Abstract
CircRNAs participate in the pathogenesis of a variety of cancers. Previous studies showed that baculoviral IAP repeat containing 5 (BIRC5) can promote tumor progression. But, the mechanisms by which circRNAs regulate BIRC5 expression in hepatocellular carcinoma (HCC) remain unknown. The clinical prognosis of BIRC5 or miR-497-5p expression in patients with HCC was assessed by TCGA RNA-seq dataset. hsa_circ_0026939 (circANKRD52) or BIRC5 was identified to bind with miR-497-5p by luciferase gene report, RIP and circRIP assays. MTT, colony formation, Transwell assays and a xenograft tumor model were used to estimate the role of miR-497-5p or circANKRD52 in HCC cells. As a result, we found that elevated expression of BIRC5 or decreased expression of miR-497-5p was linked to poor survival in HCC. Restored expression of miR-497-5p repressed cell proliferation, colony formation and invasiveness by targeting BIRC5, but its inhibitor showed the opposite results. Furthermore, circANKRD52 possessed a tumor-promoting effect by acting as a sponge of miR-497-5p and thereby upregulated BIRC5 in HCC cells. In conclusion, our findings demonstrated that circANKRD52 enhances the tumorigenesis of HCC by sponging miR-497-5p and upregulating BIRC5 expression.
Introduction
Hepatocellular carcinoma (HCC) ranks the fourth in incidence and is the second leading cause of deaths in China 1 . Despite the great progress in the detection of early-stage HCC, the advanced cases still indicate unfavorable outcomes owing to tumor infiltration and metastasis 2 . The pathogenesis of HCC is associated with the genetic mutation and dysregulation of oncogene or anti-oncogenes 3 . Identification of genetic molecules, non-coding RNAs (ncRNAs), and protein biomarkers is beneficial for early detection of HCC 4 .
Baculoviral IAP repeat containing 5 (BIRC5) as an inhibitor of apoptosis protein is associated with poor clinical outcomes and tumor growth 5 –8 . We found that increased BIRC5 expression was attributable to post-transcriptional regulation of miRNAs in HCC. They can act as oncogenic factors or anti-oncogenes by negatively regulating their targets in HCC 9 –12 . miR-203 or miR-138-5p represses the tumorigenesis by targeting BIRC5 in breast cancer 13 , ovarian cancer 14 , and bladder cancer 15 . These studies suggest that increased expression of BIRC5 may be regulated by miRNAs in HCC.
CircRNAs belong to a subgroup of endogenous ncRNAs with closed loop structure and resistance to RNase R in contrast to their corresponding linear RNAs 16 . Some studies have shown that circRNAs can act as the sponges of miRNAs in HCC 17,18 . CircRNA_100338 or circSLC3A2 expression is involved in promoting tumor metastasis, and acts as an oncogene by sponging miR-141-3p/-490-3p in HCC 19,20 , whereas circSETD3 and circMTO1 act as miR-421/-9 sponges to suppress growth and invasion of HCC cells 21,22 . Therefore, circRNAs may present novel markers for the detection of HCC.
Previous studies revealed that circDLST facilitates the tumorigenicity of gastric carcinoma (GC) 23 , but circLARP4 and circYAP1 repress GC tumorigenesis 24,25 . Herein, we found that the dysregulation of BIRC5 or miR-497-5p was linked to poor survival in HCC. Restored expression of miR-497-5p repressed GC by targeting BIRC5. Moreover, hsa_circ_0026939 (also termed as circANKRD52), predominantly localized in the cytoplasm, acted as a miR-497-5p sponge to increase BIRC5 expression, thereby contributing to HCC progression.
Materials and Methods
Identification of Differentially-Expressed Genes in HCC Samples
The differentially-expressed genes were identified in HCC tissues by GCBI and GEO dataset (https://www.ncbi.nlm.nih.gov/gds), of which BIRC5 had a significantly differential expression in HCC and was selected for further study.
Clinical Sample
The data for the pathological information of HCC and the expression of BIRC5 and 11 miRNAs (miR-135a-5p, miR-144-3p, miR-184, miR-195-5p, miR-204-5p, miR-218-5p, miR-335-5p, miR-485-5p, miR-497-5p, miR-499a-5p, and miR-501-5p) in HCC tissues were collected from TCGA dataset (http://xena.ucsc.edu/). In addition, a tissue microarray (TMA) consisting 87 tissue samples (Lot No. XT16-029) was provided by Outdo Biotech (Shanghai, China), and the detailed description of the clinicopathological information on HCC-TMA was indicated in supplemental Table S1. The study was approved by the Ethics Committee of General Hospital of Tianjin Medical University.
Cell Culture
Normal liver tissues and HCC cell lines (SMMC-7721, HepG2, SK-hep-1, Huh6, Huh7, and LO2) were stored in our laboratory and cultured in Dulbecco’s Modified Eagle medium (DMEM) medium supplemented with 10% heat-inactivated fetal bovine serum (FBS).
FISH Analysis
Biotin-labeled probe sequence for miR-497-5p (5′-ACAAACCACAGTGTGCTGCTG -3′) and digoxin-modified probe sequence for hsa_circ_0026939 (circANKRD52, 5′-GTATGGGGTTCCGCTCCCGCGGTGGAGGGC-3′) were used for FISH analysis of the expression levels and co-localization of circANKRD52 and miR-497-5p in HCC tissues or LO2 cell line. The detailed description of FISH analysis and quantification was conducted as previously reported 24 .
Quantitative Real-Time PCR (qRT-PCR)
The detailed description of qRT-PCR analysis was conducted as previously reported 24 . The primer sequences used in this study were listed in Supplemental Table S2.
Western Blot Analysis
SMMC-7721 and LO2 cell lines were harvested and extracted by using lysis buffer. The anti-BIRC5 (MAB2365, Abcam, Cambridge, MA, USA) was diluted at a ratio of 1:1000 according to the instructions and incubated overnight at 4°C. The detailed description of Western blot analysis was conducted as previously reported 24 .
Luciferase Reporter Assay
SMMC-7721 and LO2 cell lines were seeded into 96-well plates and co-transfected with a mixture of pRL-TK-pMIR luciferase reporter containing wild type (WT) and mutant (Mut) BIRC5 3’UTR or circANKRD52 3’UTR and miR-497-5p mimic or inhibitor. The luciferase activities of WT and Mut BIRC5 or circANKRD52 3’UTR were detected with a dual-luciferase reporter assay.
The Plasmid, shRNA, miR-497-5p Mimic and Inhibitor
BIRC5 or circANKRD52 plasmids (CTCCACCGCGGGAGCGGA A CCCCATACA), siRNA or shRNA vector targeting BIRC5 or circANKRD52 and miR-497-5p mimic or inhibitor were purchased from GenePharma (Shanghai, China). pcDNA3.1 and miR-NC were used as the control vector. RNAi sequences were listed as follows: si-BIRC5 GCACCACTTCCAGGGTTTATTCCCT and sh-circANKRD52: GCGGGAGCGGA A CCCCATACA.
MTT, Colony Formation and Transwell Assays
MTT, colony formation and Transwell assays were conducted as previously described 23 .
Actinomycin D and RNase R Treatment
Actinomycin D and RNase R used for the treatment of HCC cells were conducted as previously described 23 .
CircRNA In Vivo Precipitation (circRIP)
The circRIP assay was performed according to the previous report 22 .
