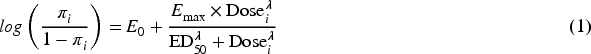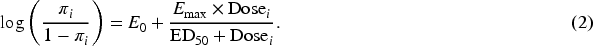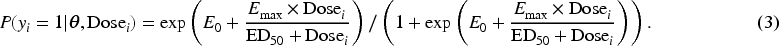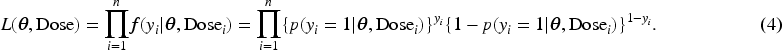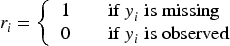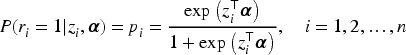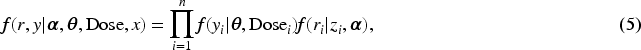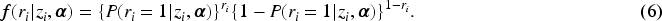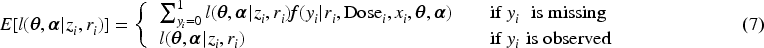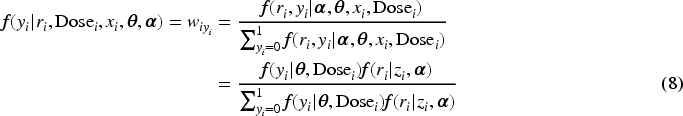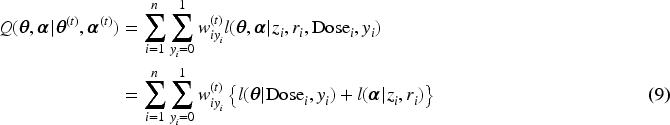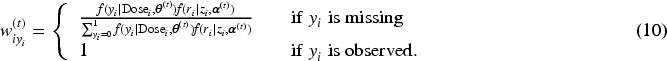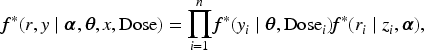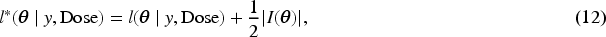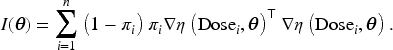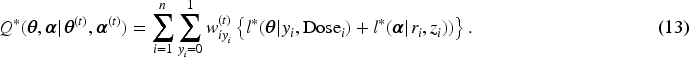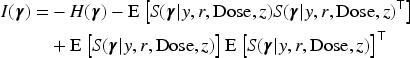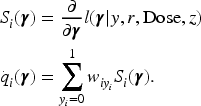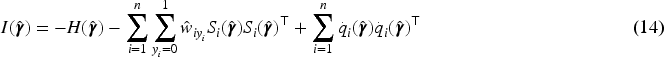Abstract
The Binary Emax model is widely employed in dose–response analysis during drug development, where missing data often pose significant challenges. Addressing nonignorable missing binary responses—where the likelihood of missing data is related to unobserved outcomes—is particularly important, yet existing methods often lead to biased estimates. This issue is compounded when using the regulatory-recommended ‘‘imputing as treatment failure’’ approach, known as non-responder imputation (NRI). Moreover, the problem of separation, where a predictor perfectly distinguishes between outcome classes, can further complicate likelihood maximization. In this paper, we introduce a penalized likelihood-based method that integrates a modified expectation-maximization (EM) algorithm in the spirit of Ibrahim and Lipsitz to effectively manage both nonignorable missing data and separation issues. Our approach applies a noninformative Jeffreys’ prior to the likelihood, reducing bias in parameter estimation. Simulation studies demonstrate that our method outperforms existing methods, such as NRI, and the superiority is further supported by its application to data from a Phase II clinical trial. Additionally, we have developed an R package, ememax (https://github.com/Celaeno1017/ememax), to facilitate the implementation of the proposed method.
Introduction
The dose–response relationship is a fundamental aspect of research in various applied statistics fields, particularly in clinical trials and bioinformatics. In real-world studies, incomplete data records are common due to reasons such as nonresponse to questionnaires, typographical errors, loss to follow-up, and data contamination. While missing data is often unavoidable, a common approach when the number of observations is large is to perform analysis using only complete records—a method known as complete case analysis (CC). However, in many scenarios, the application of CC yields biased estimates when analyzing dose–response relationships with binary outcomes.1,2 Moreover, small samples—common in randomized controlled trials—can exacerbate its variance and instability.
The challenge of missing data has been a significant area of research for decades, with numerous statistical methods developed to address it. Various missing data mechanisms, such as missing completely at random (MCAR) and missing at random (MAR), have been extensively studied, and corresponding methodologies have been proposed. It is well known that if the missing data mechanism is not appropriately modeled, parameter estimates may be biased. 3 While most related research has focused on scenarios where missing covariates and response values are assumed to be ignorable, limited attention has been given to scenarios with nonignorable missing response values, particularly within the context of modeling dose–response relationships. When the likelihood of missingness conditional on covariates depends on the unobserved values of the response, this is referred to as nonignorable missing data. 4 In this article, we explore the implications of nonignorable missing data within the context of a dose–response model with binary outcomes.
