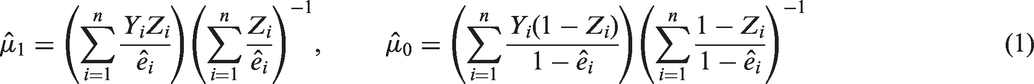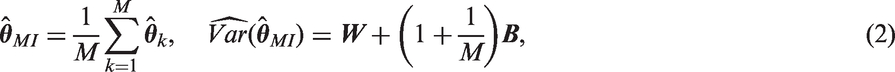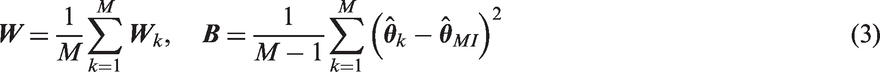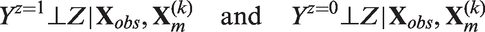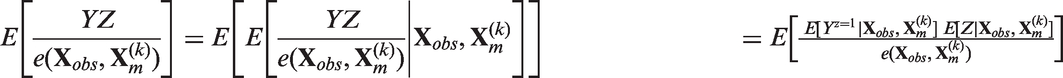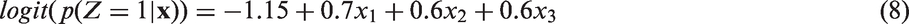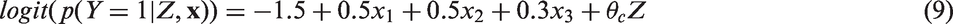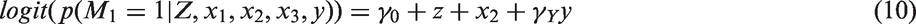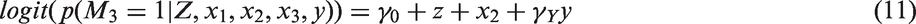Abstract
Inverse probability of treatment weighting is a popular propensity score-based approach to estimate marginal treatment effects in observational studies at risk of confounding bias. A major issue when estimating the propensity score is the presence of partially observed covariates. Multiple imputation is a natural approach to handle missing data on covariates: covariates are imputed and a propensity score analysis is performed in each imputed dataset to estimate the treatment effect. The treatment effect estimates from each imputed dataset are then combined to obtain an overall estimate. We call this method MIte. However, an alternative approach has been proposed, in which the propensity scores are combined across the imputed datasets (MIps). Therefore, there are remaining uncertainties about how to implement multiple imputation for propensity score analysis: (a) should we apply Rubin’s rules to the inverse probability of treatment weighting treatment effect estimates or to the propensity score estimates themselves? (b) does the outcome have to be included in the imputation model? (c) how should we estimate the variance of the inverse probability of treatment weighting estimator after multiple imputation? We studied the consistency and balancing properties of the MIte and MIps estimators and performed a simulation study to empirically assess their performance for the analysis of a binary outcome. We also compared the performance of these methods to complete case analysis and the missingness pattern approach, which uses a different propensity score model for each pattern of missingness, and a third multiple imputation approach in which the propensity score parameters are combined rather than the propensity scores themselves (MIpar). Under a missing at random mechanism, complete case and missingness pattern analyses were biased in most cases for estimating the marginal treatment effect, whereas multiple imputation approaches were approximately unbiased as long as the outcome was included in the imputation model. Only MIte was unbiased in all the studied scenarios and Rubin’s rules provided good variance estimates for MIte. The propensity score estimated in the MIte approach showed good balancing properties. In conclusion, when using multiple imputation in the inverse probability of treatment weighting context, MIte with the outcome included in the imputation model is the preferred approach.
Keywords
1 Introduction
Data from observational studies provide useful information to address health-related questions and notably estimate treatment effects in real settings. 1 However, because individuals are not randomised, the treatment groups are often not comparable, which may lead to confounding bias 2 if these studies are analysed without appropriate adjustment for confounding. Propensity scores (PS) have been proposed as a means to recover balance between groups on observed covariates and so obtain a consistent estimate of the causal treatment effect. 3 The PS is defined as the individual’s probability of receiving the treatment rather than the control given their baseline characteristics. 4 One popular method to achieve covariate balance between treatment groups is to weight individuals by the inverse of their PS. This approach, known as inverse probability of treatment weighting (IPTW), 5 aims to emulate the sample that would have been observed in a randomised trial.
In practice, a major issue when estimating the PS is the presence of partially observed covariates, as the PS cannot be estimated for individuals with at least one missing covariate value. Standard analyses include complete case analysis 6 or the missingness pattern approach, 7 but a popular alternative to handle missing data is multiple imputation (MI). 8 However, there remain four important unresolved questions about how to implement MI for PS analysis.
First, two approaches to combine information from the imputed datasets have been proposed in the context of PS analyses: combining the treatment effects estimated on each imputed dataset (method called MIte hereafter) or combining for each individual their PS value across the imputed datasets (method called MIps hereafter).9,10 MIte intuitively seems to adhere best to the MI philosophy by applying the full analysis strategy on each imputed dataset. Furthermore, Seaman and White 11 proved that for an infinite number of imputations, the MIte estimator is consistent. However, Mitra and Reiter 9 published recommendations on the use of MI for PS analysis, and they advocated using MIps, rather than MIte, for PS matching.
The second question is whether or not to include the outcome in the imputation model for the missing covariates. Mitra and Reiter did not include it in their study, 9 which could explain the bias they observed for both MIte and MIps. One of the advantages claimed for the PS approach in general is that it allows the investigator to use the data on the treatment and covariates to develop a well-fitting PS model without needing to look at the data on the outcome. 9 This makes it possible to avoid the temptation to search for a PS model that gives a significant treatment effect estimate, a temptation which may be experienced by an analysis which handles confounding by using regression adjustment. Intuition may therefore lead one to believe that imputation of missing covariates should also be done without using the outcome variable. 9 However, this intuition conflicts with advice to include the outcome when imputing missing covariates in a regression model whose parameters are the quantities of interest. 12
The third question is how to estimate the variance of the IPTW estimator after MI. When pooling the treatment effects (MIte), Seaman and White 11 showed that Rubin’s rule for estimating the variance performs well in practice, although theoretical justification for Rubin’s rules relies on the parameter of interest being estimated with maximum likelihood, which is not the case for the IPTW method. For MIps, there is, to our knowledge, no variance estimator that takes into account both the uncertainty due to the estimation of the PS given the complete data and that due to the imputation of the missing data.
Finally, the fourth question is how well MI performs in comparison to other popular approaches to handle missing data, namely complete case (CC) analysis or the missingness pattern (MP) approach. Qu and Lipkovitch 13 and Seaman and White 11 assessed the performance of MIte additionally including the missingness pattern indicator in the PS model, but they did not compare this approach to other combination rules after MI or to the MP approach alone.
In this article, we show that the best method when using MI with IPTW is to include the outcome in the PS model and use Rubin’s rules to combine the treatment effect estimates from the imputed datasets (the method MIte). An informal survey of recent articles using MI with IPTW revealed that suboptimal methods are commonly being used. In particular, several studies have following Mitra and Reiter’s recommendation to use MIps rather than MIte, and in several of these studies imputation is done without including the outcome in the imputation model (e.g. literature14–20). Another example of suboptimal MI is that of Hayes and Groner, 21 who randomly selected one imputation for each individual and estimated that individual’s PS based on just this single imputation. Therefore, we conducted this work to address the unresolved questions and provide new recommendations.
This article is organised as follows: we present a motivating example looking at the effect of statins on short-term mortality after pneumonia in Section 2. A description of IPTW and its underlying assumptions is given in Section 3. Section 4 presents different strategies to handle partially missing covariates for PS analysis. The consistency and balancing properties of the MI approaches are studied in Section 5 and empirically assessed in Sections 6 and 7 through a simulation study. The application of these approaches to the statin motivating example is presented in Section 8, followed by a discussion in Section 9.