RNA Immunoprecipitation (RIP)
RIP assay was conducted as previously described 23 .
In Vivo Tumorigenesis Assay
Male nude mice (6-week old) were from Shanghai SIPPR-BK Laboratory Animal Co. Ltd (Shanghai, China). All the animals used were approved by the Animal Ethics Committee of Zhengzhou University. The mice were subcutaneously inoculated with 1 × 107 of SMMC-7721 cells stably transfected with circANKRD52 or NC. The body weight and tumor size were examined, and the tumor volume was calculated according to the formula: length × width 2 /2.
Statistical Analysis
Statistical analysis was conducted as previously reported 24 .
Results
Increased Expression of BIRC5 Showed an Association with Worse outcomes in HCC
According to mRNA profiling from GSE17548 cohort, 678 differentially-expression genes were identified between HCC (n = 17) and adjacent normal tissues (n = 19), of which BIRC5 expression levels were markedly elevated in HCC tissues (P < 0.001, Fig. 1A). BIRC5 levels were further validated to be dramatically increased in pair-matched (n = 46) and unpaired tissue samples (n = 373; Fig. 1B). Additionally, BIRC5 also harbored an increased expression in HCC patients with stage II /III+IV or T3+T4 in contrast with stage I (P = 0.0024, P = 0.0002; Fig. 1C) or T1+T2 (P = 0.0034, Fig. 1D).
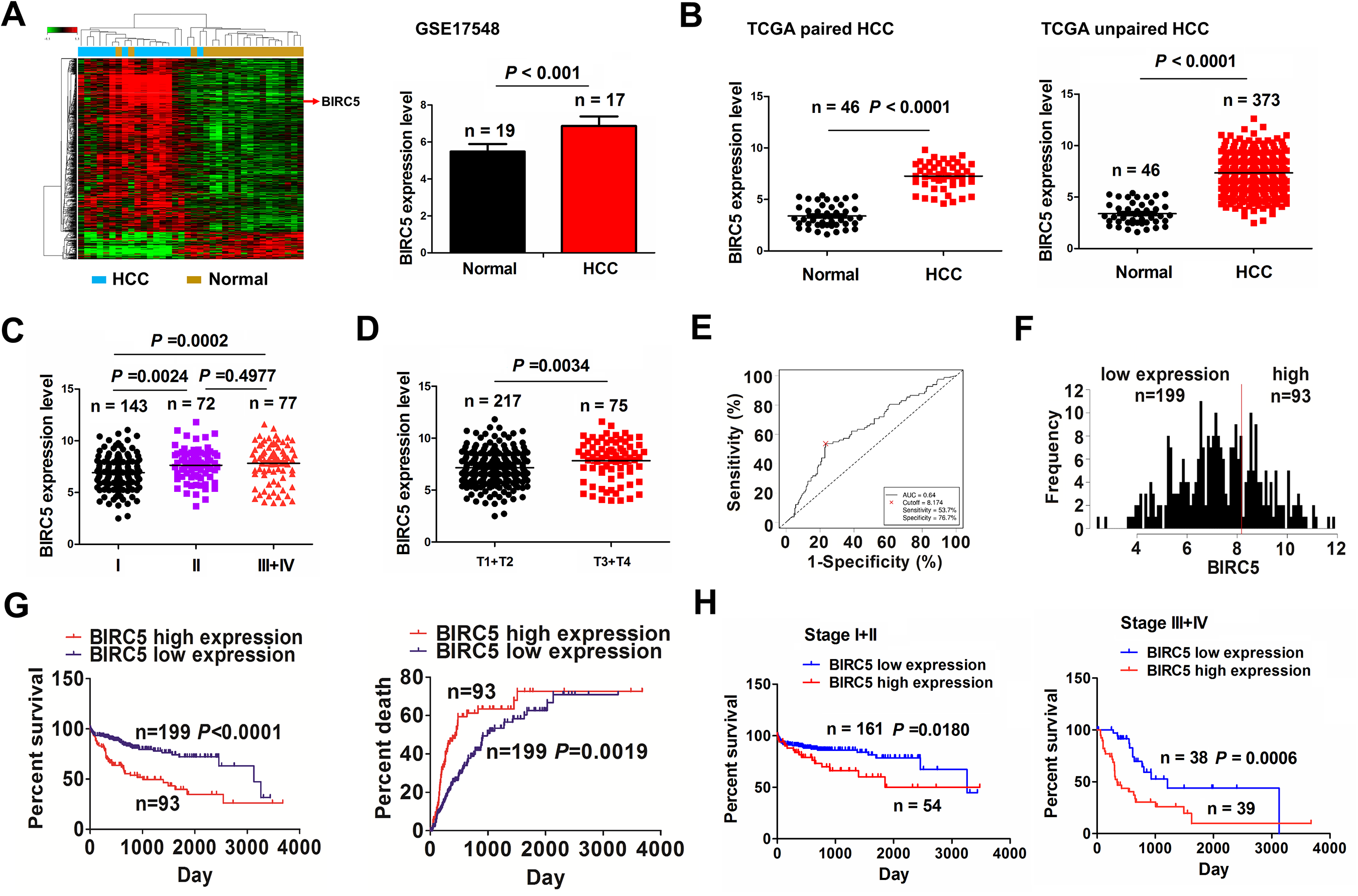
The association of BIRC5 expression with poor prognosis in patients with HCC. (A) GSE17548 analysis of the differentially-expressed BIRC5 gene between HCC (n = 17) and adjacent normal tissues (n = 19). (B) TCGA analysis of the expression levels of BIRC5 in paired (n = 46) and unpaired (n = 373) HCC tissue samples. (C, D) TCGA analysis of the expression levels of BIRC5 in HCC patients with stage I/II/III+IV and those with T1+T2/T3+T4. (E, F) ROC curve analysis of the cutoff value, sensitivity, specificity and AUC of BIRC5 in HCC patients, and its cutoff value divided the patients into high BIRC5 expression (n = 93) and low BIRC5 expression groups (n = 199). (G) Kaplan Meier analysis of the association of high or low BIRC5 expression with the overall survival and tumor recurrence in patients with HCC. (H) Kaplan Meier analysis of the association of high or low BIRC5 expression with overall survival in early-stage and late-stage HCC.
A cutoff value of BIRC5 (8.174) was obtained (Fig. 1E) and divided the cases into high and low BIRC5 groups (Fig. 1F). We found that the elevated level of BIRC5 was linked to pathological stage (P < 0.0001), tumor size (P < 0.0001) and distant metastasis (P = 0.005) in HCC (Supplemental Table S3). The cases with high BIRC5 levels harbored poorer survival (P < 0.0001) and higher tumor recurrence (P = 0.0019) in contrast with those with low BIRC5 levels (Fig. 1G). In terms of the TNM stage, the early-stage (P = 0.018) or advanced stage patients (P = 0.0006, Fig. 1 H) with increased BIRC5 levels also displayed poor survival rather than tumor recurrence (Fig. S1) in contrast with those with decreased BIRC5 levels. Furthermore, BIRC5 was demonstrated to be an independent factor of poor survival (P < 0.0001) rather than tumor recurrence (P = 0.068) in HCC (Supplemental Table S4, S5).
miR-497-5p Was Negatively Correlated with BIRC5 Expression in HCC
To explain the reason why BIRC5 was upregulated in HCC, we assessed the genetic and epigenetic changes of BIRC5, and found that BIRC5 expression had no association with its genetic (Fig. S2A) and methylation modification (Fig. S1B). Thus, whether BIRC5 expression was regulated by miRNAs at a post-transcriptional level was further investigated. A starBaseV2.0 prediction tool was used to screen 11 miRNAs which have the strongest binding potential with BIRC5 3’ UTR, and their expression levels were detected in pair-matched (Fig. 2A) and unpaired tissue samples (Fig. 2B). We found that miR-195-5p/-497-5p/-144-3p/-335-5p indicated obviously decreased levels (Fig. 2A, B), of which miR-195-5p/-497-5p/-144-3p rather than miR-335-5p were negatively correlated with BIRC5 in HCC tissues (n = 351, Fig. 2C and Fig. S3).
