Abstract
Background
Systemic lupus erythematosus (SLE) is a multisystem autoimmune disease characterized by the production of various antibodies and immune complex-mediated injury. Limited information exists about Kazakh patients, a heterogeneous group with different clinical manifestations and potentially unique genetic basis.
Objective
To describe the clinical features, associated autoantibody and cytokine profile, and the frequency of rare variants in a limited panel of genes.
Method
This study enrolled 43 Kazakh individuals: 25 with SLE and 18 healthy controls. Disease activity was assessed using the SLEDAI-2K score. Laboratory tests included C3 and C4 complement components, interleukin (IL)-6, IL-5, IL-10, IL-18, IFN and antiphospholipid IgG/IgM identified by ELISA. The antinuclear factor (ANF) on HEp-2 cells was detected using indirect immunofluorescence. Specific autoantibodies were identified by immunoblotting. A custom AmpliSeq panel of 120 genes was used on the Ion Proton Sequencer.
Results
SLE patients (SLEDAI-2K = 11,48 ± 8,7) demonstrated skin lesions (88%), joint involvement (84%), lupus nephritis (56%), and hematological disorders (40% patients). Cardiac and vascular injury was each observed in 36% of patients, while involvement of nervous system, mucous membranes, and thyroid gland each occurred in 8% of cases. Immunological tests revealed positive ANF in the majority of patients (92%) with anti-dsDNA, nucleosomes, Smith, SS-A/Ro52, SS-A/Ro60, U1-snRNP, and Rib-P0 antibodies. IL-6, IL-18, IFN levels were markedly elevated in patients with SLE relative to controls (p = .02), reflecting enhanced systemic inflammatory activity. Elevated IL-10 was found as well in patients with SLE relative to controls (p = .02). According to the results of gene panel sequencing, the most significant variants were found in genes SAMD9L, REL, IRAK1, PTPRC, TLR7, TNFAIP3, IL6ST, BLK, CCR5, TFPI, CLEC16 A, IL2RB, ITGA2B, ABCC2, KIF5A, NCF2, and CD5, none of which showed statistically important enrichment in the disease cohort.
Conclusion
Analysis of data obtained from Kazakh patients with SLE revealed diverse autoimmune profiles, including various antibodies, pro- and anti-inflammatory cytokines, as well as several variants in REL, IRAK1, PTRPC, IL6ST genes. The findings presented may contribute to the development of personalized diagnostic tools for predicting disease trajectories and guiding treatment decisions.
Introduction
Systemic lupus erythematosus (SLE) is a multi-system autoimmune disorder characterized by immune inflammation with a wide range of antibodies and clinical manifestations.
The prevalence of SLE varies from 0.3 to 31.5 cases per 100,000 individuals per year. The average age at diagnosis is 43 (±15) years. 1 A higher mortality rate has been observed among African American SLE patients compared to European ones, which may be attributed to both the severity of the condition and socioeconomic factors. 2
Published data on SLE in Kazakhstan remain limited, but several key findings have emerged from national registry analyses and regional clinical reports. The highest SLE prevalence in Kazakhstan occurred in individuals aged 45–64 years and, as in many global cohorts, women account for nearly 90% of affected individuals. 3 According to national data from 2012 to 2017, more than 4,400 patients were registered, and the incidence of SLE increased by approximately 60% during this period. 4 Beyond these registry data, detailed information on SLE in the Kazakh population remains scarce, highlighting the need for more comprehensive clinical and molecular studies.
The exact etiology of SLE remains unclear, although recent research has shed light on complex interactions between the innate and adaptive immune systems involved in disease development. A distinctive feature of SLE is the production of antibodies against several nuclear and cytoplasmic antigens, including double-stranded DNA (anti-dsDNA), Smith antibodies, Sjögren’s-syndrome-related antigens A and B (anti-SS(A), anti-SS(B).5,6 There is a gradual increase in the number and concentration of certain autoantibodies prior to the onset of lupus, such as anti-dsDNA and anti-Smith.7,8 The production of autoantibodies against self-antigens is followed by the formation of immune complexes, their accumulation in tissues and activation of the complement system. 9
Recent genomic studies have expanded the understanding of SLE as a polygenic autoimmune disease involving more than 300 susceptibility loci. 10 Most risk variants are common regulatory polymorphisms that influence immune pathways such as type I interferon signaling, antigen presentation, and B-cell activation. 2 Rare, high-impact variants contribute to a smaller subset of cases - particularly early-onset or familial SLE.11,12 Despite this progress, much of the heritability remains unexplained, highlighting the need for continued investigation in diverse and understudied populations.
Objective
To describe the clinical features, the associated autoantibodies, and cytokine profile, as well as the frequency of rare variants in a limited panel of genes related to SLE.
Materials and methods
The study cohort included 25 adult individuals of Kazakh ethnicity diagnosed with SLE due to the classification criteria, developed by EULAR/ACR 2019, 13 based on the complete clinical history and prior documented ANA positivity. Participants were recruited in 2023-2024 from the outpatient rheumatology service and inpatient departments. Clinical evaluations were conducted to assess disease activity by SLEDAI-2K.
An age- and sex-matched control group of 18 healthy volunteers of Kazakh origin, without any known history of autoimmune or connective tissue disorders, was enrolled in the study through local wellness screening programs and public health outreach initiatives.
To preserve participant confidentiality, anonymized identifiers were assigned to all study subjects. SLE patients were coded with identifiers beginning with “L,” and control participants with “C”.
Ethical local approvals
The study design, ethical considerations, and recruitment procedures received approval from the Institutional Human Research Ethics Committee (Protocol No. 085/KI-79), and all aspects of the research were conducted in compliance with the principles of the Declaration of Helsinki and the approved Research Protocol. Individuals were recruited into the study by informed consent recorded in writing form.
Immune markers determination
The level of antinuclear factor and autoantibodies to ds-DNA, Smith, SS-A/60, SS-A/52, SS-B, Scl-70, CENP-B, and RNP/Sm was determined by indirect immunofluorescence (IIF) using ANF plus (screening test) and ANA cytobead (quantitative determination of autoantibodies) reagent kits on the Aklides automatic platform from Medipan and Generic Assays GmbH (Berlin, Germany).