2 Motivating example: Effect of statin use on short-term mortality after pneumonia
To illustrate the importance of an adequate handling of missing covariates for PS analysis, we focused on a published study of the effect of statin use on short-term mortality after pneumonia. 22 We utilised the THIN database, which consists of anonymised patient records from general practitioners (GPs) in the UK. As of the end of 2015, the database represented 3.5 million unique active patients, or approximately 6% of the UK population. The database has been found to be broadly representative of the UK population, and the validity of recorded information has been established in previous studies.23,24 Douglas et al. carried out an analysis of 9073 patients who had a pneumonia episode, of whom 1398 were under statin treatment when pneumonia was diagnosed. In the statin group, 305 patients (21.8%) died within 6 months, while 2839 (37.0%) of the non-users died within 6 months. However, statin users and non users were very different in terms of characteristics, in particular on characteristics associated with mortality.
In Douglas et al., PSs were used to recover balance between groups. However, three important potential confounders were only partially observed: body mass index (BMI), smoking status and alcohol consumption, with respectively 19.2%, 6.2% and 18.5% of missing data. In the original analysis, a missing indicator method was used. This approach is similar to the missingness pattern approach described later in this article. We will explore whether handling the missing data using MI gives results that differ from those given by Douglas et al.
3 IPTW
3.1 PS estimation and assumptions
PSs have become a major tool in causal inference to estimate the causal effect of a binary treatment Z (Z = 1 if treated, Z = 0 otherwise) on an outcome in the presence of confounding.
3
The PS is the individual’s probability of receiving the treatment conditional on the individual’s values on a set of baseline covariates X.
25
The PS is usually estimated from the data using a logistic regression model, which predicts each individual’s probability of receiving the treatment from their baseline covariates.
26
The PS approach is best understood using the potential outcomes framework,
27
in which the causal effect of the treatment is defined as the difference between the two potential outcomes
The key property of the PS is that it is a balancing score. That is, if assumptions (a) to (c) are valid and the PS model is correctly specified, the variables included in the PS model are balanced between treatment groups at any level of the PS, i.e. they have the same conditional distribution in both groups given the PS. This leads to the consistency of PS-based estimators. Initially, three different PS-based approaches were proposed 3 : PS matching, subclassification and covariate adjustment. Although PS matching is the most common approach nowadays, matching often discards a substantial number of individuals from the analysis 30 and variance estimation after PS matching is not straightforward. 31 The two other approaches have drawbacks as well: residual bias due to heterogeneity within strata can remain with subclassification, 7 whereas covariate adjustment can be biased in some circumstances. 32 Thus, we focus on a fourth approach 33 : IPTW.
3.2 IPTW estimator and its variance for complete data
IPTW aims to create a pseudo-population similar to a randomised trial by weighting the individuals by the inverse of their probability of receiving the treatment they actually received (i.e.
The IPTW estimator, as with other PS-based estimators, is a ‘two-step estimator’. If the uncertainty linked to the PS estimation in the first step is not taken into account in the second step (treatment effect estimation), the repeated sampling variance of the treatment effect will be overestimated and inference will be conservative. 31 Lunceford and Davidian 35 and Williamson et al. 34 proposed a large-sample variance estimator for the IPTW treatment effect estimator in which a correction term including the variance/covariance matrix of the estimated PS parameters is applied.
4 Handling missing data in PS analysis
A major issue in PS estimation is the presence of partially observed covariates. In this section, we describe five methods for applying IPTW to incomplete data. We assume the treatment status Z and outcome Y are fully observed.
4.1 CC analysis
In CC analysis, the PS is estimated only within the subgroup of individuals with observed values for all of the variables included in the PS model, and only these individuals contribute to the estimation of the treatment effect. 36 Although the CC analysis is known to provide an unbiased estimate of the parameters of an outcome regression model when the missingness is independent of the outcome, 6 little is known about CC analysis for IPTW. Moreover, excluding individuals with missing covariates can reduce statistical power, because no use is made of partially observed records.
4.2 Missingness pattern approach
Rosenbaum and Rubin
7
defined the generalized PS
4.3 MI
The principle of MI is to generate multiple sets of plausible values for the missing variables by drawing from the posterior predictive distribution of these variables given the observed data. Variables of different types are often included in the PS model. Therefore, we focused on chained equations, in which a specific imputation model, including the outcome, is specified for each partially observed variable, rather than joint modelling to impute the missing data.
38
M complete datasets are created and analysed independently to produce estimates
Three variants of MI are MIte, MIps and MIpar. For complete data, the IPTW method involves three steps: (a) estimation of the parameters
α in the PS model; (b) calculation of individual PS’s; and (c) use of equation (1) to calculate The three approaches considered after multiple imputation (MI) of the partially observed covariates are missing values on the original dataset. 
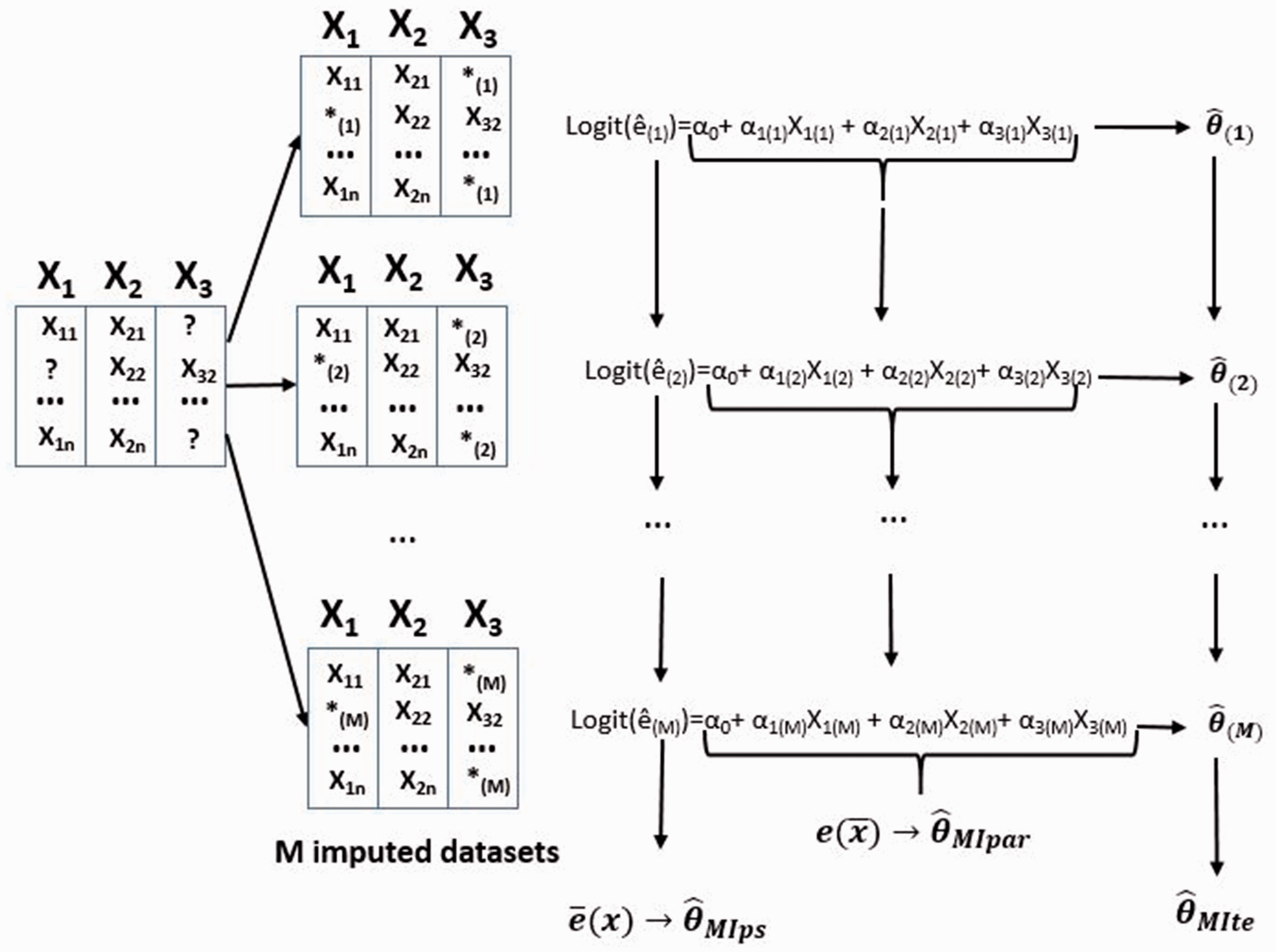
5 Balancing properties and consistency of IPTW estimator after MI
Without missing data, Rosenbaum and Rubin showed that the PS is a balancing score.