The sigmoid Emax model is commonly used in clinical trials to explore the binary dose–response relationship. Let
In clinical trials, it is common for binary outcomes to be missing for some patients receiving either an experimental or placebo dose. Given the typically small sample sizes in these studies, complete case analysis is often impractical. A standard practice, endorsed by the FDA and other regulatory agencies, is to impute all missing values as ”treatment failures.” This method, known as non-responder imputation (NRI), 8 is widely used but has well-documented limitations. NRI can introduce substantial bias into estimates and inferences, particularly when the missing data mechanism is nonignorable. 9
Another commonly used method for addressing missing data is multiple imputation (MI). This approach involves a two-stage process: first, generating several plausible imputed datasets based on assumed data distributions, and second, combining the results using a predefined pooling rule. 10 However, MI typically assumes that the missingness mechanism does not depend on the unobserved data, an assumption that is violated under nonignorable missingness. 11 Consequently, MI can also result in biased estimates and inferences. Some modified MI methods have been proposed to address missing not at random (MNAR) scenarios by model selection approach12,13 or pattern-mixture model approach, 14 but these methods often rely on assumptions of imputation model, especially the difference between distributions of respondent and non-respondent. This limits their flexibility in practical applications, where variables associated with missingness can vary widely. Moreover, the performance of MI heavily depends on the chosen pooling methods after generating and analyzing imputed datasets, which can be challenging to determine in practice. 15 Therefore, developing new approaches to fit the Emax model with incomplete data, particularly when the missing data mechanism is nonignorable, remains a critical issue in medical research.
In small or medium sample size settings for fitting binary outcome models, a common issue that can arise is the nonconvergence of estimates, a phenomenon known as “separation.” 16 Complete separation occurs when a single covariate or a linear combination of covariates perfectly predicts the outcome, leading to divergence in the estimation process. Even if complete separation does not occur, the presence of quasi-complete separation—where a subset of subjects’ responses is perfectly predicted—can still cause estimation challenges. 17 These situations, though not uncommon in biomedical datasets, are often overlooked. For example, current methods and packages that use likelihood estimation for dose–response or Emax models, such as the ClinDR package in R by Thomas 18 and the Dosefinding package by Bornkamp et al., 19 do not account for the issue of separation. Moreover, it is well known that estimation of Emax model under finite small sample sizes may encounter unstable convergence due to its strictly increasing assumption, which further complicated the model inference with binary response. 20
In the framework of generalized linear models, Heinze and Schemper 21 and Heinze 22 demonstrated that Firth’s method, 1 initially designed to reduce the bias of maximum likelihood estimates (MLE), can also effectively address the problem of separation. However, their work did not account for the presence of missing data. Extending this approach, Maiti and Pradhan, 23 and Maity, Pradhan and Das 24 applied Firth’s method to reduce bias and adjust for separation under a nonignorable missing data mechanism within the logistic regression framework.
When fitting the binary Emax model, two distinct but related separation issues can arise. The first type of separation occurs in the binary outcome y, even in the absence of any missing data. This happens when a predictor or a linear combination of predictors perfectly separates the outcome classes, leading to estimation challenges. The strictly increasing assumption of the Emax model can particularly make it prone to this type of separation, especially with small sample sizes. The second type of separation is induced by the missingness mechanism itself. In the presence of nonignorable missing data, the missing data pattern may be perfectly predicted by model variables. This further complicates the estimation process, as the missingness mechanism now introduces an additional source of separation.
In the field of dose–response relationships, no prior research has addressed the issue of nonignorable missing data within the Emax model framework. Additionally, there is no existing literature on bias reduction and separation in the Emax model, even with complete data. The key contribution of this paper is the integration of the Firth-type bias correction with the weighted EM algorithm proposed by Ibrahim and Lipsitz to address both types of separation issues within the binary Emax model framework. While previous work has dealt with the issue of nonignorable missing data and Firth-type bias correction in the context of logistic regression, our paper is the first to extend this approach to the more complex nonlinear Emax model.
The remainder of the article is organized as follows: Section 2 outlines the proposed approach and details the derivation of the estimation algorithm. Section 3 explores the simulation settings and presents the results. Section 4 discusses a real-world application, providing insights into the practical implementation of the proposed method. Finally, Section 5 offers a discussion of the findings and suggests potential directions for future research. Additionally, we have developed an R package, ememax (https://github.com/Celaeno1017/ememax), that implements our methods.
In this section, we present the proposed methodology in detail, beginning with the model formulation and assumptions that address these challenges.
Weighted EM procedure of Ibrahim and Lipsitz (IL)
Let
Firth
1
introduced a modification to the score function of the likelihood to reduce the bias of the maximum likelihood estimator (MLE) in small sample settings. Firth demonstrated that when the parameter in question is the canonical parameter of a full exponential family—such as in the logistic regression model for the missing indicator model with
Subsequently, we apply IL method with penalized joint log-likelihood, and the E-step of the EM becomes
Finally, the estimator can be obtained via any iterative procedures such as Newton–Raphson, Gauss–Newton, and Fisher scoring. Details about computational considerations, including initial value selection and computational time, are discussed in the Supplementary material.
