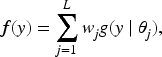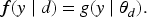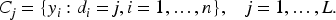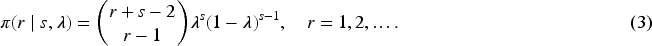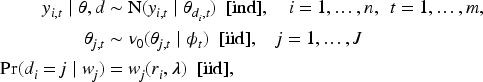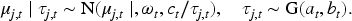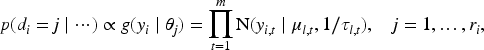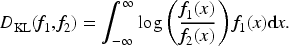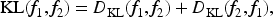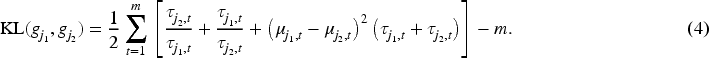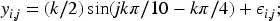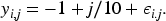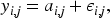Abstract
The identification of latent profile trajectories in longitudinal studies represents an important challenge for specialists since they could provide insights to better understand their problem of interest. The majority of the statistical methodologies for cluster analysis for longitudinal data are based on growth curve or mixed-effects models, and often incorporate covariates for a better adjustment. In particular, for Bayesian nonparametric methods, Dirichlet process mixture models are widely used together. We propose a clustering methodology for longitudinal data based on mixture models generated by a discrete random probability measure whose weights are decreasingly ordered by construction. Additionally, data is modeled without making use of covariates and assuming independence across time for individual measurements. Our approach also provides a straightforward procedure to merge some estimated groups, since it could happen that there are many of them, to be easily explained by experts. Our results suggest that, at least for a first analysis, this framework is enough to effectively detect groups in the data; further exploration of each group could incorporate extra information. We apply our methodology for detecting adiposity trajectories in Mexican children in a secondary analysis of the “Prenatal Omega-3 fatty acid Supplementation and Child Growth and Development” study (POSGRAD) cohort.
Keywords
Introduction
There are a number of areas with working scenarios, for example, epidemiological, medical, biological or psychological studies, leading to individual growth trajectories or longitudinal profiles.1–4 Researchers are often interested in understanding their dynamics, that is, the development or tendency over time of the resulting profiles. Additionally, there could be subsets of individuals whose growth profiles are significantly different from the overall estimates. Therefore, it might be of interest to find patterns for these heterogeneous longitudinal data sets.
One can handle this situation from a statistical learning perspective as an unsupervised classification problem, where the aim is to identify classes of trajectories. The estimated classes allow to disaggregate a larger heterogeneous population into homogeneous subpopulations, which would pinpoint meaningful groups of more similar individual profiles. These groups could, in turn, be used as predictions to either follow the change in time of the relevant variable or to propose within the latent classes other predictive or causal models.
Moreover, this approach could potentially contribute to improve the analysis of information originated from longitudinal study designs, such as cohort studies, clinical trials or interventional studies, where the latter would have at least two measurements in time for individuals. In these schemes, the aim is often to study possible changes in patterns in order to understand how particular events develop and evolve through time in different areas, 5 for example, in nutrition, health and social behavior, amongst others, enabling the identification of tendencies, changes and profile behavior in general, and improving the understanding of such events, for instance in the life course of health-disease process. Despite the striking value of such information, still only a few longitudinal studies consider in their analyses the identification of profile patterns or trajectories.6,7
The statistical modeling of longitudinal data has often been performed through growth curve models3,4,8–11 or mixed-effects models2,12 and a strong emphasis is put on the estimation of the mean function. To capture the phenomenon-driven dynamics, a very flexible mean function should be considered, therefore, there have been many efforts to generate ad-hoc proposals given a particular data behavior.13–15 Additionally, it is also common to incorporate covariates for the estimation, either for the mean or covariance functions. 16 Similarly, other models treat observations as time dependent, leading to, for example, autoregressive mean functionals.17–19 On a different perspective, longitudinal data has been also handled by computational methods.20–22
In these methodologies, the identification of classes of trajectories helps to reveal common patterns in subgroups of the data, which is achieved through the detection of the underlying clusters conforming the population. Cluster analysis for mixed-effects models is performed via latent class models, where random-effects variables or the mean functional parameters play the role of classification variables.14,23 Classical6,18,24,25 and Bayesian approaches have been developed for this purpose. For the latter, the usage of random probability measures (RPMs) is a common tool,15–17,19,23,26,27 in particular the Dirichlet process. 28 One of the advantages of this approach is that the number of latent classes is inferred from the data; furthermore, these models can be expressed as mixture models, a very well known method for cluster analysis, which also allows to characterize groups probabilistically. However, in some proposals, there are parametric assumptions, hence they need to deal with estimation challenges such as label switching, local maxima trapping, and so on.
