Abstract
Dysregulation of long intergenic non-protein coding RNA 00,641 (LINC00641) is associated with the malignancy progression of multiple cancers, including thyroid carcinoma. The current study aimed to determine the role of LINC00641 in papillary thyroid carcinoma (PTC) and the underlying mechanism. We found that LINC00641 was downregulated in PTC tissues and cells(p < 0.05), and overexpression of LINC00641 inhibited PTC cell proliferation and invasion, and induced apoptosis(p < 0.05), while silencing LINC00641 promoted the proliferation and invasion in PTC cells, and inhibited cell apoptosis(p < 0.05). Furthermore, we found that Glioma-associated oncogene homolog 1 (GLI1) expression was negatively correlated with LINC00641 expression in PTC tissues (r2 = 0.7649, p < 0.0001), and silencing GLI1 inhibited PTC cell proliferation and invasion, and induced apoptosis(p < 0.05). Meanwhile, RNA immunoprecipitation (RIP) and RNA pull-down assays confirmed that insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1) bound to LINC00641 as an RNA binding protein, and overexpression of LINC00641 destabilized GLI1 mRNA by competitively binding to IGF2BP1. Rescue experiments revealed that overexpression of GLI1 restored the inhibitory effect of LINC00641 overexpression on activation of the AKT pathway, as well as PTC cell proliferation and invasion, and counteracted the induction of cell apoptosis by LINC00641 overexpression. Finally, in vivo experimental results showed that overexpression of LINC00641 markedly suppressed tumor growth and reduced expression of GLI1 and p-AKT in xenograft tumor mice(p < 0.05). In summary, this study highlighted that LINC00641 plays a critical role in the malignant biological progression of PTC by regulating the LINC00641/IGF2BP1/GLI1/AKT signaling pathway, which may serve as a potential therapeutic target for PTC.
Introduction
Thyroid carcinoma is a type of endocrine malignancy characterized by strong invasiveness, which accounts for 1% of all human cancers. 1 The incidence rate of papillary thyroid carcinoma (PTC), a common subtype of thyroid carcinoma, is rising worldwide in recent years. 2 There are many typical treatment strategies for PTC, including thyroidectomy and radioiodine ablation, which have displayed favorable outcomes. However, the poor prognosis resulting from recurrence and distant metastasis is still a challenge. Genetic/epigenetic alteration events were involved in the progression of PTC. 3 Hence, exploration of PTC pathogenesis is essential to determine accurate prognostic markers and effective therapeutic targets.
Aberrant expression of long non-coding RNAs (lncRNAs) have emerged as an important component of tumorigenesis in multiple cancers, including PTC. Increasing evidence demonstrated that lncRNAs could regulate the malignant growth and aggressive metastasis of PTC. LncRNA CASC2 was downregulated in PTC tissues and closely correlated with poor prognosis, and overexpression of CASC2 significantly suppressed PTC cell proliferation and induced apoptosis via inactivating the AKT/ERK1/2 signaling pathway. 4 Li et al. reported that LINC00460 was upregulated in PTC tissues and cell lines, and LINC00460 could act as a competing endogenous RNA for miR-613 to upregulate the expression of Sphk2, thereby promoting PTC growth. 5 LncRNA RP11-476D10.1 were highly expressed in PTC cells and bound to miR-138-5p to promote LRRK2 expression. Silencing lncRNA RP11-476D10.1 enhanced the apoptosis and autophagy of PTC cells, while reducing cell proliferation. 6 LncRNA AB074169, as a tumor inhibitor, could increase CDKN1a expression and decrease CDK2 expression by interacting with KHSRP, thus suppressing cell proliferation in PTC. 7 These findings indicated that lncRNAs play an important role in the pathogenesis of PTC and may serve as prognostic markers and therapeutic targets for PTC.
LINC00641 was found to be downregulated in non-small cell lung cancer cells and could upregulate PLSCR4 by sponging miR-424-5p. 8 Zhang et al. reported that LINC00641 overexpression suppressed cell proliferative, migratory and invasive abilities, as well as epithelial mesenchymal transition characteristics in cervical cancer. 9 In gastric adenocarcinoma, LINC00641 participated in regulating oxaliplatin resistance by activating autophagy. 10 Bioinformatics analysis revealed that high expression of LINC00641 was associated with longer disease-free survival time in patients with thyroid carcinoma. 11 However, the regulator role of LINC00641 in PTC has not been characterized. Therefore, the biological function and molecular mechanism of LINC00641 in PTC deserve deeply illuminated.
Glioma-associated oncogene proteins 1 (GLI1) belong to the family of Krüppel-like transcription factors with C2H2-Zn finger DNA-binding domains, which is a crucial transcription factor in the hedgehog signaling pathway. 12 GLI1 protein contribute to tumor progression by inhibiting tumor cell apoptosis, stimulating cell cycle progression, and promoting tumor cell invasion and metastasis. 13 Parascandolo et al. demonstrated the association between GLI1 expression and various types of thyroid cancers. 14 They found that all thyroid tumors, regardless of their histological type, were positive for GLI1 protein, while all normal thyroid tissues were negative. Moreover, Lee et al. reported an association of the expression levels of GLI1 with lymph node metastasis in patients with PTC. 15 These studies highlight a central role of GLI1 in thyroid cancer, but the differential expression of GLI1 mRNA in cancer is not fully understood. Insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1) is a conserved RNA-binding protein acts in post-transcriptional mRNA regulation. 16 It could directly interact with GLI1 mRNA, dampening its degradation. 17
In current study, we found that the expression of LINC00641 was significantly downregulated in PTC and enhanced the stability of GLI1 mRNA by binding to IGF2BP1, which inhibited PTC cell proliferation, invasion, and induced apoptosis. Collectively, our data demonstrated that LINC00641 plays a critical role in PTC progression and might serve as a potential therapeutic target.
