Abstract
Background
Hepatocellular carcinoma (LIHC), a prevalent liver cancer with a grim prognosis due to high recurrence rates, is under scrutiny for its association with zinc finger proteins (ZNFs) in tumorigenesis. This study aims to create a prognostic model for LIHC incorporating ZNF-related genes.
Methods
By analyzing TCGA data, we identified differentially expressed genes (DEGs) between normal and LIHC samples, focusing on ZNF-related genes through univariate Cox and LASSO Cox regression. A multivariate Cox regression model was built, categorizing LIHC patients into high- and low-ZNFRS groups based on ZNF-related risk scores. Model performance was evaluated using ROC curves, with a nomogram integrating clinical data and ZNFRS. Immune microenvironment, enrichment analysis, mutations, and drug responses in LIHC were also explored.
Results
A prognostic model utilizing 10 ZNF-related genes accurately predicted LIHC survival. The low-risk group exhibited enhanced immune cell infiltration, contrasting with cell cycle and DNA replication enrichment in the high-risk group, which also displayed increased mutation rates. Promising drug candidates like SNS-314 and Decitabine warrant further investigation in LIHC treatment.
Conclusion
This study introduces impactful prognostic markers for LIHC management, emphasizing the significance of ZNFs in predicting patient outcomes and guiding treatment strategies.
Introduction
Hepatocellular carcinoma (LIHC) is a predominant form of liver cancer, ranking sixth in global cancer incidence and third in cancer-related deaths. 1 Even with considerable progress in the fight against LIHC, due to its stealth at onset and rapid advancement, as well as the absence of robust screening protocols, early-stage detection is rare, with only about 30% of patients diagnosed early.2–4 The treatment landscape for early LIHC patients is dominated by radical treatments aimed at curing the disease, including surgery and localized interventions. 5 Unfortunately, for those with later-stage LIHC, these options are often not viable, and treatment shifts to a combination of systemic therapies. 6 Despite the introduction of innovative treatments, the prognosis for these patients has not improved significantly,7,8 emphasizing the importance of discovering more accurate biomarkers and targets to enhance the effectiveness of LIHC treatments and extend patient survival.
Zinc finger proteins (ZNFs), a substantial part of the human proteome at about 5%, are involved in a variety of biological functions, such as telomere maintenance, DNA repair, chromatin remodeling, apoptosis, and autophagy.9,10 In recent decades, a mounting number of studies have exposed the crucial role of ZNFs in tumor development, progression, and metastasis. Gibbs et al. 11 have shown that ZNF165 interacts with SMAD3 to control the transcription of TGFβ-activated genes, promoting the proliferation of triple-negative breast cancer cells. In LIHC, ZNF703 overexpression is linked to faster tumor progression both in vitro and in vivo, which also induces epithelial-mesenchymal transition (EMT) by directly activating the CLDN4 gene. 12 Furthermore, the irregular expression of ZNF proteins or genes is a frequent observation in various cancers, indicating their potential as prognostic biomarkers. Poirier et al. 13 have found that ZNF768 expression is elevated in lung adenocarcinoma versus normal lung tissue, associating with clinical and pathological characteristics; depletion of ZNF768 in multiple lung cancer cell lines has been found to drastically suppress the proliferative capacity of these tumor cells. In liver cancer, elevated ZNF2 gene expression is indicative of a more severe patient prognosis, including reduced overall and disease-free survival. 14 While there has been research into the role of ZNFs in LIHC, the existing knowledge about the prognostic significance of specific ZNFs in LIHC is thin.
By using data from The Cancer Genome Atlas (TCGA), our research has developed and validated a model based on 10 ZNF-related genes for prognostic evaluation in LIHC patients. We have also probed how this model aligns with various clinical aspects of the disease, including immune profiles, functional pathways, somatic mutations, and drug sensitivities in LIHC patients, potentially unveiling new biomarkers for patient outcomes and treatment efficacy.