In this work, we consider a continuous outcome where the main interest is to identify patterns and classify the individual profiles accordingly. Unlike common methodologies for modeling longitudinal responses, we do not impose elaborated dependency structures to describe individual data, neither consider covariates. Our results suggest that we can prescind from them, at least, in a first stage when the main interest is clustering. Adopting a Bayesian nonparametric approach, patterns in the observed profiles are inferred based on a mixture model whose underlying RPM has the characteristic of having weights decreasingly ordered by construction. The first example of decreasing weights RPMs is the geometric process, 29 which has been already applied to different estimations procedures (see, e.g. Fuentes-García et al.,30,31 and Gutiérrez et al. 32 ), and has also been extended to allow a more flexible weight structure. 33 An advantage of this type of processes, when compared with the Dirichlet process, is that the order in the weights serves as a constraint diminishing non identifiability issues. Also, these processes seem to have a faster convergence under the density estimation context, c.f. Fuentes-García et al. 30 and De Blasi et al., 33 in the sense that less iterations are required to recover the mixing components.
To the best of our knowledge, there is no literature on clustering based on decreasing weights RPMs. This could be due to the fact that the geometric process tends to use a large number of components; however, the generalization provided by De Blasi et al. 33 overcomes this issue, and we wish to test it for the specific task of clustering. Additionally, since it might happen that the number of estimated classes is large to be interpreted by the expert users, we also present a straightforward procedure to fuse some groups by taking into account their profile estimated densities.
One motivation behind the proposed methodology is the better understanding of childhood obesity in developing countries, like Mexico, which would support strategies contributing its prevention. With this in mind, we consider a secondary data analysis for
Model framework
The detection of patterns in a given data set is the main goal of cluster analysis. These patterns aim to partition the data such that items gathered together, into a cluster or group, share some characteristics, that are similar among them, and at the same time, items in different clusters are different. Due to its generality, cluster analysis has received great attention in diverse fields, and as a consequence, we can find a vast amount of literature. Perhaps, mixture modeling is the most common methodology to perform cluster analysis. Mathematically, a mixture density
The choice of the value for the number of components,
Besides the Dirichlet process, there exist other RPMs widely used in Bayesian nonparametrics, for example, the two parameter Poisson–Dirichlet process,
35
generalized Gamma process,
36
which are particular cases of Gibbs-type priors (see, e.g. De Blasi et al.
37
), and others exhibiting a more complex structure, like the one presented by Gil-Leyva and Mena.
38
The election of the RPM determines, among other things, the structure of the mixing weights
Our selection uses an infinite-component mixture model whose mixing weights are ordered almost surely by construction, meaning that
The mixture model we will use is, therefore, the following
Observations are modeled through the densities
In the literature, some dependency structure is usually imposed within each observation, for example, a linear model or stochastic process. However, we follow a simpler approach since we think it is enough when the primary task is clustering. Therefore, we use an
Posterior sampling scheme
Augmenting the mixture model in Equation (1) by means of a membership variable,
Posterior samples can be obtained from a Gibbs sampler. The mixing weights are updated through their latent variables
Furthermore, the samples of these membership variables allow to infer the clustering structure underlying the data. As explained at the beginning of this section, a clustering is a partition
For every sampled clustering
Merging clusters
Although the posterior modal partition
A very widely used distance-based approach for cluster analysis is the hierarchical clustering method. In the literature, there are different measures that can be used to determine the clustering structure evolution, and the experimenter selects one by fixing a cut-off value. Some of these measures are defined in terms of the distance between pairs of data points, and others incorporate a model-based approach. In the latter, each group is modeled by a particular probability distribution, and the cluster hierarchies are built according to the distance between pairs of distributions (see, e.g. Heller and Ghahramani 39 ).