Materials and methods
Tissue samples
Twenty-four human PTC tissue samples and pair-matched adjacent non-neoplastic tissue samples were obtained from patients undergoing surgical resection at the First Affiliated Hospital of Zhengzhou University, including 14 patients were in cancer stages Ⅰ-Ⅱ, and 11 patients were in stages Ⅲ-Ⅳ. The patients were diagnosed as PTC via pathological examination and did not receive radiotherapy or chemotherapy before the surgery. All samples were snap frozen in liquid nitrogen and maintained at −80°C. The ethics committee of Zhengzhou University (Zhengzhou, China) approved the use of human tissue samples.
Cell culture
The human PTC cell lines (TPC-1, BCPAP and IHH-4) and normal thyroid epithelial cell (Nthy-ori 3-1) were purchased from the European Collection of Cell Cultures (Wiltshire, UK). BCPAP and Nthy-ori 3-1 were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium mixed with 10% fetal bovine serum (FBS; Gibco, Grand Island, NY, USA) while other cells were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone, Life Technologies, CA) containing 10% FBS. All cells were cultured in a humidified incubator at 37°C with 5% CO2.
Cell transfection
For cell transfection, LINC00641, IGF2BP1 and GLI1 were all sub-cloned into the pcDNA3.1 plasmids (pcDNA-LINC00641, pcDNA-IGF2BP1 and pcDNA- GLI1) to upregulate LINC00641, IGF2BP1 and GLI1, respectively. And the shRNAs targeting LINC00641, IGF2BP1 and GLI1 (sh-LINC00641, sh-IGF2BP1 and sh-GLI1) were separately constructed to downregulate LINC00641, IGF2BP1 and GLI1. All plasmids obtained from Invitrogen were transfected into TPC-1 and BCPAP cells by using Lipofectamine 2000 (Invitrogen; Thermo Fisher Scientific, Inc.) conforming to the guide books of the providers and then the cells were incubated for 24 h.
RNA extraction and quantitative real‐time PCR (RT‐qPCR)
Total RNA was extracted from tissues and cells with Trizol reagent (Takara). RNAs were reversed transcribed into cDNA with the reverse transcription kit (Takara). The quantitative analysis was operated by the SYBR Green PCR Kit (Takara). GAPDH was used as an internal control. Applied Biosystems Step One Plus Real‐Time PCR System (Applied Biosystems) was employed to analyze the results of RT‐qPCR with the following steps: an initial denaturation step at 95°C for 3 min, followed by denaturation at 94°C for 15 s, annealing at 55°C for 25 s and extension at 72°C for 15 s for 35 cycles. The 2−ΔΔCt method was applied to detect the relative expression levels of target genes.
Cell counting kit-8 (CCK-8) assay
CCK‐8 (Dojindo Molecular Technologies, Inc) was utilized to evaluate cell proliferation ability. The cells were seeded into 96‐well culture plates at a density of 1 × 103 cells per well. Afterwards, the cells underwent 24, 48 and 72 h of incubation in a humidified incubator at 37°C containing 5% CO2, followed by adding CCK‐8 solution (100 μL) to each well. Then the incubation continued for another 4 h. The absorbance at 450 nm was examined by employing a Multiskan Go spectrophotometer (Thermo Fisher Scientific, Inc).
Transwell assay
The invasion of cells was assessed by employing a Transwell insert chamber coated with or without Matrigel (BD Biosciences). The cells were incubated in the upper layer containing 300 μL of serum‐free medium, and the bottom chamber was filled with 10% FBS. After cultured for 48 h, cells remaining in the upper chamber were cleaned, and cells invading the bottom chambers were pictured and counted under a microscope (Leica, Wetzlar, Germany).
RNA immunoprecipitation (RIP)
RIP experiments were performed with the Magna RIP RNA‐Binding Protein Immunoprecipitation kit (Millipore). TPC cells were lysed by using RNA lysis buffer that was supplemented with protease and RNase inhibitors, and cell lysates were cultured with protein A/G magnetic beads coated with IGF2BP1 antibody (#sc-166,344) or IgG at 4°C overnight. After washing, the immunoprecipitated RNA was purified and then quantified by RT‐qPCR analysis.
RNA pull-down
RNA pull‐down assay was performed according to the suggestion of the manufacturer. In brief, The LINC00641 or antisense LINC00641 cDNA was cloned into pBluescript II vector. Biotin-labeled RNAs were transcribed in vitro and purified. The biotinylated LINC00641 and biotinylated LINC00641 antisense were, respectively, incubated with cellular protein extracted from TPC cells, and afterwards streptavidin beads were introduced. After 48 h, the recovered proteins associated with bio‐LINC00641 or bio‐LINC00641 antisense or control were resolved. The eluted solutions were analyzed by western blot assay.
Cell apoptosis
Cell apoptosis was measured by flow cytometry with Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (APOAF, Sigma, St Louis, MO, USA) according to the manufacturer’s protocols. The cells were collected and incubated with Annexin V-FITC and PI for 20 min in the dark. The apoptotic cells were analyzed by using a flow cytometer (BD Biosciences, Franklin Lakes, NJ, USA).
Western blot assay
Total proteins of cells or tissues were extracted in RIPA lysis buffer (Sigma-Aldrich, St Louis, MO, USA) containing 1% protease inhibitor. Then, the proteins were collected and quantified with BCA protein assay kit (Beyotime, Shanghai, China), followed by denaturation at 98°C for 10 min 25 μg of protein samples were separated by 10% SDS-PAGE and transferred to PVDF membranes (Millipore, Billerica, MA, USA). The membranes were blocked with 5% non-fat milk in Tris-buffer saline containing 0.1% Tween 20 (TBST) for 1 h at room temperature, and incubated with primary antibodies overnight at 4°C, following by incubated with HRP-conjugated secondary antibody (ab6721) for 2 h at room temperature. The antibodies against IGF2BP1 (#sc-166,344, 1:1000), GLI1 (#sc-515,751, 1:1000), E-cadherin (#sc-8426, 1:1000), N-cadherin (#sc-8424, 1:1000), AKT (#sc-5298, 1:1000), p-AKT (#sc-514,032, 1:1000), or GAPDH (#sc-365,062, 1:1000) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). The protein signals were visualized with Enhanced Chemiluminescence System (Amersham Pharmacia Biotech, Buckinghamshire, United Kingdom) and quantified by using Image J software.