Materials and methods
Data source
From TCGA (https://portal.gdc.cancer.gov/), RNA expression, mutation, and clinical data for LIHC (normal: 50; LIH: 374) were obtained. The gene expression datasets GSE76427 (LIHC: n = 115) and GSE16757 (LIHC: n = 97) were downloaded from the Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) database to serve as validation sets.
Our search on the UniProt (https://www.uniprot.org/) database using “Zinc finger” yielded 1275 human-expressed ZNFs with a score of 5 or above
15
(Table S1). The main research process of this study is shown in Figure 1. Study flow chart.
Differential analysis and prognostic model development and validation
Within the TCGA cohort, we identified DEGs between normal and LIHC groups using the edgeR package (version 4.0.16), 16 with a filter of |logFC| > 0.585 and FDR < 0.01. We then intersected these DEGs with ZNF-related genes to obtain ZNF-associated DEGs.
Univariate Cox regression analysis was used to identify candidate genes with prognostic value for overall survival (OS) (p-value < 0.05). The glmnet R package (version 4.1-8)
17
was utilized for LASSO Cox regression analysis to reduce multicollinearity and further select genes. A prognostic model was constructed using multivariate Cox regression analysis with the survival R package (version 3.7-0).
17
The risk score for LIHC patients (Zinc finger protein related risk score, ZNFRS) was computed using the formula below.
From TCGA and GEO datasets, LIHC samples were categorized into high- and low-ZNFRS groups at the median ZNFRS expression level. Kaplan-Meier survival curves, generated with the survival R package (version 3.7-0), delineated the survival rate differences between these groups. Moreover, the model’s predictive accuracy was evaluated through receiver operation characteristic (ROC) curve analysis using the timeROC R package (version 0.4). 18
Establishment and verification of a ZNF-based nomogram
The rms R package (version 6.8-1) 19 was used to build a nomogram that combines risk scores with clinical factors—age, gender, tumor grade, and TNM stage. Predictive accuracy was appraised by comparing the calibration plots of predicted and actual survival rates.
Gene set enrichment analysis (GSEA)
Employing GSEA (version 4.3.3) on the TCGA-LIHC dataset, we uncovered differential gene expression profiles between high- and low-ZNFRS groups, focusing on KEGG pathways with an adjusted p-value less than 0.05 as significant.
Immune-related analysis
In the TCGA-LIHC cohort, the immune infiltration score and the activation of immune-related pathways were assessed using ssGSEA in the GSVA R package (version 1.50.0). 20 We also quantified the expression of immune checkpoint molecules and presented the data with boxplots. Moreover, we derived Tumor Immune Dysfunction and Exclusion (TIDE) scores from the TIDE (https://tide.dfci.harvard.edu/) platform for both groups and visualized them using violin plots.
Cell culture
Human liver Astrocytes (LX-2), human LIHC cells HepG2 and Huh-7 were purchased from Zhejiang MeisenCTCC, China. The cells were cultured in Rosewell Park Memorial Institute 1640 (RPMI-1640) medium supplemented with 10% fetal bovine serum (FBS) and placed in an incubator containing 5% CO2 at 37°C.
Cell transfection
Sh-PYGO2 and the negative control sh-NC were designed by Beijing Tsingke Biotech Co., Ltd. (China). Transfection was performed using Lipofectamine 3000 reagent (Invitrogen, USA) according to manufacturer’s instructions. After transfection and culture for 48 h, the cells were harvested for further experiments. The infection efficiency was verified by qRT-PCR.
qRT-PCR
Primer sequences.

Immunofluorescence
The cells were fixed in 4% paraformaldehyde for 15 min and permeated with 0.1% Triton X-100. After 3 rinses with TBST, cells were sealed with a sealing solution containing 1% BSA for 1 h. The cells were incubated overnight with the primary antibody CD19 (1:500) at 4°C and then with the Alexa Fluor 488 coupled IgG secondary antibody (Invitrogen, USA) for 1 h. The nucleus was stained using DAPI (Beyotime, China). Finally, a confocal microscope was used to observe and photograph.
