Abstract
G-formula is a popular approach for estimating the effects of time-varying treatments or exposures from longitudinal data. G-formula is typically implemented using Monte-Carlo simulation, with non-parametric bootstrapping used for inference. In longitudinal data settings missing data are a common issue, which are often handled using multiple imputation, but it is unclear how G-formula and multiple imputation should be combined. We show how G-formula can be implemented using Bayesian multiple imputation methods for synthetic data, and that by doing so, we can impute missing data and simulate the counterfactuals of interest within a single coherent approach. We describe how this can be achieved using standard multiple imputation software and explore its performance using a simulation study and an application from cystic fibrosis.
Introduction
The collection of methods referred to as G-methods, developed by James Robins and co-workers, can provide valid inference for the effects of time-varying exposures or treatments in the presence of time-varying confounders – variables that affect treatment over time and the outcome of interest – even when these are affected by previous values of treatment. 1 One such method is parametric G-formula (sometimes known as G-computation). Parametric G-formula involves postulating models for the time-varying confounders and outcomes. The expected outcome under specified longitudinal treatment regimes of interest can then be estimated and contrasted. The evaluation of G-formula estimators generally involves intractable integrals. To overcome this in practice, G-formula implementations make use of Monte-Carlo integration, in which counterfactual outcomes are simulated under the treatment regimes of interest.2,3 Inference for Monte-Carlo based G-formula estimators is typically performed using bootstrapping.
A common complication in this context is missing values in the time-varying confounders, treatment and/or outcome variables. Existing implementations of G-formula in statistical software either impute such missing values once before fitting the required models
2
or fit the required models discarding person-visit observations which contain missing values in the variables concerned.
3
Alternatively, analysts may opt for ad-hoc approaches such as last observation carried forward to handle such missingness, which may make implausible assumptions about missing data.
4