Following this idea, based on a posterior estimate
For the particular mixture model detailed before, each mixture component has an
We illustrate the performance of our methodology in this section using simulated data. Qin
40
presents a method for clustering gene expression profiles, and tests it using a simulated data set which can be applied in our context. Let us define five longitudinal profiles through the following functions: A periodic function given by
A monotone increasing function
A constant function
Thus, the data set we use will be conformed by sampling
The Gibbs sampler is run using the following setup. A sample of size
In Figure 1, we only present the results for the case
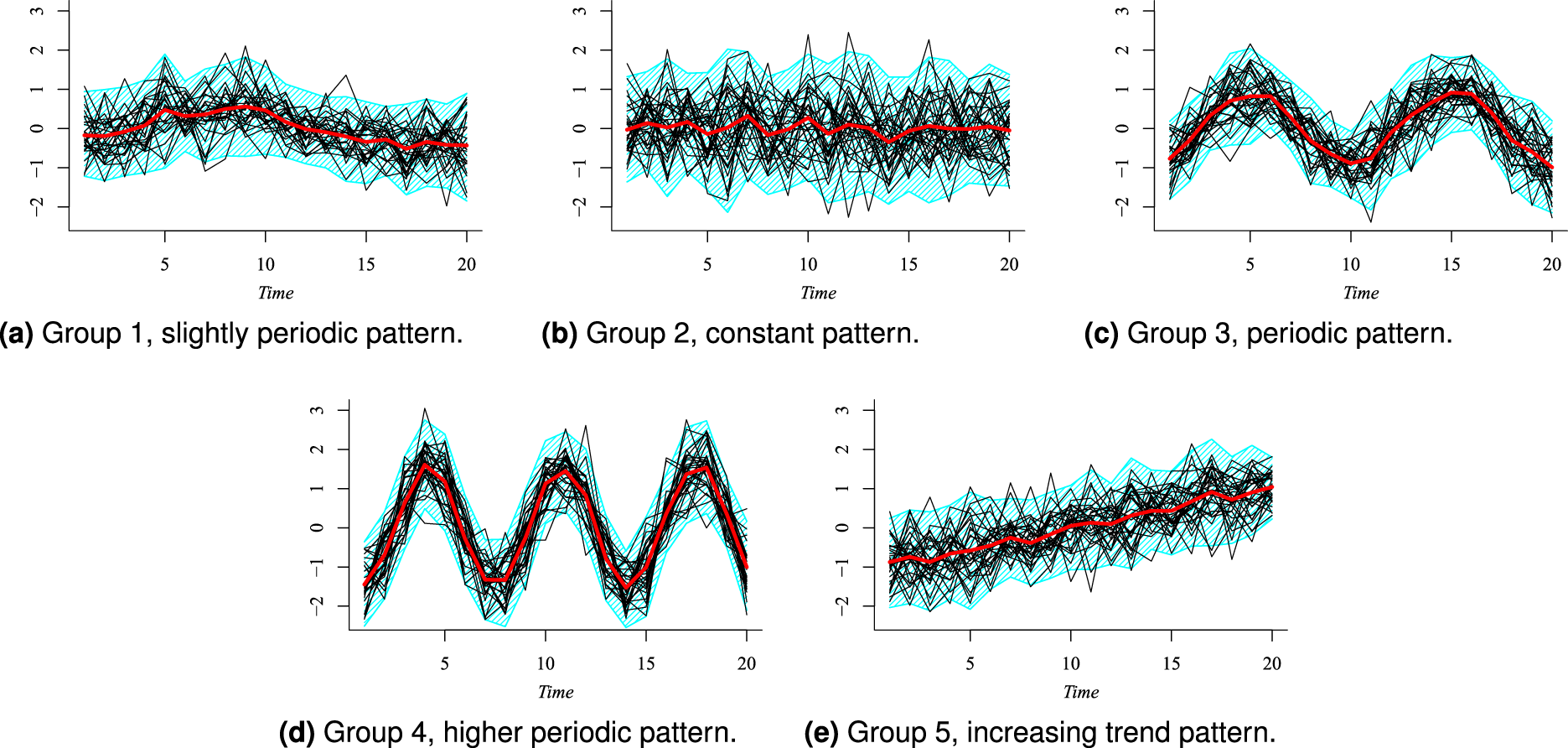
Posterior modal clustering for the simulated data, using
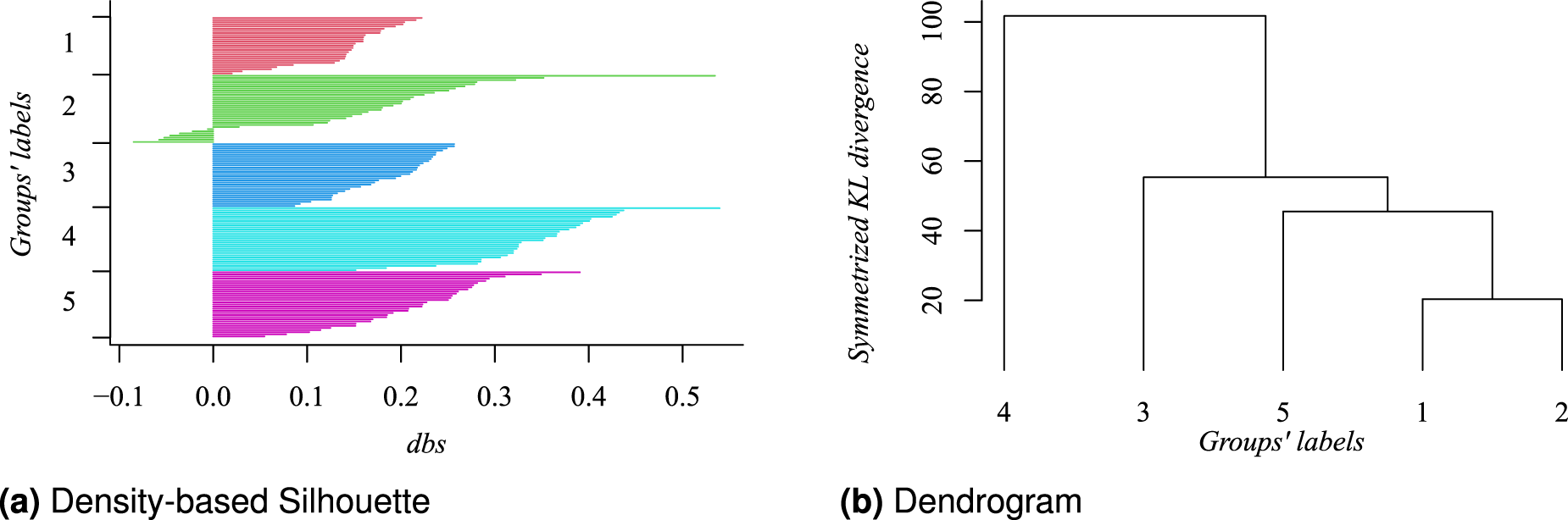
(a) Density-based Silhouette and (b) resulting dendrogram, using the symmetrized KL divergence, for the estimated clustering of the simulated data. For both panels, groups’ labels correspond to those of Figure 1. The negative values in the Silhouette plot, for Group 2, indicate that their associated observations would be better allocated in some other group. On the other hand, the dendrogram suggests that whether two groups should be merged, Group 1 and 2 would be the firsts candidates.