Cell cytoplasm/nucleus fraction isolation
The cytoplasmic and nuclear fractions were extracted from TPC-1 cells by NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific, Waltham, MA, USA), respectively. The levels of nuclear transcript and cytoplasmic transcript were detected by RT-qPCR analysis. U6 or GAPDH served as the identifiers for the nuclear and cytoplasmic fractions, respectively.
Tumor xenograft assay
BALB/c nude mice (male, four‐week-old) were bought from Shi Laike Company and reserved in the animal experimental center of our hospital after the approval by the ethics committee of our hospital. pcDNA-LINC00641 and empty vector were transfected into TPC-1 cells. Then, transfected TPC-1 cells (1 × 106) were inoculated subcutaneously in mice. The mice were observed for 4 weeks, and then euthanatized to remove the tumors.
Statistical analysis
SPSS software package (23.0 version, Chicago, IL, USA) was adopted to analyze the results. All data were presented as the mean ± standard deviation (SD). The differences among groups were estimated by one-way analysis of variance (ANOVA) and the differences between two groups were estimated by Student’st test. It was considered statistical significance when p value is less than 0.05. Each assay was performed in triplicates.
Results
LINC00641 was downregulated in thyroid carcinoma tissues and cells
Firstly, we obtained LINC00641 expression in thyroid carcinoma patients (n = 512) and healthy human individuals (n = 337) from GEPIA database (http://gepia2.cancer-pku.cn/#index). The result showed a dramatically lower expression of LIN00641 in thyroid carcinoma patients than in healthy human individuals (p < 0.05, Figure 1(a)). To further substantiate the decreased expression of LINC00641 in PTC, LINC00641 expression in PTC tissues and adjacent normal tissues were detected by RT-qPCR. We observed that the expression of LINC00641 was notably downregulated in PTC tissues compared with adjacent normal tissues (p < 0.05, Figure 1(b)). Next, we explored the expression of LINC00641 in vitro. There was a remarkable reduction in LINC00641 expression in PTC cell lines (TPC-1, BCPAP and FTC133) compared with normal thyroid epithelial cell (Nthy-ori 3-1) (p < 0.05, Figure 1(c)). The expression of LINC00641 in PTC tissues and cells. (a) GEPIA prediction of LINC00641 level in PTC. (b). The expression of LINC00641 in PTC tissues and adjacent normal tissues. N = 24 (c) The expression of LINC00641 in PTC cell lines (TPC-1, BCPAP, and FTC133) and normal thyroid epithelial cell (Nthy-ori 3-1). *p < 0.05.
LINC00641 overexpression inhibited PTC cell proliferation and promoted apoptosis
To know the biological function of LINC00641 in PTC cells, TPC-1 and BCPAP cells were transfected with LINC00641 overexpression plasmid (pcDNA-LINC00641) or negatively control (empty vector). RT-qPCR assay results showed that the introduction of pcDNA-LINC00641 induced a significant elevated of LINC00641 levels in TPC-1 and BCPAP cells compared with empty vector (p < 0.05, Figure 2(a)). We found that LINC0041 overexpression retarded TPC-1 and BCPAP cell proliferation (p < 0.05, Figure 2(b)). The results of Transwell assay suggested that the number of invaded cells was markedly reduced in TPC-1 and BCPAP cells transfected with pcDNA-LINC00641 (p < 0.05, Figure 2(c)). Flow cytometry data displayed that LINC00641 overexpression caused an increased the percentage of apoptotic cells (p < 0.05, Figure 2(d)). Furthermore, the results of western blot assays showed that the expression of E-cadherin was increased, while the expression of N-cadherin was decreased in TPC-1 and BCPAP cells transfected with pcDNA-LINC00641 (p < 0.05, Figure 2(e)). Effects of LINC00641 overexpression on PTC cell proliferation and apoptosis. TPC-1 and BCPAP cells were transfected with LINC00641 overexpression plasmid (pcDNA-LINC00641) or empty vector and incubated for 72 h. (a) RT-qPCR was applied to detect the expression of LINC00641. (b–d) CCK-8, Transwell and flow cytometry assays were used to evaluate the proliferation, invasion, and apoptosis of TPC-1 and BCPAP cells. (e) Western blot assay was used to assess the expression of E-cadherin and N-cadherin. *p < 0.05.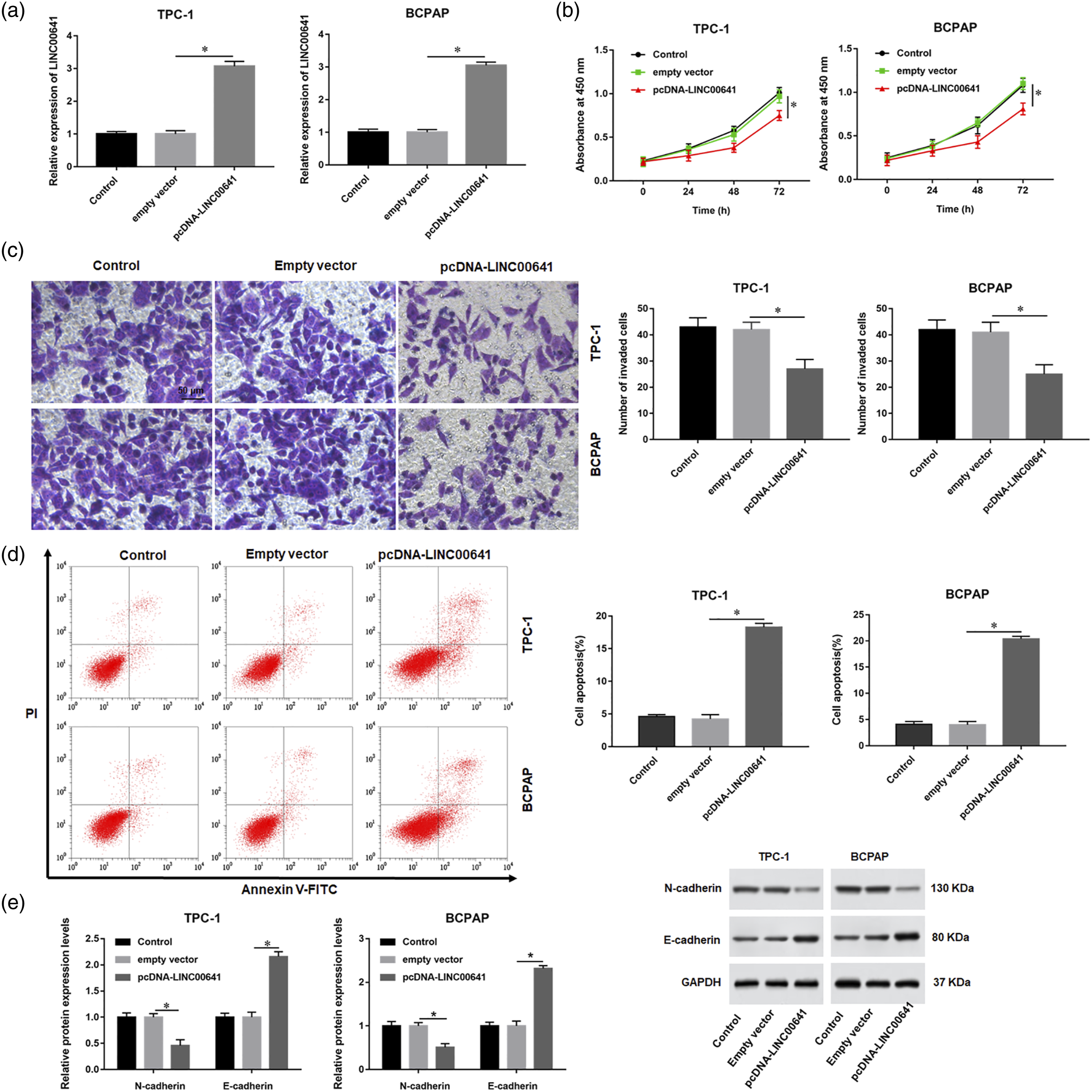