We summarize all the steps of the IL/FIL algorithm as Algorithm 1.

Let
With the variance–covariance matrix estimator
To evaluate the performance of the proposed methodology, we conducted a thorough simulation study based on a general Phase-II dose–response clinical trial setting. We investigated the effect of sample size on the estimation process by considering sample sizes of n = 150, 250, 350, and 450. The total sample size was evenly distributed across five different treatment dose arms: Dose = (0, 7.5, 22.5, 75, 225), ensuring equal sample sizes in each treatment arm. The success rate for response in the placebo arm
The response variable
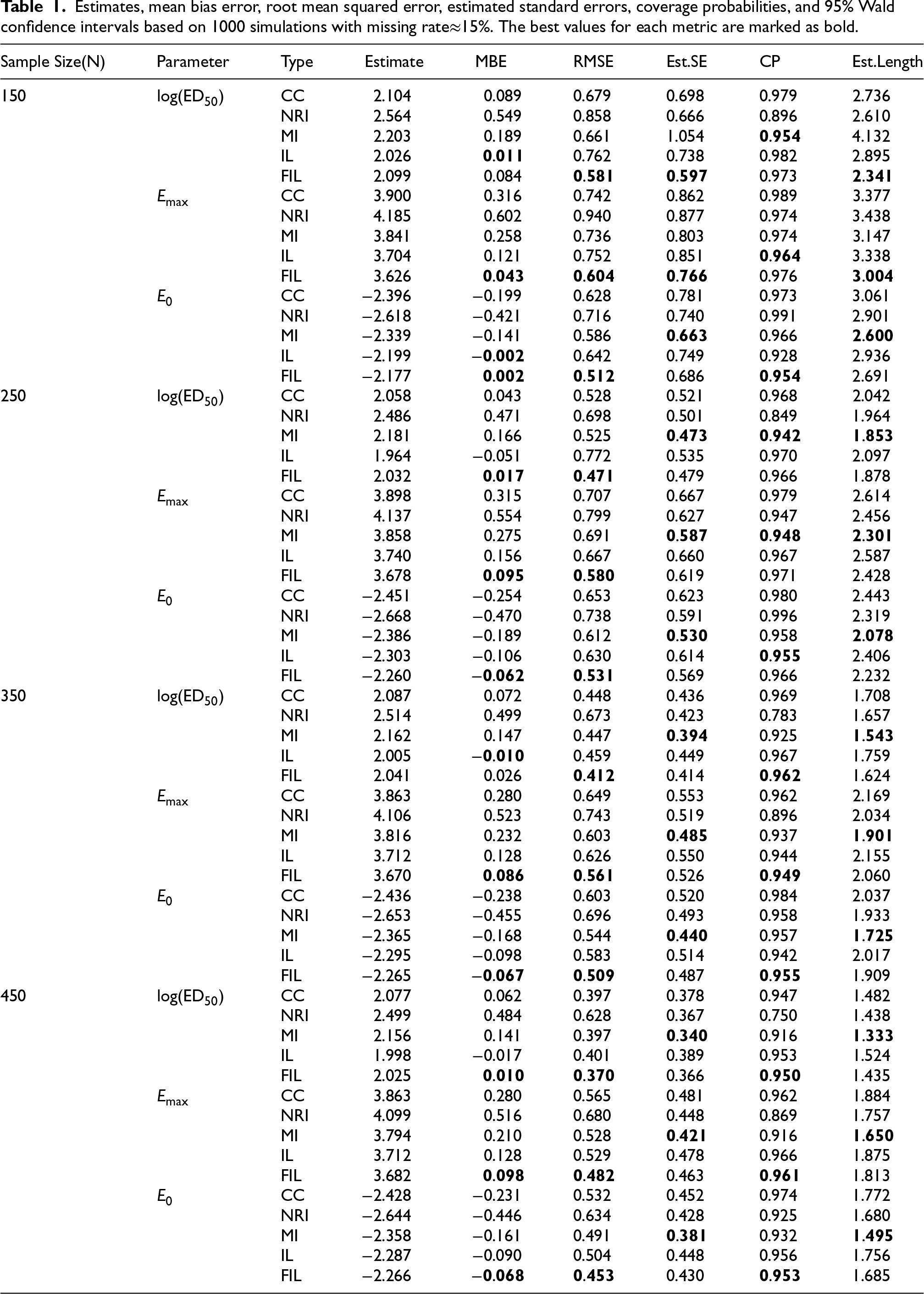
Table 1 summarizes the simulation results comparing five different estimation methods: Complete Case (CC), NRI, MI, Ibrahim–Lipsitz (IL), and Firth-Adjusted IL (FIL), based on
Estimates, mean bias error, root mean squared error, estimated standard errors, coverage probabilities, and 95% Wald confidence intervals based on 1000 simulations with missing rate
15%. The best values for each metric are marked as bold.
