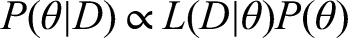Abstract
The COVID-19 pandemic has brought to the fore the need for policy makers to receive timely and ongoing scientific guidance in response to this recently emerged human infectious disease. Fitting mathematical models of infectious disease transmission to the available epidemiological data provide a key statistical tool for understanding the many quantities of interest that are not explicit in the underlying epidemiological data streams. Of these, the effective reproduction number,
Keywords
1 Introduction
In late 2019, accounts emerged from Wuhan city in China of a virus of unknown origin that was leading to a cluster of pneumonia cases. 1 The virus was identified as a novel strain of coronavirus on 7 January 2020, 2 subsequently named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), causing the respiratory syndrome known as COVID-19. The outbreak has since developed into a global pandemic. As of 3 August 2020, the number of confirmed COVID-19 cases was approaching 18 million, with more than 685,000 deaths occurring worldwide. 3 Faced with these threats, there is a need for robust predictive models that can help policy makers by quantifying the impact of a range of potential responses. However, as is often stated, models are only as good as the data that underpin them; it is therefore important to examine, in some detail, the parameter inference methods and agreement between model predictions and data.
In the UK, the first cases of COVID-19 were reported on 31 January 2020 in the city of York. Cases continued to be reported sporadically throughout February and by the end of the month, guidance was issued stating that travellers from the high-risk epidemic hotspots of Hubei province in China, Iran and South Korea should self-isolate upon arrival in the UK. By mid-March, as the number of cases began to rise, there was advice against all non-essential travel and, over the coming days, several social-distancing measures were introduced including the closing of schools, non-essential shops, pubs and restaurants. This culminated in the introduction of a UK lockdown, announced on the evening of 23 March 2020, whereby the public were instructed to remain at home with four exceptions: shopping for essentials; any medical emergency; for one form of exercise per day; and to travel to work if absolutely necessary. By mid-April 2020, these stringent mitigation strategies began to have an effect, as the number of confirmed cases and deaths as a result of the disease began to decline. As the number of daily confirmed cases continued to decline during April, May and June, measures to ease lockdown restrictions began, with the reopening of some non-essential businesses and allowing small groups of individuals from different households to meet up outdoors, whilst maintaining social distancing. This was followed by gradually reopening primary schools in England from 1 June 2020 and all non-essential retail outlets from 15 June 2020. Predictive models for the UK are therefore faced with a changing set of behaviours against which historic data must be judged and an uncertain future of potential additional relaxations.
Throughout, a significant factor in the decision-making process was the value of the effective reproduction number,
The initial understanding of key epidemiological characteristics for a newly emergent infectious disease is, by its very nature of being novel, extremely limited and often biased towards early severe cases. Developing models of infectious disease dynamics enables us to challenge and improve our mechanistic understanding of the underlying epidemiological processes based on a variety of data sources. One way such insights can be garnered is through model fitting/parameter inference, the process of estimating the parameters of the mathematical model from data. The task of fitting a model to data is often challenging, partly due to the necessary complexity of the model in use, but also because of data limitations and the need to assimilate information from multiple sources of data. 5
Throughout this work, the process of model fitting is performed under a Bayesian paradigm, where knowledge of the parameters are modelled through random variables and have joint probability distributions.
6
In full, the posterior distribution of the parameters
Adopting a Bayesian approach to parameter inference means parameter uncertainty may then be propagated if using the model to make projections. This affords models with mechanistic aspects, through computational simulation, the capability of providing an estimated range of predicted possibilities given the evidence presently available. Thus, models can demonstrate important principles about outbreaks, 8 with examples during the present pandemic including analyses of the effect of non-pharmaceutical interventions (NPIs) on curbing the outbreak of COVID-19 in the UK. 9 .
In this paper, we present the inference scheme, and its subsequent refinements, employed for calibrating the Warwick SARS-CoV-2 transmission and COVID-19 disease model
10
to the available public health data streams and estimating key epidemiological quantities such as
We begin by describing our mechanistic transmission model for SARS-CoV-2 in Section 2, detailing in Section 3 how the effects of social distancing are incorporated within the model framework. To fit the model to data streams pertaining to critical care, such as hospital admissions and bed occupancy, Section 4 expresses how epidemiological outcomes were mapped onto these quantities. In Section 5, we outline how these components are incorporated into the likelihood function and the adopted MCMC scheme. The estimated parameters are then used to measure epidemiological measures of interest, such as the growth rate (
The closing sections draw attention to how model frameworks may evolve during a disease outbreak as more data streams become available and we collectively gain a better understanding of the epidemiology (Section 7). We explore how key epidemiological quantities, in particular the reproduction number
2 Model description
Here we present the University of Warwick SEIR-type compartmental age-structured model, developed to simulate the spread of SARS-CoV-2 within regions of the UK. Matched to a variety of epidemiological data, the model operates and is fitted to data from the seven NHS regions in England (East of England, London, Midlands, North East and Yorkshire, North West, South East and South West) and the three devolved nations (Northern Ireland, Scotland and Wales). The model incorporates multiple layers of heterogeneity, through partitioning the population into five-year age classes, tracking symptomatic and asymptomatic transmission, accounting for household saturation of transmission and household quarantining.
The population is stratified into multiple compartments with respect to SARS-CoV-2 infection status (Figure 1): individuals may be susceptible (
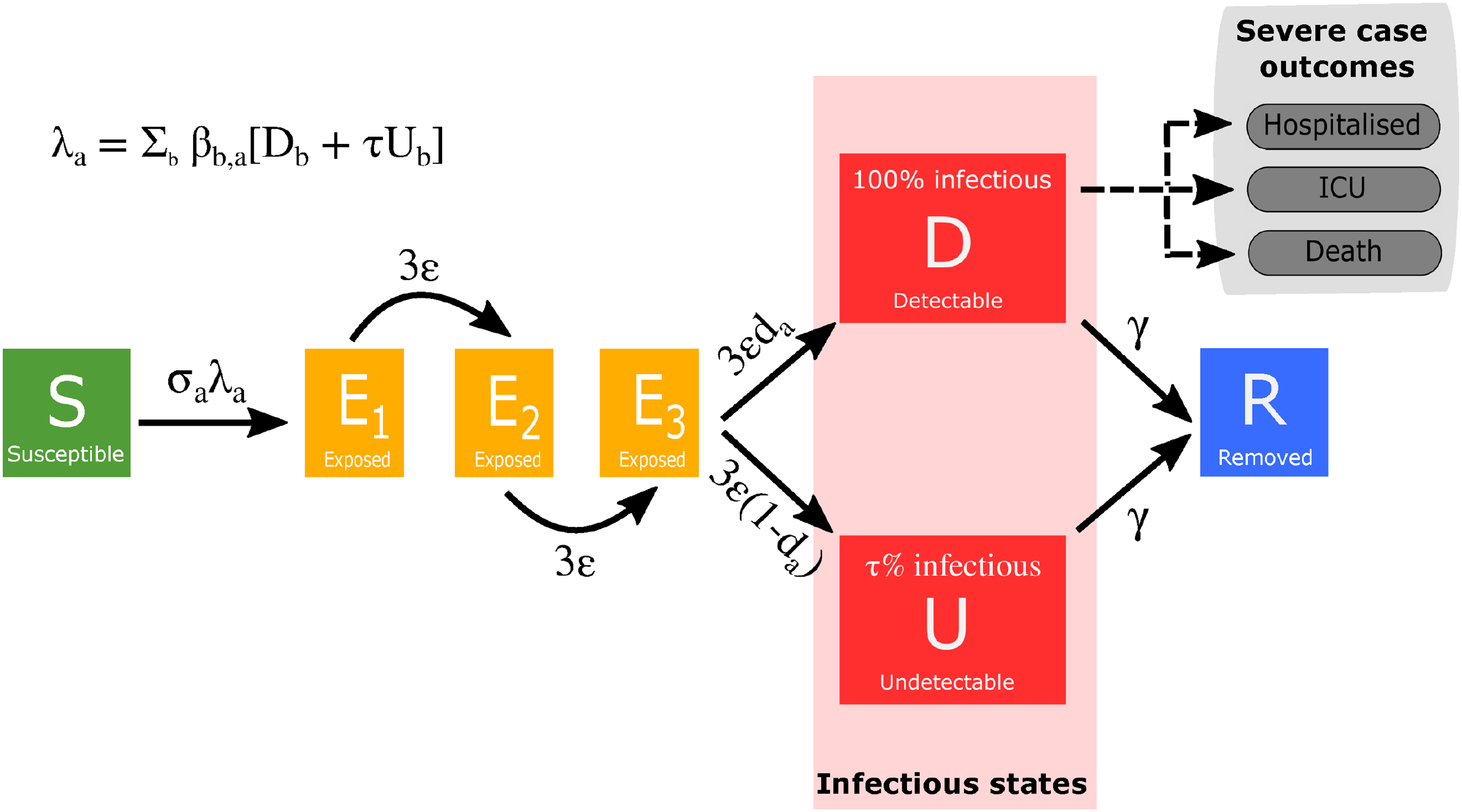
Model schematic of infection states and transitions. We stratified the population into susceptible, exposed, detectable infectious, undetectable infectious, and removed states. Solid lines correspond to disease state transitions, with dashed lines representing a mapping from detectable cases to severe clinical cases that require hospital treatment, critical care (intensive care unit (ICU)), or result in death. We stratified the population into five year age brackets. See Tables 1 and 2 for a listing of model parameters. Note, we have not included quarantining or household infection status in this depiction of the system.
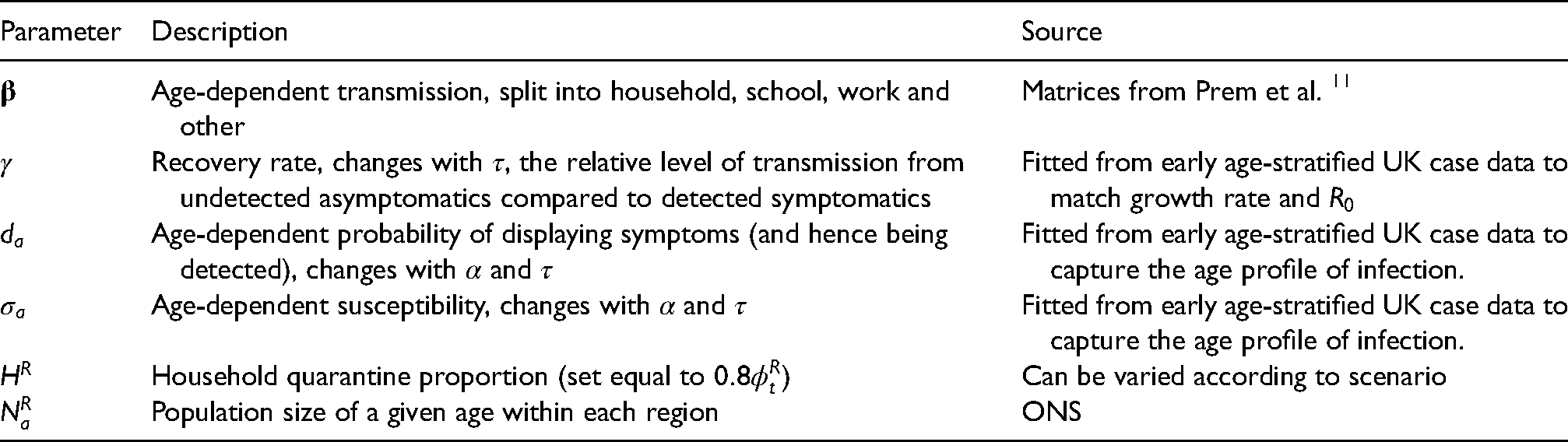
Description of key model parameters not fitted in the Markov Chain Monte Carlo (MCMC) and their source.

