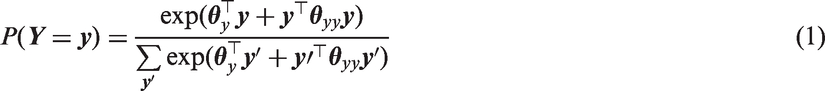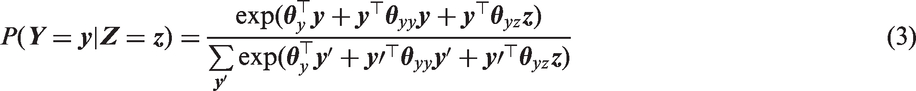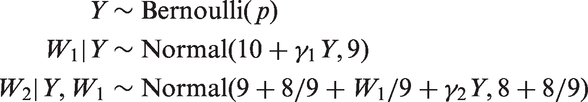Abstract
Estimating the parameters of a regression model of interest is complicated by missing data on the variables in that model. Multiple imputation is commonly used to handle these missing data. Joint model multiple imputation and full-conditional specification multiple imputation are known to yield imputed data with the same asymptotic distribution when the conditional models of full-conditional specification are compatible with that joint model. We show that this asymptotic equivalence of imputation distributions does not imply that joint model multiple imputation and full-conditional specification multiple imputation will also yield asymptotically equally efficient inference about the parameters of the model of interest, nor that they will be equally robust to misspecification of the joint model. When the conditional models used by full-conditional specification multiple imputation are linear, logistic and multinomial regressions, these are compatible with a restricted general location joint model. We show that multiple imputation using the restricted general location joint model can be substantially more asymptotically efficient than full-conditional specification multiple imputation, but this typically requires very strong associations between variables. When associations are weaker, the efficiency gain is small. Moreover, full-conditional specification multiple imputation is shown to be potentially much more robust than joint model multiple imputation using the restricted general location model to mispecification of that model when there is substantial missingness in the outcome variable.
Keywords
1 Introduction
Estimating the parameters of a regression model of interest (the ‘analysis model’) is often complicated in practice by missing data on the variables in that model. Multiple imputation (MI) is a popular method for dealing with this problem.
1
Values for the missing variables are randomly sampled conditional on the observed variables from distributions thought approximately to describe the association between these variables. The result is an imputed dataset, in which there are no missing data. This imputation is done multiple (say, M) times and the analysis model is fitted separately to each of the resulting M imputed datasets to produce M estimates of the parameters
MI methods differ in how they randomly sample values for the missing variables. The two most commonly used methods are joint model MI and full-conditional specification (FCS) MI (also known as MI by chained equations).2,3 The former involves specifying a joint model for the partially observed variables given the fully observed variables and sampling missing values from their posterior predictive distribution given the observed data. The latter involves specifying a conditional model for each of the partially observed variables given all the other variables and cycling through these models. In special cases, the two approaches are equivalent. 4 For example, when all the conditional models in FCS MI are linear regressions with main effects and no interactions, FCS MI corresponds to joint model MI using a multivariate normal joint model. Likewise, when all the variables are categorical and the conditional models are saturated logistic regressions, FCS MI is equivalent to joint model MI using a saturated log linear joint model. In general, however, FCS MI is not equivalent to joint model MI.
Liu et al.
5
(see also Zhu and Raghunathan
6
) showed that, even when FCS MI does not correspond to joint model MI, the distributions from which the two methods sample the missing values (the ‘imputation distributions’) are asymptotically the same when the conditional models used by FCS MI are compatible with a joint model. Compatibility is defined in Section 2. Although this is an important result, the ultimate purpose of MI is to enable the estimation of
When, as is commonly the situation, the partially observed variables consist of both continuous and categorical variables, the conditional models usually employed for them in FCS MI are linear regressions and multinomial logistic regressions, respectively. These are natural choices and are the default options in many statistical packages, e.g. mice and mi in R, and ice and mi in STATA. It can be shown that this set of conditional models is compatible with a restricted general location (RGL) joint model. Thus, Liu et al.'s (2014) result implies that FCS MI and joint model MI using the RGL model produce imputations that are asymptotically from the same distribution. Schafer 7 described how to carry out joint model MI using this RGL model and provided software. So, joint model MI using the RGL model and FCS MI using conditional models compatible with this model are both options for the practicing statistician.