3
A balancing score
5.1 Combining the treatment effects after MI (MIte)
Let
If
These two assumptions are the imputed-data version of what Imbens calls weak unconfoundedness.
39
Note that we do not have the analogue of the usual, stronger, assumption
5.1.1 Balancing properties
We show in Appendix 2 that
5.1.2 Consistency
Seaman and White
11
proved that for an infinite number of imputations, the estimator obtained by combining the treatment effects after MI (MIte) is consistent. To understand how this consistency relates to the SITA-type assumption above, it is helpful to consider the following expectation:
Step 6 requires the SITA-type assumption 6. Step 7 relies on PS in the kth imputed dataset being equal to the probability of being treated given the observed and imputed part of the covariates (equation 5).
5.2 Combining the PS or the PS parameters after MI (MIps and MIpar)
For PS methods, consistency comes from the ability of PS to balance covariates between groups. MIps and MIpar create a single overall PS used to estimate the treatment effect. Thus, consistency for these methods would rely on the ability of these overall PS to balance both the observed and the missing parts of the covariates in the original incomplete dataset. Rosenbaum and Rubin’s results show that this can happen only if the single PS (as estimated in MIps or MIpar) is ‘finer’ than the true (full data) PS (in other words, if the true PS is a function of the single pooled PS).
3
However, when combining the PS or the PS parameters, the overall PS used for the analysis is not a function of the observed covariates but rather is a function of the average covariates across imputed datasets, including the imputed values. Thus, the pooled PS (as estimated either in MIps or MIpar) is not ‘finer’ than the true PS according to Rosenbaum and Rubins definition (i.e. the true PS is not a function of the pooled PS). Consequently, it cannot be a balancing score. Thus, neither
6 Simulation study design
The aim of this simulation study is to assess the performance of the three MI approaches, CC analysis and missingness pattern approach for IPTW when the outcome is binary and to compare them with non-MI methods.
6.1 Data generation mechanisms
We generated datasets of sample size n = 2000, reflecting an observational study comparing a treatment Z = 1 to a control treatment Z = 0 on a fully observed binary outcome Y with three measured covariates Covariates: The three covariates Treatment assignment depends on Binary outcome: The outcome depends on the three covariates and the treatment received according to the following model:
Missingness mechanism: In this simulation study, we consider a MAR mechanism. The missingness of each partially observed covariate (X1 and X3) depends on the fully observed covariate X2, the treatment received Z and the outcome Y:
Factors used in the simulation study (
For each scenario, 5000 datasets of sample size 2000 were generated. The prevalence of the treatment was 0.3 and each partially observed covariate had 30% of data missing. Ten imputed datasets were created.
Simulation parameters used for the sensitivity analysis.

For each scenario of the sensitivity analysis, 5000 datasets were generated. The other parameter values were: RR = 2,
6.2 Methods
For each studied scenario, 5000 datasets were simulated. We compared nine methods, all of them using IPTW: treatment effect were estimated on the full dataset (before the imposition of missingness) and using CC, MP, and the three MI approaches (MIte, MIps and MIpar). For each of the three MI approaches, we considered two imputation models, including or not the outcome and we generated M = 10 imputed datasets. Simulations were performed in R and the mi package was used for MI, 41 based on full conditional specification (FCS).
6.2.1 Estimands
We focused on three estimands: the log(RR), log(OR) and the risk difference (RD). Rubin’s rules were applied to the logarithm of the RRs (or the ORs), rather than to RRs (or ORs) themselves, because the asymptotic normal approximation is likely to be better for the log RR (or log OR) than for the RR (or OR). This is important in particular when constructing confidence intervals, because log(OR) and log(RR) have unrestricted parameter spaces. 42
6.2.2 Measures of performance
For each method, we computed the absolute bias of the mean treatment effect. We also estimated the variance of the treatment effect, as well as the empirical variance, the coverage rate and the standardized differences of the covariates after IPTW.
The standardized differences for each covariate were used as a measure of method performance. In the absence of weighting, standardized differences are defined as:

7 Simulation study results
7.1 Main simulation study
Because results were similar for the three measures of interest, we present the results for relative risks (RR) on the log scale only in the main text, while results for odds ratios and risk differences are in the appendices.
7.1.1 Bias
The absolute bias of the log(RR) of the treatment, for Absolute value of the bias for the four scenarios in which 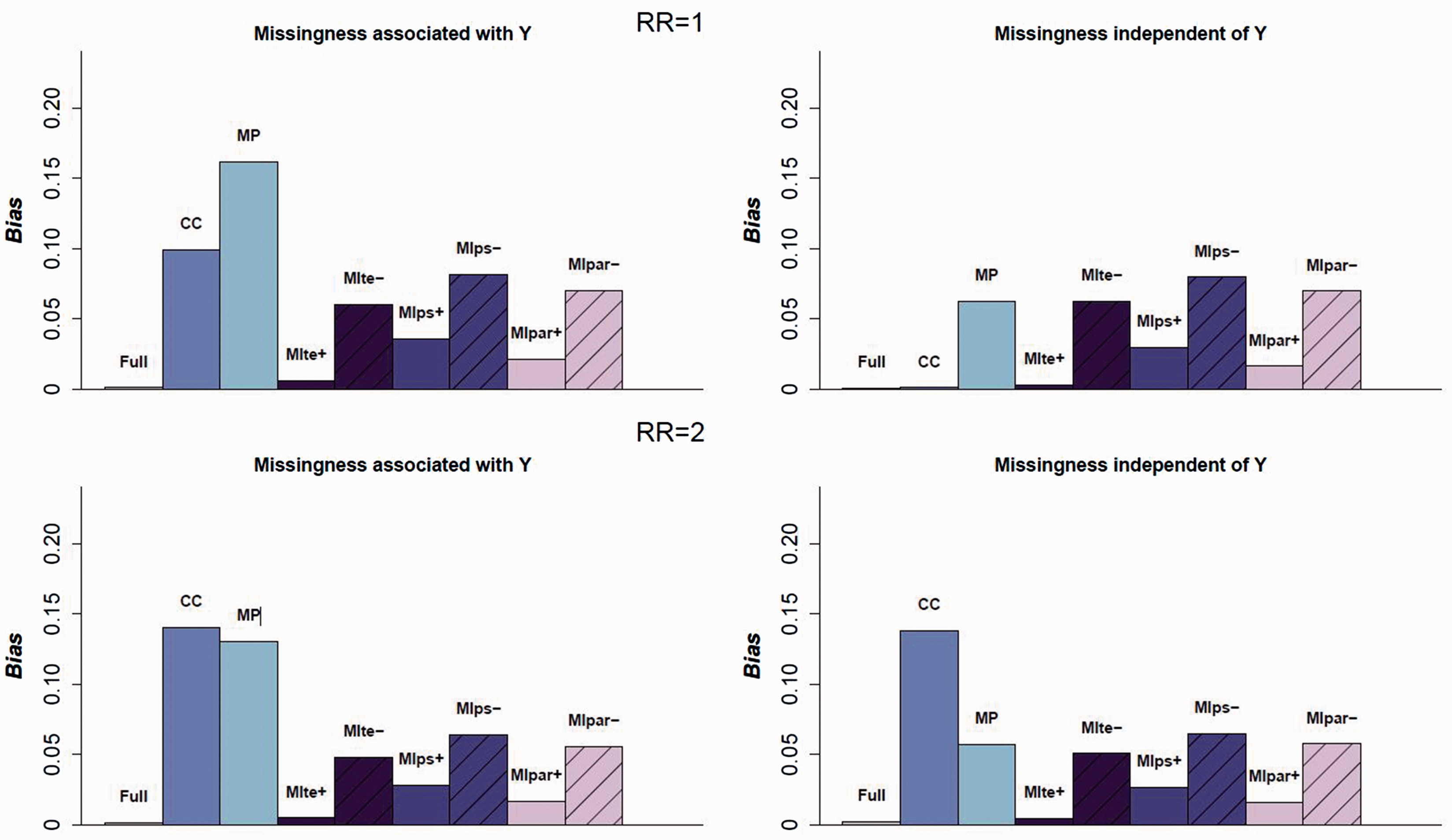
Full data, CC and MP analyses
As expected, the IPTW estimator on the full data (before generating missingness for X1 and X3) is approximately unbiased and the CC estimator is strongly biased in all scenarios except those where the outcome is not associated with missingness and there is no treatment effect. The MP approach is always biased in the situations considered, with a bias which can be even stronger than that which is observed for the CC approach. The reason for this is an incorrect PS model specification in each pattern of missingness: in the strata in which a covariate is not observed, the covariate is omitted from the model.