Description of key model parameters fitted in the MCMC.
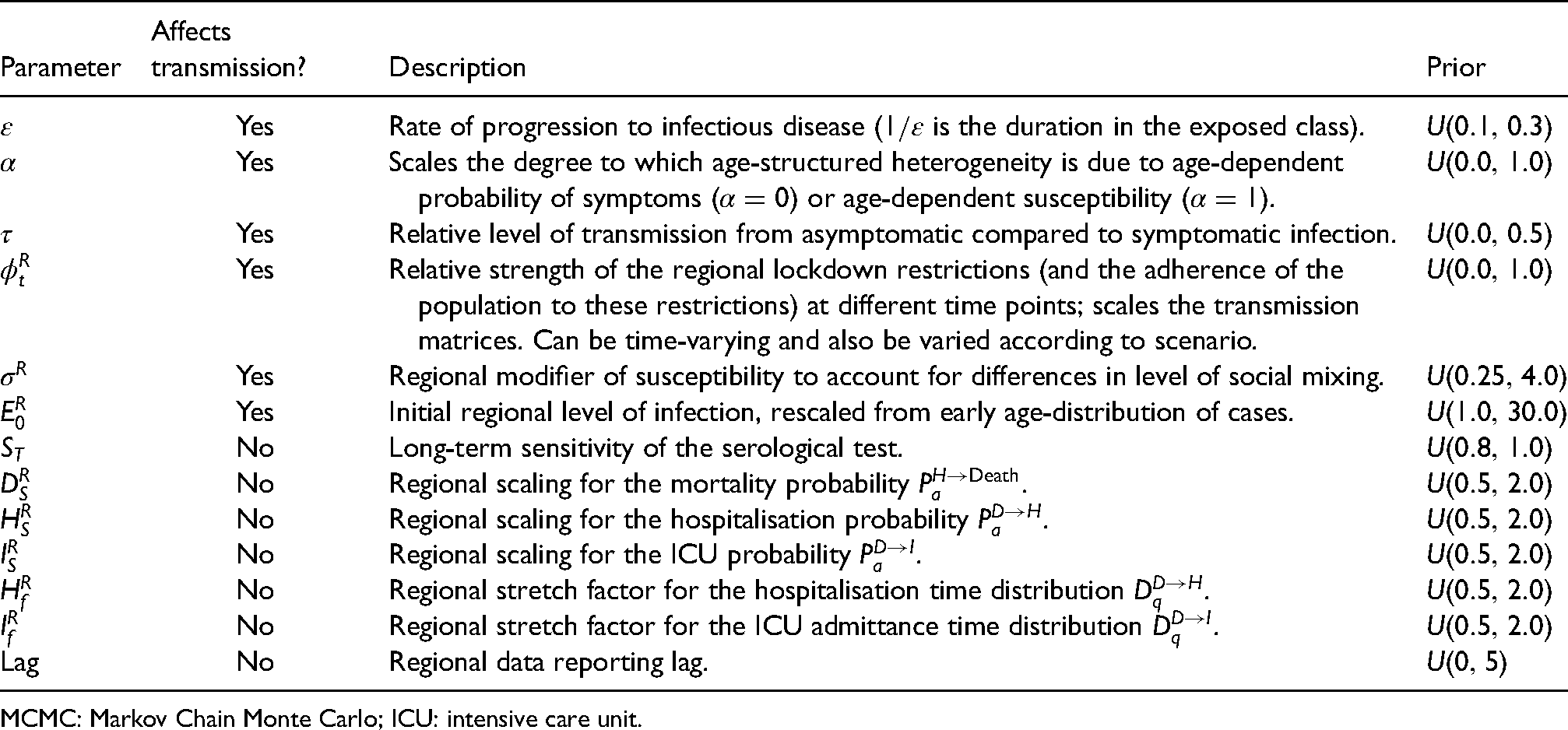

MCMC: Markov Chain Monte Carlo; ICU: intensive care unit.
The model is deterministic in structure, based on a large set of coupled ordinary differential equations (ODEs). The continuous results from these ODEs are never going to precisely match the discrete integer-valued data. We therefore assume that the observed data are an imprecise measure of the modelled quantities and represent this through a distribution (e.g. Poisson or Binomial) centred around the solution of the ODEs.
2.1 Model equations
We provide a description of the model parameters in Tables 1 and 2. The full system of ODEs for the model are given by: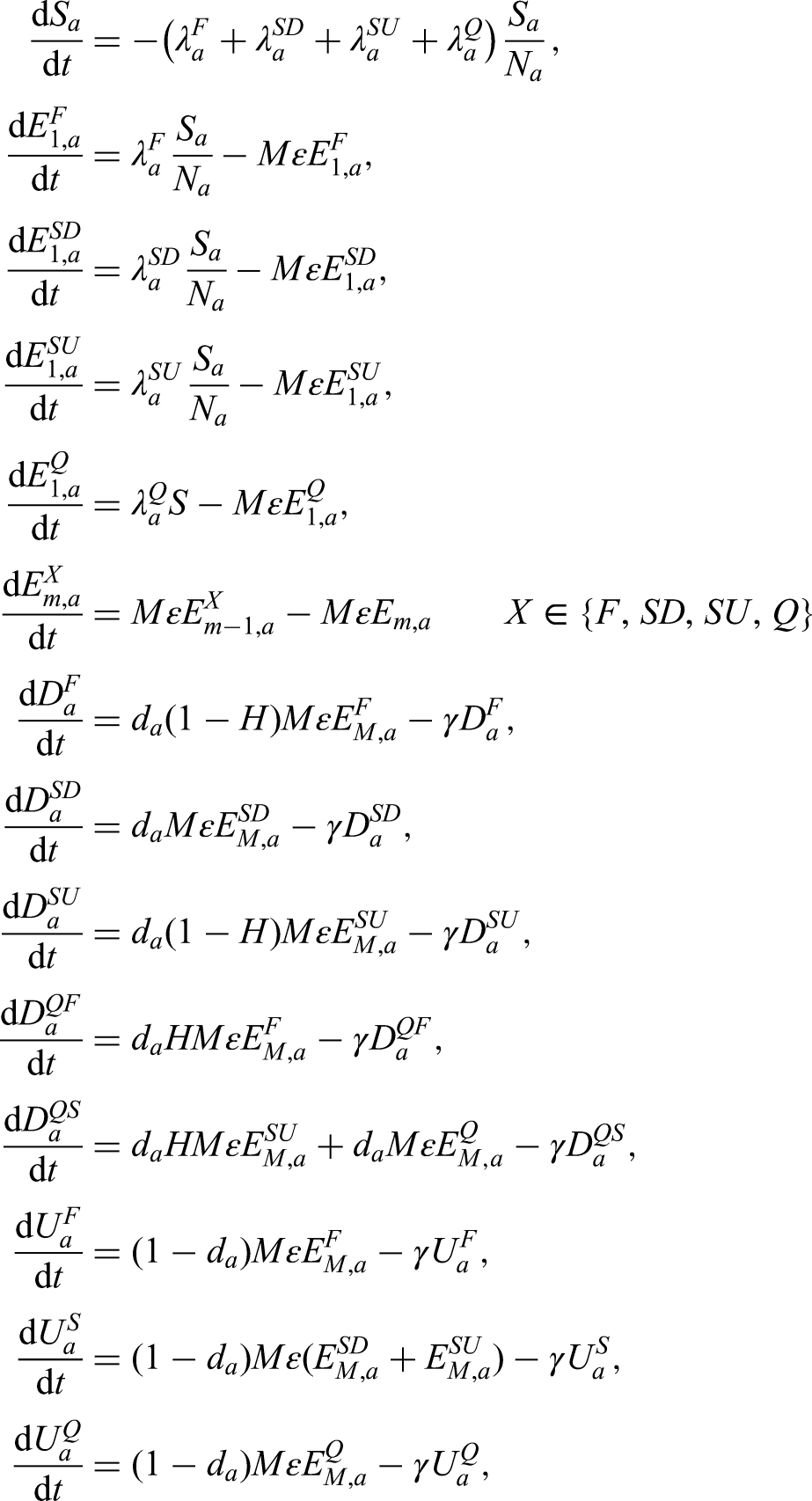
The forces of infection govern the non-linear transmission of infection. We partition the infectious pressure exerted on a given age group 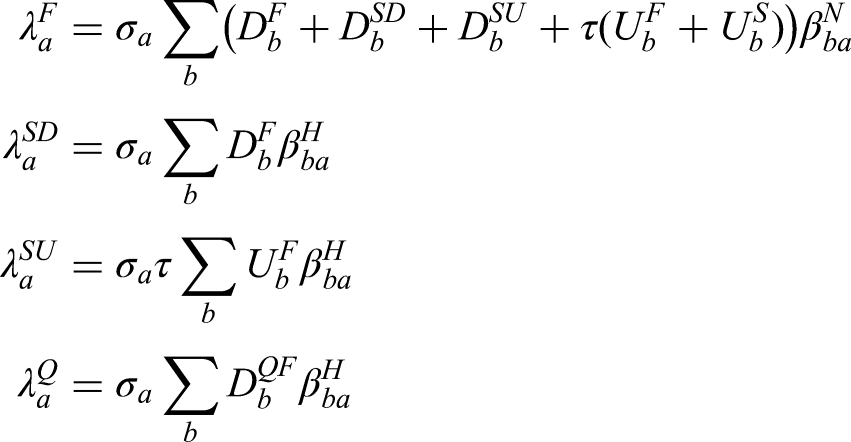
2.2 Amendments to within-household transmission
We wanted our model to be able to capture both individual-level quarantining and isolation of households with identified cases. In a standard ODE framework, the incorporation of the household structure increases the dimensionality of the system. Combined with the inclusion of other heterogeneities, such as age structure, the result can be a system whose dimensionality results in model calibration and simulation only being achievable at a large computational expense.13,14 Therefore, we instead make a number of approximations in our model to achieve a comparable effect.
We make the simplification that all within household transmission originates from the first infected individual within the household (denoted with a superscript
Given the novelty of the additional household structure that is included in this model, we clarify in more detail here the action of this formulation. We give a simpler set of equations (based on a standard 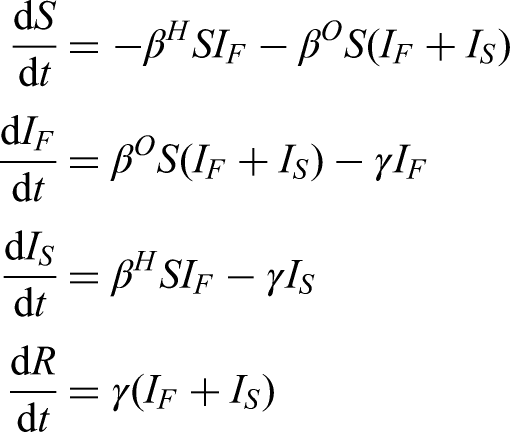
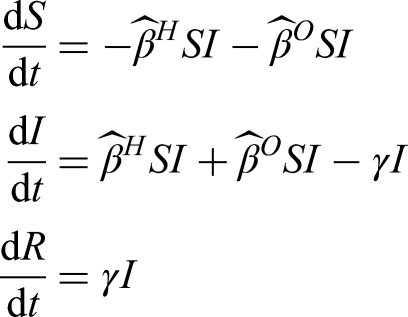
The early growth rate of the two models are
For both the simple household-structured model given here and the full COVID-19 model, the inclusion of additional household structure reduces the amount of within-household transmission compared to a model without household structure – as only the initial infection in each household (
2.3 Key model parameters
As with any model of this complexity, there are multiple parameters that determine the dynamics. Some of these are global parameters and apply to all geographical regions, with others used to capture the regional dynamics. Parameters that vary between regions are labelled with a superscript
2.4 Relationship between age-dependent susceptibility and detectability
We interlink age-dependent susceptibility,
Further, in a population that may be divided into a finite number of discrete categories according to a specific trait or traits (symptomatic and asymptomatic infection, for example), a next-generation approach can be used to relate the numbers of newly infected individuals in the various categories in consecutive generations.