Tumor mutation burden (TMB) analysis
Utilizing the maftools R package (version 2.18.0), 21 we explored the TCGA-LIHC SNV mutation data to compare the mutational burden between high- and low-ZNFRS groups. We identified the 30 genes with the highest mutation frequency in both groups and subsequently analyzed their mutation rates across the entire patient cohort.
Drug screening
Using CellMiner (https://discover.nci.nih.gov/cellminer/), we identified anti-cancer agents correlated with genes that predict treatment response and survival, focusing on drugs with lower IC50 values for greater tumor suppression.
Statistical analysis
All experimental data were analyzed using SPSS (version 22.0) (IBM, USA). All experiments were repeated at least 3 times. Data are expressed as mean ± standard deviation (SD). Differences between groups were assessed using the unmatched Student-t test or one-way analysis of variance. The image was analyzed statistically using ImageJ software. p < 0.05 was considered statistically significant.
Results
Construction and validation of a ZNF-gene-based prognostic model for LIHC
In the TCGA-LIHC cohort, comparative expression analysis was performed on normal and LIHC samples, surfacing 13,807 differentially expressed genes (DEGs) (Figure 2(A); Table S2). By focusing on ZNF genes, we filtered this list to 409 genes that were significantly associated with ZNFs (Figure 2(B); Table S3). Prognostic model construction. (A): Volcano plot of DEGs between normal and LIHC samples in the TCGA-LIHC cohort. (B): Venn diagram showing the intersection of DEGs with ZNF genes. (C and D): LASSO Cox regression analysis in the TCGA training set to select candidate genes. (E): Forest plot displaying ZNF-related genes associated with prognosis in LIHC based on multivariate Cox regression analysis results.
Subsequent univariate Cox regression identified a set of 100 genes that were significantly linked to patient prognosis (p < 0.05) (Table S4). Further refinement with LASSO Cox regression yielded a subset of 24 genes (Figure 2(C) and (D)), and ultimately, we developed a prognostic model based on 10 key genes (Figure 2(E)).
Employing the TCGA and GEO datasets (GSE76427), we determined the ZNFRS for LIHC patients, which allowed us to categorize them into high- and low-risk groups. Survival analysis within the TCGA dataset revealed that patients with higher ZNFRS had poorer OS compared to those with lower ZNFRS (Figure 3(A)). The predictive performance of our model was substantial, with AUC scores of 0.82, 0.77, and 0.73 for 1, 3, and 5 years, respectively (Figure 3(B)). The GEO dataset validation supported these results, showing improved survival for the low-risk group and AUC scores of 0.72, 0.74, and 0.88 for the 10-gene ZNF model over the same periods (Figure 3(C) and (D)). Hence, the 10-gene ZNF model demonstrates robust prognostic potential in LIHC. Model validation results in the TCGA training set and the GSE76427 validation set. (A) The Kaplan-Meier survival curves for the training set. High CDRS group in red and blue for low CDRS group. The abscissa is the survival years, and the ordinate is the survival rate. Each vertical line on the curve is for one patient. (B) ROC curves for the 1, 3, and 5-years survival rate of the training set. (C) Kaplan-Meier survival curves for the GSE76427 validation set. (D) ROC curves for 1, 3, and 5-years survival in the GSE76427 validation set. (E) Kaplan-Meier survival curves for the GSE16757 validation set. (F) ROC curves for the 1, 3, and 5-years survival rate of the GSE16757 validation set.
