectively. These pathways are associated with neuroregeneration. Therefore, these proteins are potential CSF biomarkers for stroke treatments. However, further studies are warranted to demonstrate the expression and neuroregenerative effects of these candidate proteins and pathways. Above all, the most important significance of this study is to analyze CSF to produce precise and abundant proteomic data, which is expected to be used as a valuable resource for further stroke studies.
Supplemental Material
Supplemental Material, sj-ai-1-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-ai-1-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Supplemental Material
Supplemental Material, sj-ai-2-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-ai-2-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Supplemental Material
Supplemental Material, sj-docx-1-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-docx-1-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Supplemental Material
Supplemental Material, sj-docx-2-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-docx-2-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Supplemental Material
Supplemental Material, sj-docx-3-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-docx-3-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Supplemental Material
Supplemental Material, sj-jpg-1-cll-10.1177_09636897211023474 - Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy
Supplemental Material, sj-jpg-1-cll-10.1177_09636897211023474 for Label-Free Quantitative Proteome Profiling of Cerebrospinal Fluid from a Rat Stroke Model with Stem Cell Therapy by Junseok W Hur, Min-Sik Kim, Se-Yeon Oh, Ho-Young Kang, Jingi Bae, Hokeun Kim, Hangyeore Lee, Sang-Won Lee and Dong-Hyuk Park in Cell Transplantation
Footnotes
Authors’ Contributions
JWH, MSK, SWL, and DHP designed the experiments. HYK performed animal experiments. SYO, JB, HK, HL performed mass spectrometry experiments JWH, MSK, SWL, and DHP analyzed the results. JWH, MSK, and SYO wrote the manuscript. All authors edited, reviewed and approved the manuscript.
Authors Contribution
*These authors (HUR JW, KIM MS, and OH SY) contributed equally to this work
Authors' Note
Junseok W Hur, Min-Sik Kim & Se-Yeon Oh authors contributed to this work equally and should be regarded as co-first authors.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This study was approved by the Korea University Institutional Animal Care and Use Committee (KUIACUC-20150324-3).
Statement of Human and Animal Rights
All procedures in this study were conducted in accordance with the Korea University Institutional Animal Care and Use Committee (KUIACUC-20150324-3) approved protocol.
Statement of Informed Consent
There are no human subjects in this article and informed consent is not applicable.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported in part by grants from Korea University (K1913911, K2011341); the Basic Science Research Program through the National Research Foundation (NRF) funded by the Korean Ministry of Education, Science and Technology (NRF-2013R1A1A2057994); and the Korean Health Technology R&D Project, Ministry of Health & Welfare, Republic of Korea (HI12C03370400) to Dong-Hyuk Park. This study was also supported in part by a grant from the Collaborative Genome Program for Fostering New Post-Genome Industry (NRF-2017M3C9A5031597) of the National Research Foundation funded by the Korean Ministry of Science and ICT to Sang Won Lee. We would also like to acknowledge the financial support from the Korea Healthcare Technology R&D Project (HI16C1002) and the Basic Science Research Program through the National Research Foundation (NRF-2017R1D1A1B03035760, NRF-2019R1C1C1010602, NRF-2017M3C7A1027472) funded by the Korean Ministry of Education, Science and Technology.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