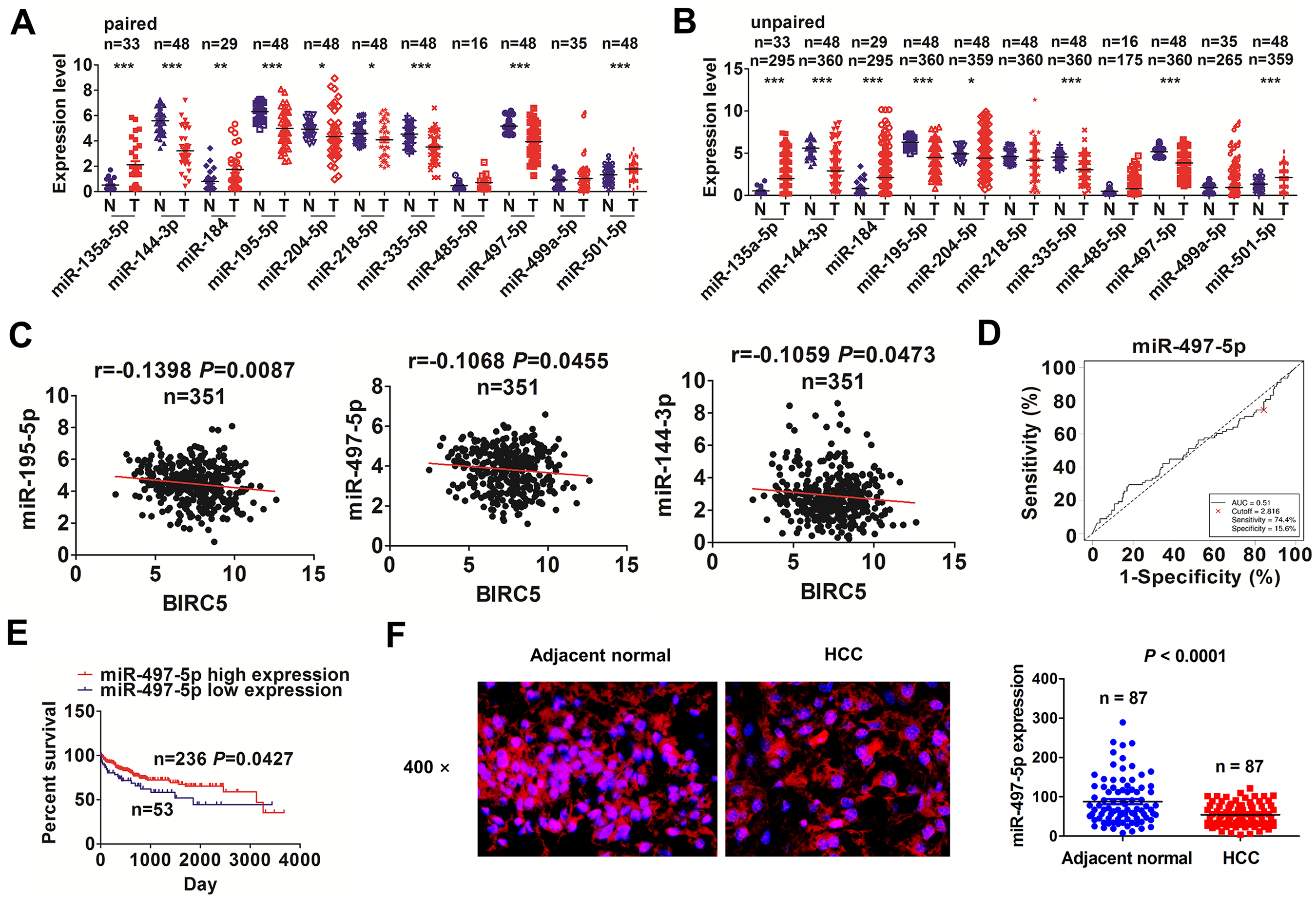
MiR-497-5p was negatively correlated with BIRC5 expression in HCC. (A, B) TCGA analysis of the expression levels of 11 miRNAs that target BIRC5 gene in paired and unpaired HCC tissues. (C) Pearson correlation analysis of the correlation of miR-195-5p/-497-5p/-144-3p with BIRC5 expression in HCC tissues (n = 351). (D) ROC curve analysis of the cutoff value, sensitivity, specificity and AUC of miR-497-5p in HCC patients. (E) Kaplan Meier analysis of the association of high or low miR-497-5p expression with overall survival in HCC patients. (F) FISH analysis of the expression levels of miR-497-5p in paired HCC tissue samples (n = 87).
Subsequently, we acquired the cutoff values of miR-195-5p/-497-5p/-144-3p in HCC patients (Fig. 2D and Fig. S4A, B), and found that miR-497-5p showed no relationship with the clinical parameters in HCC (Supplemental Table S6). The cases with decreased miR-497-5p levels instead of low miR-195-5p/-144-3p expression (Fig. S4A, B) harbored poor survival (P = 0.0427, Fig. 2E) rather than tumor recurrence (P = 0.7289, Fig. S5A) in contrast with those with increased miR-497-5p levels. Early- or late-stage cases with decreased miR-497-5p levels indicated no association with overall survival (Fig. S5B). Decreased miR-497-5p level was an independent predictor of poor survival in HCC (Supplemental Table S7). Intriguingly, FISH analysis further confirmed the decreased expression of miR-497-5p in HCC (n = 87, P < 0.0001; Fig. 2F).
BIRC5 Was Identified as a Target of miR-497-5p in HCC Cells
Having verified a negative relationship of miR-497-5p with BIRC5 in HCC, we decided whether BIRC5 was a target of miR-497-5p in HCC. A mirPath v.3 prediction tool identified the targets of miR-497-5p and their enrichment pathways, indicating that these targets were predominantly enriched in Hippo signaling pathway, where BIRC5 was a key downstream transcriptional factor of YAP by KEGG enrichment analysis (Fig. 3A and Fig. S6). Then, we assessed miR-497-5p levels in HCC cell lines by qRT-PCR, and found that miR-497-5p showed an increased level in LO2 cell line but a decreased levels in SMMC-7721 in contrast with normal liver tissues (Fig. 3B). We then found that miR-497-5p lowered BIRC5 levels in SMMC-7721 cell line (Fig. 3C), but its inhibitor had an opposite result in LO2 cell line (Fig. 3D). Luciferase reporter vectors embodying WT or Mut BIRC5 3’UTR (Fig. 3E) were pretreated with miR-497-5p in SMMC-7721 or LO2 cells. We found that the luciferase activity of WT BIRC5 3’UTR was reduced by miR-497-5p in SMMC-7721 cells, but that was elevated by miR-497-5p inhibitor in LO2 cells (Fig. 3F).
