Abstract
Background
Hyperglycemia is closely related to adverse pregnancy outcomes including pre-eclampsia (PE), a life-threatening complication with a substantial morbidity and mortality. However, the pathogenesis of abnormal placentation in gestational diabetes mellitus (GDM)-associated PE remains elusive.
Method
Here we isolated exosomes from the human umbilical vein endothelial cells (HUVECs) treated with normal level of glucose (NG) and high levels of glucose (HG). The exosomes were added to HTR-8a/SVneo cells, a trophoblast cell line. High-throughput RNA-sequencing was performed to analyzed the changed RNAs in the exosomes and exosome-treated HTR-8a/SVneo cells. HTR-8a/SVneo cell phenotypes were evaluated from the aspects of cell proliferation, cell invasion and DNA damage.
Results
After treatment with HG, the changed RNAs in exosomes was enriched in RNA stabilization and oxidative stress. The altered RNAs in the HTR-8a/SVneo cells treated with exosomes from HG-induced HUVECs were enriched in pathways related to cell adhesion, migration, DNA damage response and angiogenesis. The HG-induced exosomes impaired the proliferation and invasion of HTR-8a cells and caused the DNA damage. HG up-regulated PUM2 in the exosomes and exosome-treated HTR-8a/SVneo cells. PUM2 interacted with SOX2 mRNA, resulting in the mRNA degradation. Overexpression of SOX2 prevented the damage to HTR-8a/SVneo cells caused by the exosomes from HG-induced HUVECs.
Conclusions
We demonstrate that high glucose-induced endothelial exosomes mediate abnormal phenotypes of trophoblasts through PUM2-mediated repression of SOX2. Our results reveal a novel regulatory mechanism of hyperglycemia in development of abnormal placentation and provide potential targets for preventing adverse pregnancy outcomes.
Introduction
Gestational diabetes mellitus (GDM) or hyperglycemia is a known risk factor of adverse pregnancy outcomes (APO).1–3 As one of the most concerning APOs caused by hyperglycemia, pre-eclampsia (PE) is presented by new-onset hypertension, proteinuria and multi-organ dysfunction as progression of the disease.2,4 PE is associated with a global incidence of 4–5% and substantial mortality, leading to long-term fetal and maternal damage on circulatory and neural systems. 5
Endothelial cells, which are considered as an endocrine organ, extensively distribute in vessels in a mono-layered manner, forming barriers of substance exchange between tissue and vessels by tight junction and adherence junction. 6 Endothelial cell responses to various metabolic states to maintain homeostasis, and malfunction of endothelial cell proves to be a key contributor in metabolic disorders. 7 Owing to altered metabolic states during pregnancy, the roles that endothelial cells play in abnormal pregnancy have attracted much attention. The function of endothelial cells are especially addressed in pregnancy disorders complicated by hyperglycemia, due to the fact that diabetes has long been accepted as a vascular disease. 8 According to existing literature, endothelial cells in GDM have shown elevated pro-inflammatory state and reduced capacity of transport. 9
Extracellular vesicles (EV) including exosomes play vital roles in intercellular communication, which have also been extensively identified in feto-placental tissues. 10 EV-derived cargos, including microRNA and proteins, have been shown to involve in regulation of inflammatory responses and invasion of trophoblasts to impact on placentation. 11 However, despite of the critical roles of endothelial cells in pregnancy disorders, the function of endothelial cell-derived EVs in placentation has not yet been explored.
Pathogenesis of PE is generally considered to involve abnormal placentation, systematic vascular dysfunction and related maternal syndromes. 9 Particularly, the effect of hyperglycemia on trophoblasts and abnormal placentation is of broad interests. It has been demonstrated that trophoblasts as well as endothelial cells exhibited dysregulated responses when exposed to high-leveled glucose.12–14 However, high-leveled glucose does not directly contact with trophoblasts in actual environment of the organism, raising the question of how hyperglycemia affects trophoblasts exactly. Considering that high-leveled glucose directly interacts with endothelial cells rather than trophoblasts, we hypothesized that the endothelial cell-secreted exosomes are key mediators in the process of PE development. Taking advantage of isolated exosomes from cultured cells, here we explored the effect and regulatory mechanisms of hyperglycemia on trophoblasts, finding that pumilio RNA binding family member 2 (PUM2), a member of the PUF family of sequence-specific RNA-binding proteins functioning as a translational repressor,15–17 played a central role in modulation of cell proliferation and DNA damage through regulating RNA stability of SOX. Our data offers insights into how hyperglycemia affects trophoblasts and contributes to PE and APO, and provides potential therapeutic targets to prevent development of APOs.