MI
First, the results show that the imputation model must include the outcome, even if the outcome is not a predictor of missingness. All three MI estimators are strongly biased in all scenarios when the outcome is not included in the imputation model. Second, when the outcome is included in the imputation model, the three MI approaches lead to a decrease in bias relative to the crude analysis. However, only the MIte approach leads to an unbiased estimate in the eight main scenarios. Combining the PS parameters to estimate the PS of the average covariates (MIpar) performed better than combining the PS themselves, but both these approaches are slightly biased.
7.1.2 Standardized differences between groups
Standardized differences (in %) after IPTW for each method for one scenario: RR = 2,
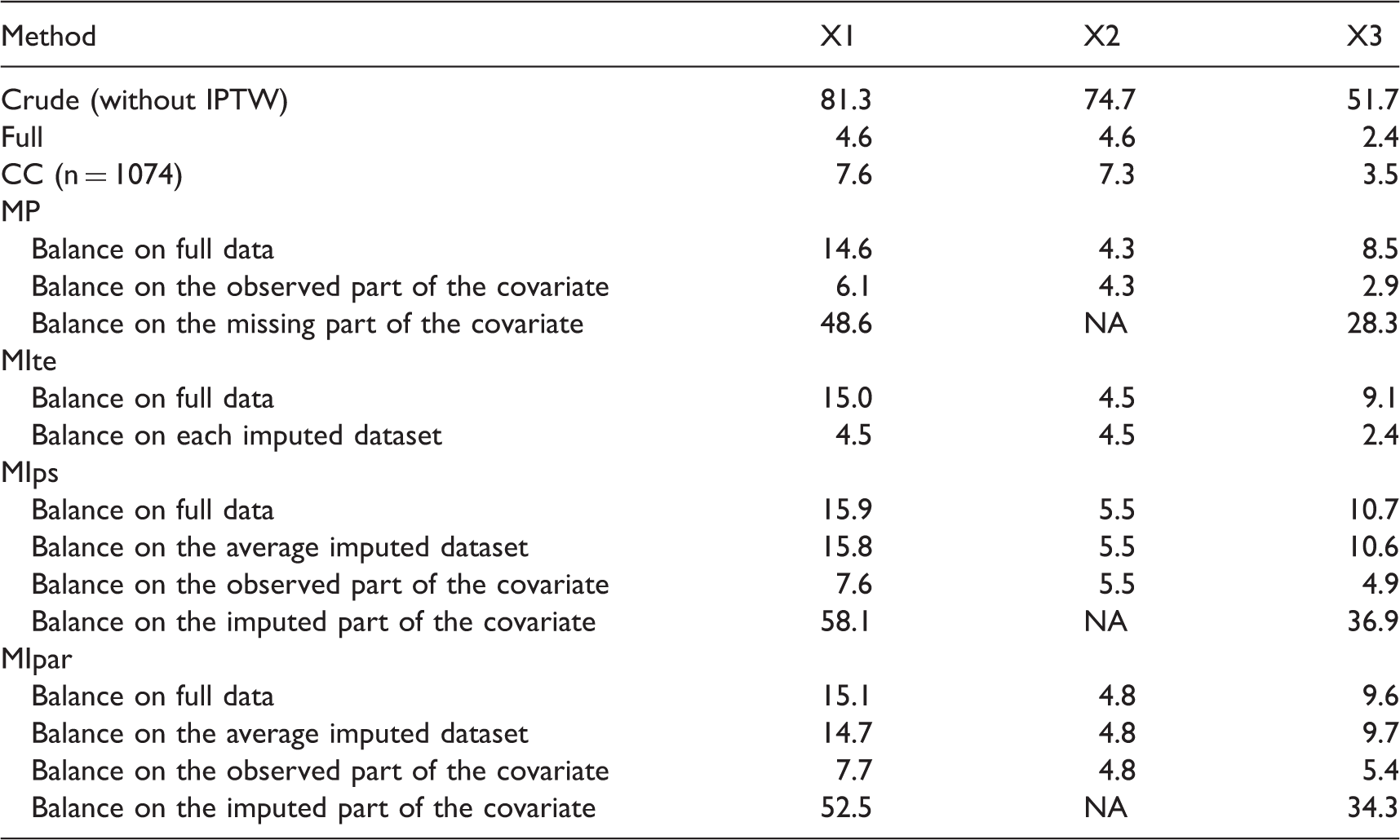
CC: complete case; MP: missingness pattern; MIte: treatment effects combined after multiple imputation; MIps: propensity scores combined after multiple imputation; MIpar: propensity score parameters combined after multiple imputation; RR: relative risk; NA: not applicable because X2 is fully observed.
7.1.3 Coverage rate and standard errors
Figure 3 displays the coverage rate for each method when the outcome is included in the imputation model. Each boxplot represents the coverage distribution for the eight main scenarios. Because the CC and MP approaches are strongly biased, their coverage rates are not relevant. The coverage rate for the MIte approach is close to the nominal value of 95%, confirming that Rubin’s rules perform well in this context provided that the within-imputation variance estimation takes into account the uncertainty in PS estimation (Table 4).