In contrast, multiple imputation (MI) of missing values is often nowadays considered an attractive approach to handling missing data, since it makes efficient use of the observed data and may in some situations make a more plausible assumption about missing data than discarding incomplete observations.
5
In principle G-formula can be used after MI: The G-formula method is applied to each imputed dataset, using bootstrapping to obtain the within-imputation variance estimate, and the estimates are then pooled using Rubin’s rules. The drawback to this approach is a very high computational cost, because the G-formula approach with bootstrapping, which itself is computationally expensive, must be repeated
This paper is organised as follows. In Section 2 we review parametric G-formula and how it is typically implemented. In Section 3 we describe how G-formula can be implemented by exploiting existing methodology for using MI to generate synthetic datasets. In Section 4 we report the results of simulation studies investigating the performance of this approach. Section 5 describes results of an illustrative analysis of data from a cystic fibrosis registry. We conclude in Section 6 with a discussion.
Review of G-formula
Suppose we have a sample of data from
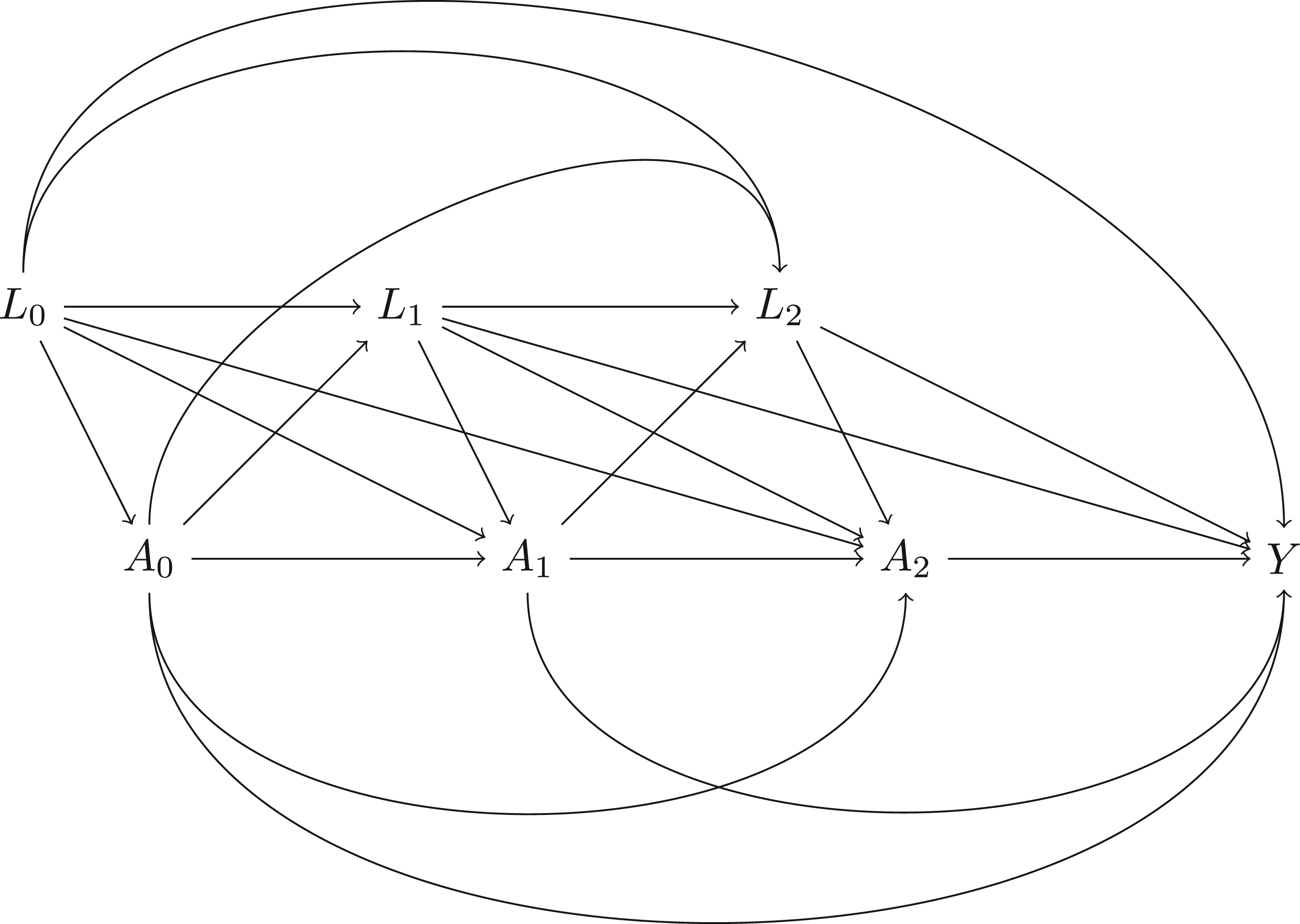
Directed acyclic graph (DAG) of a study with time-varying treatments
The G-formula estimator of 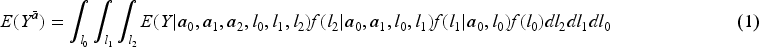

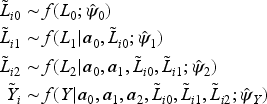
While we have stated that a model
In this section we describe how a Monte-Carlo G-formula estimator can be implemented using MI methods. In Section 3.1 we describe how the point estimator is constructed using MI. In Section 3.2 we explain why Rubin’s standard variance estimator is biased in this instance, and describe an alternative variance estimator, which was derived in the context of using MI to generate synthetic datasets by Raghunathan et al. 6 In Section 3.3 we describe how standard MI software can be used to implement the approach. Lastly, in Section 3.4 we describe how the approach readily extends to accommodate missing actual data (as opposed to missing counterfactual data).
Point estimation
To estimate
G-formula via multiple imputation (MI) data setup.
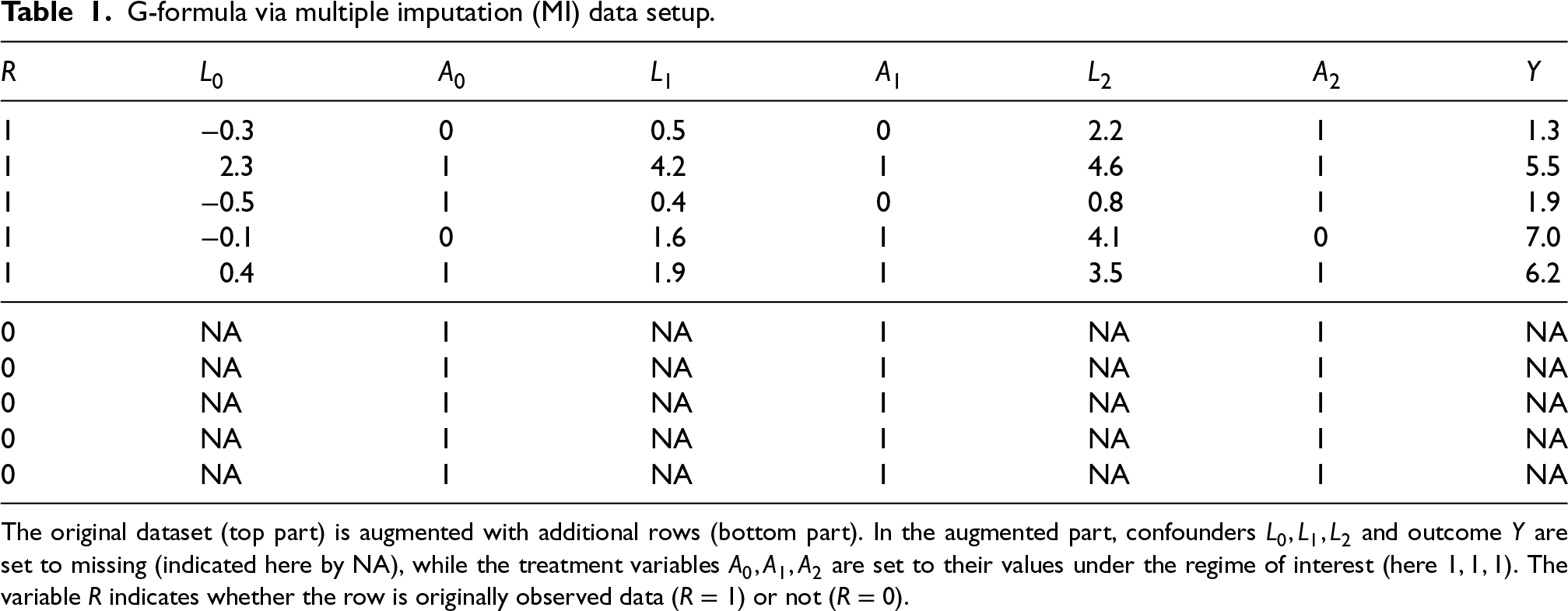
The original dataset (top part) is augmented with additional rows (bottom part). In the augmented part, confounders
and outcome
are set to missing (indicated here by NA), while the treatment variables
are set to their values under the regime of interest (here
). The variable
indicates whether the row is originally observed data (
) or not (
).