Establishment and validation of the prognostic nomogram
By developing a nomogram that accounts for both risk scores and clinical factors, we predicted the 1-, 3-, and 5-years OS for LIHC patients (Figure 4(A)). The calibration curves revealed high consistency between the predicted and actual survival rates (Figure 4(B)–(D)), validating its predictive capabilities. Nomogram development and validation. (A): Construction of a nomogram for LIHC patients that integrates risk scores with clinical variables (age, gender, stage, and TNM) to estimate 1/3/5-years survival. (B–D): Calibration plots for the 1-year (B), 3-years (C), and 5-years (D) predictions from the nomogram.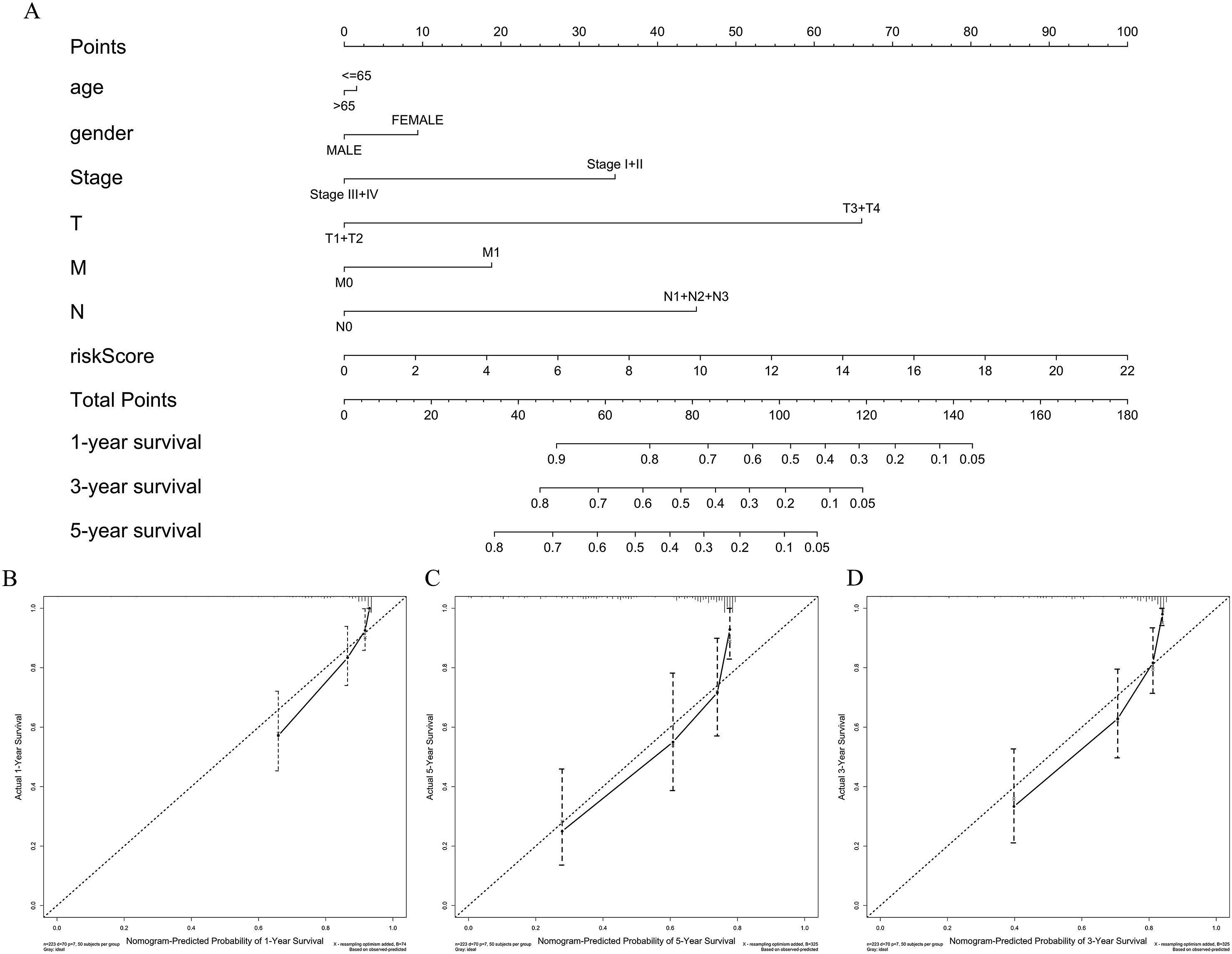
GSEA
Analyzing the TCGA-LIHC dataset with GSEA, we pinpointed key signaling pathways for high- and low-ZNFRS groups. The high ZNFRS group was notably enriched in KEGG pathways, including Cell cycle, Homologous recombination, DNA replication, Mismatch repair, and Oocyte meiosis (FDR q-val < 0.05) (Figure 5(A)). The low ZNFRS group, in contrast, was enriched for pathways such as Complement and coagulation cascades, Drug metabolism cytochrome P450, PPAR signaling pathway, Fatty acid metabolism, Glycine serine and threonine metabolism, and Retinol metabolism (FDR q-val < 0.05) (Figure 5(B)). KEGG pathway enrichment analysis. (A): Enriched Pathways in the high ZNFRS group. (B): Enriched Pathways in the low ZNFRS group.