Coverage rate of the 95% CI for each method compared. Results are pooled for the 8 main scenarios. CC: complete case; MP: missingness pattern; MIte: treatment effects combined after multiple imputation; MIps: propensity scores combined after multiple imputation; MIpar: propensity score parameters combined after multiple imputation. RR: relative risk. For the three MI methods, the outcome is included in the imputation model. Bias of the log(RR), its estimated variance and coverage rate for the three MI approaches according the sample size n for one scenario (RR = 2, MIte: treatment effects combined after multiple imputation; MIps: propensity scores combined after multiple imputation; MIpar: propensity score parameters combined after multiple imputation.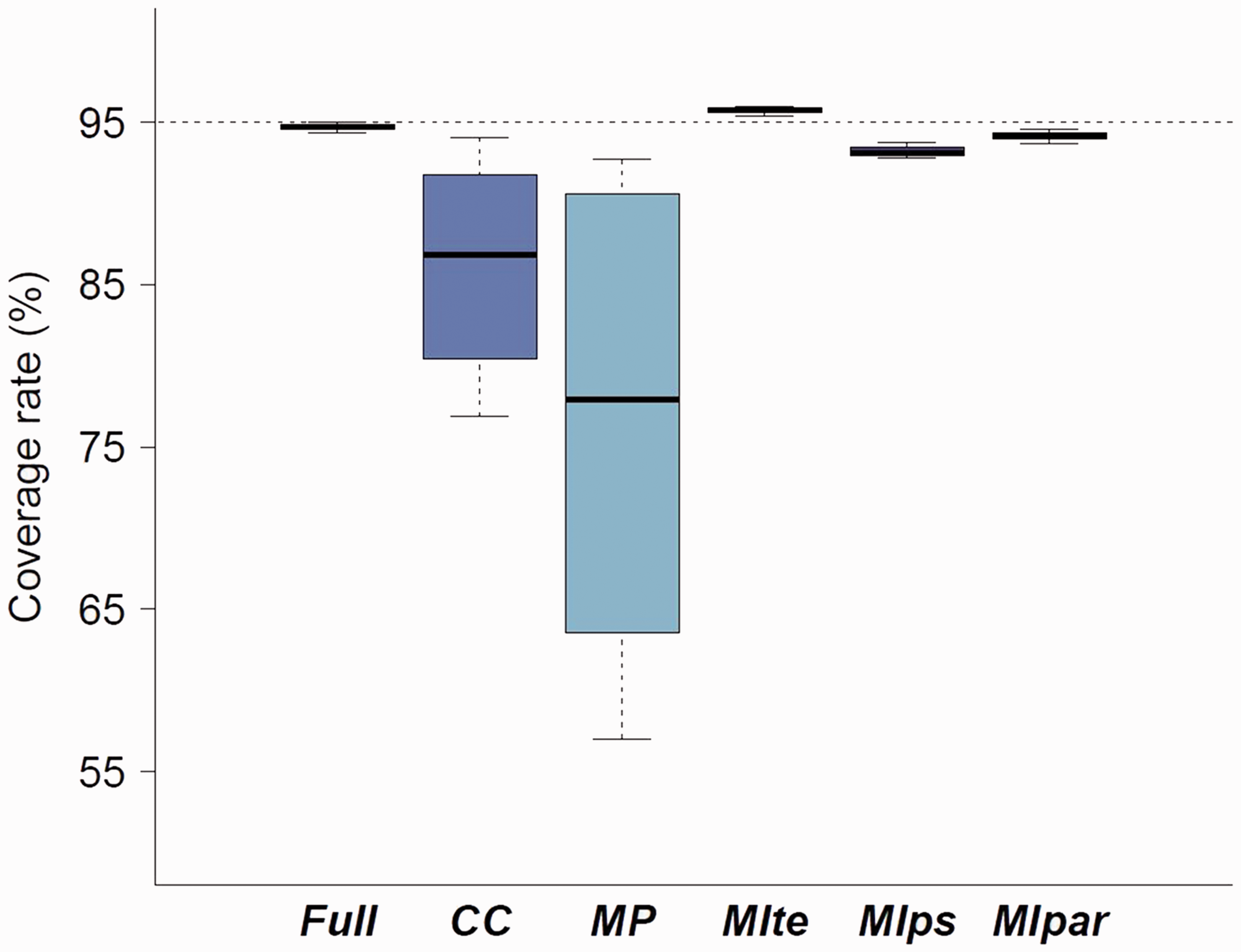
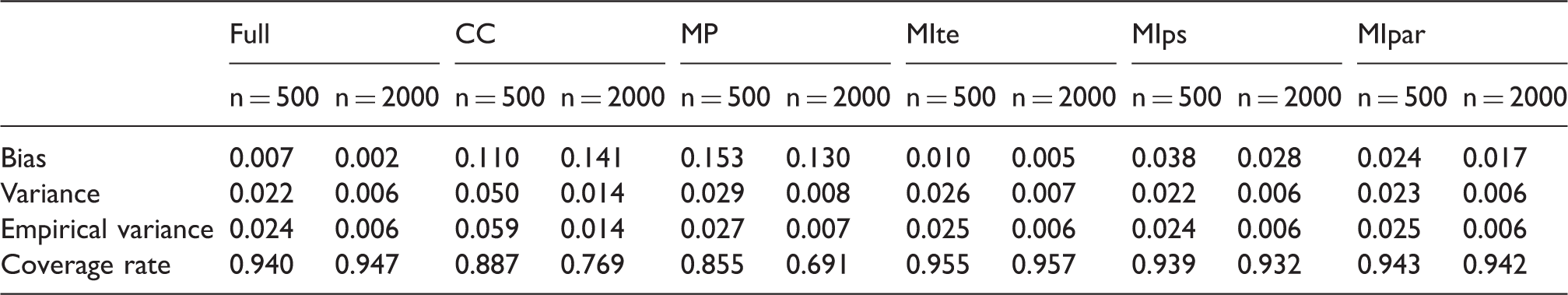
Table 1 in Appendix 5.1 shows the mean estimates from different variance estimators for each method when the outcome is included in the imputation model. For MIte, one could apply Rubin’s rules to a variance estimator that ignores the PS estimation (treating the PS weights as being known) or to a variance estimator that accounts for the PS estimation. We know that failing to account for the PS estimation results in an overestimation of the variance in the full-data context. 43 Table in Appendix 5.1 shows that for the analysis on full data and for MIte, the variances accounting for PS estimation are close to the empirical variance, whereas an estimator not taking the uncertainty in PSs estimation tends to overestimate the variance for these approaches, as expected. For MIps and MIpar, one could estimate the variance accounting for the variability linked to PS estimation but not the imputation procedure by using the pooled PS in the IPTW variance estimator or one could incorporate the PS estimation and imputation by applying the variance estimator we proposed in Appendix 1. For MIps and MIpar, there was little difference between the variance estimates accounting for the imputation and those that did not. Surprisingly, in our simulations, additionally accounting for the imputation procedure resulted in a lower variance estimator. This can be explained as follows: the within imputation variance of the PS parameters (reflecting the correlation between the covariates and treatment; the higher this is the larger gain in precision for IPTW) is higher than the between imputation variance component (noise due to missing data). However, when the missingness rate increases, the variance that correctly accounts for the imputation procedure is typically higher than the variance which ignores the imputation, because of a larger heterogeneity between imputed datasets.
7.2 Sensitivity simulation study
Appendix 5.3 presents the results of one scenario with a non-null treatment effect with a smaller sample size (n = 500). Results were similar in terms of bias for n = 500 and n = 2000. Because the variance estimator for IPTW has been developed for large samples, we observed slightly underestimated variances for the full data analysis, MIps and MIpar. This underestimation is more pronounced in the CC analysis because the sample for the analysis is even smaller (269 on average when n = 500).
Figure 4 shows the bias when 10% or 60% of each partially observed covariate is missing. Full results are presented in Appendix 5.4. For a low missingness rate, the CC and MP approaches are still biased but the three MI approaches corrected the bias. For a missingness rate of about 60% for each covariate, only the MIte approach showed good performance in terms of bias reduction, confirming the good statistical properties of this approach even with this large amount of missing data. In our sensitivity simulation study, increasing the number of imputed datasets did not strongly impact the results in terms of bias or variance (See Appendix 5.5).