15
Applying the next-generation approach to the symptomatic and asymptomatic infection states in our transmission model, the early dynamics would be specified by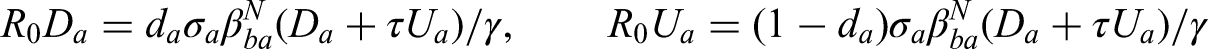
Throughout much of our work with this model, the values of
2.5 Regional heterogeneity in the dynamics
Throughout the current epidemic, there has been noticeable heterogeneity between the different regions of England and between the devolved nations. In particular, London is observed to have a large proportion of early cases and a relatively steeper decline in the subsequent lockdown than the other regions and the devolved nations. We capture this heterogeneity in our model through three estimated regional parameters that act on the heterogeneous population pyramid of each region.
Firstly, the initial level of infection in the region is re-scaled from the early age distribution of cases, with the regional scaling factor
3 Modelling social distancing
We obtained age-structured contact matrices for the United Kingdom from Prem et al.,
11
which we used to provide information on household transmission (
We assumed that the suite of social-distancing and lockdown measures acted in concert to reduce the work, school and other matrices while increasing the strength of household contacts. Two additional parameters that acted to modulate the contact structure were the relative strength of lockdown interventions,
We first capture the impact of social distancing by defining new transmission matrices (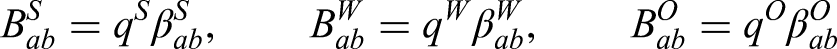
We used the assumed transmission matrices for a maximum lockdown scenario (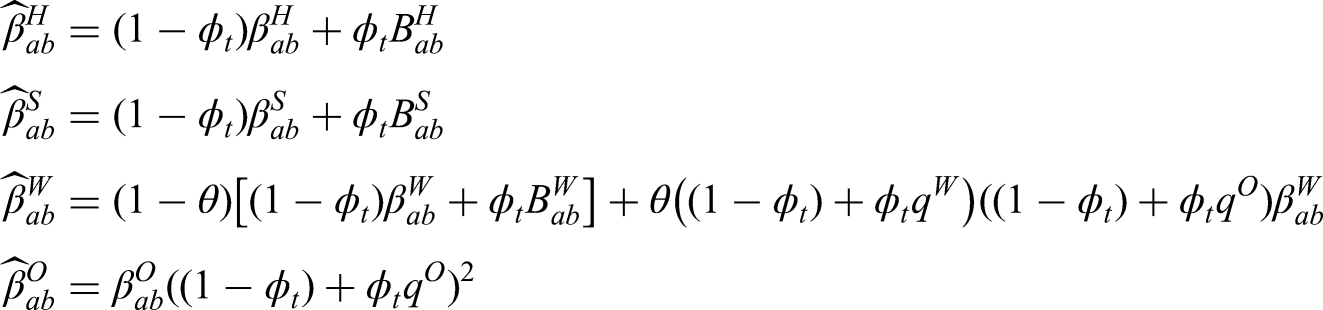
4 Public health measurable quantities
The main model equations focus on the epidemiological dynamics, allowing us to compute the number of symptomatic and asymptomatic infectious individuals over time. However, these quantities are not measured – and even the number of confirmed cases (the closest measure to symptomatic infections) is highly biased by the testing protocols at any given point in time. It is therefore necessary to convert infection estimates into quantities of interest that can be compared to data. We considered six such quantities which we calculated from the number of newly detectable symptomatic infections on a given day Hospital admissions: We assume that a fraction ICU admissions: Similarly, a fraction Hospital beds occupied: Individuals admitted to the hospital spend a variable number of days in the hospital. We therefore define two weightings, which determine if someone admitted to hospital still occupies a hospital bed ICU beds occupied: We similarly define Number of deaths: The mortality ratio Proportion testing seropositive: Seropositivity is a function of time since the onset of symptoms; we therefore define an increasing sigmoidal function which determines the probability that someone who first displayed symptoms
These nine distributions are all parameterised from individual patient data as recorded by the COVID-19 Hospitalisation in England Surveillance System (CHESS),
21
the ISARIC WHO Clinical Characterisation Protocol UK (CCP-UK) database sourced from the COVID-19 Clinical Information Network (CO-CIN),22,23 and the PHE sero-surveillance of blood donors.
20
CHESS data is used to define the probabilities of different outcomes (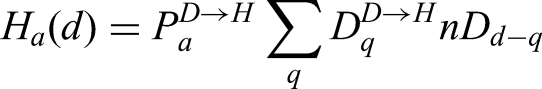
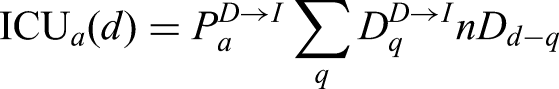
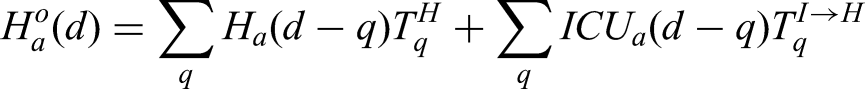
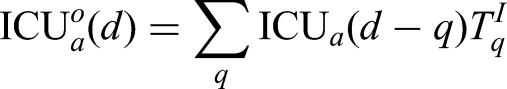
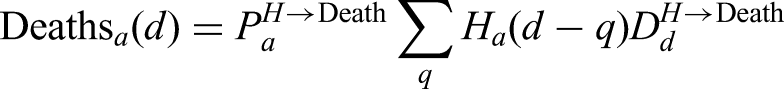
However, these distributions all represent a national average and do not therefore reflect regional differences. We therefore define regional scalings of the three key probabilities (
5 Likelihood function and the MCMC process
Multiple components form the likelihood function; most of which are based on a Poisson-likelihood. For brevity we define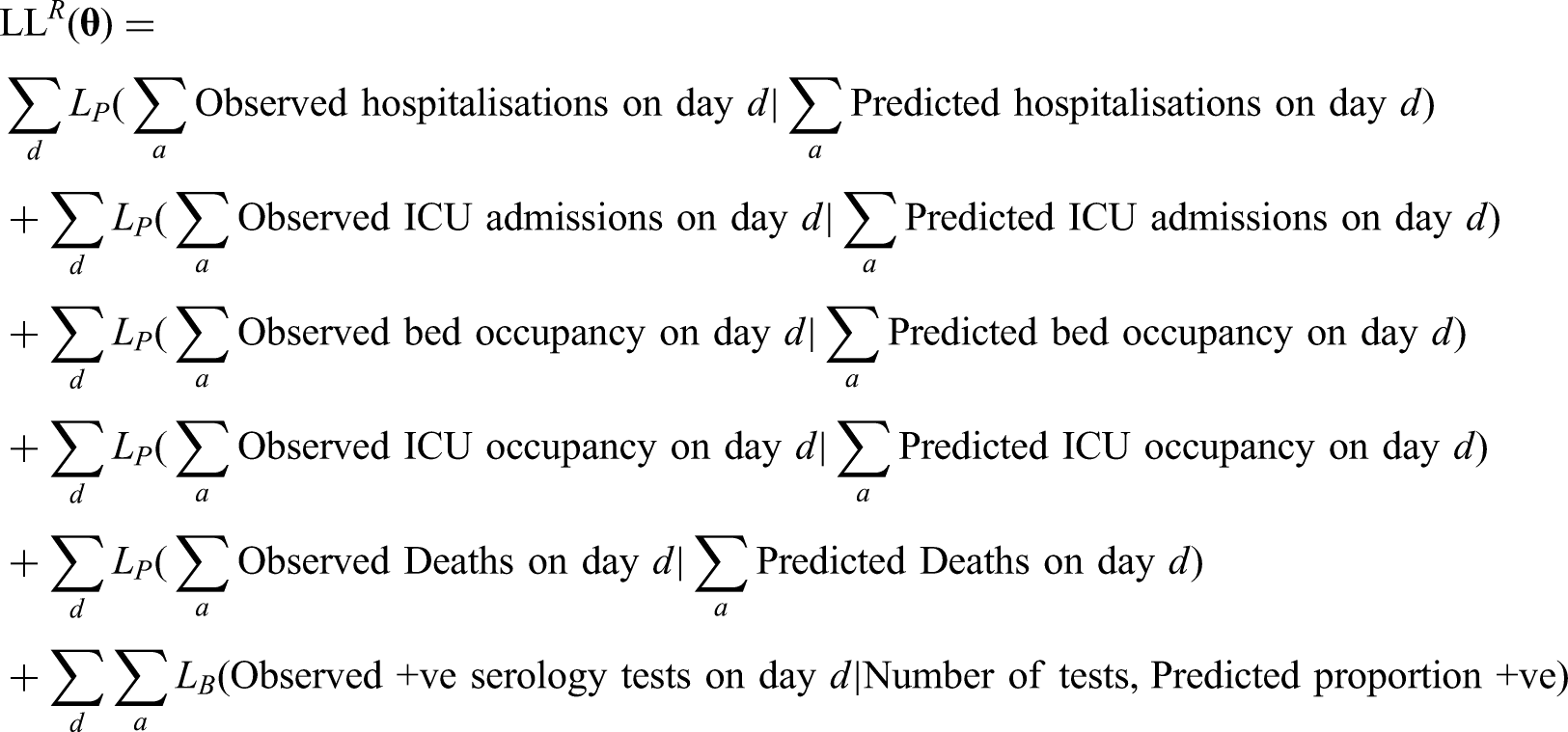 
New data are available on a daily time scale, and therefore inference needs to be repeated on a similar time scale. We can take advantage of this sequential refitting, taking random draws from the posteriors of the previous inference process to set the initial conditions for each chain, thus reducing the need for a long burn-in period.