Immunological infiltration and functional analysis for immunotherapy prognosis
Through ssGSEA analysis of the TCGA-LIHC dataset, we observed that patients with lower ZNFRS scores had a significant enrichment of immune cells including B cells, Mast cells, Neutrophils, NK cells, and T helper cells (p < 0.05), which was not the case in the high ZNFRS group (Figure 6(A)). Moreover, the low ZNFRS group had significantly higher scores in immune functions such as Cytolytic activity and IFN Responses (p < 0.05) (Figure 6(A)). Analyzing the expression of immune checkpoint genes, we found that the low ZNFRS group had a higher expression of specific immune checkpoint genes including CD14, GZMB, ADORA2A, and IDO2; in contrast, the high ZNFRS group showed a significant increase in the expression of other immune checkpoint genes like CD80, LGALS9, TNFRSF18, and VTCN1, which could be linked to the tumor immune evasion (p < 0.05) (Figure 6(B)). Furthermore, TIDE scores pointed towards a higher response to immunotherapy in the low ZNFRS group (Figure 6(C)), hinting at a possible advantage of immunotherapy for this cohort. Analysis of immune microenvironment in high- and low-ZNFRS groups. (A): Assessment of immune cell and immune function scores. (B): Levels of immune checkpoint gene expression. (C): TIDE scores. *signifies p < 0.05; **signifies p < 0.01; ***signifies p < 0.001.
Then, the expression levels of 10 characteristic genes in LIHC were predicted by TCGA database. The results showed that compared with the normal control group, KLF2, BCL11A, and ZNF385B were significantly lower expressed in LIHC (p < 0.05), while the other genes were significantly higher expressed in LIHC (p < 0.05) (Figure 7(A)). Then, the PYGO2 gene was randomly selected for preliminary experimental investigation. By qRT-PCR, it was found that PYGO2 was significantly overexpressed in HepG2 and Huh-7 cells compared with LX-2 cells (p < 0.05, Figure 7(B)). Further, we treated PYGO2 gene with low expression (Figure 7(C)). Meanwhile, immunofluorescence results showed that compared with the control group, the expression level of PYGO2 enhanced the fluorescence intensity of CD19 (Figure 7(D)). This indicated that PYGO2 was highly expressed in LIHC and reduced immune cell infiltration and inhibited cytokine secretion. Immune infiltration of model genes in LIHC. (A): The expression levels of model genes in LIHC. (B): qRT-PCR was used to verify the expression levels of model gene PYGO2 in LIHC cells. (C): Transfection efficiency of sh-NC and sh-PYGO2 was detected by qRT-PCR. (D): Immunofluorescence was used to detect the fluorescence intensity of CD19 in two groups of LICH cells.
Tumor mutation analysis
Our genetic mutation analysis results, depicted by a waterfall plot, showed TP53 and CTNNB1 as the most mutated genes in the high- and low-ZNFRS groups, respectively, with mutation rates of 44% and 28%, respectively (Figure 8(A) and (B)). Among the 10 genes in our prognostic model, BCL11A had the highest mutation rate at 28%, followed by ZNF385B at 16% (Figure 8(C)). Tumor mutation analysis. (A and B): Waterfall plots of the top 30 mutated genes in the high ZNFRS group (A) and the low ZNFRS group (B). (C): A waterfall plot displaying mutations in 10 signature genes across 25 LIHC samples.