LINC00641 negatively regulated the expression of GLI1
It is well known that lncRNAs regulate the expression of downstream genes to affect the biological processes of cells. A previous study reported that GLI1 expression level is correlated with a highly malignant phenotype of PTC.
18
To explore the biological effect of GLI1 on PTC, we assessed the expression of GLI1 and found that GLI1 expression was notably higher in the PTC tissues than that in adjacent normal tissues (p < 0.05, Figure 3(a)). Western blot assay displayed that the protein levels of GLI1 was prominently increased in PTC cell lines (TPC-1, BCPAP and FTC133) compared with normal thyroid epithelial cell (Nthy-ori 3-1) (p < 0.05, Figure 3(b)). Importantly, correlation analysis showed that GLI1 expression was negatively correlated with LINC00641 expression in PTC tissues (r2 = 0.7649, p < 0.0001) (Figure 3(c)). Besides, GLI1 expression was increased in LINC00641-depleted TPC-1 and BCPAP cells (p < 0.05, Figure 3(d)). Thereafter, transfection with a short hairpin RNA (shRNA) against GLI1 (sh-GLI1) was performed. The transfection of sh-GLI1 caused a noteworthy reduction of GLI1 mRNA levels in TPC-1 and BCPAP cells (p < 0.05, Figure 3(e)). CCK-8 and Transwell assays reflected that the proliferative and invasive abilities of TPC-1 and BCPAP cells were significantly reduced by GLI1 silencing (p < 0.05, Figure 3(f) and (g)). Similarly, we also found that cell apoptosis ratio was observably enhanced in GLI1-depleted TPC-1 and BCPAP cells (p < 0.05, Figure 3(h)). GLI1 knockdown prominently increased E-cadherin level, but conspicuously decreased N-cadherin level (p < 0.05, Figure 3(i)). In the end, downregulation of GLI1 inhibited AKT phosphorylation in TPC-1 and BCPAP cells (p < 0.05, Figure 3(j)). Effects of GLI1 knockdown on PTC cell proliferation and apoptosis. (a). The expression of GLI1 in PTC tissues and adjacent normal tissues. N = 24 (b) The protein expression of GLI1 in PTC cell lines (TPC-1, BCPAP, and FTC133) and normal thyroid epithelial cell (Nthy-ori 3-1). (c) The correlation between LINC00641 and GLI1 was measured by Pearson correlation coefficient analysis. (d) TPC-1 and BCPAP cells were transfected with LINC00641 shRNA (sh-LINC00641) or control (sh-NC). Western blot assay was applied to detect the protein expression of GLI1. TPC-1 and BCPAP cells were transfected with GLI1 shRNA (sh-GLI1) or control (sh-NC). (e) RT-qPCR was applied to detect the mRNA expression of GLI1. (F–H) CCK-8, Transwell and flow cytometry assays were used to evaluate the proliferation, invasion, and apoptosis of TPC-1 and BCPAP cells. (i–j) Western blot assay was used to assess the expression of E-cadherin, N-cadherin, and p-AKT. *p < 0.05.
IGF2BP1 served as an RNA-binding protein for LINC00641
To explore the mechanism of LINC00641/GLI1 axis, we evaluated the distribution of LINC00641 in TPC-1 cells. A subcellular fractionation assay revealed that LINC00641 was mainly located in the cytoplasm (Figure 4(a)), suggesting that LINC00641 could serve as a scaffold to participate in posttranscriptional regulation of GLI1 mRNA by directly interacting with a specific RNA-binding proteins (RBPs), as has been revealed for other lncRNAs.