6 Measuring the growth rate,
The growth rate,
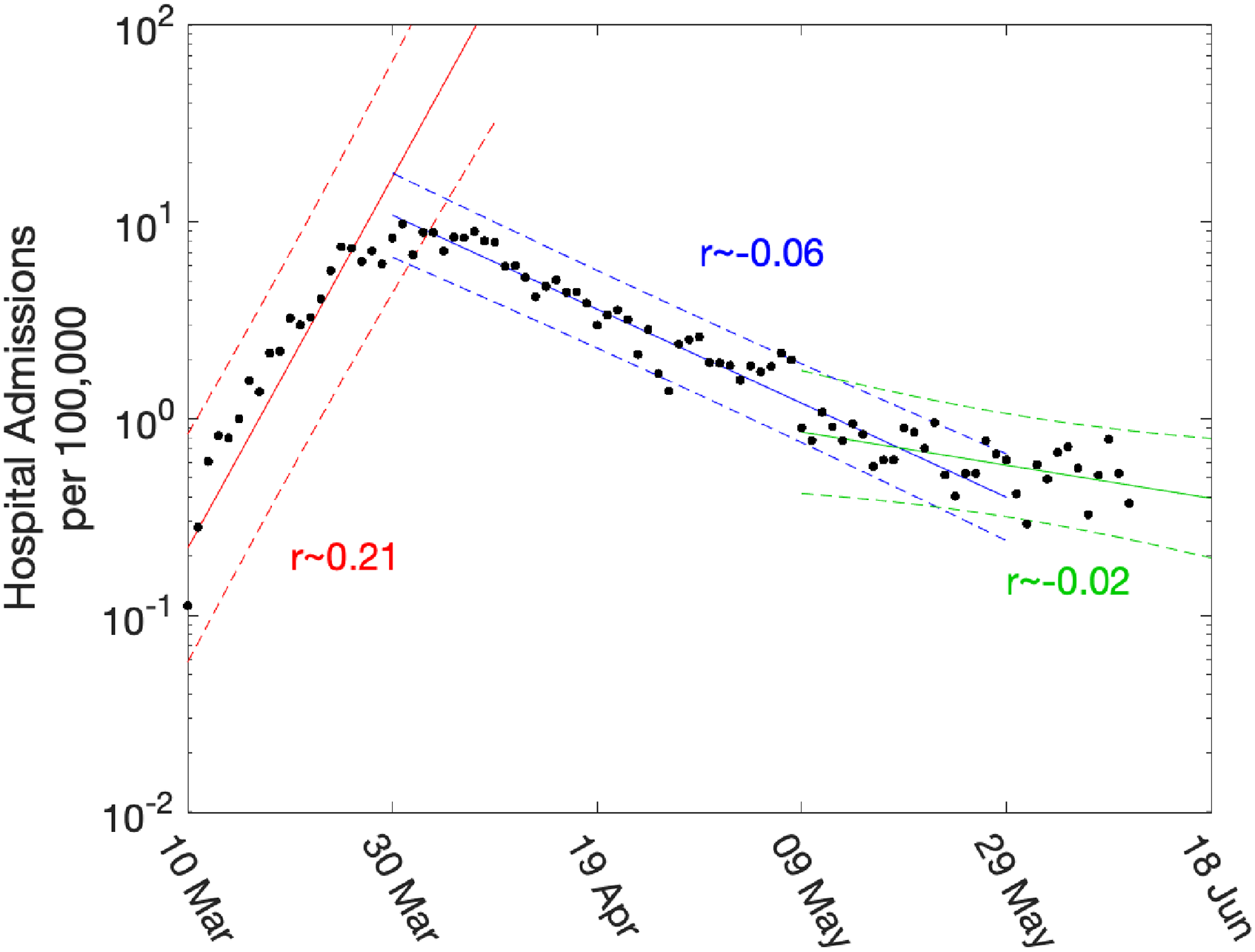
Daily hospital admissions per 100,000 individuals in London. Points show the number of daily admissions to the hospital (both in-patients testing positive and patients entering hospital following a positive test); results are plotted on a log scale. We show three simple fits to the data for pre-lockdown (red), strict-lockdown (blue) and relaxed-lockdown phases (green). Lines are linear fits to the logged data together with 95% confidence intervals, returning average growth rates of
While such statistically simple approaches are intuitively appealing, there are three main drawbacks. Firstly, they are not easily able to cope with the distributed delay between a change in policy (such as the introduction of the lockdown) and the impact of observable quantities (with the delay to deaths being multiple weeks). Secondly, they cannot readily utilise multiple data streams. Finally, they can only be used to extrapolate into the future – extending the period of exponential behaviour – they cannot predict the impact of further changes to the policy. Our approach is to instead fit the ODE model to multiple data streams, and then use the daily incidence to calculate the growth rate. Since we use a deterministic set of ODEs, the instantaneous growth rate
There has been a strong emphasis (especially in the UK) on the value of the reproductive number (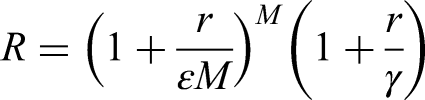
7 An evolving model framework
Unsurprisingly, the model framework has evolved during the epidemic as more data streams have become available and as we have gained a better understanding of epidemiology. Early models were largely based on the data from Wuhan and made relatively crude assumptions about the times from symptoms to hospitalisation and death. Later models incorporated more regional variation, while the PHE serology data in early May 2020 had a profound impact on model parameters.
Figure 3 shows how our short-term predictions (each of three weeks duration) changed over time, focusing on hospital admissions in London. It is clear that the early predictions were pessimistic about the reduction that would be generated by lockdown, although in part the higher values from early predictions are due to having identical parameters across all regions in the earliest models. In general later predictions, especially after the peak, are in far better agreement although the early inclusion of a step-change in the strength of the lockdown restrictions from 13 May 2020 (orange) led to substantial overestimation of future hospital admissions. Across all regions, we found some anomalous fits, which are due to changes in the way data were reported (Figures S3 and S4).
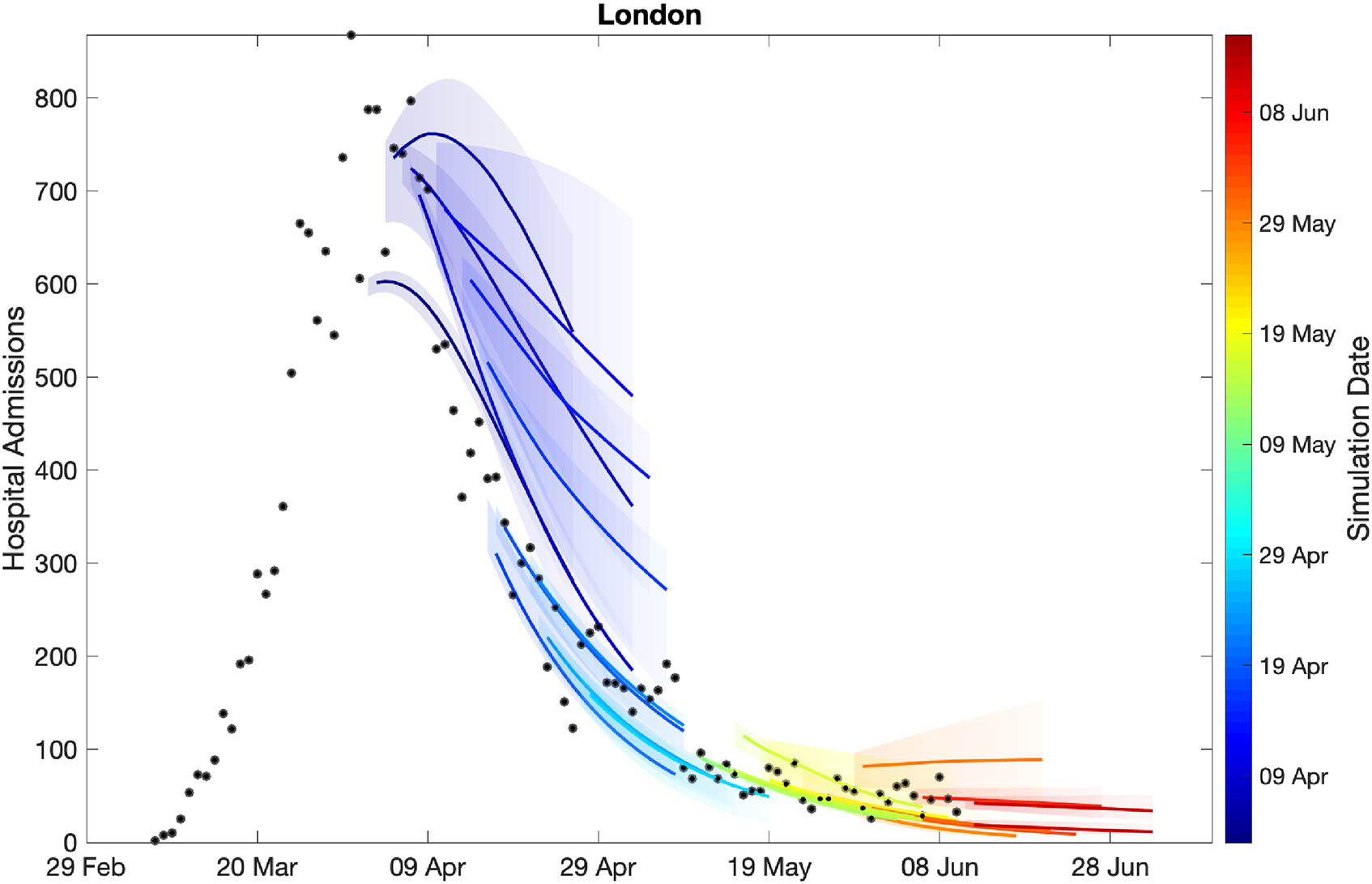
Sequential comparison of model results and data. For all daily hospital admissions with COVID-19 in London, we show the raw data (black dots) and a set of short-term predictions generated at different points during the outbreak. Changes to model fit reflect both improvements in model structure as well as increased amounts of data. The intervals represent our confidence in the fitted ODE model and do not account for either stochastic dynamics or the observational distribution about the deterministic predictions – which would generate far wider intervals.
The comparison of models and data over time can be made more formal by considering the mean squared error across the three-week prediction period for each region (Figure 4). We compare three time-varying quantities: (i) the mean value of the public health observable (in this case hospital deaths) in each region; (ii) the mean error between this data and the posterior set of ODE model predictions predicting forwards for three weeks; (iii) the mean error between the data and a simple moving average across the three time points before and after the data point. In each panel, the solid line corresponds to where the presented error statistic is equal to the mean, which is to be expected if the error originates from a Poisson distribution. The top left-hand graph in Figure 4 shows a clear linear relationship and correspondence in magnitude between the mean value and the error from the moving average, implying similarity between the variance and mean, giving support to our assumption (in the likelihood function) that the data are reasonably approximated as Poisson distributed. The other two graphs show how the error in the prediction has dropped over time from very high values for simulations in early April 2020 (when the impact of the lockdown was uncertain) to values in late May and June 2020 that are comparable with the error from the moving average.