The density-based Silhouette (DBS) information, a modification of the Silhouette information 42 for model-based clustering procedures, is a method to evaluate the quality of a particular clustering given a data set. This information is computed for each observation, and a large value indicates an agreement with the assigned group; small or negative values are indicative that such an observation could be better placed in another group.
The results in Figures 1 and 2 show that our method is able to recover the five groups, and their mean profile is close to their corresponding generating data function. The DBS summary shows, in general, a good level of agreement with the estimated groups. As expected, the estimated groups labeled as 1 and 2 seem to be the more problematic; they correspond to the periodic function with
Childhood obesity constitutes a worldwide public health problem.
43
It is crucial to study the trajectories of adiposity from an early age, it has been identified that adiposity is closely related to what happens in critical periods of development, such as the “first 1000 days” of life, and also with lifestyles during childhood.
44
Moreover, it has been shown that the patterns of change in adiposity can be heterogeneous over time and vary between children.
45
There are studies attempting to understand this heterogeneity in order to identify the patterns of change that occur naturally in populations, resulting in the identification of factors that influence an early onset and development of obesity as well as early metabolic disorders. However, most of the existing evidence comes from high-income countries where socioeconomic, cultural and nutritional conditions are different from those of middle- and low-income countries. Even more, in the latter, there are few longitudinal studies considering this kind of analysis from a longitudinal profile perspective.
7
There is a need to conduct statistical studies allowing to identify groups of individuals at higher risk of obesity, as well as their potentially associated determinants and critical windows during pregnancy, infancy and childhood, in order to guide interventions and strategies that contribute to the prevention and containment of obesity in the population. Given this, the objective of the study was to identify groups of adiposity trajectories using measurements of the body mass index (BMI)