Absolute value of the bias according to the missingness rate. CC: complete case; MP: missingness pattern; MIte: treatment effects combined after multiple imputation; MIps: propensity scores combined after multiple imputation; MIpar: propensity score parameters combined after multiple imputation; RR: relative risk. For the three MI methods, the outcome is included in the imputation model.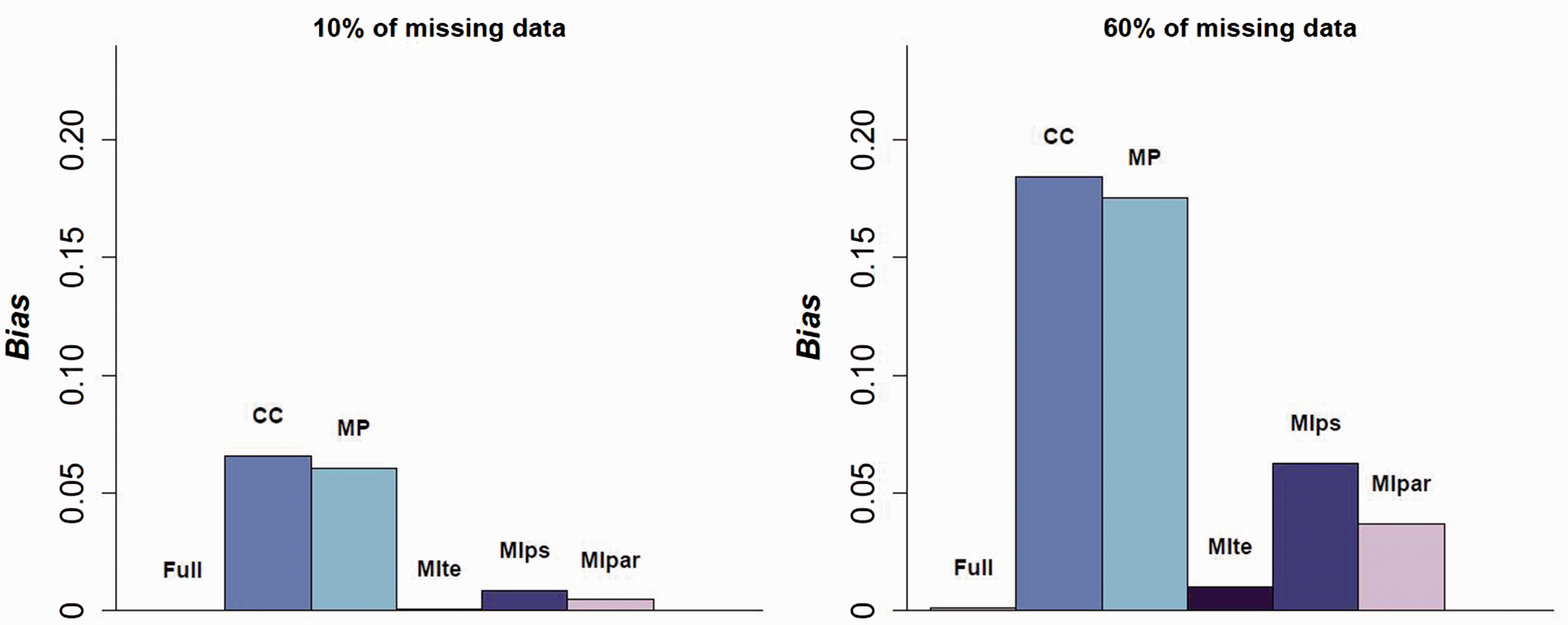
8 Application to the motivating example
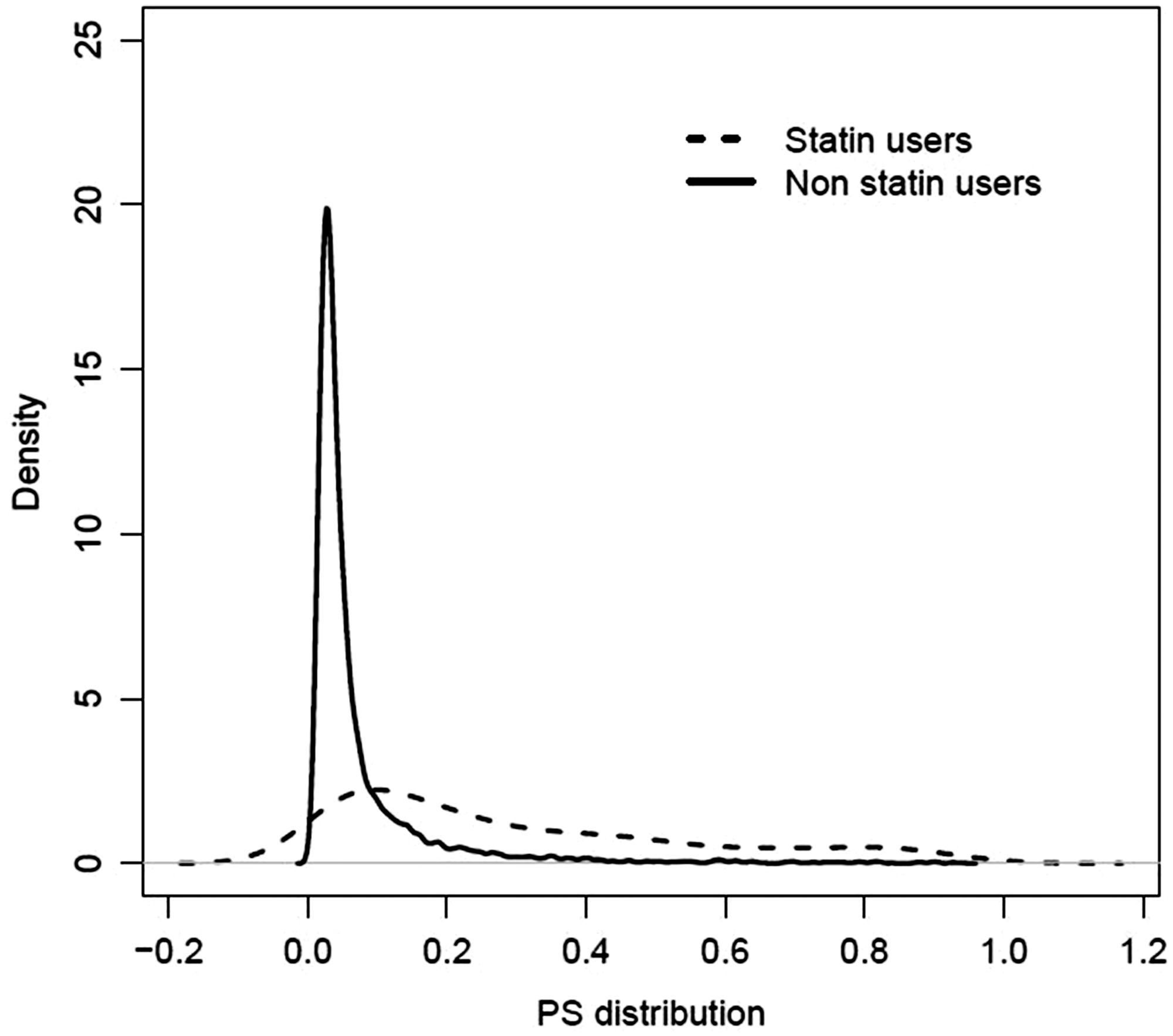
We applied CC analysis, the MP approach and the three MI strategies to estimate the effect of statin treatment on mortality after pneumonia from our motivating example dataset. For simplicity, we analysed the primary outcome, mortality within 6 months, as a binary outcome, and estimated the corresponding relative risk and its 95% confidence interval. For each approach, IPTW was used to account for the confounding. We focused the analysis on the 7158 patients without coronary heart disease. The PS was estimated from a logistic regression modelling statin use as a function of the following covariates: age, sex, body mass index (BMI), alcohol consumption, smoking status, diabetes, cardiovascular disease, circulatory disease, heart failure, dementia, cancer, hyperlipidaemia, hypertension and prescription of antipsychotics, hormone replacement therapy, antidepressants, steroids, nitrates, beta-blockers, diuretics, anticoagulants and use of antihypertensive drugs. The imbalance between the study groups is illustrated in Figure 5. Complete case (CC) analysis was conducted on the 5168 individuals with complete records. For the missingness pattern approach (MP), eight patterns were identified. However, some of these patterns were very rare. For instance, only six individuals had only the smoking status missing, and only eight had both smoking status and BMI missing. Thus, we considered only four groups:
complete records (n = 5168) for which all the covariates listed above are included individuals with only the alcohol consumption missing (n = 455) individuals with only BMI missing (n = 575) individuals with the smoking status missing (alone or in addition to BMI and alcohol consumption) and individuals with both BMI and alcohol consumption missing (n = 960) Distribution of the propensity score estimated on complete cases for statin users and non users (n = 5168).