Estimates, mean bias error, root mean squared error, estimated standard errors, coverage probabilities, and 95% Wald confidence intervals based on 1000 simulations with missing rate

When the sample size was n=150, failures or unstable estimates occurred in 1.8% (CC), 1.0% (NRI), 1.0% (MI), and 0.8% (IL) of replicates, whereas FIL always converged and yielded stable estimates. For larger sample sizes, all methods converged without issues. As shown in Table 1, NRI estimators exhibit poor MBE and RMSE compared to other methods across all scenarios, leading to lower CP despite having lower mean estimated standard errors. For CC estimators, although the RMSE occasionally performs better than IL, and they are nearly unbiased for
The FIL method for bias correction is particularly valuable when the sample size is small and separation is encountered in the missingness pattern. Additional simulations were performed where missingness was more correlated with the dose, leading to severe separation in the placebo treatment arm (results reported in Table S1). In this scenario, all methods perform worse due to the systematic bias introduced by the loss of extreme scorers (success cases) in the placebo group. However, FIL and IL still outperform CC and NRI, with FIL constantly achieving better results. Notably, IL significantly underestimates the nominal coverage, whereas FIL maintains a reasonable coverage probability. Additionally, the standard errors estimated by FIL are smaller than those from IL, demonstrating the effectiveness of the Firth-type method in mitigating the effects of separation.
Figure 1 illustrates the distribution of point estimates for different methods, along with estimates based on the full data without missingness, across varying sample sizes. The true value of
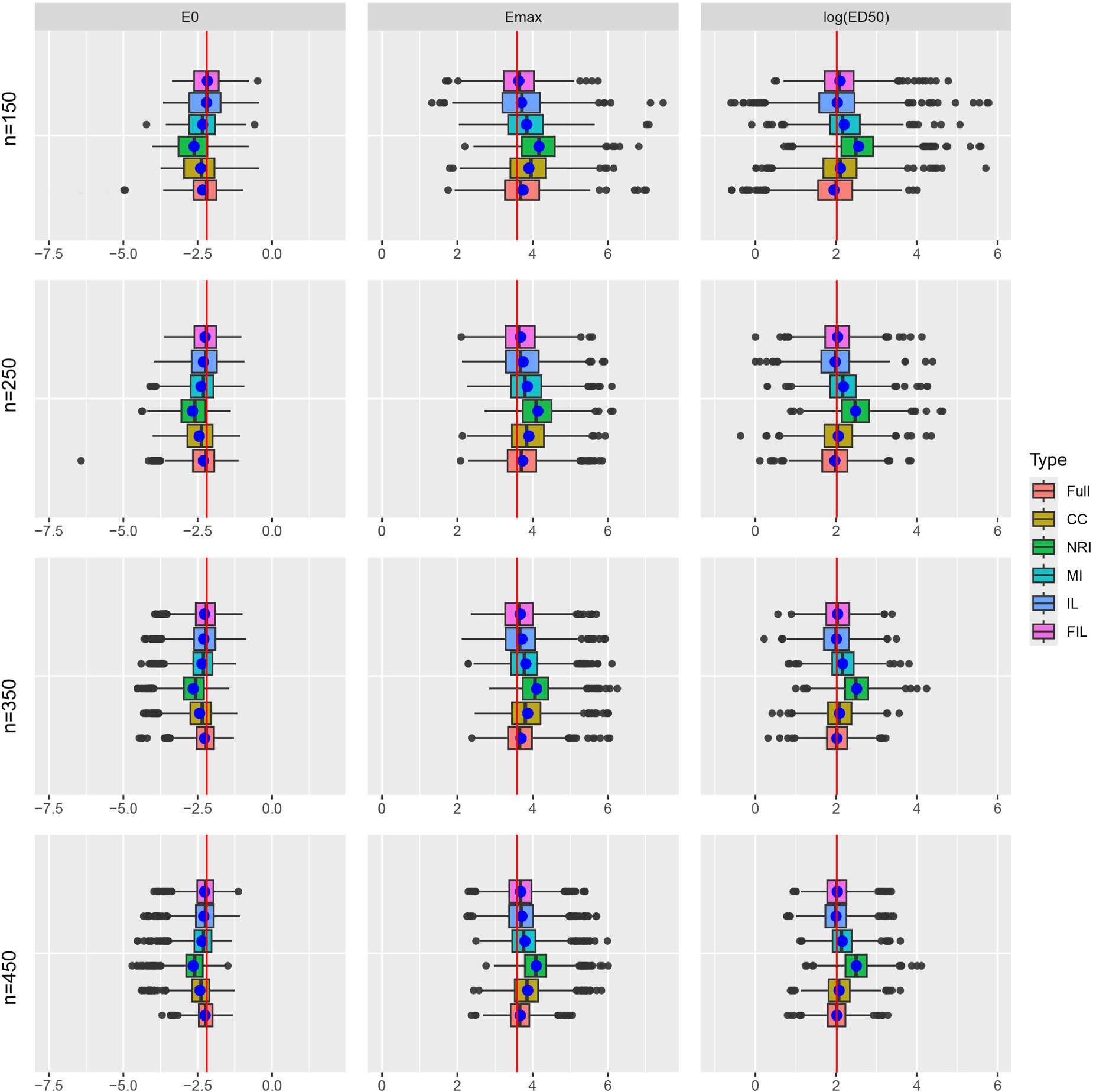
Boxplots comparing the distribution of point estimates with true parameter
Further simulations were conducted with varying missing rates and a fixed sample size of
This section presents an example of fitting a dose–response model using data from the TURANDOT study, 27 a Phase II randomized, double-blind, placebo-controlled clinical trial for ulcerative colitis in patients with moderate to severe disease. In this study, 357 patients were randomly assigned to either a placebo group or one of four active dose groups: 7.5 mg, 22.5 mg, 75 mg, and 225 mg. As reported by Vermeire et al., a non-monotone dose–response profile was observed, with lower efficacy in the highest dose group (225 mg). Among the 357 patients, 73 received placebo, 71 received 7.5 mg, 72 received 22.5 mg, 71 received 75 mg, and 70 received 225 mg. The primary endpoint was clinical remission at Week 12, which included several missing values believed to be nonignorable, as summarized in Table 2.