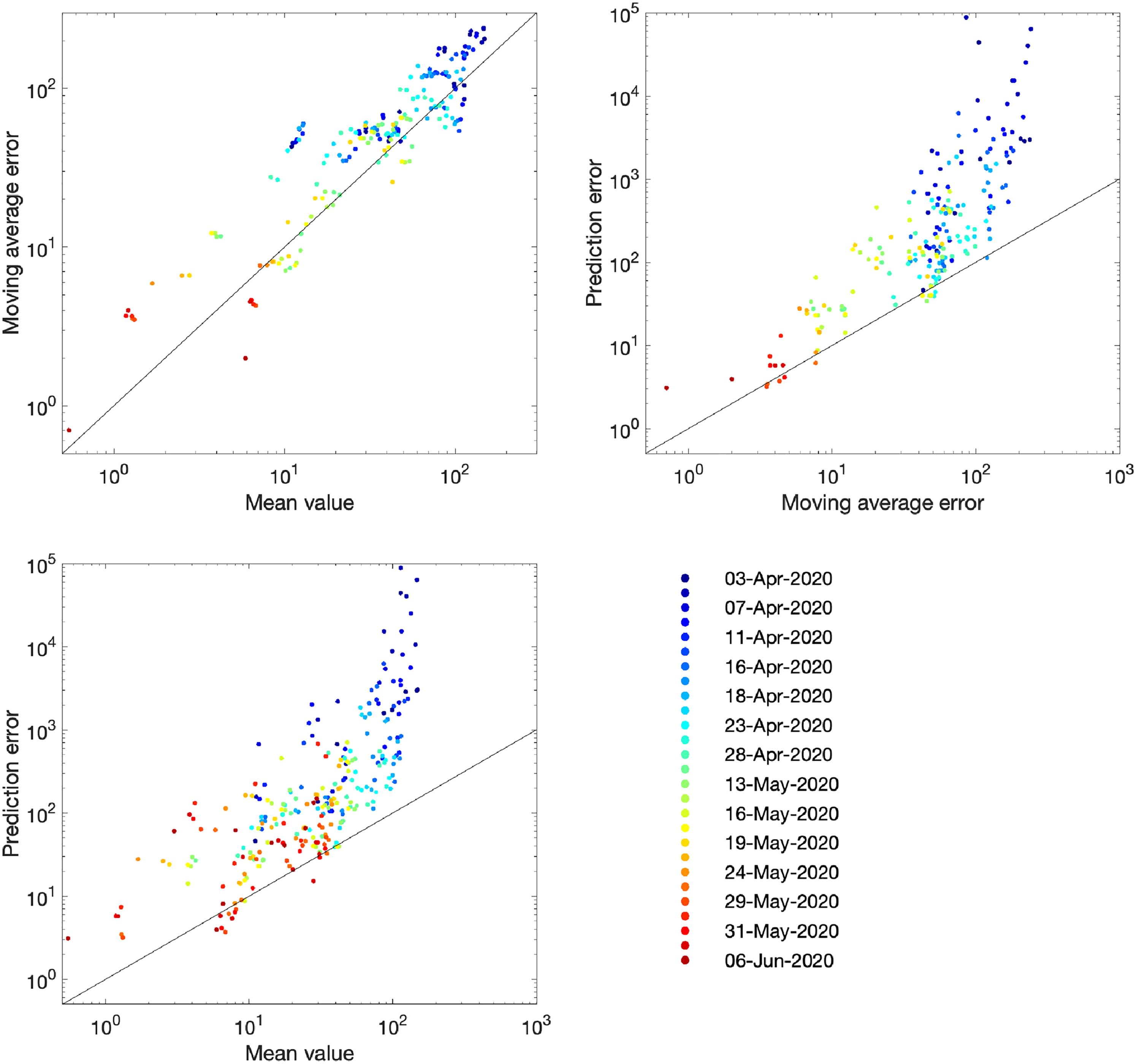
Improvement in fit over time for the number of hospital deaths. Each dot represents an analysis date (colour-coded) and region. For a data stream
8 Choice of data streams to inform the likelihood
The likelihood expression given above is an idealised measure and depends on all the observed data streams being available and unbiased. Unfortunately, ICU admission data had not been available and there were subtle differences in data streams between the devolved nations. An important question is therefore how key epidemiological quantities (and in particular the reproduction number
In high-dimensional systems with different temporal lags (see Figure S1), there are inevitably different time scales from when a change in policy or adherence occurs and when its impact is observed in key quantities. We briefly assess this problem in Figure 5, by considering the model output as surrogate data and examining how long a change in policy would take to impact the growth rate of key quantities. At time
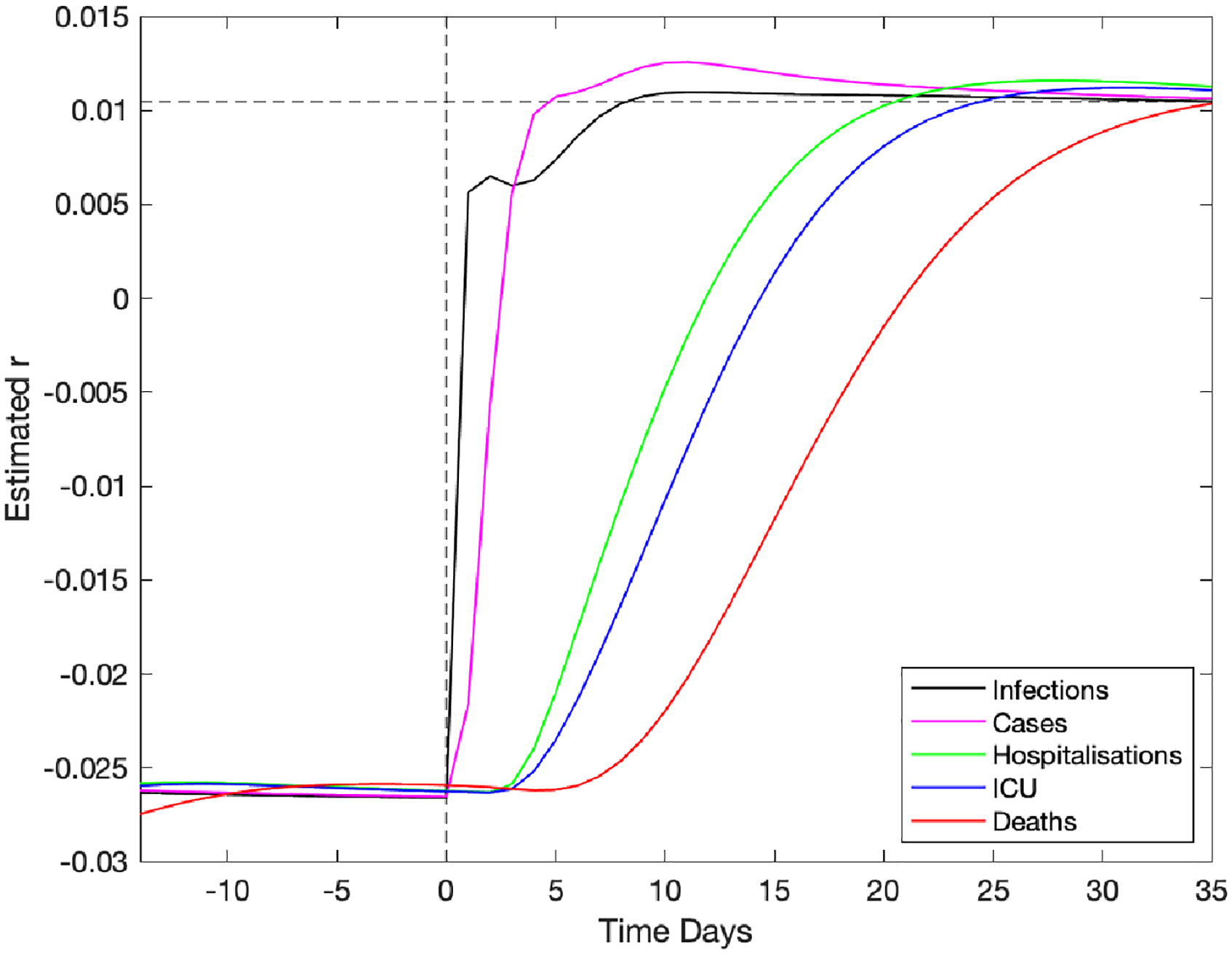
Impact of a change in underlying restrictions on the growth rate of modelled data streams. A change in the underlying restrictions occurs at time zero, taking the asymptotic growth rate
Figure 6 (left panel) shows the impact of using different observables for London (other regions are shown in Figure S5). This is achieved by only retaining a limited number of elements in the log-likelihood function, such that the model is matched to different combinations of data streams. Five different choices are shown: matching to recorded deaths only (using the date of death); matching to hospital admissions (both in-patients testing positive and admissions of individuals who have already tested positive); matching to bed occupancy, both hospital wards and ICU; matching to a combination of deaths and admissions; and finally matching to all data. Each of these different likelihood functions required an independent set of MCMC chains to be generated; from the associated posteriors, we consider an estimate of the instantaneous growth rate as the most important epidemiological characteristic. In general, we find that just using reported deaths produces the greatest spread of growth rates (
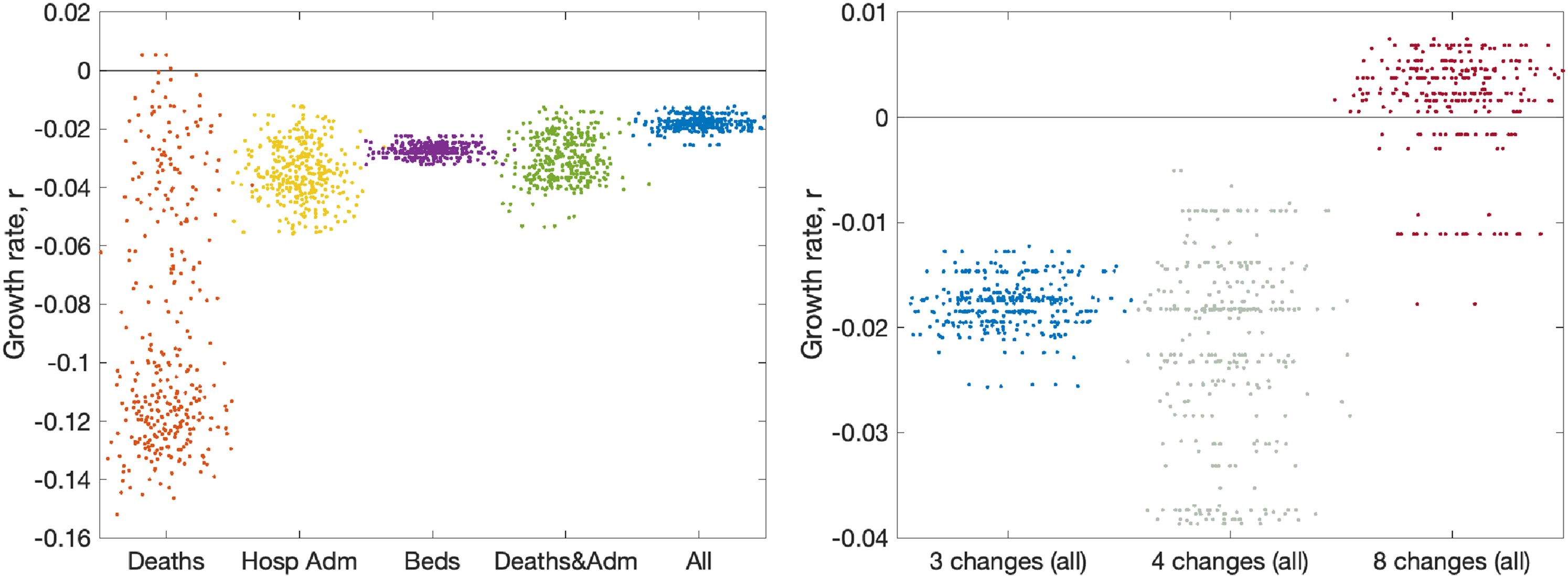
Impact of data streams and model structure on estimated growth rate. The growth rates are estimated using the predicted rate of change of new infections for London on 10 June 2020, with parameters inferred using data until 9 June 2020. The panels display posterior predictive distributions for the growth rate, where each data point corresponds to an estimate produced from a model simulation using a single parameter set sampled from the posterior distribution. To aid visualisation, we have applied a horizontal jitter to the data points. (a) The impact of restricting the inference to different data streams (deaths only, hospital admissions, hospital bed occupancy, deaths and admissions or all data); serology data was included in all inference. (b) The impact of having different numbers of lockdown phases (while using all the data); the default is three (as in Figure 2).