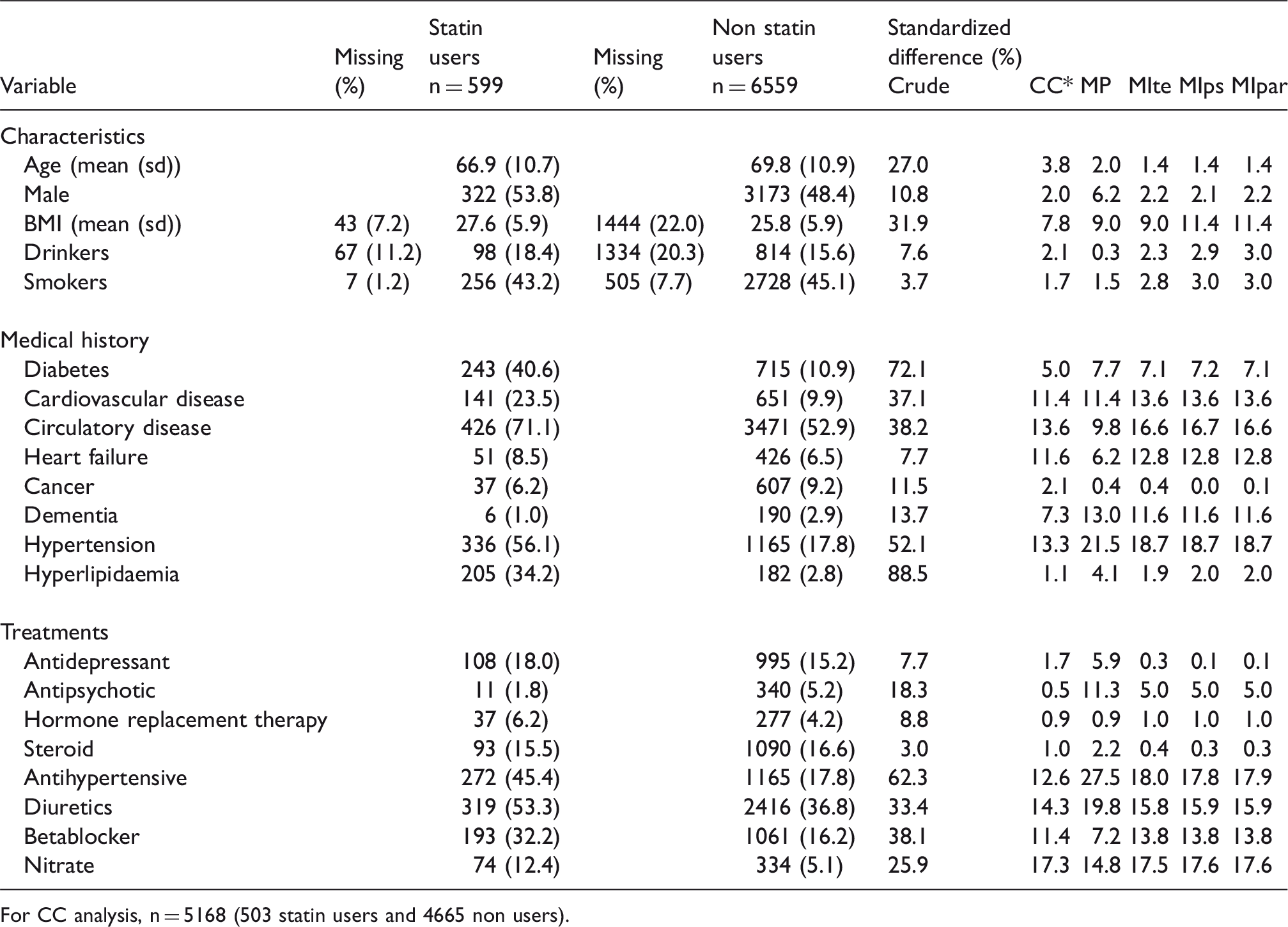
Description and comparison of statin users and non users (n = 7158).
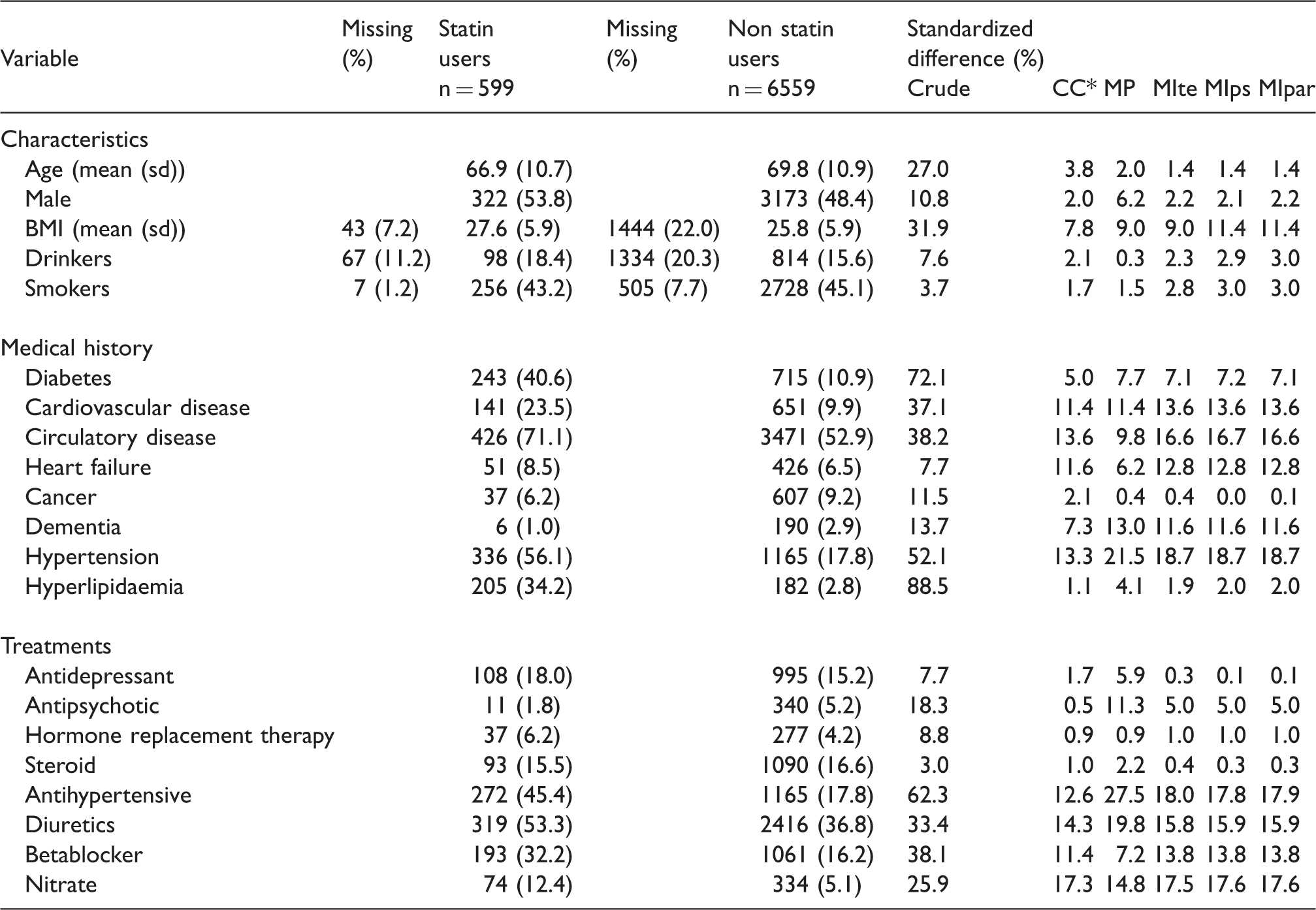
For CC analysis, n = 5168 (503 statin users and 4665 non users).
Estimate of the relative risk of mortality and its 95% confidence interval for statin vs non statin users (motivating example) (n = 7158).