G-formula via multiple imputation (MI) data setup.

The original dataset (top part) is augmented with additional rows (bottom part). In the augmented part, confounders
Next, Bayesian MI is used to generate
The resulting estimator
For the G-formula via MI approach we propose, the impact of generating each imputed dataset conditional on posterior draws of the model parameters, rather than an efficient observed data estimate, is to increase the asymptotic variance of the estimator, but this increase goes to zero as
The variance of an MI estimator is typically estimated using Rubin’s variance estimator
The G-formula via MI estimator is closely related to the use of MI to generate samples from synthetic populations, first proposed by Rubin.
12
Here the objective is to release these synthetic samples rather than the original data in order to protect the confidentiality of survey respondents’ data. For synthetic MI, Raghunathan et al.
6
developed
To build intuition for
Having generated imputed/synthetic datasets for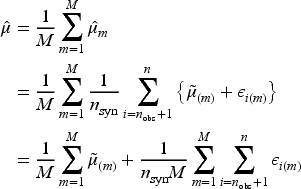 
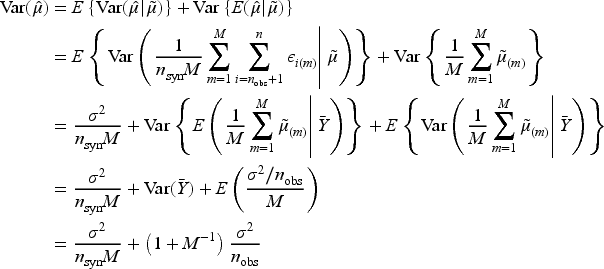
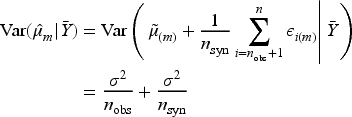
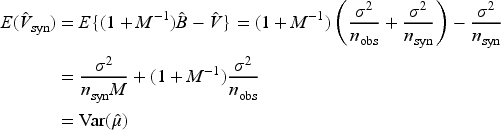
As noted by Reiter
14
and Raghunathan et al.,
6
the variance estimator
Implementation using imputation software
To implement the proposed approach, as described previously, the observed dataset of size
Ordinarily interest focuses on the contrast of potential outcome means under two (or more) different treatment regimes. To estimate the corresponding contrast in potential outcome means, we augment the observed dataset twice. In the second augmentation part, the treatment variables are set according to the second treatment regime of interest. The difference in potential outcome means can be estimated by the difference in simulated outcomes between the two augmented parts. The variance of the resulting estimator can be estimated by the sum of the variance estimator
Implementation of the preceding steps using packages such as mice in R is relatively straightforward. Nonetheless, to facilitate use of the approach, we provide the R package gFormulaMI. This augments the supplied dataset as described above and imputes missing data using the mice package. The resulting imputed datasets contain only the augmented portion of the imputations (with
As noted earlier, the standard (non-Bayesian) implementation of G-formula avoids specification of a model for
Missing data
Now suppose that there are some data missing which we want to handle by MI. Missing data could occur in either the longitudinal confounders
To multiply impute the missing values in the original dataset and the missing potential outcomes, we propose a two-stage approach where first we generate
Such a two-stage approach to impute missing values has been proposed previously for contexts where, as in the case here, there are two qualitatively different types of missing data.
17
It can be justified when (a) the missing data (here missing actual data and missing potential outcomes) are MAR, and (b) the process that divides the missing values into the two parts does not depend on the missing data.
17
The first condition follows since the missingness in the original data portion is assumed MAR and in the augmented part it is MCAR by design. The second condition holds because the division of missing data is on the basis of
A two-stage approach to handle missingness in the observed data in the context of using MI to generate synthetic samples was recently developed by Yu et al.
19
They proposed that conditional on each of the
The models required for G-formula given in equation (2) do not fully specify the joint distribution of all the variables under consideration, since they do not specify models for the treatment variables. The imputation models used to impute the missing data in the original dataset should ideally be compatible with those used to impute the augmented rows. One way to achieve this is to specify a full joint model for all the variables by, in addition to the models in equation (2), specifying models for the time-varying treatment variables. That is, for
In the setting with missing data, our R package gFormulaMI takes as input a set of
Simulations
In this section we report the results of simulations performed to examine the empirical performance of the G-formula via MI approach. We first consider, in Section 4.1, the setting where there is no missing data. Next, in Section 4.2, we consider the situation where some data are missing.