19
Subsequently, we used Starbase (http://starbase.sysu.edu.cn/) online to predict the potential proteins worked between LINC00641 and GLI1, and found that insulin‐like growth factor 2 mRNA binding protein (IGF2BP1) had the binding sites on LINC00641 (Figure 4(b)). Therefore, we further detected the association between LINC00641 and IGF2BP1 by RIP assay. The results showed that IGF2BP1 was merely pulled down by biotinylated LINC00641 (Figure 4(c)). Furthermore, RNA pull-down assay reflected that the relative enrichment level of LINC00641 was remarkably increased in the anti-IGF2BP1 group compared with the lgG group (p < 0.05, Figure 4(d)). The results of RT-qPCR showed that the expression level of LINC00641 was increased by LINC00641 overexpressed and decreased by LINC00641 knockdown (p < 0.05, Figure 4(e)). However, intervening in the expression of LINC00641 in TPC-1 cells was not found the change of IGF2BP1 protein levels (p > 0.05, Figure 4(f)). LINC00641 physically interacted with IGF2BP1 in PTC cells. (a) The location of LINC00641 was explored through subcellular fractionation. (b) The binding sites of LINC00641 and IGF2BP1 was predicted by starBase. (c–d) RNA pull-down and RIP assays showed association of IGF2BP1 with LINC00641. (e) RT-qPCR was applied to detect the expression of LINC00641. (F) Western blot was applied to detect the expression of IGF2BP1. *p < 0.05.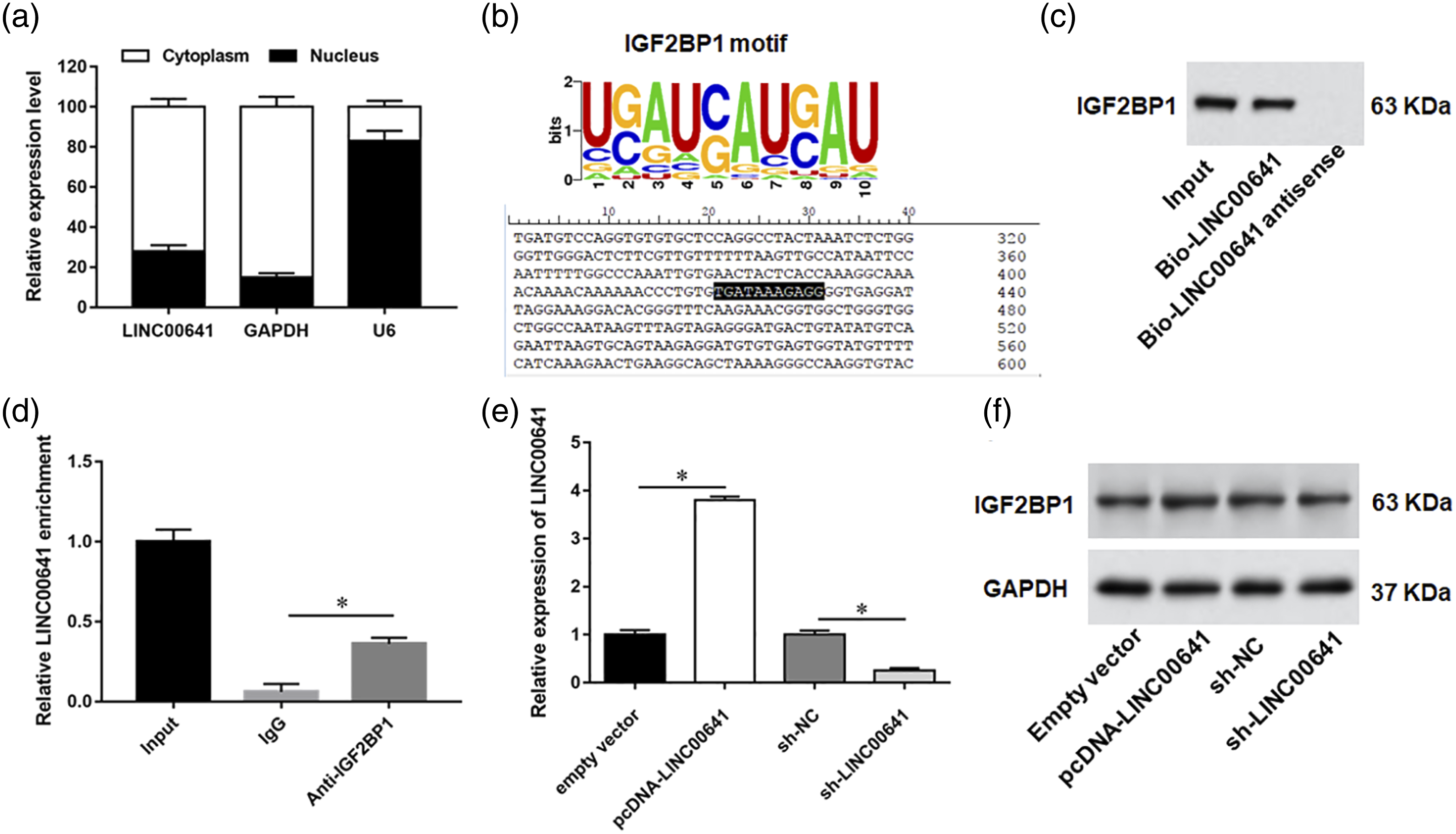
LINC00641 destabilized GLI1 mRNA by competitively binding to IGF2BP1
Interestingly, IGF2BP1 is known as a RBP, which has been found to associate with GLI1 mRNA and stabilize GLI1 mRNA in cancers cells.