Materials and methods
Cell culture and treatments
Human umbilical vein endothelial cells (HUVECs) and HTR-8a/SVneo cells were obtained from ATCC (Manassas, US) and cultured in RPMI 1640 medium/Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS) at 37°C in a humidified incubator with 95% air and 5% CO2. HUVECs were treated with normal concentration of glucose (5 mmol/l), high concentrations of glucose (20 mmol/l and 30 mmol/l), mannitol (20 mmol/l) and GW4869 (20μM) for 48 h. Mannitol was used as a control to exclude the influence of high osmotic pressure. GW4869 was used for the suppression of the secretion of exosomes. Exosomes from the culture medium of HUVECs were added to HTR-8a/SVneo cells. After 48 h, HTR-8a/SVneo cell phenotypes were assessed.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted from cells using the Trizol reagent followed by administration of chloroform. The water phase was collected. RNA was precipitated by isopropanol and washed by 75% ethanol. Quality and concentration of obtained RNA was determined using a NanoDrop Spectrophotometer (NanoDrop Technologies Inc., USA) and 2 μg RNA was reverse transcribed for template of real-time PCR amplification using TransStart Top Green qPCR SuperMix (TransGen Biotech) and gene-specific primers on an ABI 7500 system (Applied Biosystems). Following primers were used in this study (from 5′ to 3′):SOX2-F TGGACAGTTACGCGCACAT, SOX2-R CGAGTAGGACATGCTGTAGGT, CLSTN3-F CAACACGGTCCTACTGAATCC, CLSTN3-RCTCTCCTGTCGCCTTGTCA, TCF4-FTGCA AAGCCGAATTGAAGATCG,TCF4-RAGAAGGTCCAA TGATTCCATGC, EP400-FTGGTAGCATCGACAAGGC TC, EP400-RGCGAGGGACGAGGTAGAAG,PUM2-FG TCCCAGCCTATTATGGTACAGA,PUM2-RATCGCGGA GACAGTATTGCAT, PUM1-F ATGAGCGTTGCATGTG TCTTG,PUM1-RGTAGTCCACCATAGCGTCGTC, Primers U1-FGGGAGATACCATGATCACGAAGGT, PrimersU1-RCCACAAATTATGCAGTCGAGTTTCCC, β-actin-FCAT GTACGTTGCTATCCAGGC, β-actin-RCTCCTTAATGTCACGCACGAT, 18S-FCAGCCACCCGAGATTGAGCA, 18S-RTAGTAGCGACGGGCGGTGTG.
Western blot
Cells were lysed in 1% NP-40 with 0.1% SDS and 50 mM DTT, with 2 μg/ml Aprotinin, 2 μg/ml Leupeptin and 1 mM PMSF added immediately before use. Total protein was obtained by ultrasonic crushing and subsequent centrifugation. Proper amount of proteins were loaded into wells of SDS-PAGE gels and transferred onto nitrocellulose membrane (PIERCE, #88018). The membrane was then blocked with 5% nonfat milk for 2 h at 37°C, followed by incubation with primary antibodies at 4°C overnight and with secondary antibodies at room temperature for 2 h. Chemiluminescent substrates (Thermo Scientific, USA) were then used for detection of protein signals. Following primary antibodies were used in this study: CD63 (Ptgcn, 25,682-1-AP), CD9 (Ptgcn, 20,597-1-AP), TCF4 (Ptgcn, 22,337-1-AP), SOX2 (CST, #2748), CLSTN3 (Ptgcn, 13,302-1-AP), EP400 (Abcam, ab70301), PUM2 (Abcam, ab190104), β-actin (Ptgcn, 66,009-1-Ig).