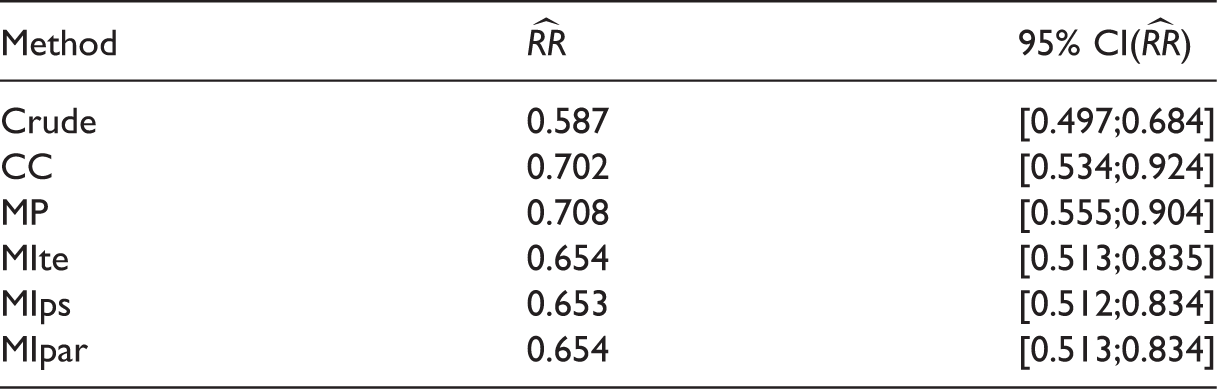
RR: relative risk; CC: complete case; MP: missingness pattern; MIte: treatment effects combined after multiple imputation; MIps: propensity scores combined after multiple imputation; MIpar: propensity score parameters combined after multiple imputation; RR: relative risk.
9 Discussion
This article aimed to address three main questions about MI in the context of IPTW: (a) does the outcome have to be included in the imputation model? (b) should we apply Rubin’s rules on the IPTW treatment effect estimates or on the PS estimates themselves? (c) how should we estimate the variance of the IPTW estimator after MI? First, results showed that the outcome must be included in the imputation model, even if the outcome is not a predictor of missingness. This is well known in the context of multivariable regression, but can be seen as counter-intuitive in the PS paradigm, because the PS model for the full data does not use the outcome data. The simulation results showed a bias in the three MI estimators when the outcome was omitted from the imputation model, even when the outcome was not a predictor of missingness. In practice, to obtain an unbiased estimate of the intervention effect when using MI, the outcome should be included in the imputation model. This may not be always possible if the outcome data has not been yet recorded and the PS is used at the design stage of the study (to find good matched controls for example), rather than at the analysis stage. In such situations, it is worth considering whether the assumptions required for CC analysis or the MP approach may be valid. If they are, one of these methods could be used instead of using MI omitting the outcome. Furthermore, the inclusion of the outcome in the imputation model may go against the principle of objective design, as described by Rubin. 44 However, when using MI, this is necessary to obtain a consistent estimator of the treatment effect. Second, we showed that combining the treatment effects after MI (MIte approach) is the preferred MI strategy in terms of bias reduction under a MAR mechanism. This estimator is the only estimator of the three MI estimators to be consistent and to provide good balancing properties. Even though MIps and MIpar are not consistent estimators of the treatment effect, they can reduce the bias of the CC analysis, if the rate of missing data is not too high. Combining the PS or the PS parameters has no clear advantage for IPTW, but may be useful in the context of PS matching: because it involves only one estimation of the treatment effect, it could provide a computational advantage for large datasets. In addition, MIte for PS matching implies that the M treatment effect estimates are estimated from different matched sets, potentially of different sample sizes, leading to a more complex variance estimation because of these different sample sizes.
Third, as long as the uncertainty in the PS estimation is taken into account in the variance estimation, 34 Rubin’s rules perform well for MIte, even for moderate sample size (n = 500). For MIpar, the proposed variance approximation (Appendix 1) showed good performance in our simulation study.
The three MI approaches differ in terms of their balancing properties. We showed that whereas the PS estimated in each dataset in the MIte approach can balance covariates between groups in each imputed dataset, this is no longer true for MIps and MIpar. However, the best method to assess covariate balance after MI remains unknown. With MIte, the aim being to estimate a treatment effect from each dataset, we require balance between groups within each imputed dataset. In contrast, for MIps and MIpar, further investigation is needed to know if we should assess the balancing properties of the pooled PS on the average covariate values across the imputed dataset or on each dataset.
Our recommendations for the optimal way to implement MI for PS analysis are in contradiction to those given by Mitra and Reiter, who suggest that pooling PSs across imputed datasets can sometimes out-perform our recommended method of pooling treatment effects. 9 In their simulation studies MIps (or MIpar) led to a bigger reduction in bias than MIte in some scenarios. This was due to a near violation of the positivity assumption in their simulated datasets 45 combined with the omission of the outcome from the imputation model. Penning de Vries and Groenwold 46 used Mitra and Reiter’s data-generating mechanism to perform a simulation study but they increased the effective sample size to ensure positivity. Under this condition and when the outcome was included in the imputation model, as we recommend, they showed MIte was superior, in accordance with our findings. While we understand the desire to adhere to the principle of objective design by omitting the outcome from the imputation model, our results show that this leads to bias in MIte and an increase in bias for MIps (and MIpar). Thus, when it is either undesirable or impossible to include the outcome in the imputation model, we would suggest considering the validity of alternative approaches, such as CC analysis or the MP approach, rather than adopting the systematically biased approach of imputation omitting the outcome from the imputation model.
The MP approach, which is widely used in practice to handle missing data for PS analysis, showed poor performance under a MAR mechanism. In our simulation study, the MP approach does not perform well, because missing data were generated under a MAR mechanism, but the MP approach relies on different assumptions about the missing data mechanism. 37 When the assumptions for the MP approach are valid, it avoids the need for an imputation model altogether, and hence avoids the need to use the outcome when calculating the PS. Because the assumptions for MI and MP are different, we believe that MP could be a promising alternative when the assumptions for MI are not valid. Further investigations are needed to understand the usefulness of the MP approach in practice. Moreover, its application to our real-life example dataset was challenging, because the sample size within each missingness pattern was not large enough to estimate the PS.
This work has some limitations. We generated only three covariates in our simulation, whereas PS are often calculated from a large number of covariates. Moreover, we studied only the common situation where the PS model only includes main effects, and we assumed as well the model including main effects only for the outcome given the covariates. When there are interactions or quadratic terms in these two models, the specification of the imputation model can be less straightforward, requiring further efforts to ensure the imputation model is compatible with the substantive (analysis) model. 46 Furthermore, we focussed on the IPTW estimator only, because of its increasing use and its application in the presence of time-varying covariates and treatment. Indeed, IPTW is widely used in marginal structural models (MSMs) to handle time-dependent confounding. Although this article is about the simpler situation of a single time-point, it is important to clarify how to use IPTW after MI correctly in this context, because otherwise it is unlikely to be done correctly in the more complex situation of time-dependent confounding. However, some issues may arise with IPTW when some estimated weights are too extreme; PS matching can be a better solution in this context. 47 Nevertheless, because the PS estimated in either MIps or MIpar is not a balancing score, PS matching (like IPTW – and like also PS stratification and PS covariate adjustment) would not be expected to perform well when used in combination with MIPs or MIPar. Conversely, because in MIte, the estimated PS can balance both the observed and imputed component of the covariates in the imputed dataset, we would expect all the PS-based methods (PS matching, stratification, covariate adjustment and IPTW) to perform well to estimate the treatment effect within each imputed dataset as long as the underlying assumptions for these methods are fulfilled.
In conclusion, for IPTW, MI followed by pooling of treatment effect estimates is the preferred approach amongst those studied, when data are MAR, and the outcome must be included in the imputation model.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Medical Research Council (Project Grant MR/M013278/1 and Unit Programme Number U105260558).
Supplemental material
Supplemental material is available for this article online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.