The level of IL-6, IL-5, IL-10, IL-18, INF were investigated by Solid-phase ELISA using monoclonal and polyclonal antibodies to IL-6 on an Alisei Quality System analyzer (Calenzano, Italy). The concentration of antiphospholipid antibodies (APLA) IgM and IgG was investigated by solid-phase ELISA, manufactured by Medipan and Generic Assays GmbH (Berlin, Germany).
Genetic assays
Genomic DNA was isolated using the GENEJET™ Whole Blood Genomic DNA Purification Mini Kit. For the sequencing of genes a customized Autoimmune Panel was developed (Supplementary methods). 14 The AmpliSeq panel targets exonic and selected regulatory regions; intronic regions without known function were not analyzed.
Multiplex PCR was performed using the ION AmpliSeq™ Library Kit Plus. Each sample was uniquely tagged using the ION Dual Barcode Kit. Prepared libraries underwent purification with AMPure XP magnetic beads. Emulsion PCR was conducted using the Ion PI™ Hi-Q™ OT2 200 System. The resulting libraries were loaded onto Ion PI™ v3 chips.
Sequencing was performed on the Ion Proton™ platform with the Ion PI™ Hi-Q™ 200 Kit. All sequencing reads were aligned to hg19/GRCh37. Variant effect prediction incorporated the following tools in Ion Reporter: PolyPhen-2, SIFT, and internal Ion Reporter predictive scoring. Each variant was assessed for potential involvement in autoimmune disease pathogenesis. Public databases such as ClinVar and dbSNP, along with predictive algorithms, were used to evaluate the functional relevance of each mutation.
This comprehensive analysis allowed for the identification of meaningful genetic alterations with potential diagnostic and prognostic value. Controls were prepared and sequenced using the same protocol and batches were balanced across cases/controls to minimize batch effects.
Statistical analysis
The statistical analyses were performed using IBM SPSS version 21.0, GraphPad Prism 8 and Python version 3.13. Data plots were expressed as means ± standard deviation (SD). Determination of statistical method was based on Gaussian distribution of data sets that were examined by Shapiro-Wilk normality test. Hierarchical clustering and heatmap visualization were performed using Ion Reporter 5.2 software (ThermoFisher Scientific, USA). Statistical evaluation of categorical variables was conducted using exact Fisher test and the Chi-square (χ2) test. Linear regression and Pearson’s correlation were performed to determine whether a relationship existed between two variables. A p-value of <0.05 was considered statistically significant.
Results
Descriptive statistics for clinical and laboratory parameters in a cohort of patients.
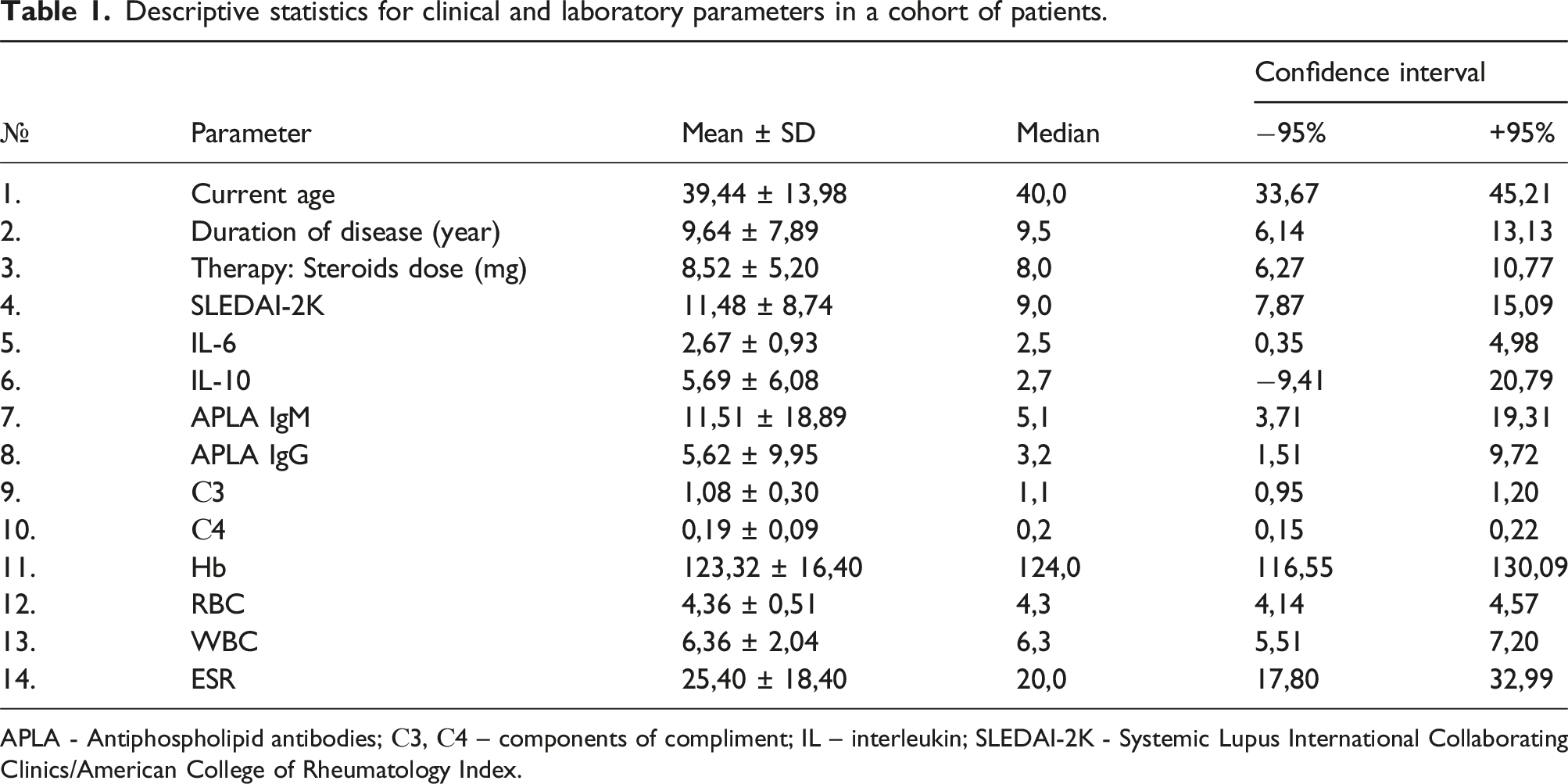
APLA - Antiphospholipid antibodies; С3, С4 – components of compliment; IL – interleukin; SLEDAI-2K - Systemic Lupus International Collaborating Clinics/American College of Rheumatology Index.