High throughput RNA-sequencing and gene ontology (GO) enrichment analysis
A mRNA library was constructed from extracted total RNA for sequencing using TruSeq®RNA Sample Preparation Kit v2 (Illumina, SanDiego, CA, USA) according to the manufacturer’s guidelines. mRNA was purified and fragmented followed by synthesize of first and second strand cDNA, and subsequently subjected to end repair, 3′ end adenylation, ligation of adapters and PCR amplification. Sequencing was then performed using a Genome Analyzer II (Illumina, San Diego, CA, USA) according the manufacturer’s recommendation. Reads per kilobase of exon model per million mapped reads (RPKM) of all transcripts were calculated to analyze gene expression. Differential expressed genes were then subjected to GO enrichment analysis (https://geneontology.org/).
RNA immunoprecipitation assay (RIP)
RIP assay was performed according to the manufacturer’s protocol of EZ-Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Catalog No. 17–701, Millipore, USA). Briefly, cell lysates from different groups were incubated with anti-PUM2 antibody (ab190104, abcam, USA) or immunoglobulin (IgG) coupled to magnetic beads at 4°C overnight. The magnetic beads were then washed for 5 times and incubated with proteinase K to digest associated proteins at 55°C for 30 min. RNA of the precipitated complex was then extracted and purified for qRT-PCR analysis.
Exosome isolation and quality evaluation
Exosomes were isolated through successively centrifuging steps. Briefly, HUVECs were centrifuged at 2000 × g, 4°C for 30 min. The supernatant was transferred to a new tube and subjected to centrifugation at 10,000 ×g 4°C for 45°min to remove large vesicles. The supernatant filtered through a 0.45 μm filter was centrifuged again at 10,000×g, 4°C for 70°min. The pellets of exosomes were suspended in 100 μL PBS. 20 μL of the PBS solution subjected to assessment of exosome morphology and distribution by HT-7700 transmission electron microscopy (Hitachi, Japan). In addition, 10 μL of the PBS solution was used for particle size analysis of exocrine in N30Enanoparticle analyzer (NanoFCM, UK).
Uptake analysis of labeled exosomes
Uptake analysis of labeled exosomes was modified from previous methods.18–20 Briefly, exosomes (200 μL) were resuspended with 200 μL PBS and mixed with 5 μL DiD fluorescent probe dye solution (AAT Bioquest, USA) in dark at 37°C for 20 min. HTR-8a cells were cultured in 6-well plates and incubated with the DiD-labeled exosomes for 2 h. Images of the red fluorescence (at 570 nm excitation wavelength) were acquired to evaluate the uptake of exosomes by HTR-8a cells.
CCK8 assay for cell proliferation
HTR-8a/SVneo cells (1 × 105) were seeded in each well of a 96-well plate. A well without cells is defined as the blank. CCK-8 solution (10 μL volume; No. CK04; Dojind, Japan) was added into each well and incubated for 1 h in the cell incubator before OD values were determined by a microplate reader at 450 nm. Cell a rate is calculated as follows: Proliferate rate = (OD value of test well – OD value of blank well)/(OD value of control well – OD value of blank well).
Immunofluorescence for DNA damage
HTR-8a cells were planted on slides in 6-well plates and were fixed with 4% paraformaldehyde (Sigma) for 20 min after attachment of the cells. The cells were then washed in PBS and blocked by normal goat serum at room temperature for 20 min. The cells were then incubated with primary antibodies against γ-H2AX (pS139) (ab215967) diluted 1:500 with 5% BSA, at 4°C overnight, washed in PBS for 3 times, and incubated with Alexa Fluor® 488 Goat Anti-Rabbit IgG (H + L) diluted at 1:100 (A11008) at 37°C for 40 min. Nuclei were lastly stained with DAPI (SC-3598), and images were acquired using a TCS-SP5 confocal microscope (Leica, Germany) at the emission wavelengths of 480 nm and 340 nm.
Trans-well assay
Cell invasion was analyzed using 6-well trans-well system (Corning) with an 8µm-pore polycarbonate filter membrane. The upper chamber was covered with 50 mg/L Matrigel diluted at 1:8 and incubated at 4°C overnight. 60–80 μl diluted Matrigel (3.9 µg/μl) was then added into the upper chamber and incubated at 37°C for 30 min. The upper chamber was then seeded with HTR-8a cells and inserted into the lower chamber containing complete medium. After incubation for the required period of time, the cells on the interior of upper chamber were removed, and the cells in the lower chamber were fixed for 15 min and stained with hematoxylin for 10 min. The number of cells was counted and the images were aquiredunder 200× microscopic fields.