ZNF-gene signatures for predicting LIHC drug response
As chemotherapy remains a central part of LIHC treatment, we investigated the implications of certain drugs for LIHC patients. The outcomes showed that an upregulation of KLF2 and ZNF385B was associated with increased resistance to chemotherapeutic drugs SNS-314 and Megestrol acetate. Conversely, a rise in BCL11A and KDM5B expression resulted in sensitivity to AZD-8055 and Decitabine (Figure 9). These correlations might guide future LIHC treatment decisions. Correlation between signature gene expression and drug sensitivity.
Discussion
LIHC is a tenacious malignancy within the digestive system, typically presenting at a late stage with a poor prognosis, as reported in medical research. 22 The absence of a robust prognostic model hampers clinical decision-making. The scientific community is increasingly recognizing the importance of ZNFs, with a focus on C2H2-ZNFs, in the advancement and metastasis of LIHC. Their role in biological functions, including EMT, cell proliferation, apoptosis, and chemoresistance, suggests that these proteins might be key in both treating LIHC and predicting patient outcomes. 23 Despite scattered evidence on the role of specific ZNFs in LIHC, a detailed investigation of these genes has been missing. This study pioneers the examination of the prognostic significance of ZNFs in LIHC and constructs a risk prognosis model based on these genetic signatures. It introduces new avenues for prognostic assessment and treatment in LIHC patients, potentially leading to better patient outcomes.
We developed a prognostic model for LIHC that incorporates 10 key genes: PYGO2, MAGEB2, TRIM39, ZNF746, KDM5B, SAP30, KLF2, BCL11A, ZDHHC22, and ZNF385B. Pygopus-2 (PYGO2), a significant part of the Wnt/β-catenin signaling, is overexpressed in LIHC and is associated with poor prognosis, including larger tumor size, vascular invasion, and tumor differentiation.24,25 The potential of PYGO2 as a therapeutic target is highlighted by its ability to inhibit cell proliferation, invasion, and angiogenesis when downregulated in LIHC. 26 ZNF746, a ZNF and a known substrate of the E3 ligase Parkin (PARIS), is implicated in cancer progression and EMT.27,28 The study by Kim et al. 29 reveals that ZNF746 overexpression promotes LIHC progression by inhibiting PGC-1α transcription, making ZNF746 a potential therapeutic target for LIHC treatment. Lysine-specific demethylase 5B (KDM5B) is a key player in the advancement of multiple cancers, promoting tumor growth and metastasis.30,31 Within LIHC, elevated KDM5B is associated with poorer patient outcomes 32 and contributes to the development of chemoresistance and cancer stem cell properties. 33 SAP30, integral to the Sin3/HDAC repressive complex, influences cellular activities. 34 Evidence from LIHC research points to a correlation between elevated SAP30 levels and reduced OS.35,36 KLF2, a member of the Krüppel-like factor family, is gaining recognition for its ability to combat LIHC. Li et al. 37 have uncovered the tumor-suppressive activity of KLF2 through its interaction with the TGF-β pathway, offering new insights into LIHC biology. Moreover, the long non-coding RNA GHET1 is now known to drive LIHC proliferation by downregulating KLF2, and its elevated levels are associated with an unfavorable prognosis. 38 TRIM39, also known as ring finger protein 23, is a significant factor in the modulation of immune responses and the regulation of cell cycle and apoptosis.39,40 It has been linked to the promotion of cell proliferation and migration in LIHC. 41 The protective role of TRIM39 in LIHC, however, remains unclear, possibly due to the heterogeneity of the disease and the variability in patient samples. There is a significant knowledge gap regarding the functions of MAGEB2, BCL11A, ZDHHC22, and ZNF385B in LIHC. Future research could bridge this gap, potentially uncovering new therapeutic targets and strategies for LIHC.