The frequency of clinical manifestations of systemic lupus erythematosus in the studied Kazakh cohort.
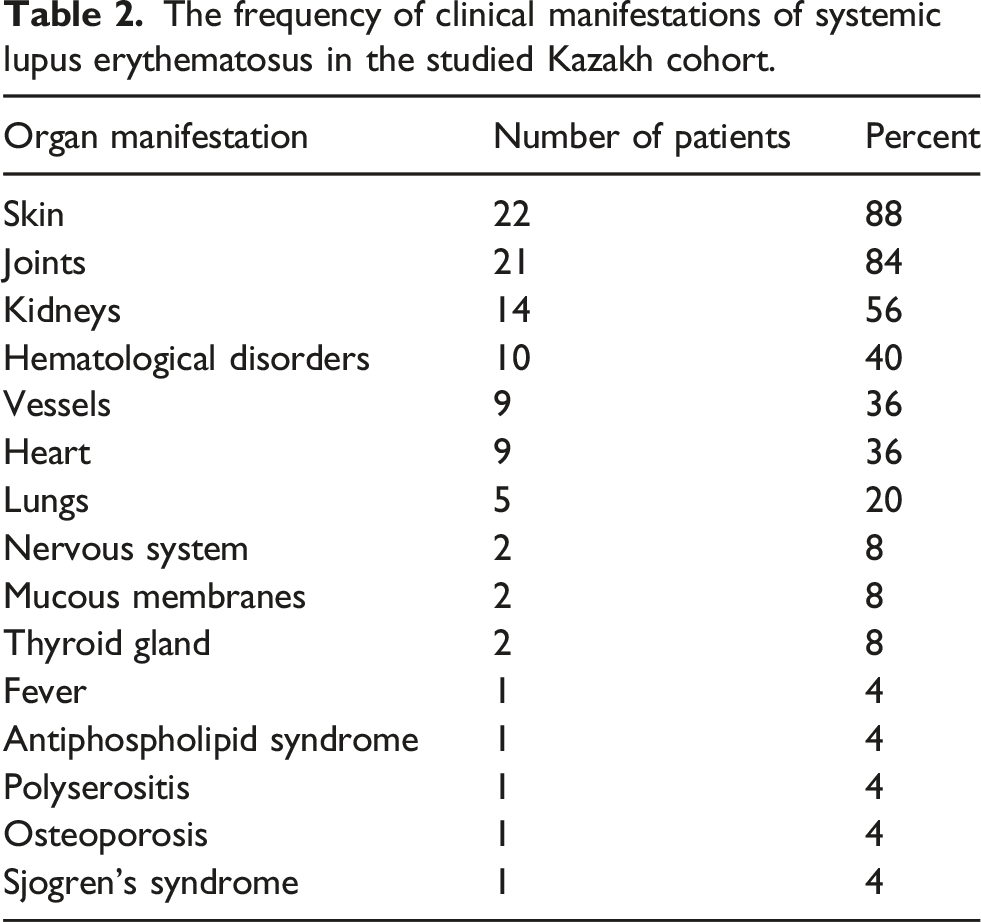
In addition, aseptic necrosis of the femoral heads was identified in one patient (4%), with low bone mineral density (osteoporosis and osteopenia) occurred in four patients (12%). Cushing syndrome was documented in four patients (16%), while kidney and heart insufficiency - in two patients, and respiratory failure - in one (4%).
The ANF on HEp2 cells was positive in 23 patients. The most common antibodies were anti-SS-A/Ro60, SS-A/Ro52, and anti-ds-DNA, known as diagnostic criteria for SLE (Figure 1). Anti-nucleosome, anti-Smith, anti-U1-snRNP, anti-Histone, anti-rib-P0 were positive in 16-24% of cases (Figure 1(c)). IgM APLAs were within the reference range in 88% individuals, IgG APLAs - in 96% (Figure 1(d)–(e)). None of the antibodies mentioned above appeared in control group. Antinuclear antibody, antibodies to extractable nuclear antigens and antiphospholipid antibodies in the main group of patients: (a) Antinuclear factor on HEp2 cells estimated by IELISA, titer 1:1280, nuclear homogeneous (AC-1) of glow; positive antibodies to ds-DNA, nucleosome, histone; (b) Antinuclear factor, titer 1:320, nuclear speckled (AC-4/5) and cytoplasmic dense fine speckled (AC-19/20) type of luminescence; positive antibodies to ds-DNA, SS-A/60, SS-A/52, U1-snRNP, RNP/Sm, ribP0, Histon, antibodies to Nucleosome and Smith showed a borderline result.; (c) Spectrum of antibodies (percentage of ANA/ENA) revealed in SLE patients; (d) Antiphospholipid antibodies (APLA) of IgM isotype; (e) APLA of IgG isotype. The concentration of antibodies was categorized based on reference intervals: within the reference interval (0.0–10.0 pg/mL) and above the reference interval (>10.0). n – number of patients; SLE – systemic lupus erythematosus.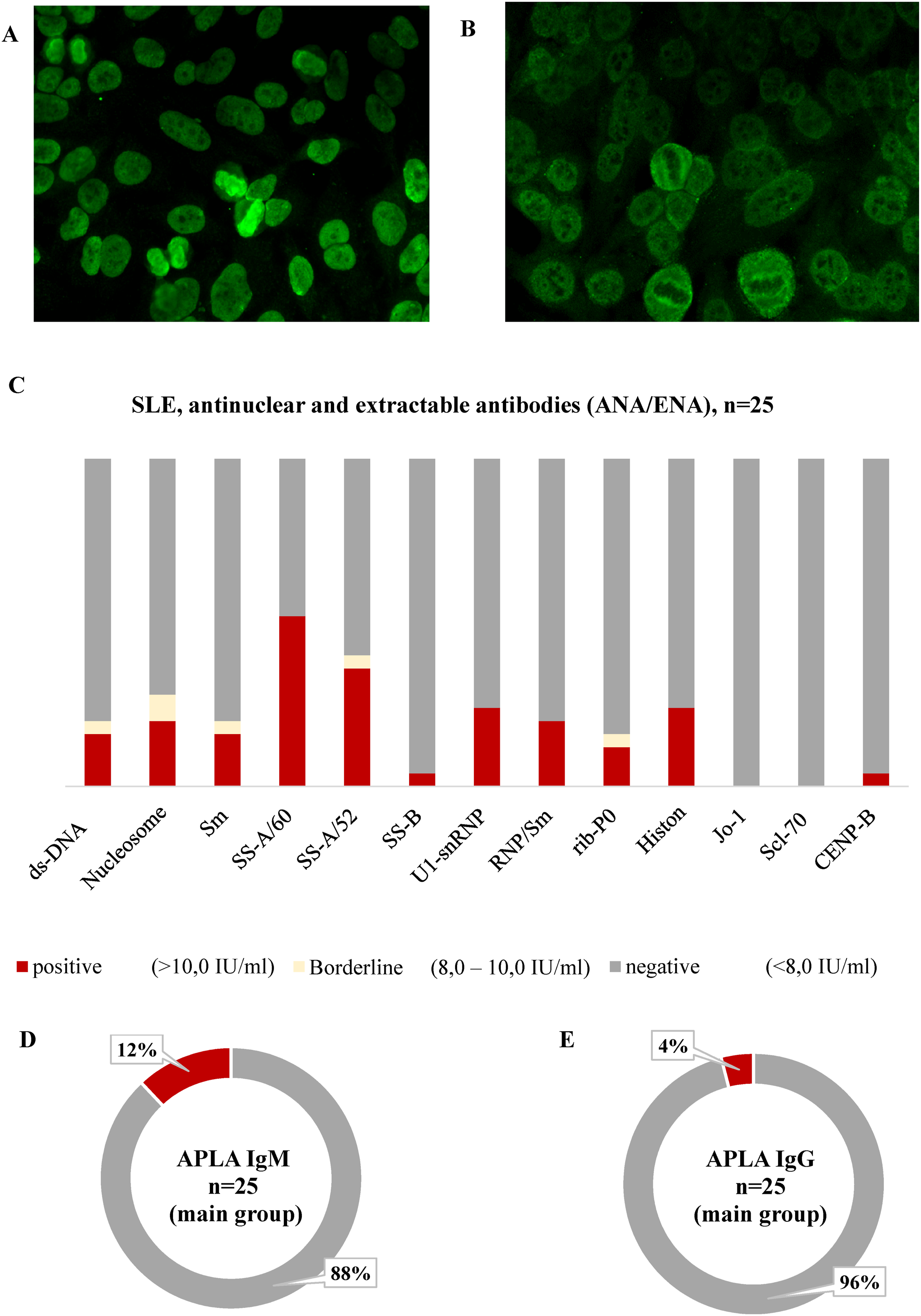
Evaluation of immunological-clinical associations using the χ2 test with Yates’s correction revealed significant associations between anti-dsDNA, nucleosomes, Smith, SS-B, RNP/Sm, U1-snRNP, ribosomal P0, CENP-B and cutaneous and joint manifestations (χ2 = 7–36, p < .001). Anti-dsDNA was associated with neurolupus, while anti-nucleosomes and anti-Smith antibodies - with glomerulonephritis (p < .04).