RNA stability assay
HTR-8a/SVneo cells treated with 5 μg/mL Actinomycin D for 0, 2, 4, 6, 8 or 12 h were collected and lysed for RNA extraction. Messenger RNA (mRNA) levels of SOX2 were analyzed by quantitative polymerase chain reaction (qPCR), using 18S-RNA or β-Actin as the internal control. Half-life of mRNA was calculated as previously described. 21
Statistical analysis
With the help of the Prism software (GraphPad, USA), quantification results were statistically analyzed by two-tailed Student’s t tests for comparison between two groups and by one-way ANOVA for analysis of multiple groups. Results are shown as means (average value from three repeated experiments) ±SEM. Values of p < 0.05, 0.01 and 0.001 were considered significant and marked as *, ** and ***, respectively.
Results
RNA-sequencing of exosomes secreted by high glucose-induced HUVEC cells revealed altered pathways of RNA modulation and PUM1/2 levels
To explore the impact of high glucose on endothelial cells, HUVEC cells were treated with high glucose (HG) at 30 mmol/L and normal glucose (NG) at 5 mmol/L. Exosomes were isolated from the supernatant of cultured cells and morphology and particle sizes of the exosomes were evaluated, showing that the exosomes were of sufficient quality for further analysis (Figure 1(A)). Both of NG-treated cells and HG-treated cells showed significant higher protein levels of CD63 and CD9 in exosomes than those in total cell lysates, which also indicated successful isolation of exosomes (Figure 1(B)). Gene expression profiles of exosomes isolated from NG or HG treated HUVECs. (A) Morphology (upper panel) and particle sizes (lower panel) of isolated exosomes from NG or HG treated HUVECs. Scale bar represents 200 nm (B) Representative illustration of western blotting and statistical analysis of protein levels of CD63 and CD9 in total cell lysates or isolated exosomes of NG or HG treated HUVECs. β-Actin is regarded as the internal control. (C) Volcano plotting of differential expressed genes (DEGs) screened by fold change and p-value according to RNA-sequencing results comparing isolated exosomes of NG and HG treated HUVECs. (D) GO analysis of significantly up-regulated genes in isolated exosomes of HG treated HUVECs compared to those of NG treated HUVECs. (E) Heat map of significantly up-regulated genes in isolated exosomes of HG treated HUVECs compared to those of NG treated HUVECs. (F) Illustration of mRNA levels of PUM1/2 detected by quantitative PCR from isolated exosomes of NG or HG treated HUVECs. *p < 0.05, ***p < 0.001 vs. Negative control (n = 3, Student’s t tests).
GO analysis of up-regulated DEGs of HG-treated HUVECs.

SOX2 was identified as a potential target of PUM2 in HTR-8a cells treated by HG-HUVEC-derived exosomes
To investigate whether trophoblasts are affected by exogenous exosomes, exosomes isolated from supernatant of HG or NG treated HUVEC cells (designated as HG-HUVEC-Exo or NG-HUVEC-Exo in following contents) were co-cultured with HTR-8a/SVneo cells, a cell-line developed from first-trimester extra-villous trophoblasts.