Sample size, the number of missing response cases, the number of remission cases, and baseline statistics for each dosage group in TURANDOT study.
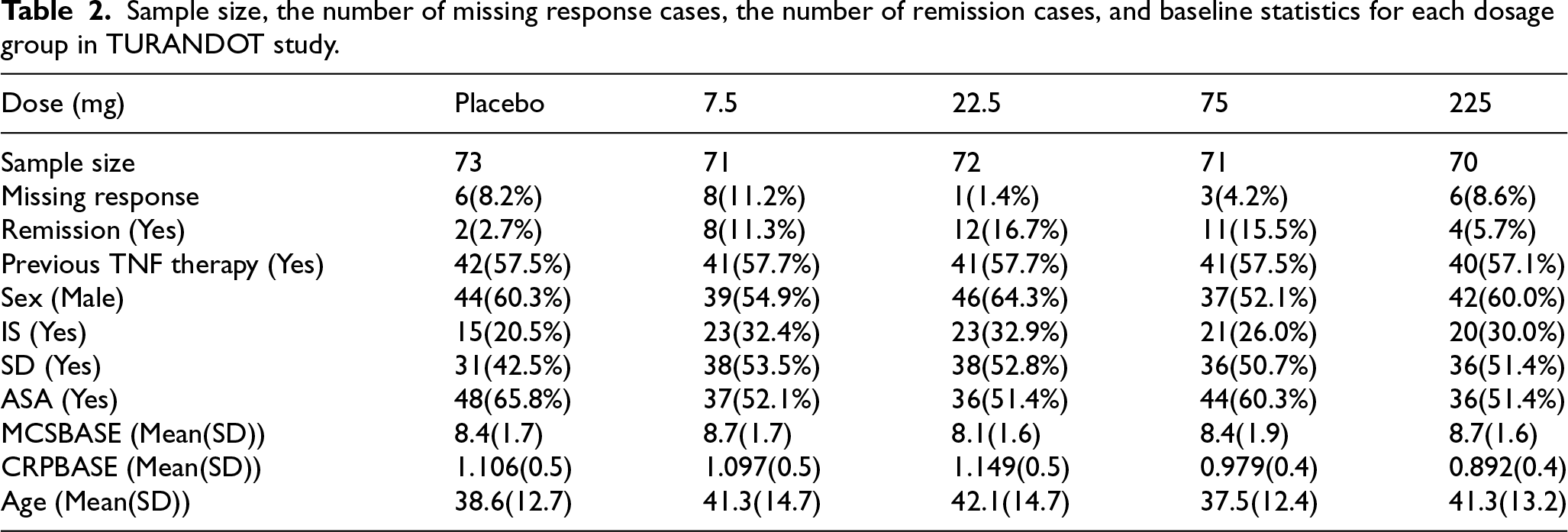
Sample size, the number of missing response cases, the number of remission cases, and baseline statistics for each dosage group in TURANDOT study.

We fitted the Emax model, assuming all missing values in remission were nonignorable. For modeling the missingness indicator, we performed model selection based on the Akaike Information Criterion (AIC) with a lot of covariates and their interactions, and the final model included the following covariates: remission response (y), dose, Mayo score at baseline (MCSBASE), C Reactive Protein score at baseline (CRPBASE), age, sex, immune suppressant status (IS), steroid use history (SD), and Acetylsalicylic acid use history (ASA). We compared our proposed method to complete case (CC) analysis, NRI, and MI. Note that for MI, we used the imputation model identical to the one employed in IL and FIL, excluding the response y. The results of the dose–response relationship using different methods are shown in Table 3.