17
.RIP assay confirmed the binding of IGF2BP1 and GLI1 (Figure 5(a)). Western blot assay showed that GLI1 expression was increased in IGF2BP1-upregualted TPC-1 cells but reduced in IGF2BP1-depleted TPC-1 cells (p < 0.05, Figure 5(b)). To further confirm the stability of GLI1 mRNA is associated with LINC00641 mediated IGF2BP1 recruitment, an RIP assay by using IGF2BP1 antibody was conducted, and the results showed that the LINC00641 overexpression reduced the interaction of IGF2BP1 with GLI1 mRNA (p < 0.05, Figure 5(c)), while LINC00641 silencing strengthened the association between IGF2BP1 and GLI1 mRNA (p < 0.05, Figure 5(d)). Additionally, upregulation of IGF2BP1 could reverse LINC00641 overexpression-induced the downregulation of GLI1 (p < 0.05, Figure 5(e)). Afterward, RT-qPCR assay results displayed that GLI1 mRNA degradation induced by actinomycin D (a transcriptional inhibitor) treatment was inhibited by LINC00641 knockdown and accelerated by IGF2BP1 silencing (p < 0.05, Figure 5(f)). Enforced expression of IGF2BP1 restored the LINC00641 overexpression-mediated GLI1 mRNA degradation after treatment with actinomycin D (p < 0.05, Figure 5(g)). LINC00641 destabilized GLI1 mRNA by interacting with IGF2BP1. (a) RIP assay showed association of IGF2BP1 with GLI1. (b) Western blotting was applied to detect the protein expression of GLI1. (c–d) TPC-1 cells were transfected with LINC00641 overexpression plasmid (pcDNA-LINC00641) or shRNA (sh-LINC00641), and RIP assay was applied to analyze the enrichment of GLI1 mRNA in RNA-protein complexes pulled down by anti-IGF2BP1 antibody. (e) Western blotting was applied to detect the protein expression of GLI1. (f–g) The degradation rate of GLI1 mRNA was evaluated by RT‐qPCR. *p < 0.05.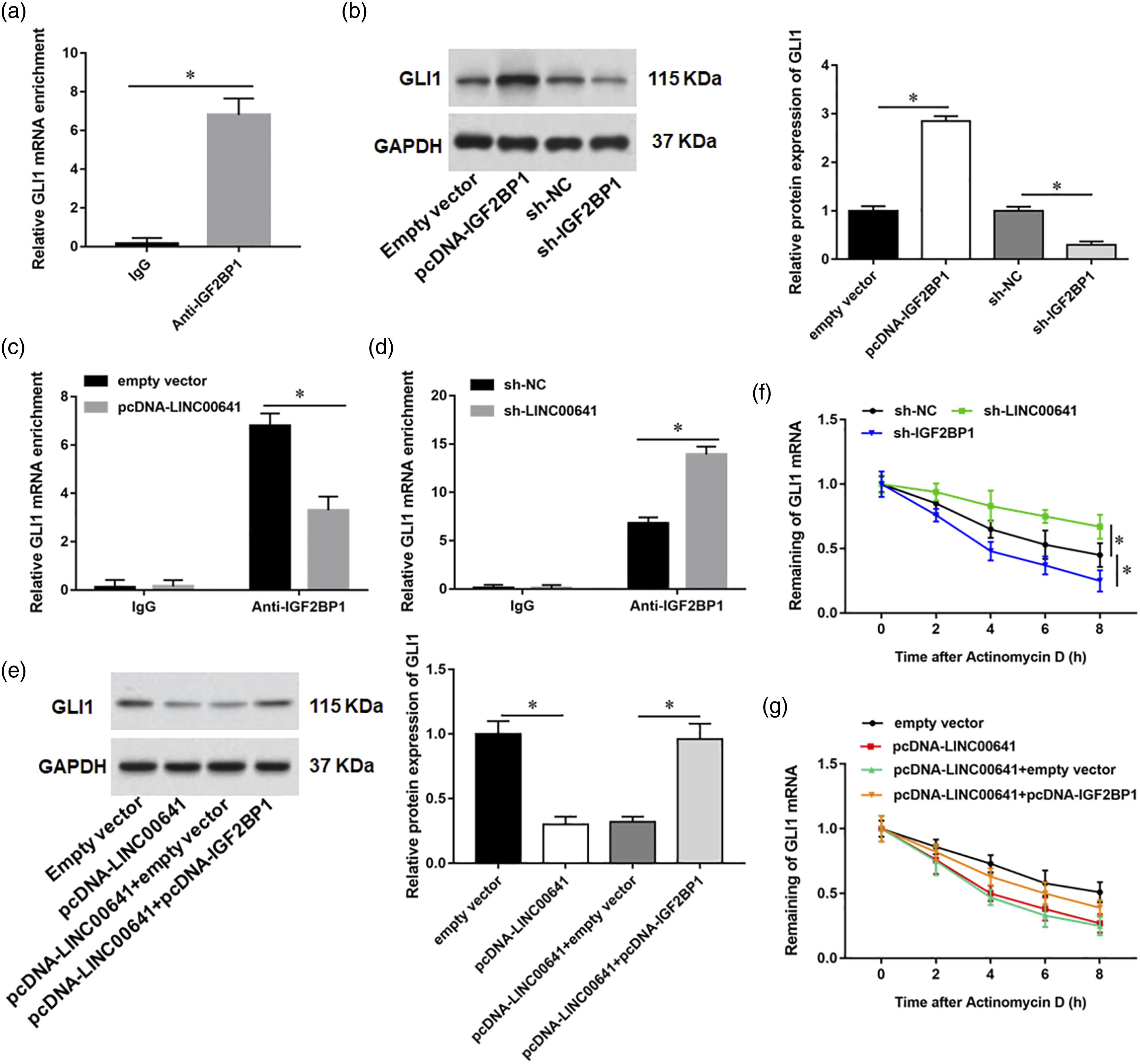
GLI1 is involved in LINC00641-mediated PTC cell proliferation and apoptosis
We next investigated whether GLI1 could rescue the inhibitory effect of LINC00641 overexpression on the malignant progression of PTC cells. TPC-1 and BCPAP cells were transfected with pcDNA-LINC00641 alone or co-transfected with pcDNA-GLI1. Elevated of GLI1 significantly enhanced the capacity of cell proliferation and invasion of LINC00641-upregualted TPC-1 and BCPAP cells (p < 0.05, Figure 6(a) and (b)). GLI1 overexpression also abolished the inductive effect of apoptosis by LINC00641 overexpression (p < 0.05, Figure 6(c)). In addition, the upregulation of E-cadherin and the downregulation of N-cadherin by LINC00641 overexpression were abrogated by GLI1 overexpression (p < 0.05, Figure 6(d)). Likewise, GLI1 overexpression reversed the effect of LINC00641 overexpression on p-AKT protein levels (p < 0.05, Figure 6(e)). Effects of GLI1 overexpression on LINC00641 overexpressed PTC cells. TPC-1 and BCPAP cells were transfected with pcDNA-LINC00641 alone or together with pcDNA-GLI1. (a–c) CCK-8, Transwell and flow cytometry assays were used to evaluate the proliferation, invasion, and apoptosis of TPC-1 and BCPAP cells. (d–e) Western blot assay was used to assess the expression of E-cadherin, N-cadherin, and p-AKT. *p < 0.05.