SS-A/Ro60 demonstrated associations with vascular lesions, cardiac involvement, central nervous system (CNS) and pulmonary manifestations (p < .05). In comparison, SS-A/Ro52 positivity was confined to cutaneous, joint, and CNS involvement (χ2 = 7, p < .01). CENP-B and ribosomal P0 antibodies correlated with renal pathology (p = .008).
Serum cytokine and complement measurements revealed several significant differences between SLE and control groups (Figure 2). IL-6 and IL-18 levels were elevated in SLE relative to controls (p < .004). Interferon (IFN) and IL-10 showed a modest elevation in SLE in comparison with healthy controls (p = .02). In contrast, IL-5 concentrations did not differ between groups, suggesting that Th2-mediated pathways are less prominent in this cohort. Comparative analysis of serum cytokines IL-6, IL-18, IFN, IL-10, C3, and complement components (C3, C4) levels in patients with SLE and healthy controls (*p value style: p = .1234 (ns), 0.0332 (*), 0.0021 (**), 0.0002 (***).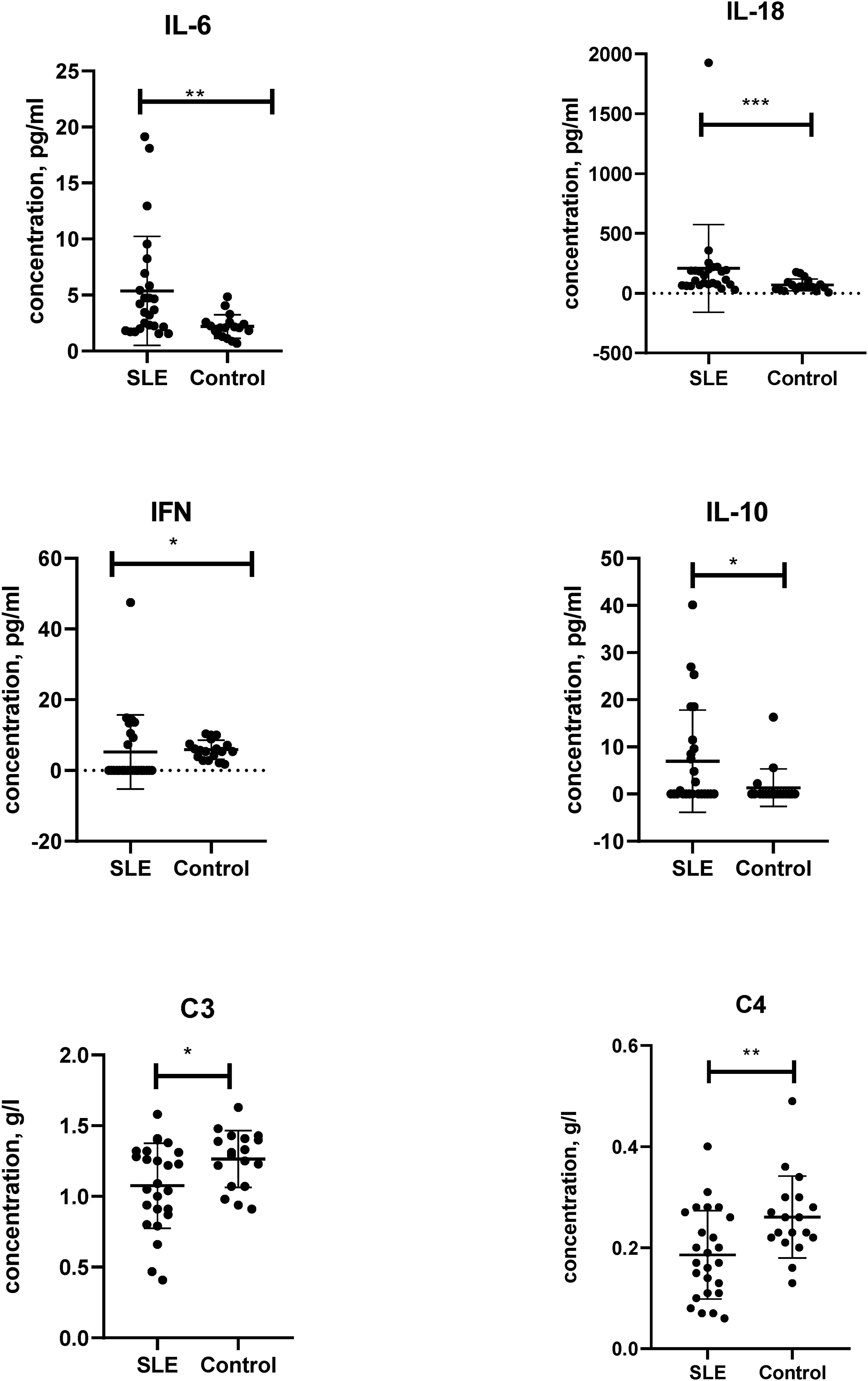
Complement analysis demonstrated characteristic reductions in C3 and C4 among SLE patients compared with healthy controls. C3 levels were significantly decreased (p = .03), while C4 exhibited an even more pronounced reduction (p = .007).
All cytokines and components of compliment were normal in all participants in the control group. Notably, IL-5, IL-6, IL-10 and IL-18 concentrations were correlated with SLEDAI-2K scores (p < .04).
A comprehensive analysis of genetic variants revealed both likely pathogenic and variants of uncertain clinical significance (VUS) across a range of immune-related genes with no difference between the SLE group and healthy individuals (Suppl.Tab.2). Likely pathogenic variants in REL, PTPRC, CD40, TNFAIP3, IL6ST, and TLR7 harbored more commonly in patients than in controls, suggesting their potential involvement in disease pathogenesis. For example, a variant in TNFAIP3 (chr6:138199775 T/TC), a critical negative regulator of NF-κB signaling, was detected in 5 patients and only once in the control group. In the PTPRC gene two likely pathogenic variants were found: chr1:198682102 AT/A and chr1:198687328 AT/A, which was absent in the control group.