Analysis result of TURANDOT data with different missing data handling methods and the proposed method.
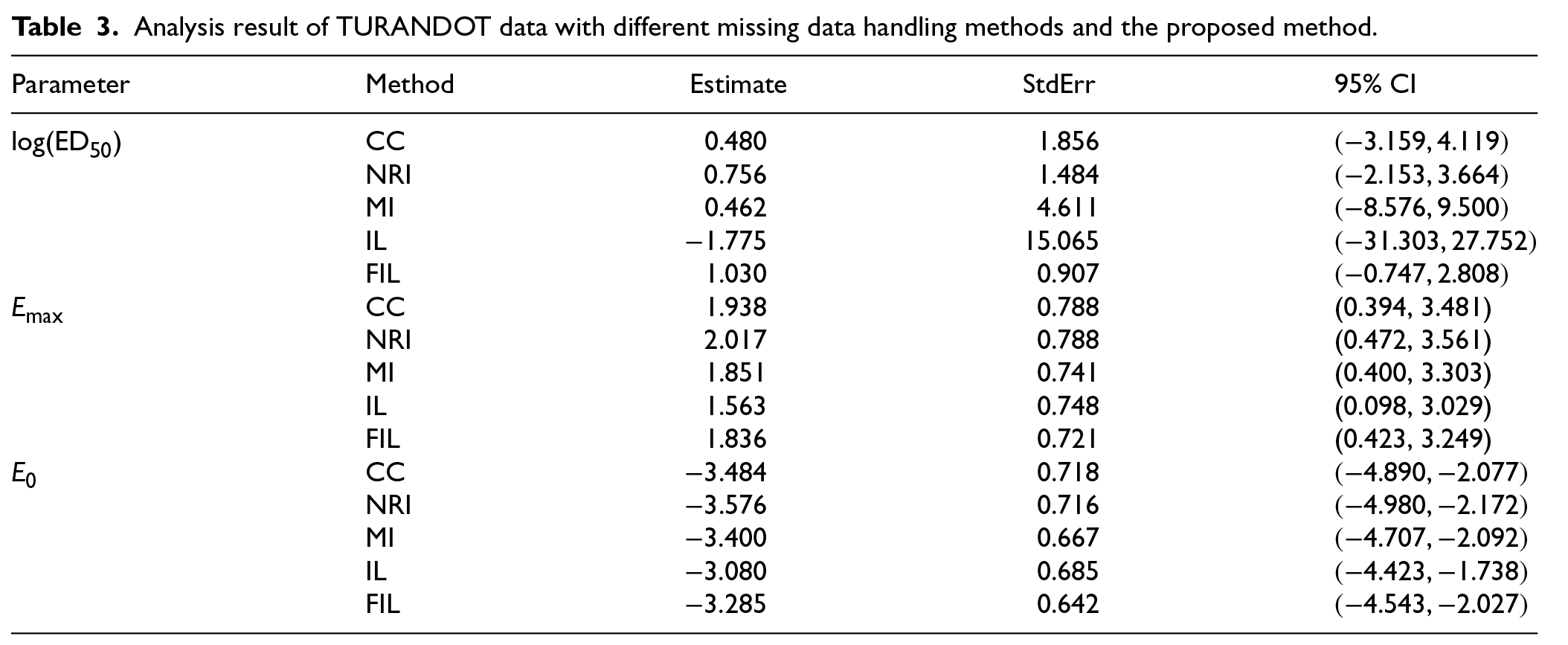
We observe that the IL method yields an unstable estimate for log(
To further evaluate the performance of these methods, we estimated the probabilities of remission using the coefficient estimates of the fitted Emax model. To quantify uncertainty of probabilities, we use a parametric bootstrap with
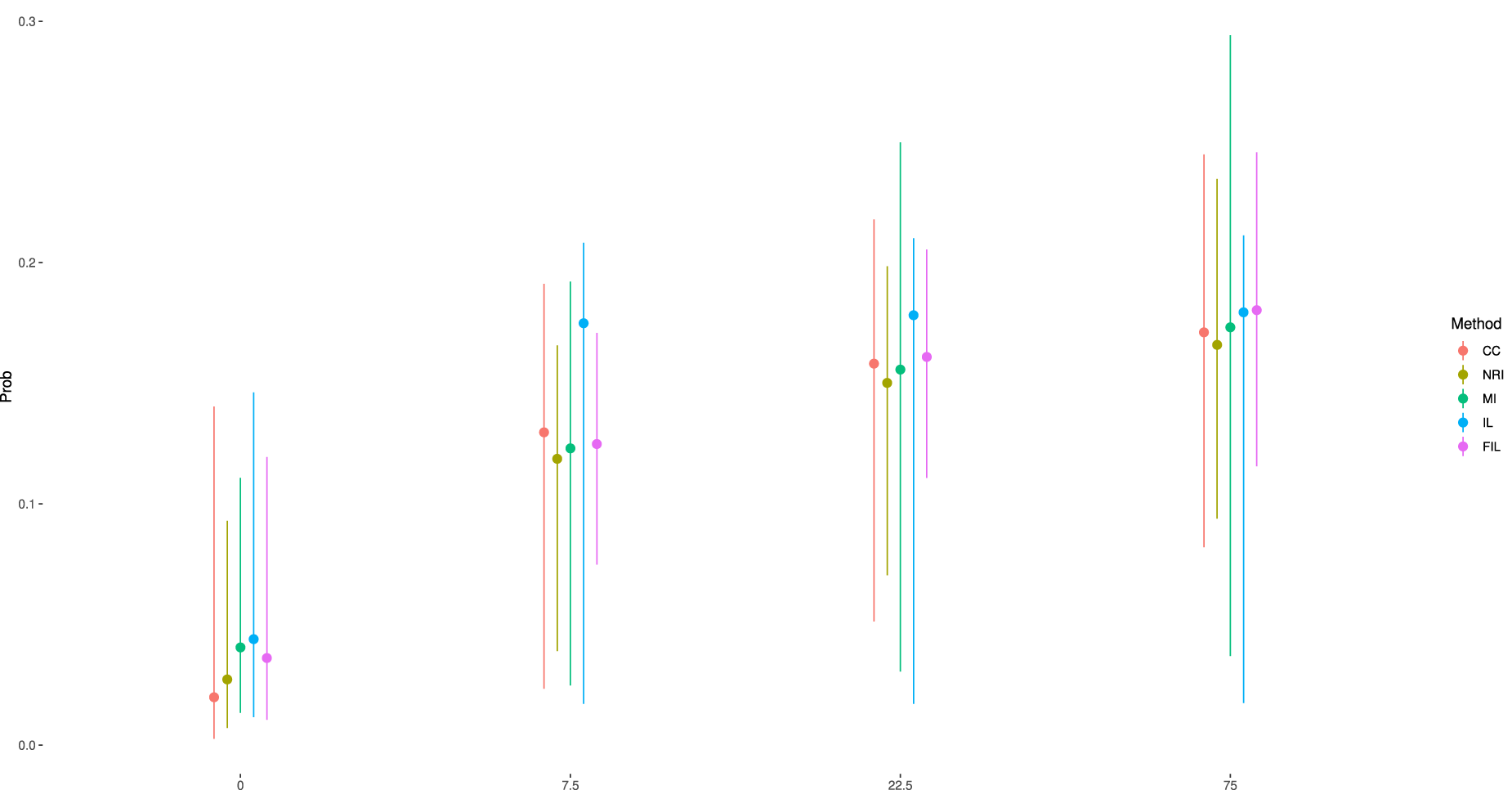
Estimated dose response remission probabilities based on different methods with their bootstrapped 95% confidence intervals.
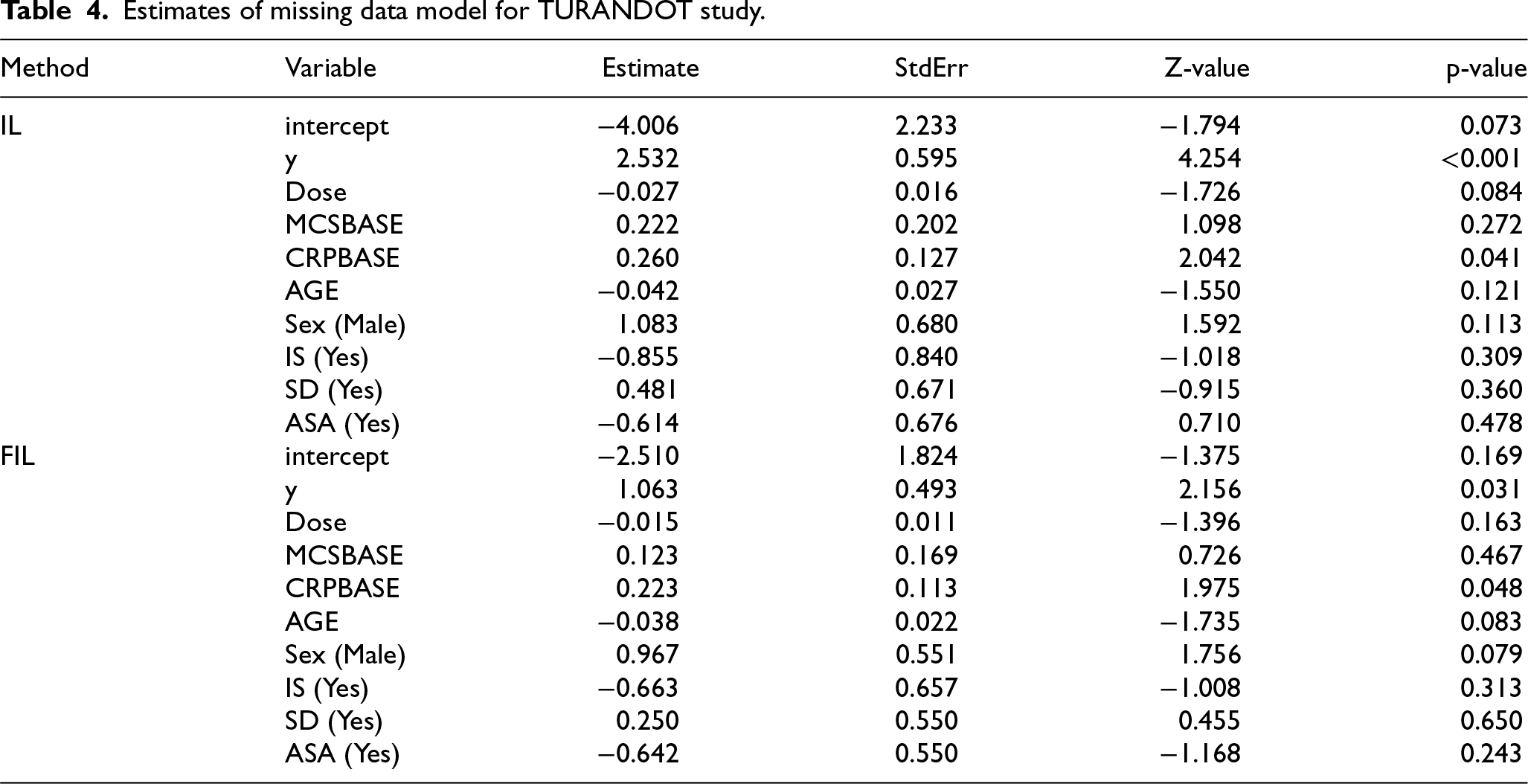
Table 4 presents the parameter estimates from the logistic regression of the missingness indicator, including their standard errors, Z-values, and p-values. The response variable clinical remission, coded as y, is significant at the 5% significance level while fitting the model in both the Ibrahim-Lipsitz (IL) and Firth-adjusted IL (FIL) methods, indicating that the probability of a response being missing depends on the response itself. This result provides supportive evidence that the missing data mechanism in this dataset may be nonignorable, while it is impossible to distinguish from MAR with the observed data only. 28
Estimates of missing data model for TURANDOT study.