No missing data
We simulated datasets for
Table 2 shows results based on 10,000 simulations per scenario. As expected since the imputation models were correctly specified, the G-formula via MI estimator for
Simulation results for G-formula via multiple imputation ( MI) without any missing data.
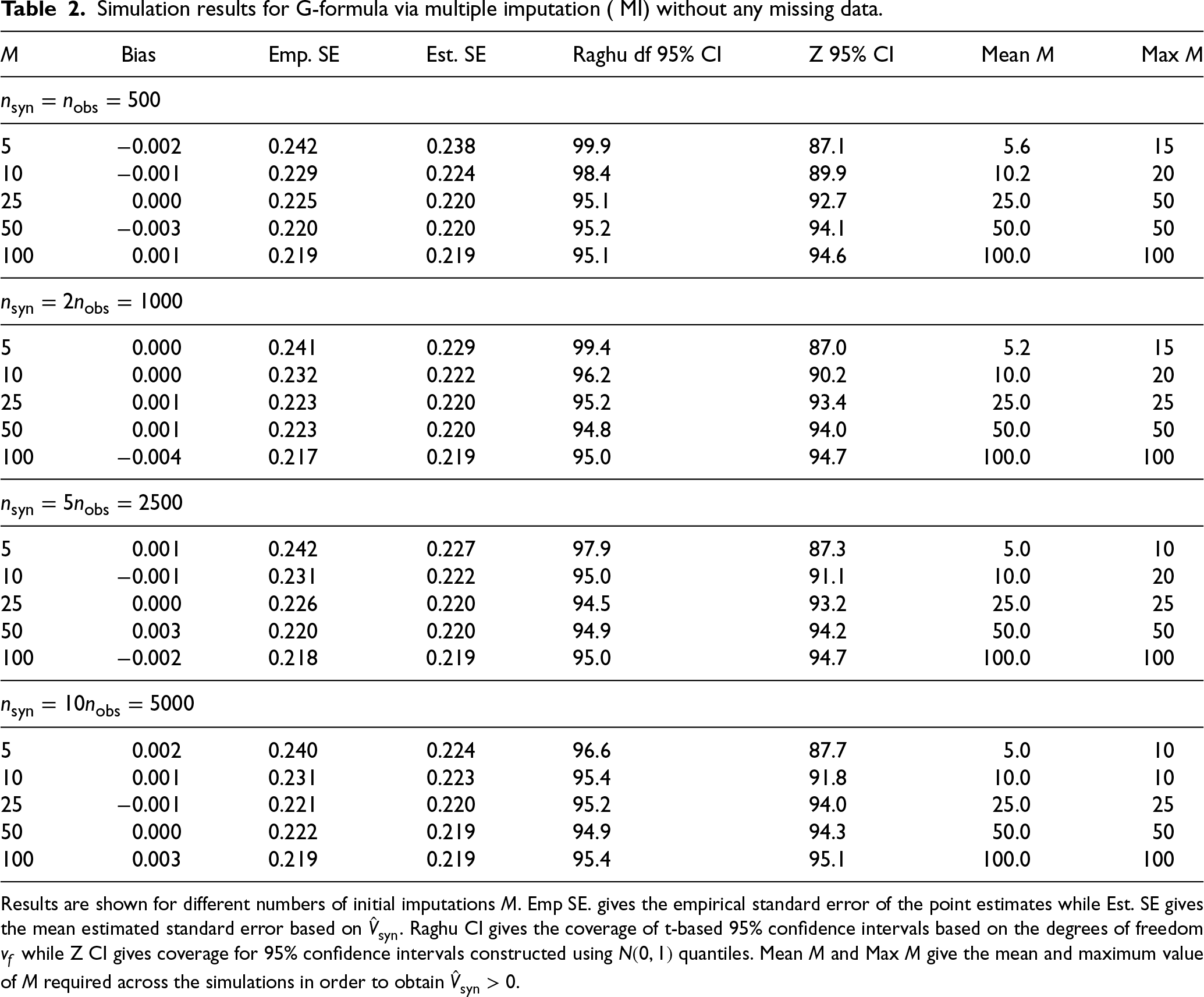
Results are shown for different numbers of initial imputations
. Emp SE. gives the empirical standard error of the point estimates while Est. SE gives the mean estimated standard error based on
. Raghu CI gives the coverage of t-based 95% confidence intervals based on the degrees of freedom
while Z CI gives coverage for 95% confidence intervals constructed using
quantiles. Mean
and Max
give the mean and maximum value of
required across the simulations in order to obtain
.
Simulation results for G-formula via multiple imputation ( MI) without any missing data.