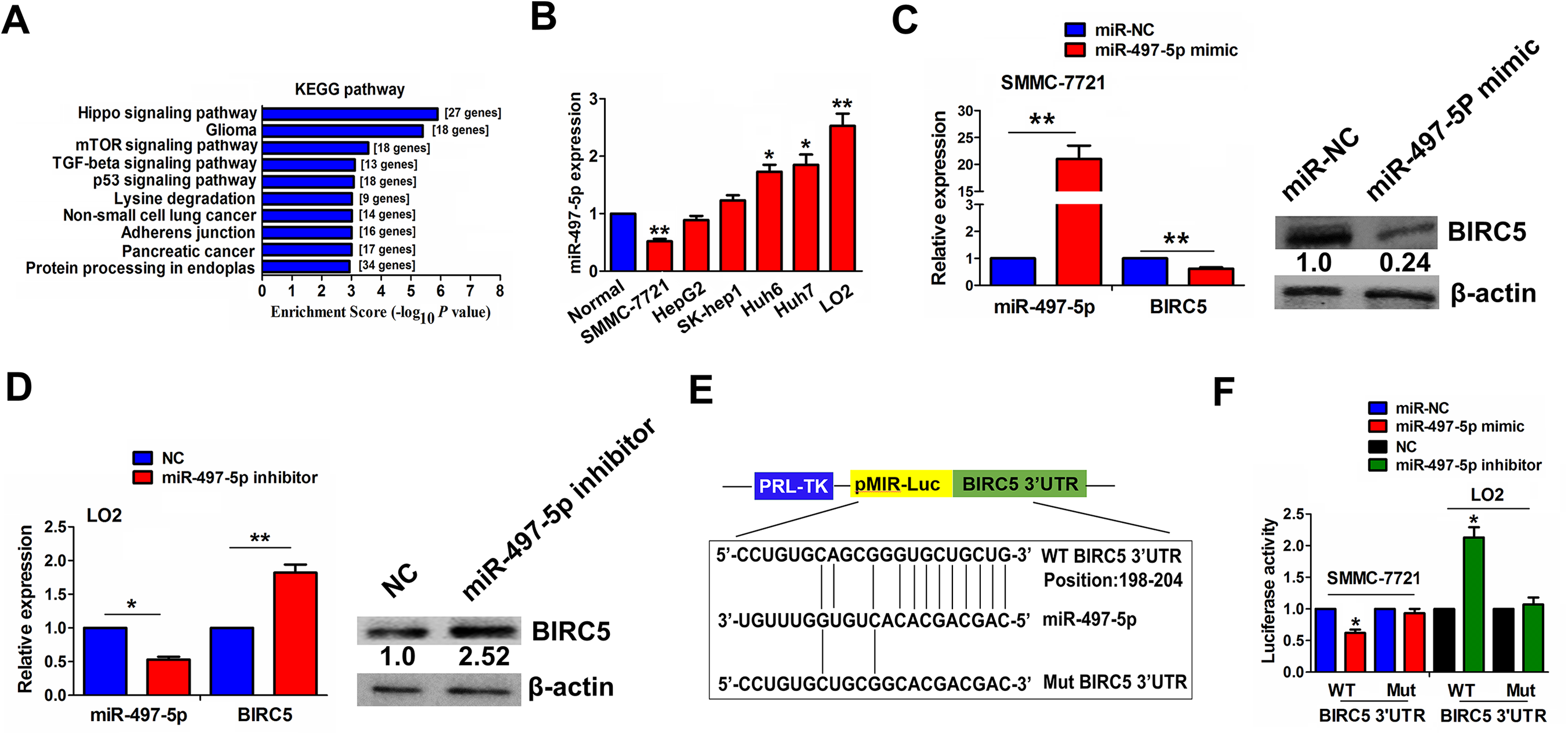
BIRC5 was identified as a direct target of miR-497-5p in HCC cells. (A) KEGG analysis of the involvement of target genes of miR-497-5p in Hippo signaling pathway by using a mirPath v.3 prediction tool. (B) qRT-PCR analysis of the expression levels of miR-497-5p in various HCC cell lines and normal liver tissues. (C, D) qRT-PCR and Western blot analysis of the effects of miR-497-5p mimic or inhibitor on BIRC5 expression in SMMC-7721 or LO2 cell line. (E, F) The binding sites of miR-497-5p with WT or Mut BIRC5 3’UTR, and the luciferase activity of WT or Mut BIRC5 3’UTR after transfection with miR-497-5p mimic or inhibitor in SMMC-7721 or LO2 cell line. Data are the means ± SEM of three experiments. *P < 0.05, **P < 0.01.
BIRC5 Counteracted the Anti-Oncogenic Role of miR-497-5p in HCC
The functional interaction between miR-497-5p and BIRC5 was further assessed in HCC cells. The knockdown or overexpression efficiency of BIRC5 or si-BIRC5 in HepG2 or LO2 cell lines was determined (Fig. 4A). Cell proliferation, colony formation number and invasive potential were enhanced by BIRC5 or miR-497-5p inhibitor, but weakened by BIRC5 silencing or miR-497-5p in SMMC-7721 or LO2 cell lines (Fig. 4B–F). The antitumor activities induced by miR-497-5p were reversed by BIRC5 overexpression in SMMC-7721 cells, but miR-497-5p inhibitor-induced tumor proliferation was attenuated by BIRC5 knockdown (Fig. 4B–F).