Several immune signaling genes, including PTPRC, CD40, TNFAIP3, IL6ST, and TLR7, harbored likely pathogenic variants more commonly in patients than in controls. A likely pathogenic variant in the TNFAIP3 gene (chr6:138199775 T/TC) was found in 5 patients diagnosed with SLE and in one control individual. The variant in BLK gene chr8:11405631AG/A - classified as a VUS - was identified in 7 SLE patients and one control.
A variant of uncertain clinical significance in the TFPI gene (chr2:188361619 AT/A) was identified exclusively in the patient group. The same was with pathogenic variant in IL2RB (chr22:37528499 CT/C) and likely pathogenic variant in NCF2 gene (chr1:183532590 C/TG ref CG). Three variants in the ABCC2 gene were identified: chr10:101578956 CA/C (pathogenic), chr10:101563821 G/C (VUS), and chr10:101603641 CA/C (likely pathogenic), which was not registered in the control group. CD58, SAMD9L, ABCC2, CLEC16 A had multiple variants per gene, illustrating the presence of several distinct INDEL or SNV events within the same gene. Multiple likely pathogenic variants in SAMD9L (chr7:92764567 CA/C, chr7:92762089 C/T) were identified prevalently in SLE patients (Suppl.Tab.2).
A rare genetic variant located at chrX:153278833 (GCC/GCCG), currently classified as a variant of uncertain significance (VUS), was identified exclusively within the SLE cohort (n=4) and was absent in healthy controls. This variant resides within the IRAK1 locus, a gene encoding a critical kinase that orchestrates signaling through Toll-like receptor (TLR) and IL-1 signaling pathways.
Three variants (chr5:55247857 GT/G, chr5:55265588 AT/A) were prevalent in SLE patients than in controls (5 SLE vs 1 control). IL6ST mediates IL-6 signaling, promotes B cell maturation and Th17 differentiation, implicated in autoantibody production and tissue damage.
Two variants chr3:46415285 GC/G, chr3:46414784 G/T were present exclusively in SLE cohort. The clinical significance of these findings is underscored by experimental data indicating that CCR5 deficiency exacerbates nephritis in murine lupus by increasing ligand availability for CCR1.
Two variants of uncertain significance chr16:11217782 A/G, chr16:11260331 G/A were SLE-specific. These variants reside within the CLEC16A, which modulates autophagy, mitochondrial dynamics, and MHC class II presentation - processes linked to SLE.