Results are shown for different numbers of initial imputations
For comparison with performance of G-formula based on the usual implementation approach, Table 3 shows results from 10,000 simulations under the same data generating mechanism obtained using the gfoRmula package. Here pooled models are fitted to the data in long form, as opposed to in wide form in our G-formula via MI implementation using mice. The results shown in Table 3 are based on assuming a normal linear model for the single continuous time-varying confounder, and included the past measurements of treatment and the confounder, plus visit time, as covariates. The results show the estimates were also unbiased, and had empirical SE very slightly below that achieved by the G-formula via MI approach when using
Simulation results for G-formula using gfoRmula package, based on Monte-Carlo sample sizes of
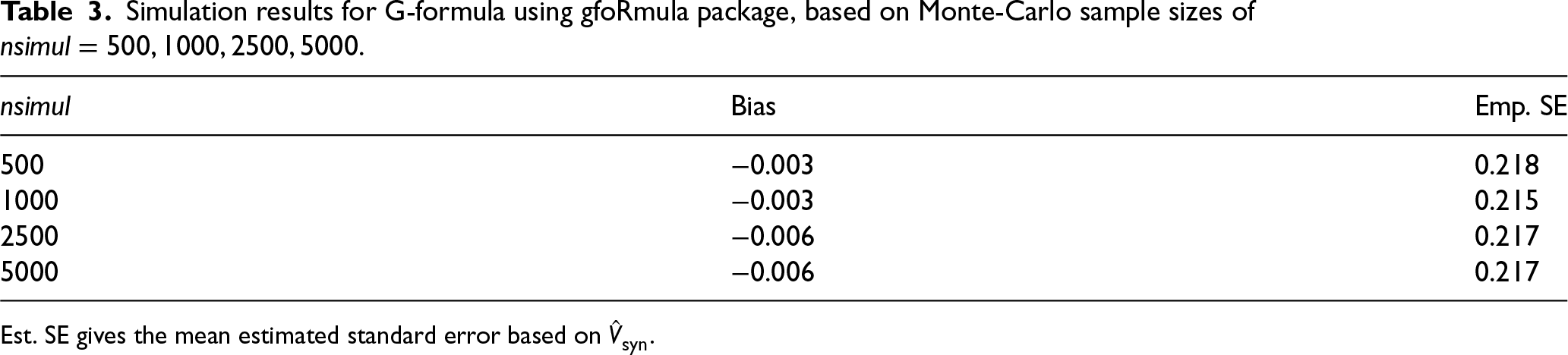
Est. SE gives the mean estimated standard error based on
Table 4 shows results of simulations for G-formula via MI performed using a smaller sample size of
Simulation results for G-formula via MI without any missing data, but using
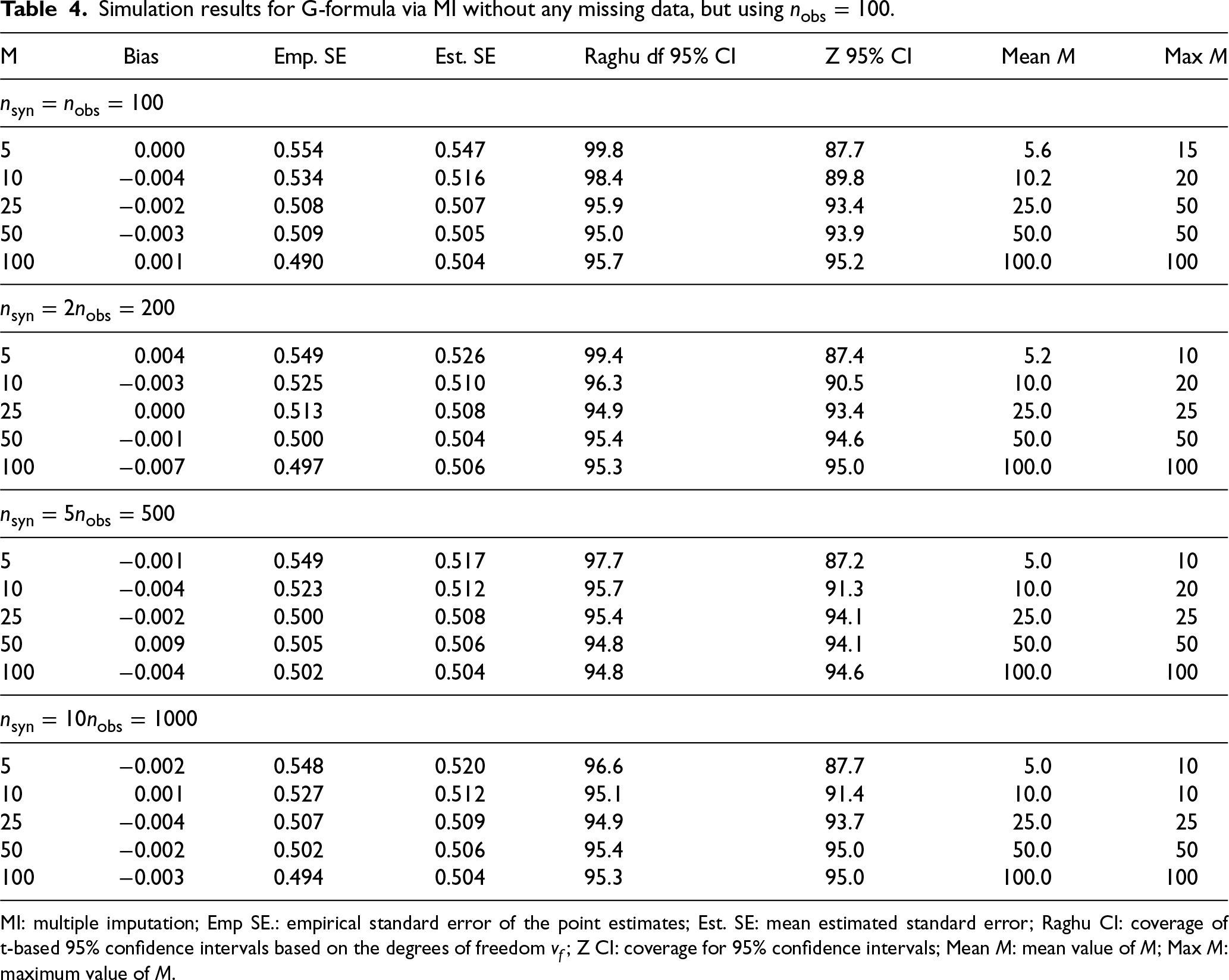
MI: multiple imputation; Emp SE.: empirical standard error of the point estimates; Est. SE: mean estimated standard error; Raghu CI: coverage of t-based 95% confidence intervals based on the degrees of freedom
We additionally ran 10,000 simulations with
Next we performed simulations where some data were missing. Data in each of
To implement G-formula via MI we used an initial call to mice to impute the missing values
Table 5 shows the results based on 10,000 simulations per value of
Simulation results for G-formula via MI with missing data.