In this longitudinal study, a secondary analysis performed to data in the POSGRAD birth cohort,
34
the BMI
Children with BMI
Therefore, the variable of interest
The posterior modal partition
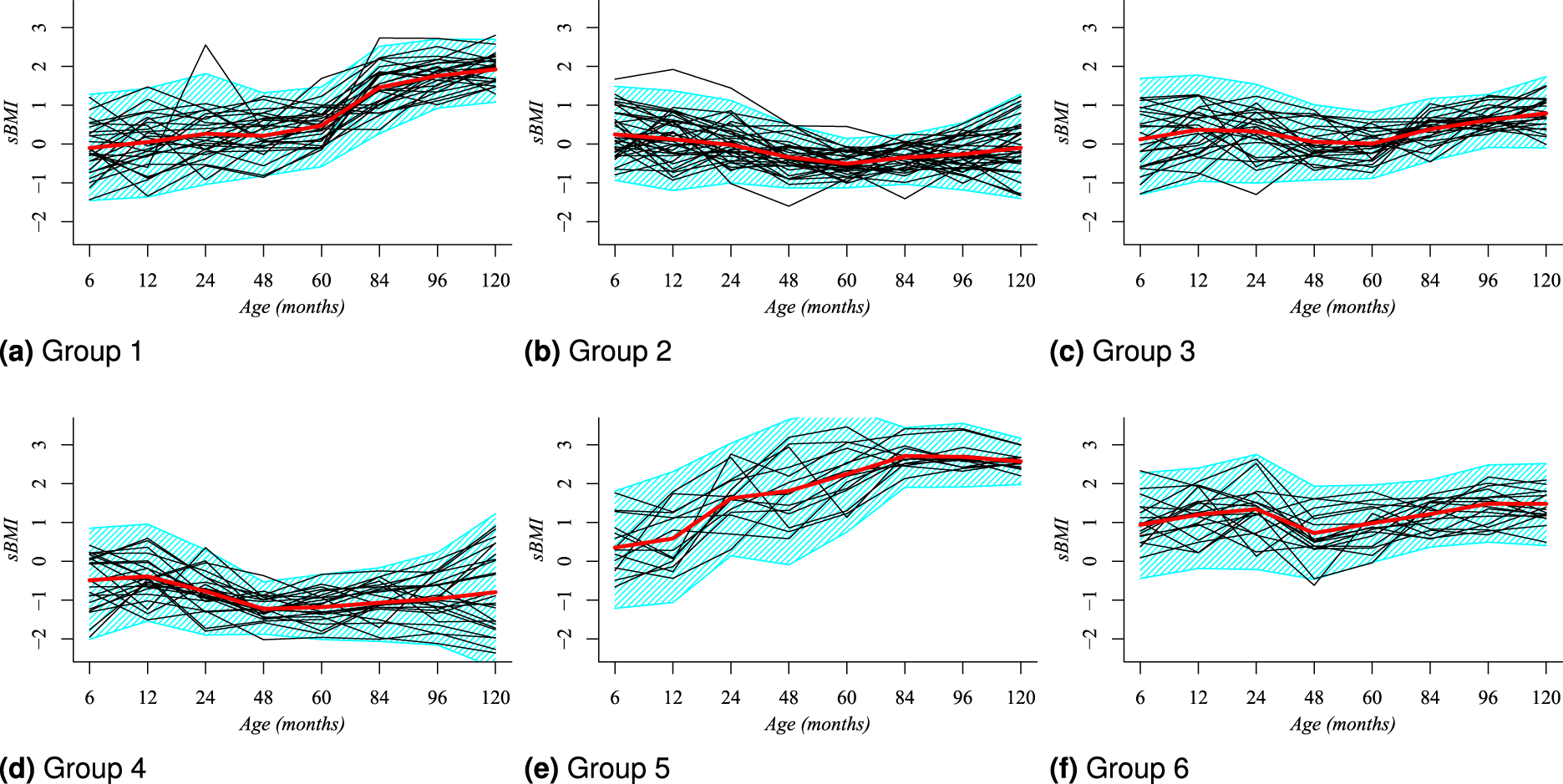
Posterior modal clustering,
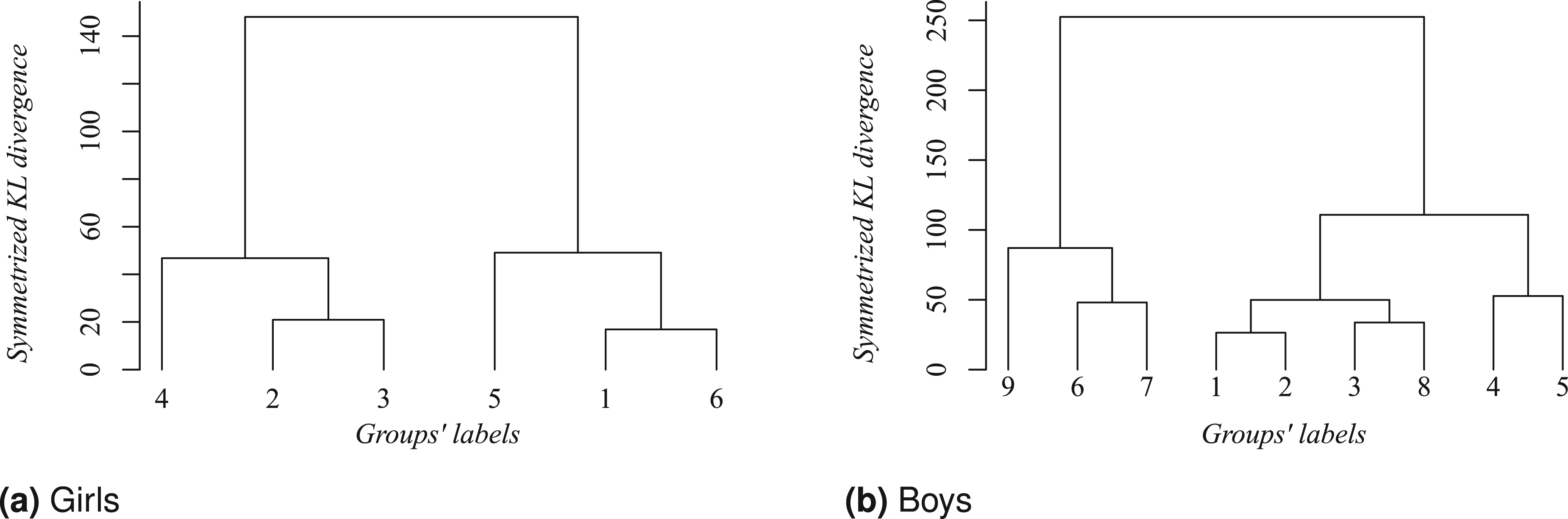
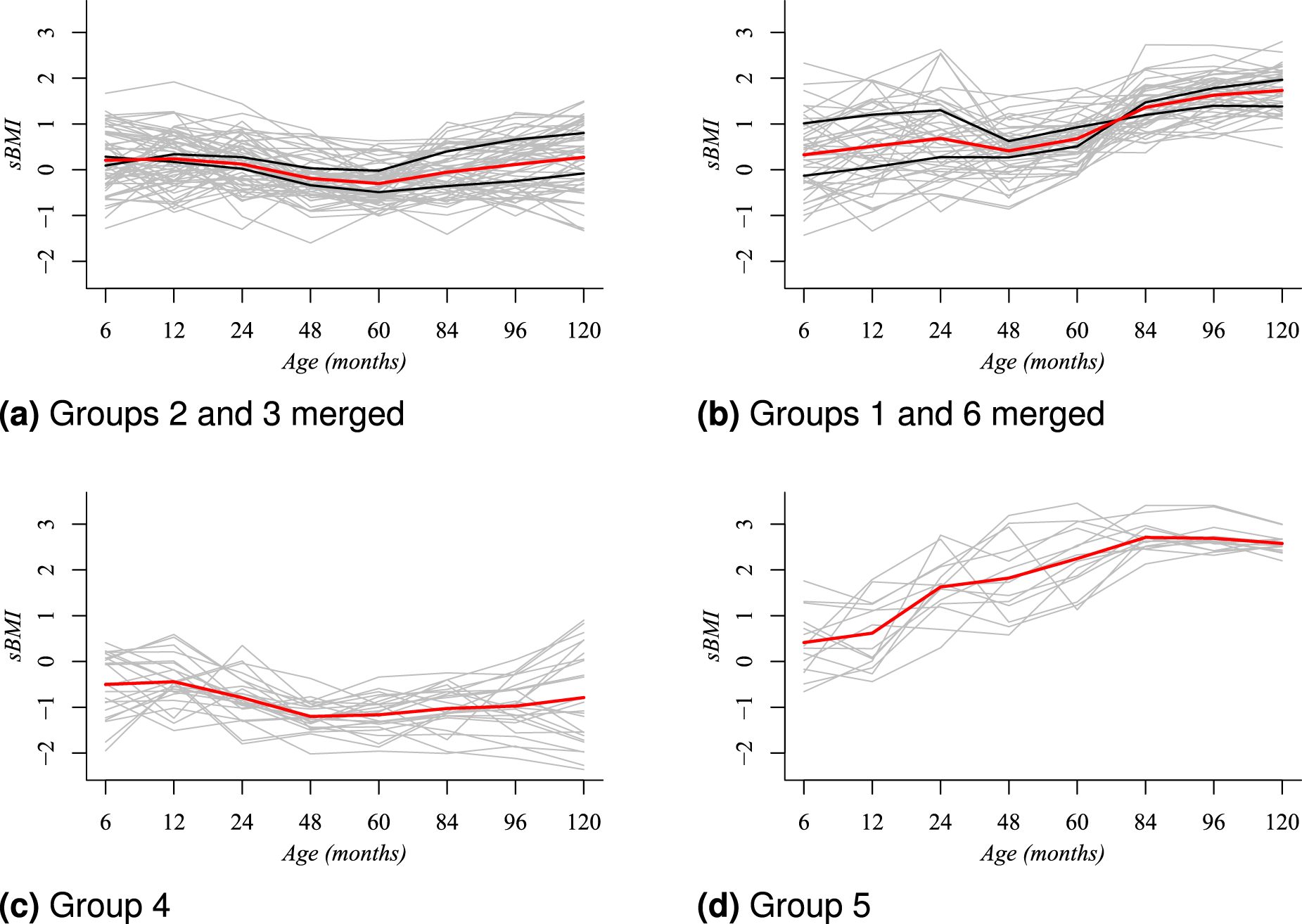
Simplified clustering structure for girls based on the posterior modal partition
Our result in Figure 5 identifies that girls in Panel 5(a), from clusters labeled as 2 and 3, who started at six months with a BMI
In the case of boys, the posterior modal partition
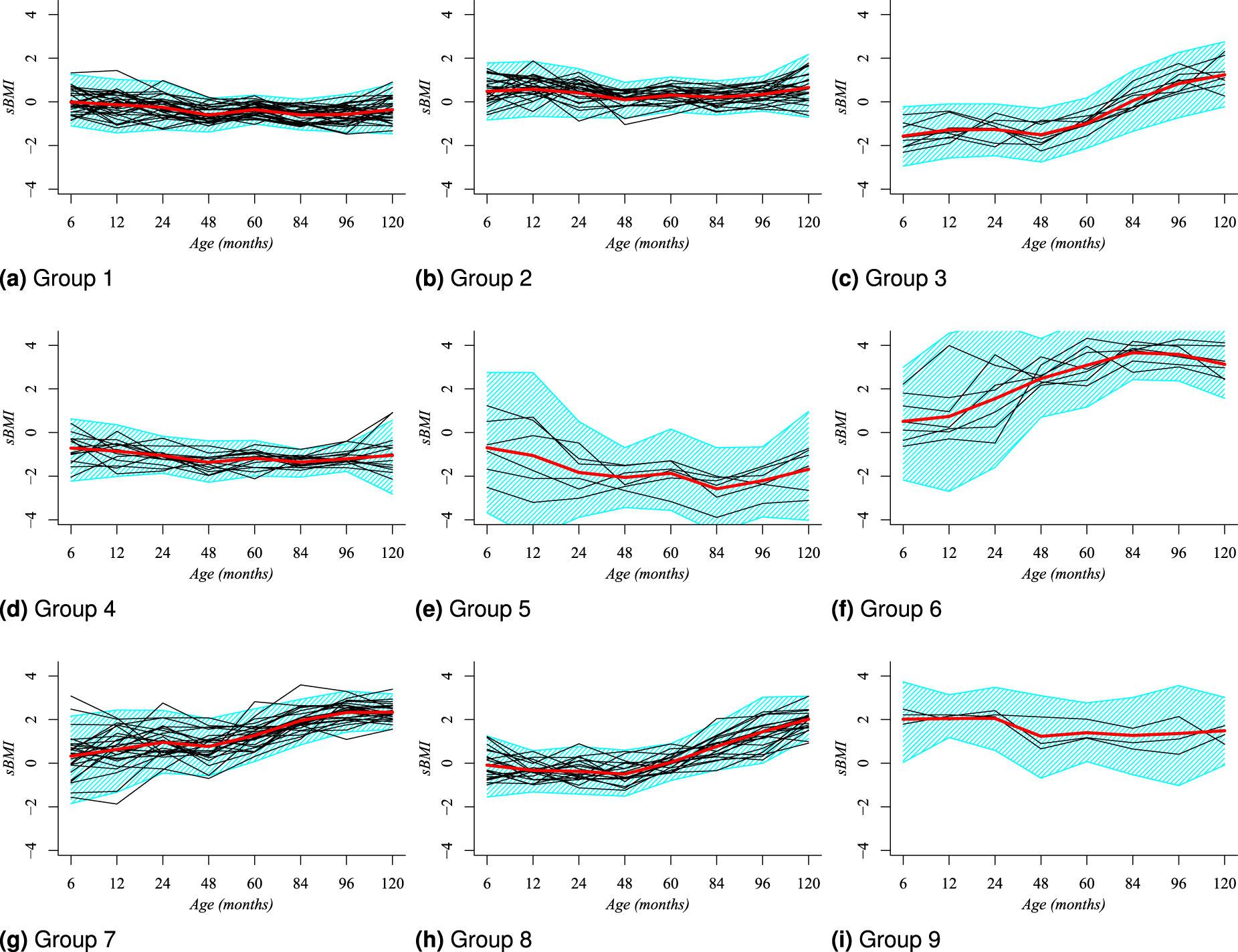
Posterior modal clustering,
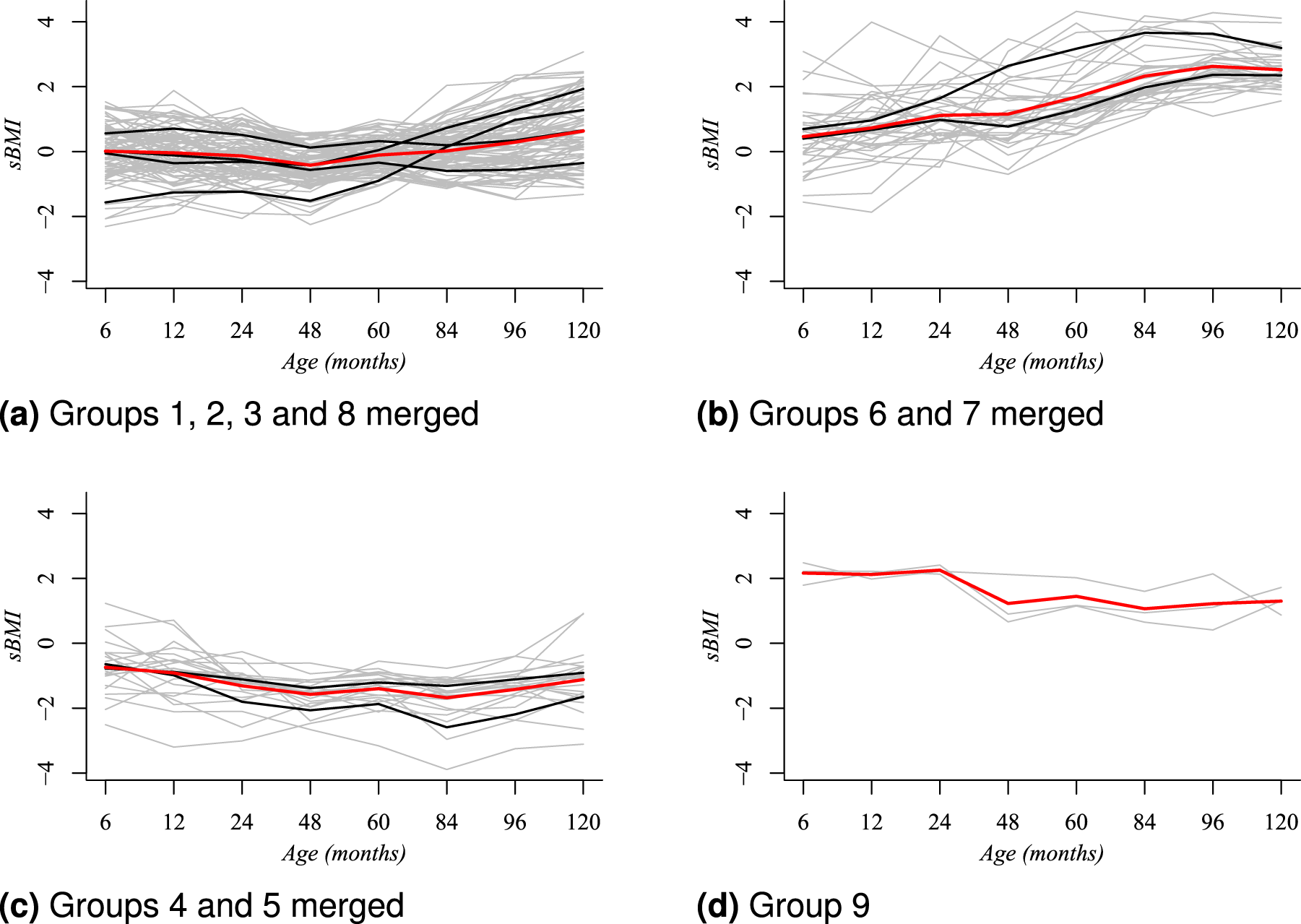
Simplified clustering structure for boys based on the posterior modal partition
Cluster analysis is an extremely common task in applied disciplines. It allows to discover patterns which could help to explain the heterogeneity in some population of interest. We focus in scenarios where the data of interest come from longitudinal studies, often appearing in cohort studies, and have presented a fully