LINC00641 overexpression impeded tumor growth in xenograft model
To evaluate the effects of LINC00641 on PTC in vivo, nude mice were inoculated with TPC-1 cells transfected with the empty vector or pcDNA-LINC00641 to establish a model of subcutaneous xenograft tumor. Enforced expression of LINC00641 remarkably restrained the size and weight of tumors at the end of experiments (p < 0.05, Figure 7(a) and (b)). The results of RT-qPCR and Western blot assay showed that the expression of LINC00641 was upregulated, and the protein expression of GLI1 and p-AKT was decreased in the pcDNA-LINC00641 group (p < 0.05, Figure 7(c) and (d)). Effects of LINC00641 overexpression on tumor growth in a nude murine model. TPC-1 cells transfected with pcDNA-LINC00641 or empty vector were injected subcutaneously into nude mice. (a–b) The tumor volume and weight were measured in mice. (c) The expression of LINC00641 was detected with RT‐qPCR. (d) The protein levels of GLI1 and p-AKT were evaluated by western blot assay. *p < 0.05.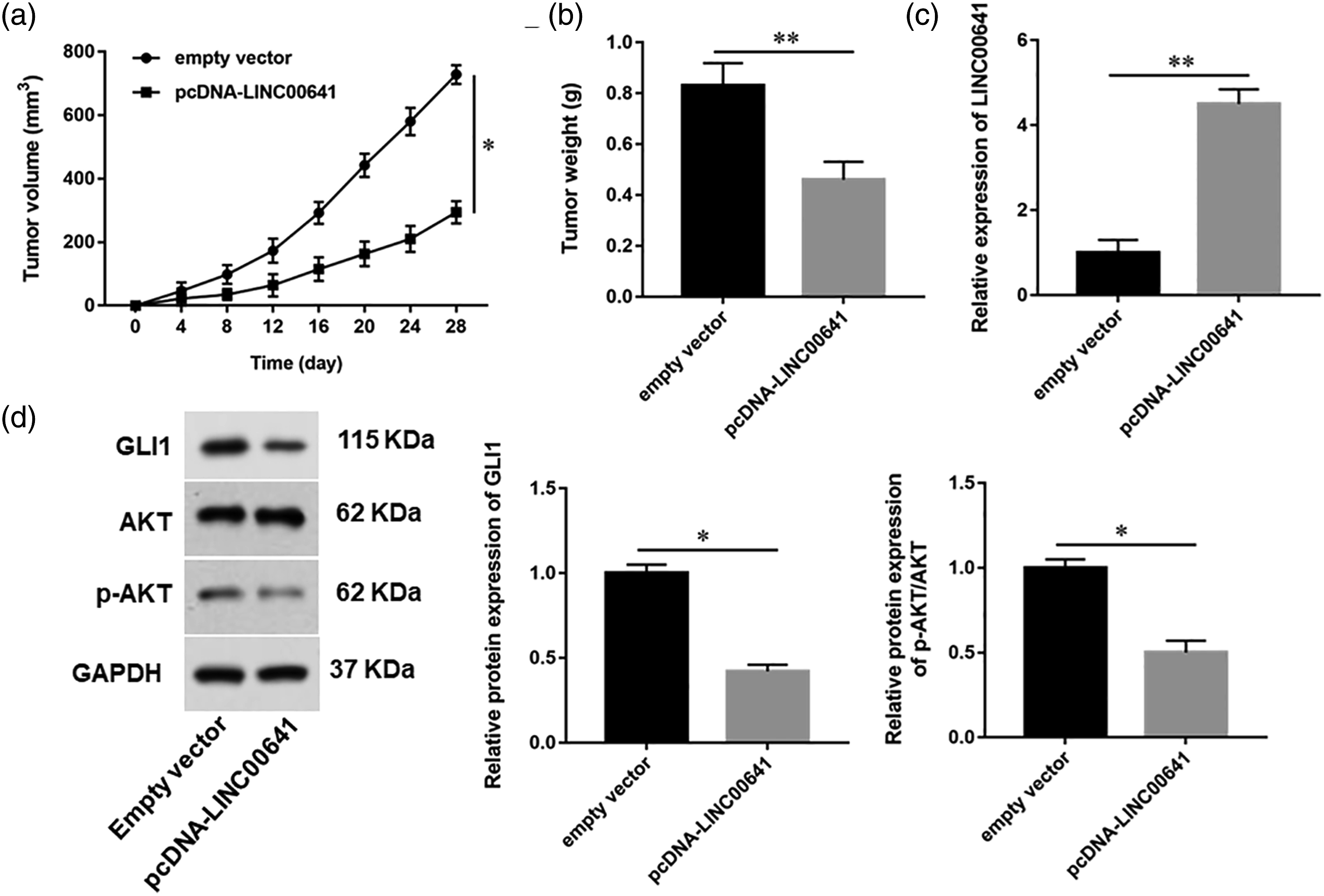
Discussion
LINC00641 is a long non coding RNA localized on chromosome 14q11.2, and is aberrantly expressed in malignant tumors.9,20 A number of studies have demonstrated that LINC00641 silencing or overexpression was able to inhibit tumor growth, which highlights that LINC00641 may be a promising therapeutic target to induce cell death in malignant tumors. Mao et al. suggested that LINC00641 was negatively correlated with tumor volume, lymph-node metastasis and clinical stage in breast cancer, which could suppress breast cancer cell proliferation, migration and invasion by downregulating miR-194-5p. 21 In bladder cancer, absence of LINC00641 in patients predicted a poor prognosis, and LINC00641 overexpression significantly restrained the malignant progression of bladder cancer cells in vivo and in vitro by modulating the miR-197-3p/KLF10/PTEN/PI3K/AKT pathway. 22 Liu et al. uncovered that LINC00641 acted as a competitive endogenous RNA to inhibit growth and invasion of prostate cancer cells by sponging miR-365a-3p to elevate VGLL4 expression. 23 Consistent with previous reports, in this study, our results revealed that LINC00641 was downregulated in PTC tissues and cells, and upregulation of LINC00641 inhibited proliferation, invasion, and induced apoptosis of PTC cells.