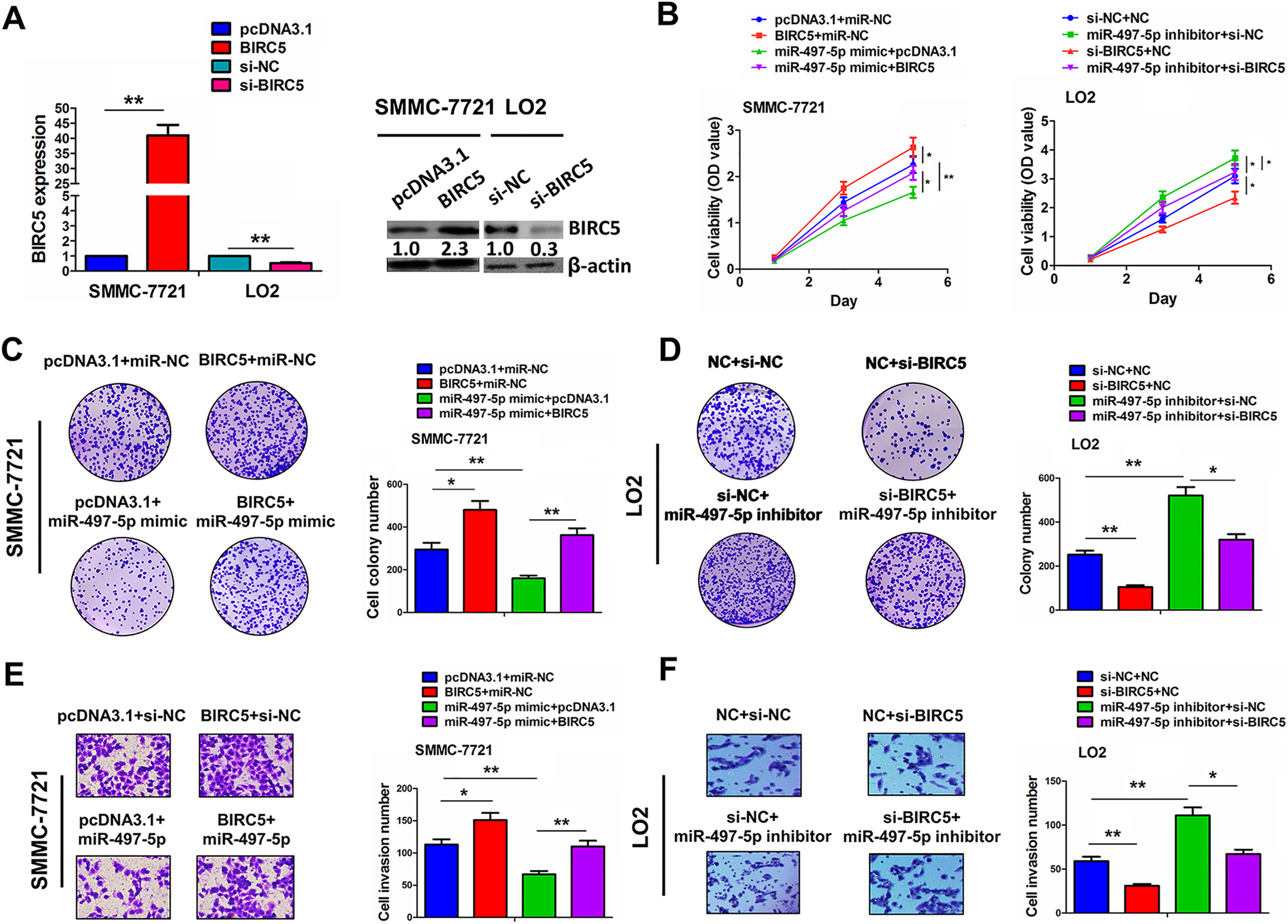
MiR-497-5p inhibited HCC cell growth and invasion by targeting BIRC5. (A) qRT-PCR and Western blot analysis of the transfection efficiency of BIRC5 plasmid or si-BIRC5 in SMMC-7721 or LO2 cell line. (B) MTT, (C, D) colony formation and (E, F) Transwell assay were conducted for assessing the cell viability, colony formation number, and invasive potential after the co-transfection with miR-497-5p mimic and BIRC5 plasmid in SMMC-7721 cells or miR-497-5p inhibitor and si-BIRC5 in LO2 cells. Data are the means ± SEM of three experiments. *P < 0.05; **P < 0.01.
A Novel circANKRD52 Was Identified in HCC
To further screen the circRNAs that may bind with miR-497-5p, we utilized a starBaseV2.0 prediction tool to identify 20 circRNAs that can bind with miR-497-5p (Supplemental Table S8), of which hsa_circ_0026939 has the most binding sites with miR-497-5p. The circRNA hsa_circ_0026939 is originated and formed from exon 14, 19 regions within ankyrin repeat domain 52 (ANKRD52) locus by back-splicing, and termed as circANKRD52 (Fig. 5A). We found that circANKRD52 yielded a resistance to RNase R treatment in contrast with linear ANKRD52 in SMMC-7721 and LO2 cell lines (Fig. 5B). After treatment with Actinomycin D, the half-life (more than 24 h) of circANKRD52 was longer than that (less than 6 h) of linear ANKRD52, indicating that circANKRD52 was stable in SMMC-7721 and LO2 cells (Fig. 5C). We found that circANKRD52 was substantially localized in the cytoplasm of HCC cells and tissues in contrast with ANKRD52 (Fig. 5D, E).
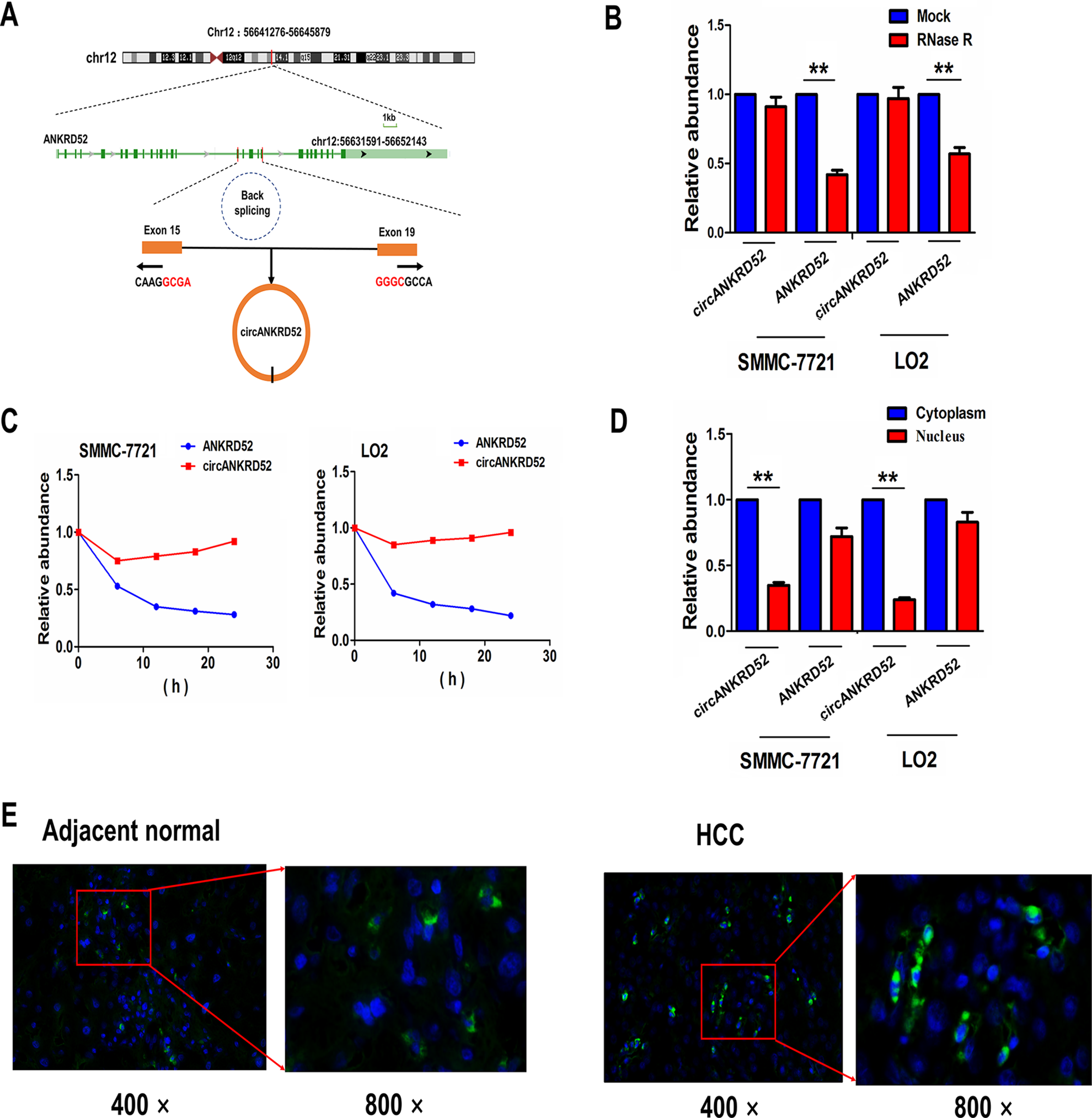
Identification of a novel circANKRD52 in HCC cells. (A) The genomic loci of circANKRD52 and its origin from the back-splicing of linear gene ANKRD52. (B) qRT-PCR analysis of the expression levels of circANKRD52 and ANKRD52 after exposure to RNase R in SMMC-7721 or LO2 cell line. (C) qRT-PCR analysis of the expression levels of circANKRD52 and ANKRD52 after exposure to Actinomycin D at indicated time points in SMMC-7721 or LO2 cell line. (D) qRT-PCR analysis of the enrichment levels of cytoplasmic and nuclear circANKRD52 and ANKRD52 in SMMC-7721 or LO2 cell line. (E) FISH analysis of the cellular localization of circANKRD52 in HCC and adjacent normal tissues. The nuclei were stained with DAPI for blue color, and circANKRD52 in the cytoplasm was stained for green color. Data are the means ± SEM of three experiments. **P < 0.01.