23
Differential expressed genes were screened from transcriptomic data of RNA-sequencing of the exosome-treated HTR-8a/SVneo cells (Figures 2(A)–(C)). Down-regulated genes were subjected to GO clustering analysis (Figure 2(C) and Table 2), and we found that biologic processes including adhesion, migration, angiogenesis, cell cycle and regulation of transcription exhibited prominent alteration. Gene expression profiles of HTR-8a/SVneo cells treated by NG-HUVEC-Exo or HG-HUVEC-Exo. (A) Volcano plotting of DEGs screened by fold change and p-value according to RNA-sequencing results comparingHTR-8a/SVneo cells treated by NG-HUVEC-Exo or HG-HUVEC-Exo. (B) GO analysis of significantly down-regulated genes in HTR-8a/SVneo cells treated by HG-HUVEC-Exo compared to those treated by NG-HUVEC-Exo. (C) Heat map of significantly down-regulated genes in HTR-8a/SVneo cells treated by HG-HUVEC-Exo compared to those treated by NG-HUVEC-Exo. (D) Venn diagram illustration of target genes of PUM2 according to Lin et al. (blue), Zhang et al. (yellow) and down-regulated DEGs in HTR-8a/SVneo cells treated by HG-HUVEC-Exo (green). Four common elements are listed. (E) Validation of mRNA levels of TCF4, SOX2, CLSTN3 and EP400 in HTR-8a/SVneo cells treated by HG-HUVEC-Exo compared to those treated by NG-HUVEC-Exo. (F) Representative illustration of western blotting and statistical analysis of protein levels of TCF4, SOX2, CLSTN3 and EP400 in HTR-8a/SVneo cells treated by HG-HUVEC-Exo as compared to those treated by NG-HUVEC-Exo. β-Actin is regarded as the internal control. *p < 0.05, **p < 0.01, ***p < 0.001 vs. Negative control (n = 3, Student’s t tests). GO analysis of down-regulated DEGs of HG-HUVEC-Exo treated HTR-8a cells.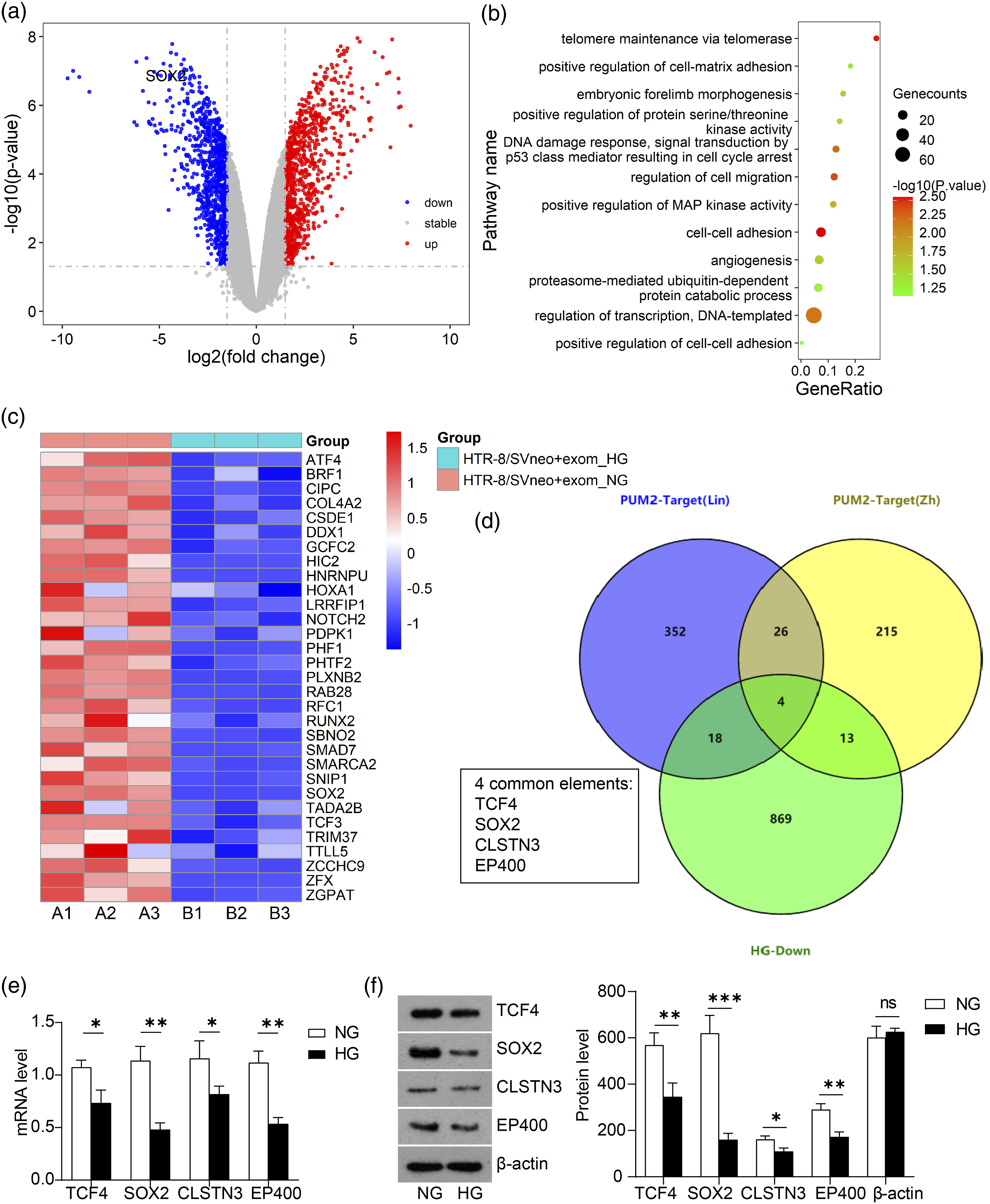

Considering the function of gene and protein repression mediated by PUM (PMR), PUM from HG-HUVEC exosomes might repress gene expression in trophoblasts. Therefore, we screened for potential targets of PUM in HG-HUVEC-Exo treated HTR-8a cells, mainly focusing on down-regulated genes in transcriptomic data. Two previous studies of Zhang et al. 17 and Lin et al. 16 reported hundreds of PUM2-targeting genes with 30 shared genes. Aligning with our list of down-regulated genes from RNA-sequencing analysis, we obtained four common elements, TCF4, SOX2, CLSTN3 and EP400 (Figure 2(D)). We validated the protein and transcriptional levels of the four targets by PCR and Western blot. Compared to TCF4 and CLSTN3, SOX2 and EP400 declined more prominently in mRNA levels as well as protein levels upon HG-HUVEC-Exo treatment (Figures 2(E) and (F)). As