As mentioned in Section 7, the number of phases used to describe the reduction in transmission due to lockdown has changed as the situation, model and data evolved. The model began with just two phases; before and after lockdown. However, in late May 2020, following the policy changes on 13 May 2020, we explored having three phases. Having three phases is equivalent to assuming the same level of adherence to the lockdown and social-distancing measures throughout the epidemic, with changes in transmission occurring only due to the changing policy on 23 March 2020 and 13 May 2020. However, a different number of phases can be explored (Figure 6, right panel). Moving to four phases (with two equally spaced within the more relaxed lockdown) increases the variation, but does not have a substantial impact on the mean. Allowing eight phases (spaced every two weeks throughout lockdown) dramatically changes our estimation of the growth rate as the parameter inference responds more quickly to minor changes in observable quantities.
Lastly, it was noted in late May 2020 that one of the quantities used throughout the outbreak (number of daily hospital admissions) could lead to biased results. Hospital admissions for COVID-19 are comprised of two measures:
In-patients who test positive; this includes both individuals entering the hospital with COVID-19 symptoms who subsequently test positive, and hospital-acquired infections. Given that both of these elements feature in the hospital death data, it is difficult to separate them. Patients arriving at the hospital who have previously tested positive. In the early days of the outbreak, these were individuals who had been swabbed just prior to admission; however, in the later stages, there are many patients being admitted for non-COVID-related problems that have previously tested positive.
It seems prudent to remove this second element from our fitting procedure, although we note that for the devolved nations this separation into in-patients and new admissions is less clear. Removing this component of admissions also means that we cannot use the number of occupied beds as part of the likelihood, as these cannot be separated by the nature of admission. In Figure 7, we therefore compare the default fitting (used throughout this paper) with an updated method that uses in-patient admissions (together with deaths, ICU occupancy and serology when available). We observe that restricting the definition of hospital admission leads to a slight reduction in the growth rate
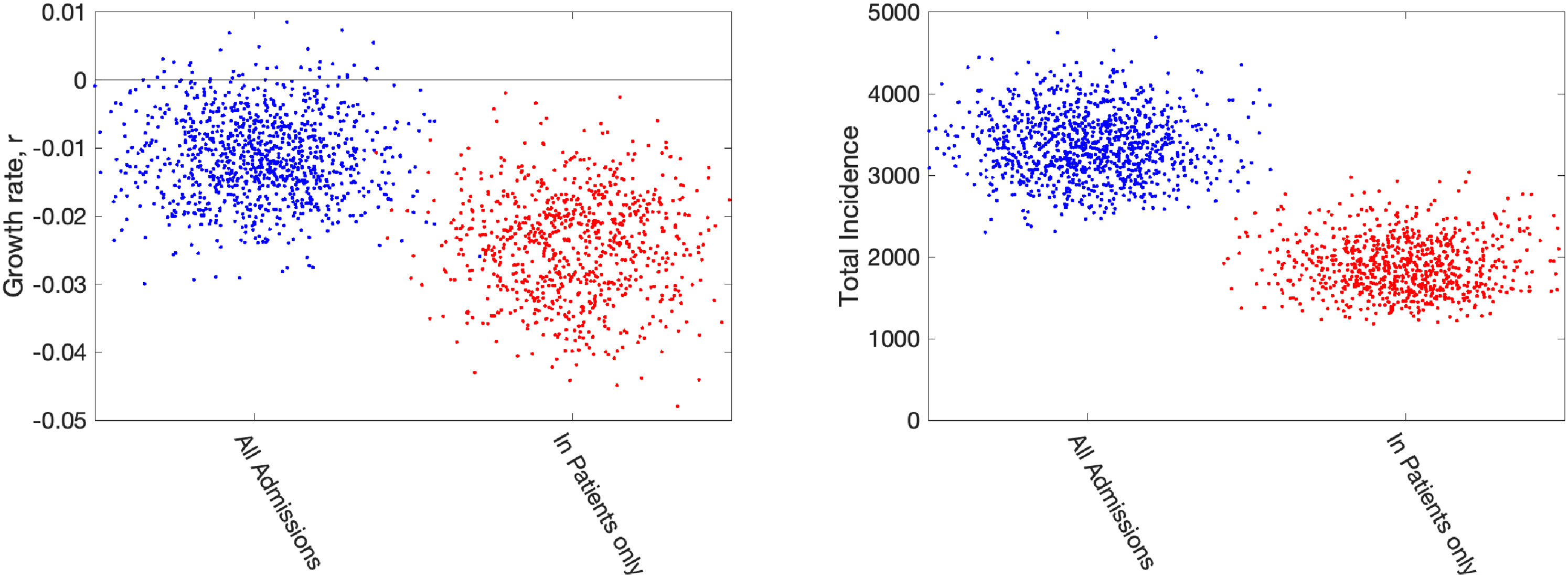
Impact of including different types of hospital admission in parameter inference. Growth rates and total incidence (asymptomatic and symptomatic) estimated from the ODE model for 10 June 2020 in London. The panels display posterior predictive distributions for the stated statistic, where each data point corresponds to an estimate produced from a model simulation using a single parameter set sampled from the posterior distribution. To aid visualisation, we have applied a horizontal jitter to the data points. In each panel, blue dots (on the left-hand side) give estimates when using all hospital admissions in the parameter inference (together with deaths, ICU occupancy and serology when available); red dots (on the right-hand side) represent estimates obtained using an alternative inference method that restricted to fitting to in-patient hospital admission data (together with deaths, ICU occupancy and serology when available). Parameters were inferred using data until 9 June 2020, while the growth rate
9 Fits and results at mid-June 2020
We now wish to compare how the fits made weekly (or more frequently) from late March to early June 2020 compare to later results. We note that this period also saw considerable development of the model structure as more data streams became available.
We used a fit to the data performed on 14 June 2020 (which matched to in-patient data, ICU occupancy, date of death records and serological results) to infer the change in NPIs and adherence,
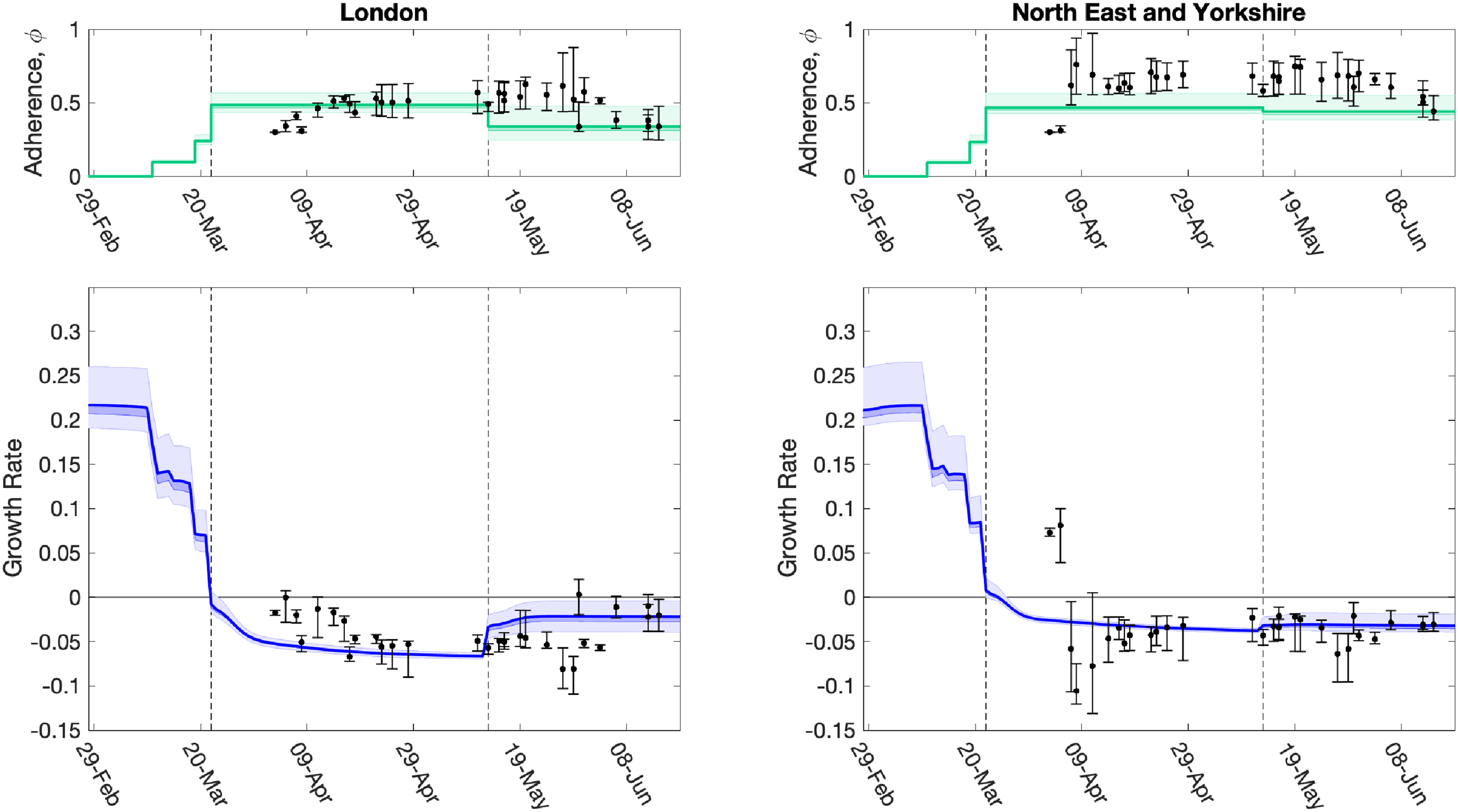
Evolution of the strength of interventions and adherence values (
The time profile of the predicted growth rate illustrates how the imposition of lockdown measures on 23 March 2020 led to
The relative strength of lockdown restrictions parameter,
Using parameters drawn from the posterior distributions, the model produces predictive posterior distributions for multiple health outcome quantities that have a strong quantitative correspondence to the regional observations (Figure 9). We recognise there was a looser resemblance to data on seropositivity, though salient features of the temporal profile are captured. In addition, short-term forecasts for each measure of interest have been made by continuing the simulation beyond the date of the final available data point, assuming that behaviour remains as of the final period (starting 13 May 2020).