In the current article, we focus on the situation where the aim is to estimate the parameters of the analysis model and MI is used to handle missing data in the variables of that model. We elucidate the relation between joint model MI and FCS MI using compatible conditional models. We focus on the important case where joint model MI uses the RGL model and the (compatible) analysis model is a linear or logistic regression with parameters
The structure of the article is as follows. In Section 2, we describe FCS and joint model MI in general and discuss how they are related when the conditional models are compatible. This relation can be one of equivalence in finite samples or asymptotic equivalence. The RGL model is introduced in Section 3. In Section 4, the asymptotic RE of inference from FCS MI with compatible conditional models versus that from the corresponding joint model MI is explored in depth for simple cases of the RGL model: one with a binary and two continuous variables, and one with four binary variables. In addition, the RE of the two MI methods is explored in a more complex situation using data simulated from a realistic data-generating mechanism based on the Barry Caerphilly Growth Study (BCGS). 8 In Section 5, we discuss and illustrate, using simulated data and data from the National Childhood Development Study (NCDS), 9 the relative robustness of FCS MI and joint model MI to misspecification of the joint model for the covariates implied by the RGL model. Section 6 contains a discussion.
2 Relation between FCS MI and joint model MI
Let
In joint model MI, a model
In FCS MI, a set of K conditional models
Hughes et al.
4
noted that FCS MI and the Gibbs sampler algorithm (and hence joint model MI) are equivalent when, for each k, the parameters
The non-informative margins condition cannot hold unless
An important theoretical result about the asymptotic relation between FCS and joint model MI was provided by Liu et al.
5
This result can apply even when the non-informative margins condition is not satisfied. They defined the set of conditional models
Theorem 1 of Liu et al. 5 says that if (i) the set of conditional models is compatible with a joint model, (ii) this joint model is correctly specified and (iii) the data are missing at random (MAR), then the total variation distance between the distribution of the imputed data obtained from FCS MI and the distribution of the imputed data obtained from joint model MI tends to zero in probability as the sample size tends to infinity. More informally, we can say that the distribution of the imputed data is asymptotically the same whether one imputes by FCS MI or by joint model MI using the corresponding joint model. Liu et al. 5 say that ‘iterative imputation [i.e. FCS MI] and joint Bayesian imputation [i.e. joint model MI] are asymptotically the same’ (p. 161).
Three comments are worth making. First, asymptotic equivalence of the imputation distributions of FCS MI and joint model MI does not mean that the two resulting Rubin's Rules estimators of
3 The RGL model
Let
This RGL model implies that the conditional distribution of any element of
As mentioned at the end of Section 2, if some elements
Note that, unlike the RGL model, the CRGL(
Higher order interactions can be added to the log linear models of expressions (1) and (3). The conditional models of FCS MI then require additional interaction terms to remain compatible with this more general RGL or CRGL model. However, we focus on the log linear model with just main effects and pairwise interactions (expressions (1) and (3)) and study the impact of the absence of higher-order terms on the RE of FCS MI and joint model MI for inference about
4 Asymptotic RE of RGL versus FCS MI
4.1 Information in the marginal distribution
In Section 2, we noted that when the marginal distribution of
First, expressions (1) and (2) imply that the marginal distribution of
Second, suppose for simplicity that there are no continuous variables
In the remainder of this section, we study how much this information in the margins affects the asymptotic RE of the Rubin's Rules estimator of
4.2 One binary and two continuous variables
Suppose that data are generated by the RGL model used by Hughes et al.