Our research into LIHC has revealed that the level of immune cell infiltration varies among patient groups, potentially impacting their immunotherapy outcomes. Those with lower ZNFRS scores show a higher presence of immune cells like B cells, mast cells, neutrophils, NK cells, and Th cells, indicating a more engaged tumor microenvironment that might be better at recognizing and attacking tumor cells. B cells are pivotal in the immune response against cancer, capable of both directly attacking tumor cells with antibodies and supporting T-cell responses through antigen presentation and cytokine secretion. 42 Their role also extends to the formation of tertiary lymphoid structures (TLS) within tumors, which could aid in the activation and proliferation of immune cells. 43 Mast cells are key modulators of antitumor immunity, deploying mediators that activate and shape the immune cell landscape. 44 They amplify the immune response to cancer by recruiting and activating immune cells, including NK cells, dendritic cells, and T cells, and, by interacting with them, increase the activity of these immune cells.45,46 Neutrophils can directly target tumor cells with cytotoxic molecules like ROS and RNS. 47 They also differentiate into APC-like cells, presenting tumor antigens to T cells and thus strengthening the antitumor immune response.48,49 Equipped with a range of receptors, NK cells are adept at recognizing and eliminating cancer cells. They form immunological synapses with tumor cells and use perforin and granzymes to induce cell death.50,51 Moreover, NK cells communicate with dendritic cells (DCs), a crucial interaction that modulates the overall antitumor response. By secreting cytokines like IFN-γ and TNF-α, NK cells can stimulate DC maturation, enhancing the adaptive immune response and tightening immune surveillance against cancer. 52 Th1 cells, a subset of T helper cells, secrete cytokines that activate cytotoxic T cells (CTLs) and natural killer (NK) cells, thus strengthening their ability to eliminate cancer cells. 53 Our analysis indicates that LIHC patients with lower ZNFRS levels are more responsive to immunotherapy, suggesting that a stronger immune response could be the key to unlocking the full potential of treatment. The research emphasizes a possible correlation between immune cell infiltration levels and the effectiveness of immunotherapy in LIHC patients, which could be pivotal for developing new treatment strategies. Future investigation should delve deeper into the mechanisms of action of these immune cells in LIHC and their role in shaping immunotherapy responses.
Our research has zeroed in on SNS-314, Megestrol acetate, AZD-8055, and Decitabine as promising candidates for LIHC treatment. SNS-314 as a pan-inhibitor of Aurora kinases, stands out for its potent anti-tumor activity and lower IC50 values. It has been shown to deplete YAP and accumulate P21, driving LIHC cell polyploidy and apoptosis. 54 Megestrol acetate (MA), as a 17alpha-hydroxyprogesterone derivative, has been found to substantially enhance the survival rate of LIHC patients, increasing it from 7 to 18 months, and to mitigate tumor growth in non-resectable cases. 55 The possibility that MA could be affecting LIHC through other hormone receptors adds to its therapeutic impact.56,57 In this study, the expression levels of SNS-314 and MA were positively correlated with KLF2 and ZNF385B, respectively. At the same time, both KLF2 and ZNF385B were lowly expressed in LIHC. Therefore, we speculate that KLF2 and ZNF385B may play a role in the chemotherapy resistance of LIHC by inhibiting the proliferation, migration and invasion abilities of cancer cells and promoting apoptosis.58,59 Research has pointed to the anti-LIHC potential of AZD-8055, a dual inhibitor of mTORC1/2, showing that AZD-8055 exhibits anti-tumor activity in preclinical LIHC models, and this activity is further enhanced when AZD-8055 is combined with C6 ceramide. 60 The PI3K/AKT/mTOR pathway, a key regulator in LIHC, is targeted by Decitabine, leading to partial apoptosis. 61 The addition of Vorinostat to this regimen has been shown to enhance its antitumor activity in HepG2 human LIHC cells, indicating a promising direction for combinatorial therapies. 62 In pancreatic cancer, KDM5B is upregulated and promotes drug resistance. 63 KDM5B is also highly expressed in nasopharyngeal carcinoma (NPC), and silencing its expression makes cells sensitive to cisplatin. 64 In this study, KDM5B was highly expressed in LIHC, and the IC50 value of Decitabine was significantly negatively correlated with the expression level of KDM5B. This may be because the types of cancers and drugs that the studies looked at were different. Therefore, further experiments are needed to verify the potential mechanism of action between the model gene and the drug. These drugs could be the next frontier in LIHC treatment. In addition, most of the current studies on these drugs in LIHC are combined drugs. Therefore, the efficacy and safety of each drug and how it is administered still need to be further validated through clinical trials.