Heatmaps with the distribution of gene variants demonstrated a notable enrichment of likely pathogenic and VUS variants in immunoregulatory genes among SLE patients (Figure 3). Comparative Heatmaps of Germline Mutations in 120 Target Genes for both the control and SLE groups. (a) Heatmap for 18 Control sample; (b) Heatmap for 25 Systemic lupus erythematosus samples. Rows represent 120 targeted genes, and columns - individual samples, which are hierarchically clustered based on mutation presence and predicted impact. The color scale indicates variant classification: green - variant of unknown impact, orange - missense variant, red - nonsense variant, maroon - splice variant, white - no detected variant.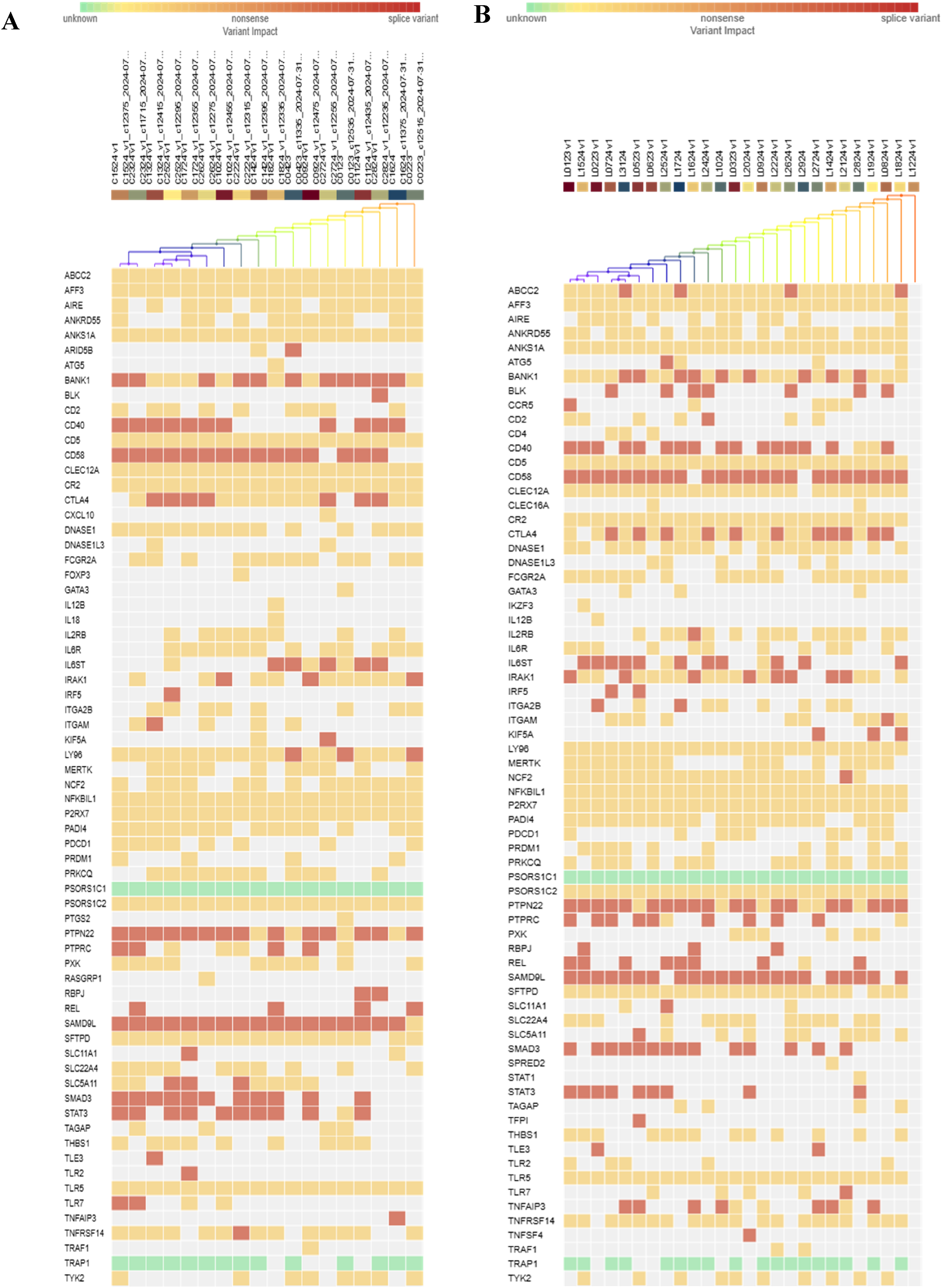
To explore differences in the frequency of genetic variants Fisher test and Yates-corrected chi-square (χ2) test were applied, with no significances detected. Notably, a variant in ABCC2 (chr10:101578956) demonstrated p = .0742, hinting at a possible trend toward differential distribution. Similar observations were made for IL6ST (chr5:55247857; p = .0730) and BLK (chr8:11405631; p = .1113). Overall, loci with p-values below 0.2 like ABCC2, IL6ST, IRAK1, and BLK do not meet criteria for significance, they may represent biologically relevant signals that warrant re-evaluation in an expanded dataset.
Analysis of genotype–phenotype associations revealed multiple relationships between immune-related gene variants and specific organ manifestations. TLR7, SLC5A11, CCR5, IL2RB, ITGA2B, NCF2, CD5, and CR2 demonstrated the largest chi-square values (χ2 = 28.9–32.2; p < .0001), indicating strong enrichment in patients with skin and joint pathology.
Several variants of uncertain significance and likely pathogenic have been linked to the risk of developing lupus nephritis. A subset of genes demonstrated significant associations with kidney manifestations, including SLC5A11, IRF5, BLK, CD58, SAMD9L, REL, CD40, TLR7, CCR5, TFPI, CLEC16 A, ABCC2, IL2RB, ITGA2B, KIF5A, NCF2, CD5, CR2 (χ2 11-14.0; p < .002). The most frequent variants in this dataset are in PTPN22 (chr1:114381166 CT/C) and BANK1 (chr4:102951316 GA/G). Several patients carry variants such in BANK1 which is known to be associated with B-cell signaling abnormalities. The combination of PTPN22 and BANK1 variants appears in multiple patients.
Variants in IL6ST (chr5:55247857 GT/G and chr5:55265588 AT/A detected in L0724 and L2824) and TNFAIP3 (chr6:138199775 T/TC in L1624 and L2024 patients) appear in patients with complex mutation profiles, suggesting potential links to more severe or fibrotic manifestations like lupus nephritis.
The most pronounced associations with CNS involvement were observed for PTPN22, CD58, and CD40 (χ2 = 14.0-16.6; p ≤ .004). SAMD9L also showed a weaker association (χ2 = 9.5; p = .002). Significant associations were found for PTPN22, CD58, and CD40 (p < .003) and pulmonary involvement.