4
If Y is fully observed,
When Y is partially observed,
Percentage asymptotic REs of LDA versus logistic regression analysis when using complete data.
These complete-data results suggest RGL MI may often not be much more efficient than FCS MI when Y is partially observed. To investigate this, we assumed that Y is missing with probability 0.5, either completely at random or at random with probability
Percentage asymptotic REs of RGL MI versus FCS MI for four different analysis models.

E(Y): marginal mean of Y; lr: logistic regression; ld: linear discriminant analysis; ln: linear regression; (V): the regression coefficient associated with variable V (with (0) meaning the intercept); MAR: missing at random; MCAR: missing completely at random.
It is interesting that FCS MI is less efficient than RGL MI when the analysis is LDA. This shows that, for the RGL model, asymptotic efficiency is lost by the imputer assuming less than the analyst. Meng 12 gave a different example of an imputer assuming less than an analyst and showed there was no loss of asymptotic efficiency in that case. It is also interesting that RGL MI is more efficient than FCS MI when the analysis is logistic regression. This is an example of the imputer assuming more than the analyst. Since FCS MI with an infinite number of imputations followed by logistic regression analysis is asymptotically equivalent to logistic regression analysis using only complete cases (because individuals with missing outcome provide no information in logistic regression when all covariates are fully observed), the greater efficiency of RGL MI followed by logistic regression analysis is an illustration of ‘super-efficiency’.12,14
Next, we investigated whether the asymptotic REs in Table 1 reflect the REs in finite samples. Table 6 in the online Appendix shows, for two scenarios, the RE of LDA using complete data versus logistic regression using complete data for a variety of sample sizes. The REs were estimated using 10,000 simulated datasets. Table 7 in the online Appendix shows, for the same two scenarios (and using the same 10,000 simulated datasets), the finite-sample RE of RGL MI versus FCS MI (using M = 50 imputations) for the four analyses of Table 2. The finite-sample REs are similar to the asymptotic REs. Note that, as expected, the Rubin's Rules point estimators were approximately unbiased for all methods (data not shown).
4.3 Four binary variables
Now suppose that data are generated by the log linear model with L = 4 binary variables, P(Y1, Y2, Y3.Y4) ∝ exp
Detailed results are given in the online Appendix. In summary, we found that analysing the complete data by fitting the log linear model was hardly any more efficient than using logistic regression: all REs were less than 107%. Likewise, analysing RGL MI was not much more efficient than FCS MI: all REs were less than 116% when analysis was by logistic regression and all were less than 108% when analysis was by fitting the log linear model. These maximum REs required strong associations between variables, i.e.
4.4 Simulation study based on BCGS
To investigate RE of RGL MI versus FCS MI in a realistic setting, we carried out a simulation study based on real data from the BCGS. The BCGS is a follow-up study of a dietary intervention randomised controlled trial of pregnant women and their offspring. 8 Participants in the original trial were followed up until offspring were five years old. When aged 25, these offspring were invited to participate in a follow-up study. There were 951 offspring in the trial, of whom 712 participated in the follow-up study.
For the simulation study, we considered eight variables: sex, childhood weight (at age 5), adult overweight (a binary indicator of BMI
Simulated datasets were created as follows. First, we fitted the RGL model of equations (1) and (2) to the BCGS data. Then, the fitted model was used to generate complete data on the eight variables for each of 951 hypothetical individuals independently. Missingness was then imposed using missingness models whose parameters were estimated from the BCGS data.
The simulation study was in two parts. In Part I, three of the continuous variables (father's, mother's and childhood weights) were treated as auxiliary variables, i.e. they were included in the imputation model but not in the analysis model; the other variables were included in both models. Using auxiliary variables in the imputation model may increase efficiency and make the MAR assumption more plausible. 15 With auxiliary variables, it may be worth imputing a missing outcome even if the covariates in the analysis model are fully observed.