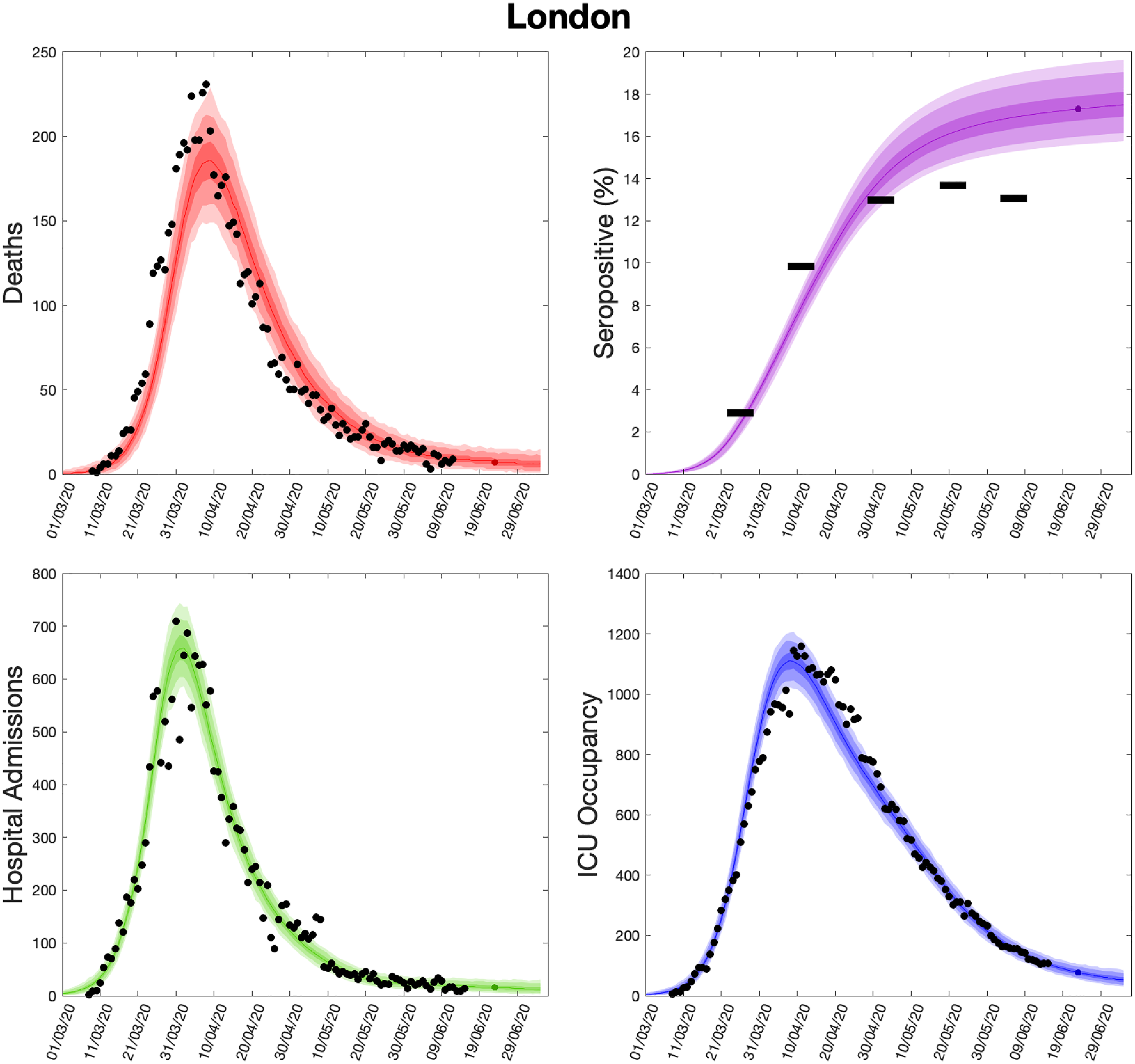
Health outcome predictions of the SARS-CoV-2 ODE transmission model from the beginning of the outbreak and three weeks into the future for London. (Top left) Daily deaths; (top right) seropositivity percentage; (bottom left) daily hospital admissions; (bottom right) ICU occupancy. In each panel: filled markers correspond to observed data, solid lines correspond to the mean outbreak over a sample of posterior parameters; shaded regions depict prediction intervals, with darker shading representing a narrower range of uncertainty (dark shading – 50%, moderate shading – 90%, light shading – 99%). The intervals represent our confidence in the fitted ODE model and do not account for either stochastic dynamics or the observational distribution about the deterministic predictions – which would generate far wider intervals. Predictions were produced using data up to 14 June 2020. SARS-CoV-2: severe acute respiratory syndrome coronavirus 2; ODE: ordinary differential equation; ICU: intensive care unit.
10 Discussion
In this study, we have provided an overview of the evolving MCMC inference scheme employed for calibrating the Warwick COVID-19 model 10 to the available health care, mortality and serological data streams. We have focused on the period May–June 2020, which corresponds to the first wave of the outbreak; a brief account of further refinements is given below. The work we describe was performed under extreme time pressures, working from limited initial knowledge and with data sources of varying quality. There are therefore assumptions in the model that with time and hindsight we have refined and compared to other more recently available data sources; similarly, the focus on hotspots of infection during the summer and the rise in cases into autumn 2020 has shaped much of our methodology. This article relates the model formulation that was used to understand the dynamics, predict cases and advise policy during the first wave.
A comparison of model short-term predictions and data over time (i.e. as the outbreak has progressed) demonstrated an observable decline in the error – suggesting that our model and inference methods have improved. We have considered in some detail the choice of data sets used to infer model parameters and the impact of this choice on the key emergent properties of the growth rate
It is important that uncertainty in the parameters governing the transmission dynamics, and its influence on predicted outcomes, be robustly conveyed. Without it, decision-makers will be missing meaningful information and may assume a false sense of precision. MCMC methodologies were a suitable choice for inferring parameters in our model framework, since we were able to evaluate the likelihood function quickly enough to make the approach feasible. However, a fundamental part of assessing whether your empirical estimates of the posterior distributions is valid and robust, which we have omitted to discuss thus far, is the use of MCMC convergence diagnostics. In the early stages of code development, we were checking convergence using the Gelman–Rubin test, 24 comparing intra-chain with inter-chain variability (a computable convergence diagnostic as throughout the fitting process we were generally using at least 15 independent chains). Yet, in the midst of a global public health emergency such as a pandemic, there are extremely short time scales over which the results need to be generated. In this instance, the speed of the epidemic meant that results needed to be produced every 2–3 days. Results were required even if full convergence had not been achieved, precluding any regular assessment of convergence and mixing – on the basis that a reasonable but imperfect answer was better than none. 25 These reflections signpost that attention should be paid to ensuring adequate research support is provided to permit the design of more robust and efficient ways of performing statistical inference for complex models in real-time.
Nevertheless, for some model formulations and data, it may not be possible to write down or evaluate the likelihood function. In these circumstances, an alternative approach to parameter inference is via simulation-based, likelihood-free methods, such as approximate Bayesian computation.26–28 We also recognise the appropriate mathematical structure of the model is also uncertain. Our methodology is formulated around deterministic differential equations that work well for large populations and significant levels of infection. On the other hand, stochastic effects are ignored and stochastic approaches may be needed when modelling low infection level regimes. In addition, a subset of our parameters had fixed values throughout our analyses, which means we may have underestimated the overall amount of parameter uncertainty.
As we gain a collective understanding of the SARS-CoV-2 virus and the COVID-19 disease it causes, the structure of infectious disease transmission models, the inference procedure and the use of data streams to underpin these models must continuously evolve. The evolution of the model through the early phase of the epidemic (up to June 2020) is documented here (Figures 3 and 8) and we feel it is meaningful to show this evolving process rather than simply present the final finished product. A vast body of work exists describing mathematical models for different infectious outbreaks and the associated parameter inference from epidemiological data. In most cases, however, these models are fitted retrospectively, using the entire data that have been collected during an outbreak. Fitting models with such hindsight is often far more accurate than predictions made in real-time. In the case when models are deployed during active epidemics, there are also additional challenges associated with the rapid flow of detailed and accurate data. Even if robust models and methods were available from the start of an outbreak, there are still significant delays in obtaining, processing and inferring parameters from new information. 5 This is particularly crucial as new interventions are introduced or significant policy changes occur, such as the relaxation of multiple NPIs during May, June and July of 2020 or the introduction of the nationwide ‘test and trace’ protocol. 29 Predicting the impact of such changes will inevitability be delayed by the lag between deployment and the effects on observable quantities (Figure 5) as well as the potential need to reformulate model structure or incorporate new data streams.
Multiple refinements to the model structure and approaches have been realised since June 2020 and more are still possible. The three biggest changes have been forced by external events: the rise and spread of the Alpha (B.1.1.7) variant during the latter part of 2020; the rise and spread of the Delta (B.1.617.2) in April and May of 2021 30 ; and the development and delivery of vaccines from December 2020 onwards. 31 The two variants have necessitated an increase in the dimension of the ODEs as at least two variants need to be modelled simultaneously (either Alpha out-competing wild type, or Delta out-competing Alpha); the two new variants also require the estimation of variant-specific parameters governing their relative transmission rates and the proportion of infected individuals that require hospital treatment or die from the disease. 32 The spread of these two variants is captured by looking at ‘S-gene failures’: the TaqPath system used to perform polymerase chain reaction (PCR) tests in many regions of the country fails to detect the S-gene of Alpha due to a point mutation. The rise of Alpha is therefore determined by the increase of S-gene failures, while the decline of Delta is captured by the decline of S-gene failures. Vaccination also requires a large number of parameters: in particular, the vaccine efficacy after one and two doses against infection, symptoms, severe illness and hospitalisation, death, and against both Alpha and Delta variants are needed within the model. We treat these additional parameters as inputs to the model, based on the estimations made by Public Health England. 33
Other changes to the model structure include using the proportion of community (known as Pillar 2) PCR samples that are positive rather than the number of positive tests. We feel that this proportion is less likely to be biased by changes in testing behaviour, and so provide a more stable estimate of the level of infection in the community. We also no longer use serology data from blood-donors, as again this is likely to suffer from a number of confounding factors. Instead, data from the national REACT 2 study
34
is incorporated into the likelihood and helps to anchor the total number of previously infected individuals in each region. More consideration has been given to detecting changes in the strength of social distancing (
Despite all of these improvements over the last year, there are still aspects that could be further improved. The understanding that infection may be partially driven by nosocomial transmission,35,36 while significant mortality is due to infection in care homes37,38 suggests that additional compartments capturing these components could greatly improve model realism if the necessary data were available throughout the course of the epidemic. Similarly, schools, universities and some workplaces pose additional risks, so there is merit in considering how these amplifiers of community infection could be incorporated within the general framework.39–41 Additionally, if in a regime with much lower levels of infection in the community, it may be prudent to adopt a stochastic model formulation at a finer spatial resolution to capture localised outbreak clusters, although the potential heterogeneity in local parameters may preclude accurate prediction at this scale.
In summary, if epidemiological models are to be used as part of the scientific discussion of controlling a disease outbreak it is vital that these models capture current biological understanding and are continually matched to all available data in real-time. Our work on COVID-19 presented here highlights some of the challenges with predicting a novel outbreak in a rapidly changing environment. Probably the greatest weakness is the time that it inevitably takes to respond – both in terms of developing the appropriate model and inference structure, and the mechanisms to process any data sources, but also in terms of delay between real-world changes and their detection within any inference scheme. Both of these can be shortened by well-informed preparations; having the necessary suite of models supported by the latest most efficient inference techniques could be hugely beneficial when rapid and robust predictive results are required.
Supplemental Material
sj-pdf-1-smm-10.1177_09622802211070257 - Supplemental material for Fitting to the UK COVID-19 outbreak, short-term forecasts and estimating the reproductive number
Supplemental material, sj-pdf-1-smm-10.1177_09622802211070257 for Fitting to the UK COVID-19 outbreak, short-term forecasts and estimating the reproductive number by Matt J. Keeling, Louise Dyson, Glen Guyver-Fletcher, Alex Holmes, Malcolm G Semple, , Michael J. Tildesley and Edward M. Hill in Statistical Methods in Medical Research
Footnotes
Acknowledgements
We acknowledge the support of Jeremy J Farrar, Nahoko Shindo, Devika Dixit, Nipunie Rajapakse, Piero Olliaro, Lyndsey Castle, Martha Buckley, Debbie Malden, Katherine Newell, Kwame O’Neill, Emmanuelle Denis, Claire Petersen, Scott Mullaney, Sue MacFarlane, Chris Jones, Nicole Maziere, Katie Bullock, Emily Cass, William Reynolds, Milton Ashworth, Ben Catterall, Louise Cooper, Terry Foster, Paul Matthew Ridley, Anthony Evans, Catherine Hartley, Chris Dunn, Debby Sales, Diane Latawiec, Erwan Trochu, Eve Wilcock, Innocent Gerald Asiimwe, Isabel Garcia-Dorival, J Eunice Zhang, Jack Pilgrim, Jane A Armstrong, Jordan J Clark, Jordan Thomas, Katharine King, Katie Alexandra Ahmed, Krishanthi S Subramaniam , Lauren Lett, Laurence McEvoy, Libby van Tonder, Lucia Alicia Livoti, Nahida S Miah, Rebecca K Shears, Rebecca Louise Jensen, Rebekah Penrice-Randal, Robyn Kiy, Samantha Leanne Barlow, Shadia Khandaker, Soeren Metelmann, Tessa Prince, Trevor R Jones, Benjamin Brennan, Agnieska Szemiel, Siddharth Bakshi, Daniella Lefteri, Maria Mancini, Julien Martinez, Angela Elliott, Joyce Mitchell, John McLauchlan, Aislynn Taggart, Oslem Dincarslan, Annette Lake, Claire Petersen, and Scott Mullaney.