is the probability that each of
,
,
,
and
are missing.
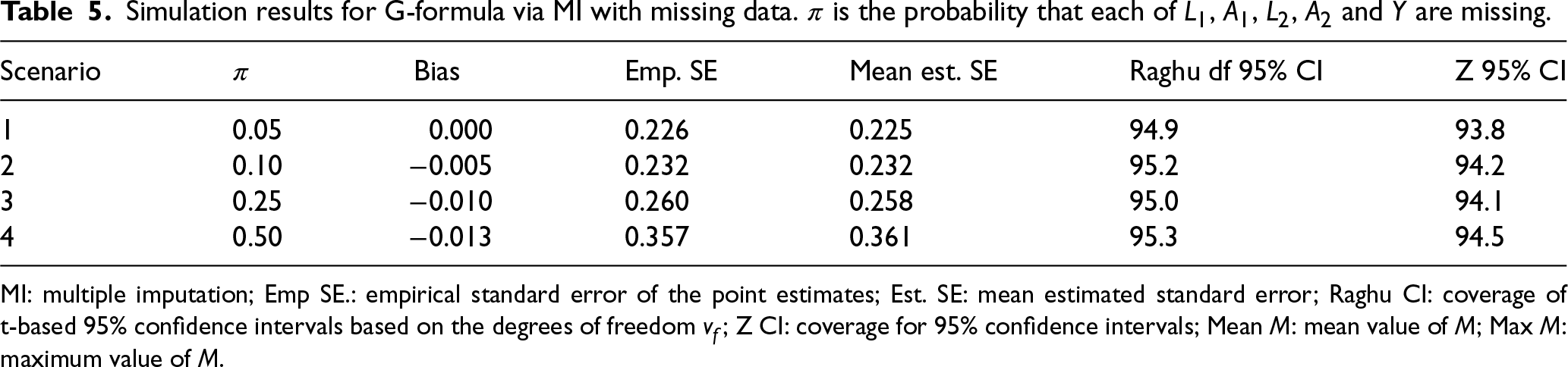
MI: multiple imputation; Emp SE.: empirical standard error of the point estimates; Est. SE: mean estimated standard error; Raghu CI: coverage of t-based 95% confidence intervals based on the degrees of freedom
; Z CI: coverage for 95% confidence intervals; Mean
: mean value of
; Max
: maximum value of
.
Simulation results for G-formula via MI with missing data.

MI: multiple imputation; Emp SE.: empirical standard error of the point estimates; Est. SE: mean estimated standard error; Raghu CI: coverage of t-based 95% confidence intervals based on the degrees of freedom
In this section we provide an illustrative example of the use of the G-formula via MI approach to investigate the effects of multiple treatments on lung function in people with cystic fibrosis (CF). Many people with CF are prescribed at least one mucoactive treatment to help improve lung function. In the UK, the most commonly prescribed nebulised mucoactive treatment is dornase alfa (DNase), and many patients already using DNase may later add or switch to hypertonic saline. Existing research investigates the effects of taking DNase or hypertonic saline alone, but the effects of using both treatments in combination are less well understood. Here we investigate the following question: for people with CF who are already established on DNase, does adding hypertonic saline have any additional benefit for lung function? In a recent study this question was investigated using marginal structural models estimated using inverse probability of treatment weighting to address time-dependent confounding. 20
Our example uses data from the UK Cystic Fibrosis Registry, which collects longitudinal data on almost all people with CF in the UK. 21 Longitudinal data are collected annually, when CF patients are seen at an outpatient clinic for a comprehensive review. The review data includes evaluation of clinical status, lung function, chronic medications, hospital admissions and health complications.