Discussion
This study provides valuable insights into the clinical, immunological, and genetic characteristics of SLE within Kazakh cohort. Population-level studies from Central Asia and neighboring regions remain relatively sparse, but available epidemiological and clinical reports suggest broadly similar phenotypic patterns. The observed rates of irreversible organ damage in our cohort - particularly osteoporosis, kidney and cardiopulmonary failure - align with data from large international studies. 15 Cumulative glucocorticoid exposure has been identified as a major factor associated with increased risk of osteoporosis and avascular necrosis in the Hopkins Lupus Cohort. 16 Similarly, a report from the Kyoto Lupus Cohort documented a notable burden of vertebral fractures and emphasized their association with cumulative organ damage and glucocorticoid exposure. 17
Our findings can be placed in the context of the previously published large-scale description of SLE in Kazakhstan, included 4,448 patients across the country and demonstrated a steady rise in SLE incidence from 2012 to 2017, with a marked female predominance. 4 Similar to the registry, our patients were predominantly women and displayed a wide range of multisystem involvement. 4 However, our group shows a higher proportion of organ-specific complications, particularly cutaneous (88%), joint (84%), and renal manifestations (56%).
Treatment patterns in our cohort also align with national trends. 18 In the registry, nearly all patients received glucocorticoids (98%), and more than 40% were treated with cytotoxic agents or mycophenolate mofetil; hydroxychloroquine was prescribed in 39%. 18
In the main group most patients were positive for ANF, except two - with a clinical diagnosis of SLE established prior to the start of the study. This represents a well-recognized clinical phenomenon: ANA titters may fluctuate over time.
Multiple relationships between specific autoantibodies and organ involvement have been found. Antibodies to dsDNA, nucleosomes, Smith, SS-B, RNP/Sm, U1-snRNP, ribosomal P0, and CENP-B showed strong associations with cutaneous and joint manifestations. Anti-dsDNA was correlated with “butterfly” rash, arthralgia/arthritis and neuropsychiatric involvement. Up-to-date research suggested that they exhibit high specificity for SLE (96%), but their diagnostic sensitivity remains relatively low (52–70%). 19 According to the available literature, the detection of these antibodies may indicate the development of lupus nephritis. 20 However, our research and recent findings challenged this view. Our data supports the other research where anti-dsDNA were associated with higher prevalence of musculoskeletal involvement. 21
About 20% of Kazakhs patients demonstrated anti-nucleosome antibodies known for the association with glomerulonephritis that was revealed in our research as well (χ2 = 4.08, p = .04). The prevalence of anti-nucleosome antibodies in SLE varies from 50% to 100%, they can be used to diagnose SLE. 22 The probability that a patient with a positive result for anti-nucleosome antibodies will have SLE is 41 times higher than that of a patient with a negative result. 23
Anti-Smith antibodies appeared in 16% of patients and demonstrated a statistically significant association with the development of lupus nephritis (p = .04). Given their high specificity, these autoantibodies serve as critical longitudinal predictors of renal compromise and poor outcomes of SLE. 19
Moderate associations observed for histone, U1-snRNP antibodies with rash and arthritis (p < .001), without significant links to visceral involvement. 24% of our patients had anti-histone antibodies, typical for SLE, RA, Sjögren’s syndrome, inflammatory myositis, and mixed connective tissue diseases. 24
More than a half of Kazakh patients with SLE had anti-SS-A/Ro antibodies, which could be found in SLE, Sjögren’s syndrome, as well as other autoimmune diseases.25,26 SS-A/Ro60 demonstrated significant associations with vascular lesions, cardiac manifestations, pulmonary and CNS involvement (p < .05). SS-A/Ro52 positivity was confined to cutaneous, joint, and CNS injury, while SS-B exhibited an additional association with renal involvement.
Anti-RNP antibodies were detected in 24% Kazakh patients with SLE, though in other research 30–40% of SLE patients and in nearly all patients with mixed connective tissue disease demonstrated them. 27 These antibodies displayed significant associations with cutaneous and joint lesions (p < .0001), weaker with renal involvement (p < .04), indicating its broad multisystem activity.
The prevalence of anti-Rib-P was 12% in Kazakh patients, associated with erythema, arthritis, lymphopenia, neuropsychiatric symptoms, lupus nephritis, liver involvement, and juvenile SLE. 28 In Kazakh patients strong association was revealed between anti-Rib-P and cutaneous and joint manifestations (p = .0002).
Anticentromere autoantibodies were found in one patient, which is rare in SLE, more often for systemic sclerosis, Raynaud’s phenomenon, and CREST syndrome. 29 CENP-B and ribosomal P0 antibodies correlated with renal pathology.
The inflammatory interleukins (IL-6, IL-18), interferon, and hypocomplementemia were consistent with reports from large international SLE cohorts. 30 Serum IL-6, IL-18 correlated with disease severity and renal involvement,31,32 reliable with our patients which demonstrated IL-6 and IL-18 correlation with SLEDAI-2K scores (p < .04). IL-18 is known to promote IFN-γ production by T- and NK-cells and supports proliferation of activated T cells. 32 We have found correlation between the level of immunoregulatory IL-10 and SLEDAI-2K as well (p = .002).