CircANKRD52 Facilitated HCC Cell Growth and Invasion
CircANKRD52 overexpression sequence or shRNA sequence against the back-splicing site of circANKRD52 was indicated in Fig. 6A. The knocking or overexpressing efficiency of circANKRD52 or sh-circANKRD52 in SMMC-7721 or LO2 cell line was decided (Fig. 6B). Further observations indicated that overexpression of circANKRD52 enhanced cell growth and invasiveness in SMMC-7721 cells, while knockdown of circANKRD52 reverse these effects in LO2 cell line (Fig. 6C–E). We also constructed a circANKRD52 or NC-stably transfected SMMC-7721, and the tumors in circANKRD52 group displayed a faster growth rate than the NC (Fig. 6F), and the tumor volume and weight in circANKRD52-transfected group were larger than those in NC transfected group (P = 0.0011, P = 0.0001; Fig. 6G).
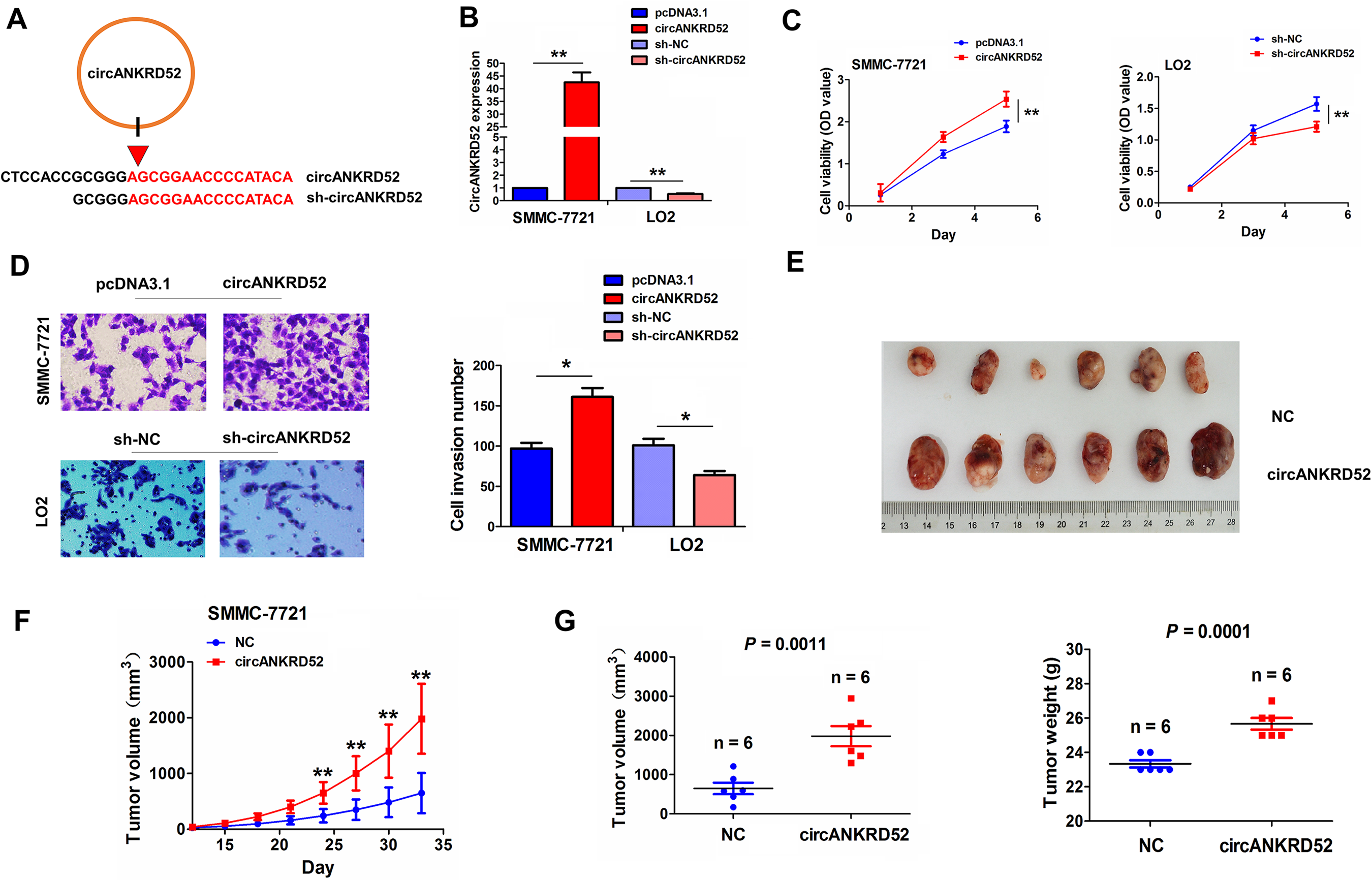
CircANKRD52 promoted HCC cell proliferation and invasion in vitro and in vivo. (A) Schematic representation of circANKRD52 overexpression sequences and its target sequences specific to the back-splicing junction of circANKRD52. (B) qRT-PCR analysis of the transfection efficiency of circANKRD52 plasmid or sh-circANKRD52 in SMMC-7721 or LO2 cell line. (C) MTT, (D) colony formation and (E) Transwell assays were used for determining the cell viability, colony formation number and invasive potential after the transfection with circANKRD52 plasmid in SMMC-7721 cells or sh-circANKRD52 in LO2 cells. (F) A growth curve analysis of the tumor growth in circANKRD52 or NC transfected SMMC-7721 group. (G) Comparison of the tumor size and weight of xenograft tumors induced by circANKRD52 or NC transfected SMMC-7721 cells. Data are the means ± SEM of three experiments. *P < 0.05; **P < 0.01.