Using data from 2007 to 2018, we included individuals with CF, aged 6 years or older, who had been prescribed DNase, but not hypertonic saline, for at least two consecutive years. Organ transplant recipients, and people prescribed certain treatments (mannitol, ivacaftor, lumacaftor/ivacaftor, tezacaftor/ivacaftor) were excluded. Time zero was defined as the date of the most recent annual review at which the inclusion and exclusion criteria were met, but which allowed for the maximum possible follow-up time up to 5 years. The outcome of interest is lung function, and this is measured at the annual review as forced expiratory volume in one second (FEV
Treatment effect estimates were obtained using the standard implementation of G-formula and using G-formula via MI. Standard implementation was done using the R package gfoRmula, with
Four thousand, seven hundred fifty-nine individuals were eligible for inclusion in the analysis. Details on how the study sample was selected, baseline characteristics by treatment group, and amount of missing data by year, are provided in Section D, Tables 1 and 2, and Figure 1 of the Supplemental material. Of the 4,759 individuals included in the study, 2,255 were complete cases. Table 6 shows the results from each analysis along with the total running time.
Results using data from the UK CF registry.
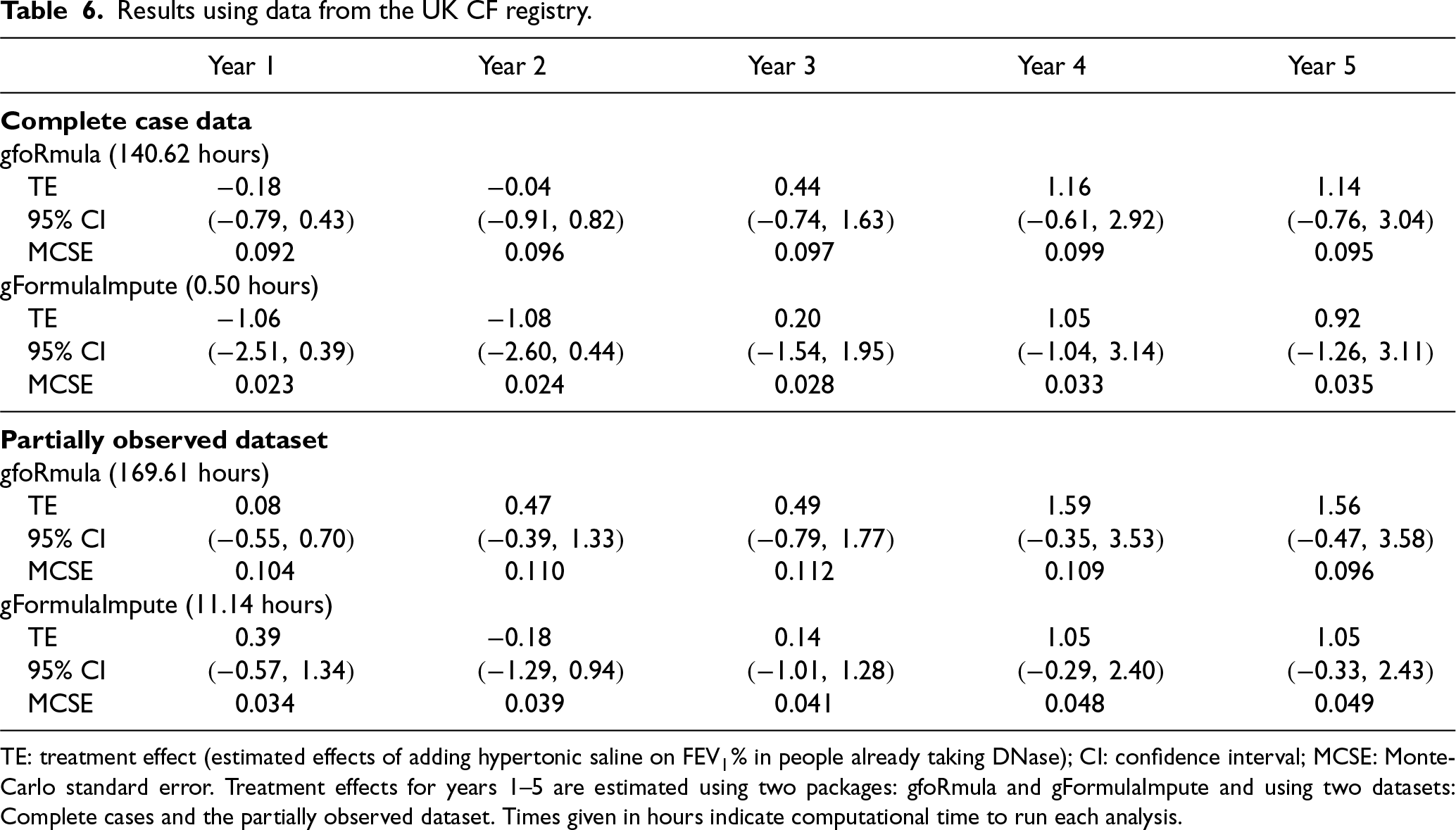
TE: treatment effect (estimated effects of adding hypertonic saline on FEV
% in people already taking DNase); CI: confidence interval; MCSE: Monte-Carlo standard error. Treatment effects for years 1–5 are estimated using two packages: gfoRmula and gFormulaImpute and using two datasets: Complete cases and the partially observed dataset. Times given in hours indicate computational time to run each analysis.