In this article, we addressed the challenge of estimating the coefficients for the binary Emax model in the presence of missing responses under a nonignorable missing data mechanism. Our simulation studies demonstrate that the proposed Firth-type corrected weighted EM procedure of Ibrahim and Lipsitz (FIL) outperforms commonly used missing data handling strategies such as NRI and MI. Additionally, when fitting the binary Emax model with small or medium sample sizes, maximum likelihood-based approaches often face convergence issues due to complete separation or produce unstable estimates with significant bias and variation due to quasi-separation. In such scenarios, the FIL method offers a robust and reliable solution, effectively addressing the complexities and challenges posed by the data.
For both the IL and FIL methods, it is crucial to select an appropriate model for describing the likelihood of the missingness indicator,
The rationale for choosing an EM approach rather than multiple imputation (MI) to address missingness may warrant further explanation. As discussed in the introduction, MI is primarily designed for the MAR mechanism, which does not apply when the missing data are nonignorable. While recent literature, such as Im & Kim 29 and Galimard et al., 13 has proposed methods to handle nonignorable missingness, the selection of an appropriate imputation model remains a challenge and is often determined through sensitivity analysis. In contrast, with the IL and FIL methods, model selection can be performed using standard techniques, making these methods more straightforward to implement and interpret.
In dose–response analysis, the Multiple Comparison Procedure-Modeling (MCP-Mod) approach, developed by Bretz, Pinheiro, and Branson,30,31 combines hypothesis testing and modeling with Type I error control. MCP-Mod uses AIC to select the best model for fitting the dose–response relationship, making it a natural extension to apply our proposed methods within the MCP-Mod framework, given its likelihood-based foundation. Indeed, Diniz et al. have recently developed a Firth-type MCP-Mod for Weibull regression with time-to-event data in small sample sizes. 32 Further research could explore the implementation of these methods for binary or count dose–response models. In fact, MCP-mod contains a four-parameter Emax model as potential, thus extending to MCP-mod can capture situations when the three-parameter Emax model is misspecified. Additionally, penalization methods could be applied to the potential models within MCP-Mod to control the risk of separation, offering another avenue for future investigation.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802251403356 - Supplemental material for Robust Emax model fitting: Addressing nonignorable missing binary outcome in dose–response analysis
Supplemental material, sj-pdf-1-smm-10.1177_09622802251403356 for Robust Emax model fitting: Addressing nonignorable missing binary outcome in dose–response analysis by Jiangshan Zhang, Vivek Pradhan and Yuxi Zhao in Statistical Methods in Medical Research
Footnotes
Acknowledgements
The authors are grateful to an Associate Editor and referees, whose insightful comments have helped improve the work and presentation.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.