In Part II, there were no auxiliary variables: all eight variables were included in the analysis model. In the absence of auxiliary variables, Von Hippel 15 recommended including all individuals in the imputation step but then excluding those with imputed outcomes before fitting the analysis model to the imputed data, in order to reduce bias caused by a possibly misspecified imputation model. This approach is valid when (i) the data are MAR, (ii) the model for the conditional distribution of outcome given covariates implied by the imputation model is the same as the analysis model and (iii) the analysis model is correctly specified. We therefore analysed imputed data both before and after excluding imputed outcomes. As the proportion of missing covariate values among offspring with observed outcome was small in the BCGS dataset, we increased this proportion for the simulation (Part II only).
For both parts, we checked that the RGL model was not an obvious poor fit to the BCGS data by comparing the LORs from a complete-case logistic regression of adult overweight on the other variables with the corresponding LORs implied by the fitted RGL model. The estimates were similar, providing some reassurance.
We considered several simulation scenarios, by varying the strength of association between the auxiliary variables and outcome (in Part I) and the amount of missingness in the covariates (in Part II). For each scenario, we simulated 1000 datasets using R. RGL MI was performed using the mix library in R; FCS MI used the ice package in STATA; M = 100 imputations were used. Full details are given in the online Appendix.
To summarise the results, in Part I the maximum RE of RGL MI versus FCS MI was 104% (this was for LOR of ex-smoker). In Part II, the covariate with the highest RE was ex-smoker; this RE was 105% when 33% of ex-smoker values were missing and 111% when 54% of ex-smoker values were missing. When imputed outcomes were excluded before fitting the analysis model, these maximum REs decreased to, respectively, 102% and 108%. Full results are in the online Appendix.
5 Robustness of FCS and joint model MI
In Section 4, we demonstrated that RGL MI can be more efficient than FCS MI but that the gains seem to be small unless associations between variables are very strong. In this section, we show that these efficiency gains can come at the price of bias when the RGL model is misspecified. In Section 5.1, we modify the RGL model used in Section 4.2 so that W1 is not normally distributed given Y. It is now a CRGL(W1) model but not a RGL model. We show that when W1 is fully observed and Y is partially observed, logistic regression gives unbiased estimation when imputation is by FCS MI but not when RGL MI is used. In Section 5.2, we modify the log linear model of Section 4.3 by introducing a third-order interaction between Y2, Y3 and Y4. The log linear model with only main effects and pairwise interactions is now misspecified. We show that this causes bias when RGL MI is used but not when FCS MI is used, because FCS MI makes no assumption about the distribution of fully observed variables. In the online Appendix, we present a realistic analysis of data from the NCDS, 9 which illustrates that use of RGL MI can lead to serious bias in a situation where FCS MI does not.
5.1 One binary and two continuous variables
We simulated data from the following modification of the RGL model of Section 4.2.