Bayesian nonparametric model for identifying profile patterns. Our approach diverges from most frameworks where the Dirichlet process is used, and adopts a class of RPMs whose realizations are such that their weights are decreasingly ordered, producing simple yet effective mixture models for clustering.
Decreasing weight RPMs have been successfully applied for density estimation (cf. De Blasi et al. 33 ), as well as for other estimation problems, showing a competitive performance, also when compared with more standard nonparametric models like the Dirichlet process. As far as we know, the present work is pioneering in applying this class of RPMs for clustering, also obtaining good results. Moreover, and at least in real data applications, when the estimated clustering contains many groups, some of them can be merged with the help the additional information provided by the methodology; in our case, we utilized the estimated profile densities.
With respect to the modeling of longitudinal data, we also considered an uncomplicated approach avoiding the usage of covariates and any dependency structure for describing the evolution in the measurements in the same individual. The appealing features of decreasing weight RPMs and the simple data model work well together, outperforming more common approaches. As the results in the adiposity application suggest, our approach is able to identify population with differentiated patterns, particularly allowing to pinpoint the higher risk groups. Once identified, it would allow to design specific studies to detect potential factors and events associated with this groups.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802251414594 - Supplemental material for Cluster analysis for longitudinal data and its application in the detection of adiposity trajectories
Supplemental material, sj-pdf-1-smm-10.1177_09622802251414594 for Cluster analysis for longitudinal data and its application in the detection of adiposity trajectories by Asael Fabian Martínez, Ivonne Ramírez-Silva and Ruth Fuentes-García in Statistical Methods in Medical Research
Footnotes
Funding
The authors recieved no financial support for the authorship and/or publication of this article: Asael Fabian Martínez was supported by PEPADI project 12601018. Ruth Fuentes-García would like to acknowledge the support of PAPIIT IN100823, UNAM.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.