Yet, our work is not without limitations. The limited clinical data in public cohorts may conceal the model’s potential correlations with certain clinical variables, underscoring the need for more exhaustive data collection. Second, the TCGA-LIHC dataset used in this study was drawn primarily from patients in the United States and may not fully reflect the characteristics of LIHC patients from different regions and ethnicities around the world. Therefore, the applicability of the findings may be limited. Although we performed validation in TCGA and GEO datasets, the sample size is still limited. This can result in some analyses having insufficient statistical power to detect some potentially significant differences. Third, in constructing the prognostic model, we used LASSO Cox regression analysis, which assumes that only a few genes have a significant impact on prognosis. While this approach helps reduce overfitting, it can also miss some important features. Fourth, when screening DEGs, we used |logFC| > 0.585 and FDR < 0.01 as thresholds. While these thresholds help screen out significant DEGs, they may also miss some biologically important genes. Fifth, although we predict potential small-molecule drugs for LIHC, these predictions need to be validated through in vitro drug screening and trials. Future studies could consider more comprehensive analyses using more relaxed thresholds or in combination with other bioinformatics tools. Future studies need to be validated in multi-center, large-scale datasets to reduce the effects of dataset bias and sample size limitations. At the same time, the combination of multiple computational methods and experimental verification will contribute to a more comprehensive understanding of genetic characteristics and drug resistance mechanisms. Finally, despite these limitations, our findings provide useful insights into risk assessment and treatment of LIHC.
In conclusion, this study constructed a risk-prognostic model for 10 ZNF-related feature genes, which can accurately predict the prognosis of LIHC patients. At the diagnostic stage, clinicians can perform genetic tests on tumor tissue from LIHC patients, including expression levels of ZNF-related characteristic genes. According to the test results, the risk score of patients was calculated using the risk score model developed in this study, and the patients were divided into high and low risk groups. The prognosis was then predicted using a nomogram combined with risk scores and clinical features. Patients in the low ZNFRS group may respond better to immunotherapy due to their higher levels of immune cell infiltration. Therefore, immunotherapy regimens can be prioritized. For patients in the high ZNFRS group, due to their higher rate of gene mutation, they may be resistant to certain chemotherapy drugs or targeted drugs. Therefore, more appropriate treatment options need to be selected based on drug sensitivity analysis.
For some complex cases, combination treatment strategies, such as the combination of immunotherapy with chemotherapy or targeted therapy, may be considered to improve treatment effectiveness. In the course of treatment, clinicians can regularly carry out genetic testing on the tumor tissue of patients to monitor the expression changes of ZNF-related characteristic genes and then adjust the treatment plan in time. According to the follow-up data, the patient’s risk score was updated to provide reference for follow-up treatment. With these strategies, the ZNF related prognosis model can provide clinicians with a powerful tool to help more accurately assess patient prognosis and develop personalized treatment plans.
Supplemental Material
Supplemental Material - Exploring the utility of zinc finger protein-related genes in predicting hepatocellular carcinoma prognosis, immune responses, and drug efficacy
Supplemental Material for Exploring the utility of zinc finger protein-related genes in predicting hepatocellular carcinoma prognosis, immune responses, and drug efficacy by Pengtao Zhai, Mei Li and Yuan Cheng in Human & Experimental Toxicology
Footnotes
Ethical statement
Ethical approval
TCGA and GEO belong to public databases. The patients involved in the database have obtained ethical approval. Users can download relevant data for free for research and publish relevant articles. Our study is based on open source data, so there are no ethical issues.
Author contributions
Conception and design of study: Pengtao Zhai.
Acquisition of data: Pengtao Zhai, Mei Li.
Analysis and interpretation of data: Yuan Cheng.
Drafting of the manuscript: Pengtao Zhai, Mei Li.
Revising the manuscript critically for important intellectual content: Yuan Cheng.
Approval of the version of the manuscript to be published: Pengtao Zhai, Mei Li, Yuan Cheng.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data and materials in the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.