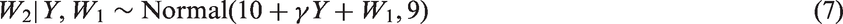
Now, W1 is no longer normally distributed given Y. This is a CRGL(W1) model but not a RGL model. We considered eight scenarios defined by the value of γ (1 or 3), by which variables were partially observed (either just Y or both Y and W2), and by whether data were missing completely at random (MCAR) or MAR. The probability that each partially observed variable was observed was 0.5 if data were MCAR and
Missing data were imputed using either FCS MI or RGL MI, with M = 50 imputations. Four analyses were carried out: estimating E(Y) by the sample mean of Y; estimating parameters β0, β1 and β2 of
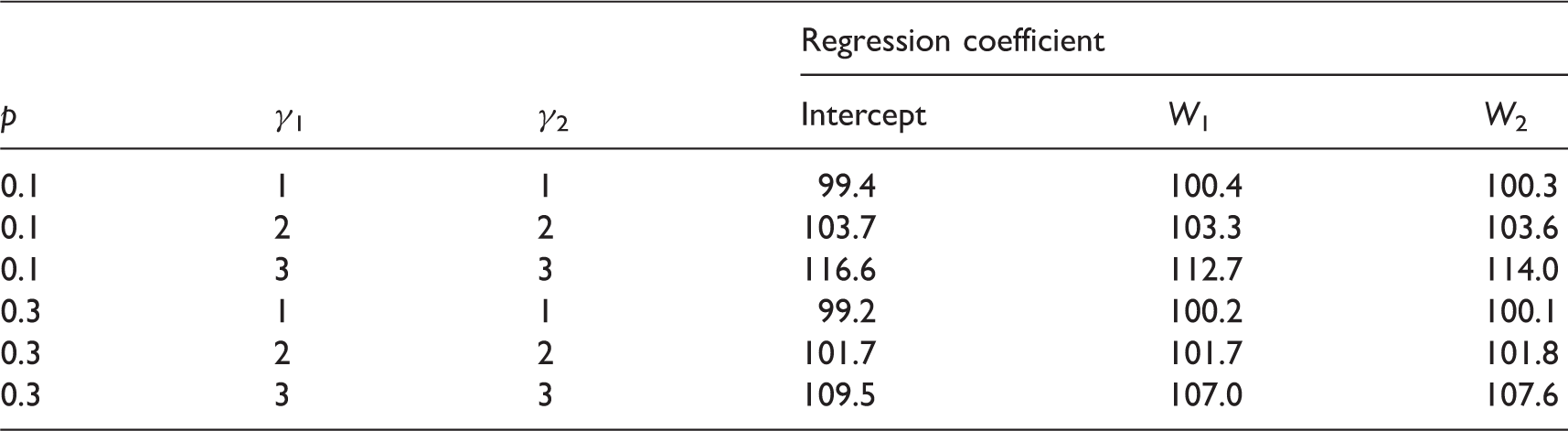
Mean estimates when RGL model is misspecified and γ = 1.

Lr: logistic regression; ld: linear discriminant analysis; ln: linear regression; (V): the regression coefficient associated with variable V (with (0) meaning the intercept); MCAR: missing completely at random; FCS: full-conditional specification; RGL: restricted general location; MAR: missing at random.
For the scenarios where Y and W2 were both partially observed, we also applied logistic regression to the datasets imputed by RGL MI after excluding the individuals whose Y value had been imputed. Similarly, we applied linear regression to the imputed datasets after excluding the individuals whose W2 value had been imputed. This strategy of excluding imputed outcomes before analysing the data has been advocated by Von Hippel 15 as a way of reducing bias caused by a possibly misspecified imputation model. Table 9 in the online Appendix shows the results. Most or all of the bias has been removed for logistic regression but none has been removed for linear regression. Note that Von Hippel did not recommend this approach when MI is done using strong auxiliary variables.
Finally, in scenarios where Y1 and W2 were both MCAR, we additionally imposed 10% missingness on W1. Tables 3 and 8 show that although there is some bias for logistic regression analysis when FCS MI is used, this is much less than when RGL MI is used.
5.2 Four binary variables
Consider again the data generating mechanism of Section 4.3 but now suppose that the log linear model contains an additional third-order interaction:
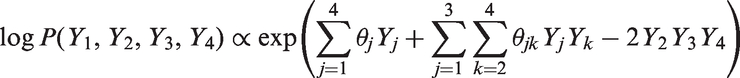
The log linear model with only main effects and pairwise interactions is now misspecified. Table 10 in the online Appendix shows, for four true values of
6 Discussion
FCS and joint model MI yield imputed data with the same asymptotic distribution when the conditional models used by FCS MI are compatible with the joint model. However, we have shown that this asymptotic equivalence in terms of the imputation distribution does not imply that FCS and joint model MI yield equally asymptotically efficient estimates of the parameters in the analysis model. Moreover, FCS MI can be more robust than joint model MI to misspecification of the joint model. We focussed on the RGL model. The efficiency gain from using joint model MI with this model (RGL MI) rather than the corresponding FCS MI appears to be small, except when the outcome is categorical and has a large proportion of missingness and very strong associations exist between the outcome and covariates. On the other hand, we have shown that if the RGL model is misspecified, RGL MI can be much more biased than FCS MI in this same situation, even when covariate-outcome associations are weaker.