CircANKRD52 Could Sponge miR-497-5p in HCC Cells
FISH showed that circANKRD52 exhibited a co-localization with miR-497-5p in LO2 cell line, and indicated an opposite relationship with miR-497-5p in tissue samples (r = - 0.315, P = 0.003; Fig. 7A). Overexpressing circANKRD52 decreased miR-497-5p expression in SMMC-7721, but raised by silencing circANKRD52 in LO2 (Fig. 7B), but miR-497-5p exerted no effect on circANKRD52 expression in SMMC-7721 and LO2 cells (Fig. S7). The luciferase reporter vector containing WT or Mut circANKRD52 3’UTR (Fig. S8) was pretreated with miR-497-5p in SMMC-7721 or LO2, indicating that the luciferase activity of WT circANKRD52 3’UTR was lowered by miR-497-5p in SMMC-7721, but elevated by miR-497-5p inhibitor (Fig. 7C). Moreover, we performed a RIP for Ago2 in liver cancer cells and assessed endogenous levels of circANKRD52 and miR-497-5p and found that circANKRD52 and miR-497-5p were mainly increased in Ago2 pellet (Fig. 7D). To further validate the direct interaction between circANKRD52 and miR-497-5p in HCC cells, we used a circANKRD52 probe to perform a RNA in vivo precipitation assay. CircANKRD52- probes were used to purify the RNA pulled down from the circANKRD52 and we found that the enrichment levels of circANKRD52 and miR-497-5p, examined by qPCR were increased as compared with the controls in SMMC-7721 and LO2 cells (Fig. 7E).
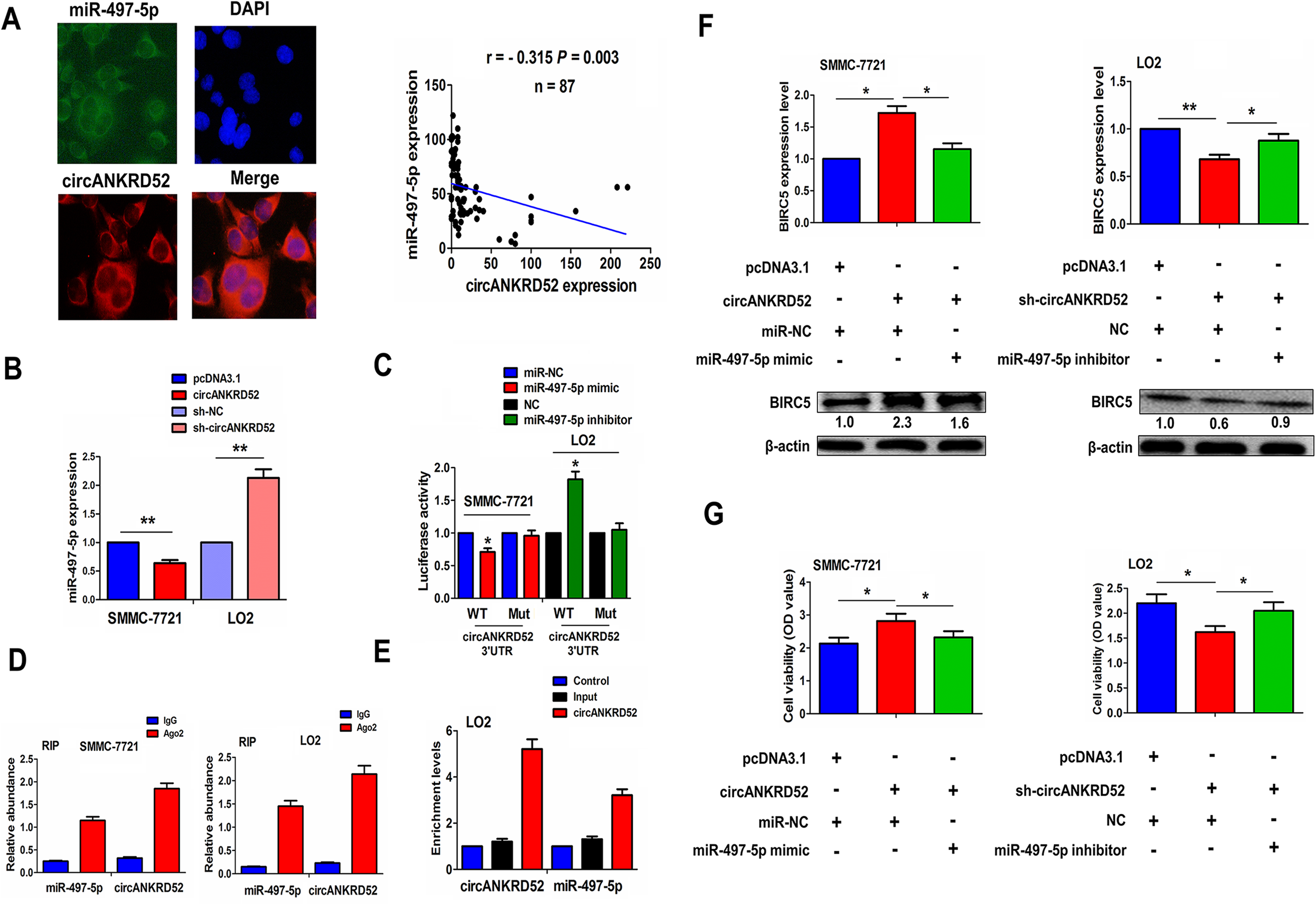
CircANKRD52 acted as a sponge of miR-497-5p in HCC cells. (A) FISH analysis of the colocalization of circANKRD52 with miR-497-5p in LO2 cells and their correlation in HCC tissues. (B) qRT-PCR analysis of the expression levels of miR-497-5p after transfection with circANKRD52 in SMMC-7721 cells or sh-ANKRD52 in LO2 cells. (C) Analysis of the luciferase activity of WT or Mut circANKRD52 3’UTR after transfection with miR-497-5p mimic in SMMC-7721 cells or miR-497-5p inhibitor in LO2 cells. (D) RIP assay of the amount of circANKRD52 and miR-497-5p in SMMC-7721 or LO2 cells by qRT-PCR analysis. (E) circRIP analysis of the purified RNA pulled down by a circANKRD52-specific probe and qRT-PCR analysis of the enrichment levels of circANKRD52 and miR-497-5p in LO2 cells. (F) Western blot analysis of the expression levels of BIRC5 after co-transfection with circANKRD52+miR-497-5p mimic in SMMC-7721 cells or sh-circANKRD52+miR-497-5p inhibitor in LO2 cells. (G) MTT analysis of cell proliferation after co-transfection with circANKRD52+miR-497-5p mimic in SMMC-7721 cells or sh-circANKRD52+miR-497-5p inhibitor in LO2 cells. Data are the means ± SEM of three experiments. *P < 0.05; **P < 0.01.
We further found that ectopic expression of circANKRD52 increased BIRC5 expression, but this result was counteracted by miR-497-5p; Likewise, knockdown of circANKRD52 decreased BIRC5 expression, but this result was reversed by miR-497-5p inhibitor (Fig. 7F). Additionally, the proliferation-promoting effect induced by circANKRD52 was attenuated by miR-497-5p in SMMC-7721, while the anti-proliferative effect caused by silencing circANKRD52 was reversed by miR-497-5p inhibitor in LO2 cells (Fig. 7G).
Discussion
Accumulating studies indicated that BIRC5 expression levels are increased in human tumors and display tumor invasiveness and worse prognosis 5 –8 . Herein, BIRC5 expression was elevated in HCC tissue samples, and its increased levels were linked to pathological stage, tumor size and distant metastasis. BIRC5 indicated poor prognosis and functioned as an oncogene in HCC. In coinciding with previous studies 26 , our results revealed that BIRC5 might be a predictor in HCC.