Results using data from the UK CF registry.

TE: treatment effect (estimated effects of adding hypertonic saline on FEV
Overall, and in line with previous results,
20
we found little evidence that adding hypertonic saline has any effect on FEV
G-formula via MI is an attractive approach for implementing parametric G-formula, that enables imputation of missing data and simulation of counterfactuals under the desired treatment regime(s) of interest. Moreover, it avoids the need to use bootstrapping for inference, which is particularly attractive in the context of combining MI for missing data with G-formula for causal inference. This is achieved by exploiting existing results for using MI to create synthetic datasets. The simulation results presented here suggest the G-formula via MI approach can perform well, requiring a relatively small number of imputations for reliable inference in the setup we used.
One alternative to imputing (actual) missing data when implementing G-formula is to fit each of the models required using the subset of records for which the variables involved in each model are fully observed, as is implemented for example in the gfoRmula package in R. 3 These complete case model fits yield consistent estimates of the respective conditional model parameters provided the probability of having all the variables involved in the model is independent of the dependent variable conditional on the covariates. 23 When the pattern of missingness in the longitudinal dataset is complex, consisting of both intermittent missingness and missingness due to dropout, such an assumption can sometimes be deemed more plausible than missing at random, whose meaning becomes complex in such settings. 24 Thus an alternative possible version of the G-formula via MI approach when some data are missing is to fit each of the required models using their respective complete case fits. Such an approach would be more efficient and plausibly less likely to be biased than applying the method to the subset of individuals who have all variables at all time points fully observed.
To obtain point estimates and inferences with sufficiently small Monte-Carlo error, existing simulation based implementations of G-formula may require both the Monte-Carlo sample size to be large and also a large number of bootstrap samples to be used. Our simulations and data analysis suggest that reliable inferences can be obtained via MI methods using smaller Monte-Carlo sample sizes (
We note that while standard G-formula is typically implemented using Monte-Carlo simulation as we have described, an alternative version based on iterative conditional expectations can be used. 25 This approach requires models for a series of conditional mean functions, rather than for the full distribution of the time-varying confounders and outcome. This makes it potentially less prone to model misspecification, particularly in the case of several time-varying confounders, where the standard approach requires a choice of factorisation to be made to specify the distribution of the time-varying confounders at each time point. Moreover, it does not require the use of simulation, and closed form variance estimators can be constructed based on estimating equation theory. 26
In this paper we have focussed on G-formula where the outcome is a variable
Implementations of G-formula (e.g. the G-formula packages in Stata 2 and R 3 ) often fit models pooled across time points for each variable. This is achieved by formatting the data in so-called long form. Doing so permits borrowing of information across time points in the estimation of regression parameters, but of course relies on the validity of the assumption that the conditional distribution of confounders given earlier variables is homogeneous across time points. Although this approach could be implemented via the MI approach we have outlined, we do not believe it is possible using standard imputation software such as mice in R. This is because having transformed the data into long form, it is not (currently) possible to update values from one row of the data frame from another within the algorithm.
While our focus in this paper has been on static treatment regimes, G-formula can be used to estimate the effects of dynamic treatment regimes, where the exposure or treatment at a given time point is assigned dependent on the longitudinal history observed up to that time. The G-formula via MI approach can be extended to this case, by setting the treatment variables to missing in the augmented part of the dataset and then specifying how they should be imputed based on the preceding (in time) variables. This can be achieved for example in the mice package through the use of user specified deterministic (or indeed stochastic) imputation methods. Moreover, we believe the basic approach we have described can be extended and applied to more general and complex causal structures specified by a DAG.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802251316971 - Supplemental material for G-formula with multiple imputation for causal inference with incomplete data
Supplemental material, sj-pdf-1-smm-10.1177_09622802251316971 for G-formula with multiple imputation for causal inference with incomplete data by Jonathan W Bartlett, Camila Olarte Parra, Emily Granger, Ruth H Keogh, Erik W van Zwet and Rhian M Daniel in Statistical Methods in Medical Research
Footnotes
Acknowledgements
The authors thank people with CF and their families for consenting to their data being held in the UK CF Registry, and NHS teams in CF centres and clinics for the input of data into the Registry. We also thank the UK Cystic Fibrosis Trust and the Registry Steering Committee for access to anonymised UK CF Registry data.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Bartlett, Olarte Parra and Daniel were supported by a UK Medical Research Council Grant (MR/T023953/1 and MR/T023953/2). Keogh and Granger were supported by a UKRI Future Leaders fellowship (MR/S017968/1) awarded to Keogh.
Data availability
The gFormulaMI R package is available from CRAN. R code for the simulations and CF analysis are available from https://github.com/jwb133/gFormulaViaMultipleImputationPaper. Researchers can apply for access to the CF data from ![]() .
.
ORCID iDs
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