The targeted sequencing dataset did not reveal statistically robust differences in variant frequencies between patients with SLE and the control group. A number of loci with well-established immunological relevance - such as TNFAIP3, PTPRC, CD40, IL6ST, TLR7 and SAMD9L - showed modest numerical enrichment among cases, but these shifts remained below thresholds typically interpreted as biologically decisive. Taken together, the patterns hint at perturbed NF-κB and TNF-linked pathways, particularly through alterations in TNFAIP3, TNIP1, IL6ST, and TRAF family members. Earlier studies have connected variants in TNFAIP3, including rs5029939, with renal or neuropsychiatric involvement in SLE, though population-specific discrepancies complicate generalization. 33
Polymorphisms in genes implicated in B-cell co-stimulation and tolerance - CD40, CD58, BANK1, BLK, PTPN22, PDCD1 - have been reported previously in SLE and other autoimmune diseases. 34 Lower expression of PTPRC (CD45) in SLE immune cell subsets 35 aligns with the presence of several variants in this gene in our cohort. Exclusive detection of ABCC2 variants in SLE cases is notable given the transporter’s broader roles in cellular stress responses. 36 Observed combinations of BANK1, BLK, and PTPN22 variants may further reflect shifts in B-cell activation thresholds, consistent with prior data linking BLK to renal disease susceptibility. 37
A cluster of mutations within genes central to type I interferon signaling - IRF5, TLR7, TLR9, TYK2, STAT1/4, IL12 B, IRAK1 - suggests sustained perturbation of interferon-dependent circuits, a long-recognized hallmark of lupus. 35 A variant in TFPI, identified solely in the patient group highlights the increasingly appreciated connections between coagulation pathways and inflammatory signaling. Exclusive or recurrent variants in IL2RB and CLEC16 A point toward involvement of T-cell homeostasis and autophagy-linked immune regulation. 38 Likewise, variants in SAMD9L are compatible with recent work indicating that alterations in this gene can amplify interferon-driven immunopathology. 39
Mutations in IRF5, TLR7, TLR9, TYK2, STAT1, STAT4, IL12 B, IL12RB, and IRAK1 suggest dysregulation of type I interferon signaling. 35 Activation of IFN response has been known as a hallmark of lupus, driving both innate and adaptive immune activation. 35 Variants IRF5, ITGAM, FCGRs, PTPN22, and BANK1 are known to elevate the risk of lupus nephritis. 40 Multiple PTPRC alterations and isolated mutations in CTLA4 underscore the convergence of antigen receptor signaling irregularities and impaired T-cell tolerance. 41 Variations in TNFAIP3, IL6ST, TLR7, and IRAK1 were more common among individuals with broader variant burdens, reflecting their association with severe or multi-system forms of SLE reported in other populations.42,43 Emerging phenotype-linked variation in SLC5A11 and ITGA2B may reflect ancestry-specific signals requiring further exploration. 44
Repeated variants in CLEC16 A across patients reinforce its role as an emerging autoimmune susceptibility gene, through effects on autophagy, mitophagy, and B-cell immune regulation. 38 Recent data demonstrate that non-coding CLEC16 A variants (rs17673553) can modulate enhancer activity and alter autophagy in B cells. 38
Similarly, SAMD9L variants appeared across multiple patients in our cohort. New evidence showed that SAMD9L mutations can impair hematopoiesis, promote immune dysregulation, and synergize with inflammatory stimuli such as interferons. 39
Several genetic polymorphisms are thought to influence susceptibility to lupus nephritis. 40 IRF5, ITGAM and FCGR, PTPN22 in chr1:114381166 CT/C and BANK1 in chr4:102951316 GA/G were found in Kazakh cohort. The combination of PTPN22 and BANK1 variants appears in multiple patients.
Several patients demonstrated PTPRC variant chr1:198682102 AT/A. This gene encoded CD45, a critical regulator of antigen receptor signaling in lymphocytes. 41
Variants in IL6ST and TNFAIP3 appear in patients with more complex mutation profiles, suggesting potential links to more severe or fibrotic manifestations like lupus nephritis. TNFAIP3 encodes A20, a ubiquitin-editing enzyme that suppresses NF-κB signaling. IL6ST encodes gp130, a signal transducer for IL-6 family cytokines, tied to inflammation and fibrosis. The lack of variant enrichment in our cohort is consistent with the well-established notion that SLE has a heterogeneous and highly polygenic genetic architecture, in which rare coding variants generally account for only a small fraction of the overall genetic risk. 45
The absence of statistically significant enrichment supports the widely accepted view that SLE is a polygenic disorder in which rare coding variants explain only a narrow fraction of the genetic contribution. 45 Monogenic forms do occur, they almost invariably present early in life or with unusual clinical patterns. 46 Even in cohorts enriched for childhood-onset disease, deleterious rare variants are identified in a minority of patients. 47 In this context, clinical and serological phenotyping continues to provide more practical insight for disease stratification than incomplete rare-variant profiling, especially in our adult population. 3
Limitations of this study include the relatively small sample size and the cross-sectional design. Our cohort predominantly reflects the adult Kazakh SLE population encountered in routine clinical practice, 3 which may exhibit a lower burden of early-penetrant rare variants. The utility of the current sequencing panel is shaped by its design timeline. Because it was finalized in early 2022, several gene classes consolidated in later high-impact reviews were not included. Nevertheless, the panel encompasses many of the mechanistic modules now considered central to rare-variant risk - genes involved in nucleic-acid clearance (DNASE1, DNASE1L3, TREX1), interferon signaling (IRF5/8, TYK2, STAT1/3/4, USP18, ISG15), autophagy (ATG5, ATG7, IRGM), TAM-receptor pathways (MERTK, TYRO3, GAS6), and regulators of B- and T-cell activation and tolerance (FOXP3, CD247, CTLA4, PTPN22, PRKCQ, BLK, BANK1, PRDM1). Recent work demonstrates that rare, high-effect variants typically map onto these same backbone pathways as common risk alleles, effectively bridging monogenic and polygenic disease architectures.12,45
These observations underscore the necessity for broader and deeper genetic approaches in future studies - whole-exome or whole-genome sequencing deployed in ancestrally diverse and earlier-onset cohorts. Non-coding and structural variants, which are invisible to targeted designs, are likely to refine our understanding of SLE susceptibility. Integrative approaches combining genomic, transcriptomic and epigenomic datasets may ultimately allow more precise molecular sub-classification of SLE and inform personalized therapeutic decision-making.
Conclusions and future perspectives
This first immunogenetic examination of SLE within a Kazakh cohort reveals heterogeneous autoantibody landscapes and a subset of variants in REL, IRAK1, PTPRC, IL6ST and related genes. Although no single variant reached statistical significance, the overall profile reflects the polygenic and pathway-layered nature of SLE. As molecular biomarkers become progressively intertwined with clinical decision-making, these findings provide an early framework for future personalized management strategies. Planned transcriptomic work, coupled with whole-genome sequencing and longitudinal clinical follow-up will help clarify how genetic and molecular features influence disease trajectories in this patient population.
Supplemental material
Supplemental Material - Clinical, serological, and targeted genetic analysis of systemic lupus erythematosus in Kazakhstan
Supplemental Material for Clinical, serological, and targeted genetic analysis of systemic lupus erythematosus in Kazakhstan by Lina Zaripova, Аbay Baigenzhin, Alyona Boltanova, Zhanna Zhabakova, Maxim Solomadin, Diana Makimova, Larissa Kozina in Lupus
Footnotes
Acknowledgments
The authors acknowledge Dr Natalya Krivoruchko for her expert consultation and contributions during the preparation of the revised manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Science Committee of the Ministry of Science and Higher Education of the Republic of Kazakhstan; Grant No. BR28713159.
Supplemental material
Supplemental material for this article is available online.