Apart from sponging and sequestering specific microRNAs, lncRNAs could also regulate the expression of targeting genes via binding to RNA‐binding proteins (RBPs). IGF2BP1 is a conserved RBP and plays an essential role in carcinogenesis.24,25 In a variety of cancers, an increasing number of lncRNAs have been identified as direct regulators of IGF2BP1 during tumor cell proliferation and invasion. For example, lncRNA THOR directly bound to IGF2BP1 to stabilize IGF2 mRNA, which in turn affected the MEK-ERK signaling pathway to facilitate tongue squamous cell carcinomas. 26 LINC01093 interacted with IGF2BP1, which weakened the interaction between IGF2BP1 and GLI1 mRNA, thus inhibiting the progression of hepatocellular carcinoma. 17 LncRNA KB-1980E6.3 contributed to breast cancer stem cell stemness via interacting with IGF2BP1 to facilitate c-Myc mRNA stability. 27 In our study, we utilized the bioinformatics software Starbase prediction and found that IGF2BP1 had the binding sites on LINC00641, and the binding of LINC00641 to IGF2BP1was validated by RIP and RNA pull-down assays.
Additionally, IGF2BP1, as a key post transcriptional regulator, has been reported to regulate the mRNA stability, leading to an increase in its expression and ultimately the presentation of the cancerous phenotypes.28,29 In our study, we found that IGF2BP1 bound to GLI1 mRNA, and overexpression of IGF2BP1 promotes GLI1 protein expression. Furthermore, LINC00641 overexpression reduced the interaction of IGF2BP1 with GLI1, and LINC00641 silencing strengthened the association between IGF2BP2 and GLI1 mRNA. As shown above, we found that LINC00641 competitively binds to IGF2BP1, preventing GLI1 mRNA binding to IGF2BP1 and promoting its degradation.
The Hedgehog (Hh) signaling pathway is crucial for the development and progression of multiple malignancies, including PTC. 15 GLI1, as a key transcriptional activator of Hh signaling pathway, has been revealed to be an oncogene in a variety of cancers. Upregulation of GLI1 was correlated strongly with tumor aggressiveness, which could lead to epithelial-to-mesenchymal transition in gastric cancer. 30 A Study showed that GLI1 promoted colorectal cancer cells metastasis in a Foxm1-dependent manner by activating EMT and the PI3K-AKT signaling. 31 Multilayered microRNAs and lncRNAs have been reported to affect the expression of GLI1 by regulating its transcription, translation, and activity. MiR-361 reduced the expression of GLI1 via binding to its 3′UTR, thus inhibiting esophageal carcinoma progression. 32 LINC01106 could activate the transcription of GLI1 through recruiting FUS to GLI1 promoter. 33 In this study, we uncovered that GLI1 silencing inhibited PTC cell proliferation, invasion and promoted apoptosis, as well as increased E-cadherin and decreased N-cadherin expression by activating AKT pathway. Besides, rescue assay showed that GLI1 overexpression reversed the inhibitory effect of LINC00641 overexpression on the malignant progression of PTC cells.
Conclusion
In conclusion, our study identified that LINC00641 was downregulated in PTC tissues and cells. Overexpression of LINC00641 inhibited the IGF2BP1-mediated mRNA stabilizing of GLI1 and ultimately disrupted the malignant progression of PTC. The present study provided a new insight into the regulatory mechanism of LINC00641 in PTC progression. However, it is an extremely complex regulatory system in vivo, and this study only explored the effect of overexpressing LINC00641 on mice with transplanted tumors, therefore, the specific regulatory mechanism of LINC00641 on TPC progression in vivo needs further investigation.
Footnotes
Appendix
Acknowledgements
The authors would like to express his gratitude to the First Affiliated Hospital of Zhengzhou University who hosted the study.
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article
Ethical Approval
The present study and the associated experimental protocols were performed in compliance with ethical guidelines and approved by the Institute Research Medical Ethics Committee of the First Affiliated Hospital of Zhengzhou University (approval number: 2020-KY-152). All PTC tissue samples and pair-matched adjacent non-neoplastic tissue samples were used in accordance with the Helsinki declaration.
Informed Consent
Written informed consent was obtained from the guardians of all subjects.
Data Availability
All data generated or analyzed during this study are included in this published article. Data will be made available from the corresponding author on reasonable request.