Robustness of RGL MI can be improved by including additional interactions in the model (this could have been done in, e.g. the analysis of the NCDS data in the online Appendix) or by conditioning on fully observed variables
Although careful assessment of goodness of fit of an imputation model could detect poor fit of that model, we suspect that this may often not be done in practice. For this reason, FCS MI may be safer than RGL MI when a large proportion of outcomes are missing, unless imputed outcomes are excluded from the subsequent analysis. Since, as Sullivan et al. 16 noted, this exclusion can itself induce another bias, we suggest that a good approach may be to use FCS MI imputing the outcome last and including imputed outcomes in the analysis. Our results indicate that the efficiency loss from using FCS rather than joint model MI is unlikely to be significant in practice.
Several comparisons of joint model MI with FCS MI have been published (e.g. Lee and Carlin 17 and Kropko et al. 18 and references therein). However, these have focussed on joint model MI using a multivariate normal model. The distributions of the imputed data from this joint normal model MI and from FCS MI are not asymptotically equivalent, unless all variables are continuous. These published comparisons have generally noted little difference in efficiency, and relative robustness depended on how categorical variables were handled by joint normal model MI.
An alternative to RGL (or CRGL) MI and FCS MI is joint model MI under the latent normal model of Goldstein and Carpenter.2,19 This can be implemented using REALCOM-IMPUTE or the jomo package in R. This software allows conditioning on fully observed variables. This approach also extends to multi-level data by using random effects. Unlike the RGL model, the latent normal model does not imply conditional distributions that are linear or logistic/multinomial regressions. This is why we compared FCS MI with RGL MI rather than with joint model MI under the latent normal model. It also means that, in general, there is incompatibility (sometimes known as ‘uncongeniality’ 12 ) between a latent normal imputation model and a linear or logistic regression analysis model. Nevertheless, while some forms of incompatibility (uncongeniality) between imputation and analysis models (e.g. an imputation model that ignores an interaction present in the analysis model 12 ) may cause substantial bias in the estimates of the parameters of the analysis model, other forms may often not matter in practice. 7 Moreover, MI under the latent normal model may be more robust than RGL MI to model misspecification; more research on this is needed.
A limitation of our work is that, because it was not feasible to study every possible data-generating mechanism, we cannot rule out the possibility that there are scenarios in which large efficiency gains are possible without requiring strong associations between variables, although this seems unlikely. We focussed on parameter estimation. It is plausible that many of our conclusions could apply when the model of interest is used for prediction, since in that case the linear predictor is a weighted average of the individual parameters. However, further research is warranted into the RE and relative robustness of joint model MI and FCS MI when the ultimate aim is prediction, classification or clustering. Another direction of future research would be to compare FCS and RGL MI when data are missing not at random, the CRGL model is misspecified or FCS MI does not use compatible conditional models. 20 It is possible that FCS MI using incompatible conditional models may be more efficient than joint model MI using a misspecified joint model, especially when those conditional models have been chosen to fit well to the observed data.
In conclusion, FCS MI may be preferable to joint model MI using the compatible joint model, viz. the RGL model: it is more robust and is usually only slightly less efficient.
Footnotes
Acknowledgements
We thank Prof Yoav Ben-Shlomo and Dr Anne McCarthy, from the School of Social and Community Medicine, University of Bristol, for granting access to the BCGS data; Prof Chris Power for assistance in obtaining the NCDS data, and the Centre for Longitudinal Studies for providing these data; and Drs Finbarr Leacy, Ian White and Jonathan Bartlett for helpful discussions.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: SRS and RAH were supported by Medical Research Council [grant numbers U105260558, MR/J013773/1].