Declaration of conflicting interest
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: MGS reports grants from DHSC NIHR UK, MRC UK, HPRU in Emerging and Zoonotic Infections, the University of Liverpool during the conduct of the study; other from Integrum Scientific LLC, Greensboro, NC, USA, outside the submitted work; the remaining authors declare no competing interests; no financial relationships with any organisations that might have an interest in the submitted work in the previous three years; and no other relationships or activities that could appear to have influenced the submitted work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Institute for Health Research (award CO-CIN-01), the Medical Research Council (grant MC_PC_19059) and by the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Emerging and Zoonotic Infections at the University of Liverpool in partnership with PHE, in collaboration with Liverpool School of Tropical Medicine and the University of Oxford (NIHR award 200907), Wellcome Trust and Department for International Development (215091/Z/18/Z), and the Bill and Melinda Gates Foundation (OPP1209135). The views expressed are those of the authors and not necessarily those of the DHSC, DID, NIHR, MRC, Wellcome Trust or PHE. Study registration ISRCTN66726260. MJK, LD, AH and MJT were supported by the Engineering and Physical Sciences Research Council through the MathSys CDT (grant number EP/S022244/1); MJK, LD, MJK and EMH were supported by the Medical Research Council through the COVID-19 Rapid Response Rolling Call (grant number MR/V009761/1); GG-F was supported by the Biotechnology and Biological Sciences Research Council through the MIBTP (BB/M01116X/1); MJK, LD and MJT were supported by UKRI through the JUNIPER modelling consortium (grant number MR/V038613/1). MJK is affiliated with the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Gastrointestinal Infections at the University of Liverpool in partnership with the UK Health Security Agency (UKHSA), in collaboration with the University of Warwick. MJK is also affiliated to the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Genomics and Enabling Data at the University of Warwick in partnership with UKHSA. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR, the Department of Health and Social Care or the UK Health Security Agency. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author contributions
Ethical considerations
Ethical approval was given by the South Central – Oxford C Research Ethics Committee in England (Ref. 13/SC/0149), the Scotland A Research Ethics Committee (Ref 20/SS/0028), and the WHO Ethics Review Committee (RPC571 and RPC572, 25 April 2013). Data from the CHESS database were supplied after anonymisation under strict data protection protocols agreed between the University of Warwick and Public Health England. The ethics of the use of these data for these purposes was agreed by Public Health England with the Government’s SPI-M(O) / SAGE committees.
Data availability
This work uses data provided by patients and collected by the NHS as part of their care and support #DataSavesLives. We are extremely grateful to the 2,648 frontline NHS clinical and research staff and volunteer medical students, who collected this data in challenging circumstances; and the generosity of the participants and their families for their individual contributions in these difficult times. The CO-CIN data was collated by ISARIC4C Investigators. ISARIC4C welcomes applications for data and material accessible through our Independent Data and Material Access Committee (https://isaric4c.net).
Data on cases were obtained from the CHESS data set that collects detailed data on patients infected with COVID-19. Data on COVID-19 deaths were obtained from Public Health England. These data contain confidential information, with public data deposition non-permissible for socioeconomic reasons. The CHESS data resides with the National Health Service (www.nhs.gov.uk) whilst the death data are available from Public Health England (![]() ).
).
Patient and public involvement
This was an urgent public health research study in response to a Public Health Emergency of International Concern. Patients or the public were not involved in the design, conduct, or reporting of this rapid response research.
ISARIC 4C Investigators
Consortium lead investigator: J Kenneth Baillie.
Chief investigator: Malcolm G Semple.
Co-lead investigator: Peter J M Openshaw.
ISARIC clinical coordinator: Gail Carson.
Co-investigators: Beatrice Alex, Benjamin Bach, Wendy S Barclay, Debby Bogaert, Meera Chand, Graham S Cooke, Annemarie B Docherty, Jake Dunning, Ana da Silva Filipe, Tom Fletcher, Christopher A Green, Ewen M Harrison, Julian A Hiscox, Antonia Ying Wai Ho, Peter W Horby, Samreen Ijaz, Saye Khoo, Paul Klenerman, Andrew Law, Wei Shen Lim, Alexander, J Mentzer, Laura Merson, Alison M Meynert, Mahdad Noursadeghi, Shona C Moore, Massimo Palmarini, William A Paxton, Georgios Pollakis, Nicholas Price, Andrew Rambaut, David L Robertson, Clark D Russell, Vanessa Sancho-Shimizu, Janet T Scott, Louise Sigfrid, Tom Solomon, Shiranee Sriskandan, David Stuart, Charlotte Summers, Richard S Tedder, Emma C Thomson, Ryan S Thwaites, Lance CW Turtle, Maria Zambon. Project Managers Hayley Hardwick, Chloe Donohue, Jane Ewins, Wilna Oosthuyzen and Fiona Griffiths.
Data analysts: Lisa Norman, Riinu Pius, Tom M Drake, Cameron J Fairfield, Stephen Knight, Kenneth A Mclean, Derek Murphy and Catherine A Shaw.
Data and information system manager: Jo Dalton, Michelle Girvan, Egle Saviciute, Stephanie Roberts Janet Harrison, Laura Marsh and Marie Connor.
Data integration and presentation: Gary Leeming, Andrew Law and Ross Hendry.
Material management: William Greenhalf, Victoria Shaw and Sarah McDonald.
Outbreak laboratory volunteers: Katie A Ahmed, Jane A Armstrong, Milton Ashworth, Innocent G Asiimwe, Siddharth Bakshi, Samantha L Barlow, Laura Booth, Benjamin Brennan, Katie Bullock, Benjamin WA Catterall, Jordan J Clark, Emily A Clarke, Sarah Cole, Louise Cooper, Helen Cox, Christopher Davis, Oslem Dincarslan, Chris Dunn, Philip Dyer, Angela Elliott, Anthony Evans, Lewis WS Fisher, Terry Foster, Isabel Garcia-Dorival, Willliam Greenhalf, Philip Gunning, Catherine Hartley, Antonia Ho, Rebecca L Jensen, Christopher B Jones, Trevor R Jones, Shadia Khandaker, Katharine King, Robyn T Kiy, Chrysa Koukorava, Annette Lake, Suzannah Lant, Diane Latawiec, L Lavelle-Langham, Daniella Lefteri, Lauren Lett, Lucia A Livoti, Maria Mancini, Sarah McDonald, Laurence McEvoy, John McLauchlan, Soeren Metelmann, Nahida S Miah, Joanna Middleton, Joyce Mitchell, Shona C Moore, Ellen G Murphy, Rebekah Penrice-Randal, Jack Pilgrim, Tessa Prince, Will Reynolds, P Matthew Ridley, Debby Sales, Victoria E Shaw, Rebecca K Shears, Benjamin Small, Krishanthi S Subramaniam, Agnieska Szemiel, Aislynn Taggart, Jolanta Tanianis, Jordan Thomas, Erwan Trochu, Libby van Tonder, Eve Wilcock and J Eunice Zhang.
Local principal investigators: Kayode Adeniji, Daniel Agranoff, Ken Agwuh, Dhiraj Ail, Ana Alegria, Brian Angus, Abdul Ashish, Dougal Atkinson, Shahedal Bari, Gavin Barlow, Stella Barnass, Nicholas Barrett, Christopher Bassford, David Baxter, Michael Beadsworth, Jolanta Bernatoniene, John Berridge, Nicola Best, Pieter Bothma, David Brealey, Robin Brittain-Long, Naomi Bulteel, Tom Burden, Andrew Burtenshaw, Vikki Caruth, David Chadwick, Duncan Chambler, Nigel Chee, Jenny Child, Srikanth Chukkambotla, Tom Clark, Paul Collini, Catherine Cosgrove, Jason Cupitt, Maria-Teresa Cutino-Moguel, Paul Dark, Chris Dawson, Samir Dervisevic, Phil Donnison, Sam Douthwaite, Ingrid DuRand, Ahilanadan Dushianthan, Tristan Dyer, Cariad Evans, Chi Eziefula, Chrisopher Fegan, Adam Finn, Duncan Fullerton, Sanjeev Garg, Sanjeev Garg, Atul Garg, Jo Godden, Arthur Goldsmith, Clive Graham, Elaine Hardy, Stuart Hartshorn, Daniel Harvey, Peter Havalda, Daniel B Hawcutt, Maria Hobrok, Luke Hodgson, Anita Holme, Anil Hormis, Michael Jacobs, Susan Jain, Paul Jennings, Agilan Kaliappan, Vidya Kasipandian, Stephen Kegg, Michael Kelsey, Jason Kendall, Caroline Kerrison, Ian Kerslake, Oliver Koch, Gouri Koduri, George Koshy, Shondipon Laha, Susan Larkin, Tamas Leiner, Patrick Lillie, James Limb, Vanessa Linnett, Jeff Little, Michael MacMahon, Emily MacNaughton, Ravish Mankregod, Huw Masson , Elijah Matovu, Katherine McCullough, Ruth McEwen , Manjula Meda, Gary Mills , Jane Minton, Mariyam Mirfenderesky, Kavya Mohandas, Quen Mok, James Moon, Elinoor Moore, Patrick Morgan, Craig Morris, Katherine Mortimore, Samuel Moses, Mbiye Mpenge, Rohinton Mulla, Michael Murphy, Megan Nagel, Thapas Nagarajan, Mark Nelson, Igor Otahal, Mark Pais, Selva Panchatsharam, Hassan Paraiso, Brij Patel, Justin Pepperell, Mark Peters, Mandeep Phull, Stefania Pintus, Jagtur Singh Pooni, Frank Post, David Price, Rachel Prout, Nikolas Rae, Henrik Reschreiter, Tim Reynolds, Neil Richardson, Mark Roberts, Devender Roberts, Alistair Rose, Guy Rousseau, Brendan Ryan, Taranprit Saluja, Aarti Shah, Prad Shanmuga, Anil Sharma, Anna Shawcross, Jeremy Sizer, Richard Smith, Catherine Snelson, Nick Spittle, Nikki Staines , Tom Stambach, Richard Stewart, Pradeep Subudhi, Tamas Szakmany, Kate Tatham, Jo Thomas, Chris Thompson, Robert Thompson, Ascanio Tridente, Darell Tupper-Carey, Mary Twagira, Andrew Ustianowski, Nick Vallotton, Lisa Vincent-Smith, Shico Visuvanathan, Alan Vuylsteke, Sam Waddy, Rachel Wake, Andrew Walden, Ingeborg Welters, Tony Whitehouse, Paul Whittaker, Ashley Whittington, Meme Wijesinghe, Martin Williams, Lawrence Wilson, Sarah Wilson, Stephen Winchester, Martin Wiselka, Adam Wolverson, Daniel G Wooton, Andrew Workman, Bryan Yates and Peter Young.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.