Subsequently, we found that increased BIRC5 expression was duo to a post transcriptional regulation of miRNAs in HCC. Previous studies revealed that BIRC5 expression can be negatively regulated by miR-542-3p in HCC 27 , miR-203 in breast cancer 13 and ovarian cancer 14 , miR-485-5p in breast cancer 28 , and miR-218 in colorectal cancer 29 . In the present study, BIRC5 was negatively related with miR-195-5p/-497-5p/-144-3p expression in HCC, and only miR-497-5p decreased levels showed an association with a worse survival in HCC. miR-497-5p levels were reduced in melanoma 30 , lung cancer 31 , and osteosarcoma 32 , suppresses the cell proliferation and induces cycle arrest and cell apoptosis 30 –32 . Herein, miR-497-5p levels were lowered in HCC and its low expression was linked to poor survival. Restored expression of miR-497-5p suppressed HCC cell viability and cell invasion, and these effects could be reversed by BIRC5. BIRC5 was found to be a target of miR-497-5p, and miR-497-5p might inhibit HCC cells by targeting BIRC5.
Accumulating data indicated that circRNAs function as oncogenes 18 –20 or anti-oncogenes in HCC 21,22 . A novel circANKRD52 was herein identified and executed the capability to sponge miR-497-5p. Previous findings indicated that ANKRD52 favors the melanoma metastases 33,34 . In addition, MALAT1 accelerates renal cell carcinoma by sponging miR-203 and upregulating BIRC5 35 and circRNA-miRNA-BIRC5 axis is associated with prognosis of gastric cancer 36 . Herein, we found that circANKRD52 produced an RNase R resistance and a stable expression in HCC cells and possessed a co-localization with miR-497-5p. Further investigations showed that circANKRD52 sponged miR-497-5p to upregulated BICR5, thereby contributing to HCC tumorigenesis (Fig. 8).
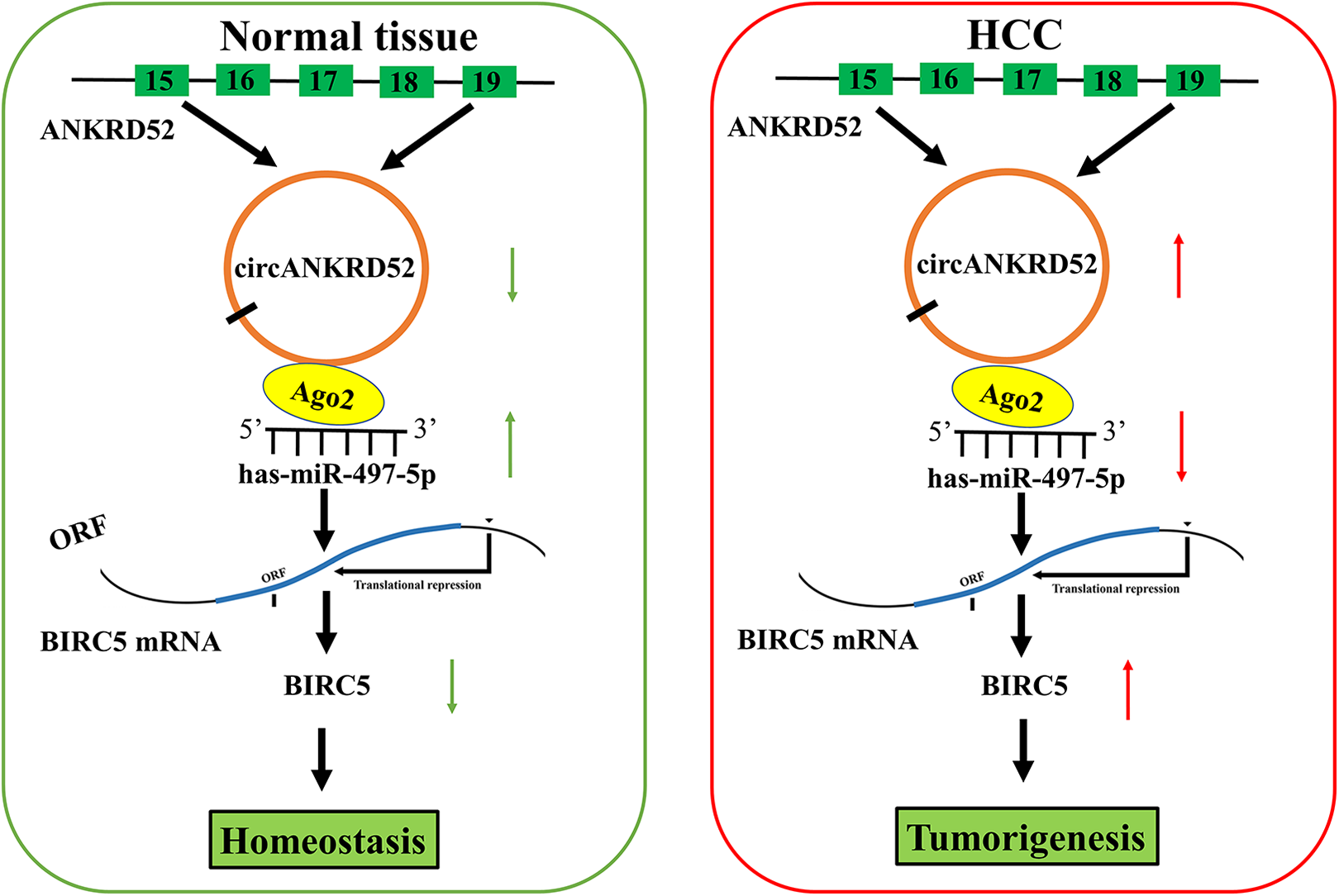
Schematic representation of the proposed mechanism of circANKRD52 in HCC cells. CircANKRD52 acted as a miR-497-5p sponge to upregulate BIRC5 expression, contributing to the tumorigenesis of HCC.
Taken together, our findings demonstrated that the elevated levels of BIRC5 or decreased levels of miR-497-5p were linked to worse outcomes in HCC. Restoration of miR-497-5p inhibited HCC cell growth by targeting BIRC5. CircANKRD52 was identified to act by sponging miR-497-5p and upregulating BIRC5 expression, and might predict poor survival in early-stage HCC.
Supplemental Material
Supplemental Material, sj-docx-1-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-docx-1-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-png-1-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-png-1-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-1-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-1-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-2-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-2-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-3-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-3-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-4-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-4-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-5-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-5-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-6-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-6-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Supplemental Material
Supplemental Material, sj-tif-7-cll-10.1177_09636897211008874 - CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression
Supplemental Material, sj-tif-7-cll-10.1177_09636897211008874 for CircANKRD52 Promotes the Tumorigenesis of Hepatocellular Carcinoma by Sponging miR-497-5p and Upregulating BIRC5 Expression by Mingzhi Zhang, Xinxin Yan, Peihao Wen, Wenkun Bai and Qingyu Zhang in Cell Transplantation
Footnotes
Authors’ Contributions
QZ, PW, and WB designed the present experiments and MZ was responsible of drafting the paper. MZ and XY showed the equal contributions to this study. MZ and XY conducted the studies. PW conducted statistical analyses and WB collected the data. All the authors checked the final manuscript.
Mingzhi Zhang and Xinxin Yan, are contributed equally to this work.
Ethical Approval
The research protocol was approved by the Ethics Committee of General Hospital of Tianjin Medial University, Tianjin, China.
Statement of Human and Animal Rights
All procedures with human subjects in this study were conducted in accordance with the Human and Animal Ethics Committee of General Hospital of Tianjin Medial University, Tianjin, China.
Statement of Informed Consent
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Our work was supported by the grants from Joint Funds of the National Natural Science Foundation of China (No. U1504804).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