an important transcription factor required for pluripotency, 24 the role of SOX2 in PE or GDM remain unexplored. Among the biologic processes we identified through GO clustering, SOX2 participated in regulation of transcription and embryonic forelimb morphogenesis. Considering that protein levels of SOX2 showed the most obvious variation between NG and HG conditions among the 4 genes. SOX2 was selected for our following study of molecular mechanisms.
HTR-8a cells treated by HG-HUVEC-Exo demonstrated cell damage and instability of SOX2
Glucose concentrations between 20 to 30 mmol/l have been used to mimic high-glucose conditions in previous studies.25–28 Our experiments were conducted in cells treated by NG (5 mmol/l), HG (20 mmol/l), HG (30 mmol/l) and mannitol as an osmotic control for more rigorous study design and careful conclusions. We labeled HUVEC-derived exosomes with DiD fluorescent probes and uptake of the labeled exosomes by HTR-8a/SVneo cells was certified by immuno-fluorescent imaging. Compared to labeled NG-exosomes, the red fluorescence of labeled HG-exosomes showed a higher quantity (Figure 3(A)). A classic exosome-inhibitor, GW4869, was used as previously described
29
for pre-treatment of HUVEC cells at a concentration of 20 μM, before harvesting HG-exosomes from HUVEC cells. The application of GW4869 significantly reduced secretion of HG-exosomes from HUVEC cells, which thereby led to reduced uptake of exosomes in HTR-8a cells (Figure 3(A)). These data reinforce that HG induces exosome secretion and the exosomes are thereby internalized by HTR-8a cells. HTR-8a cells treated by HG-HUVEC-Exo demonstrated cell damage and decreased levels of SOX2. (A) Illustration of uptake of DiD-labeled exosomes (red) in HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. Nucleus was counterstained by DAPI (blue). (B) Immunofluorescence of γ-H2AX pS139 (green) in HTR-8a cells treated by treated by exosomes harvested from HUVECs subjected to conditions as indicated. Nucleus was counterstained by DAPI (blue). (C) Illustration of trans-welled cells for evaluating invasiveness of HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. (D) Statistical analysis of cell counts in γ-H2AX pS139 immunostaining (left panel) and trans-well assay (right panel) of HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. (E) Illustration of proliferate rate evaluated by CCK8 assay of HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. (F) Statistical analysis of mRNA levels of PUM2 and SOX2 in HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. (E) Representative illustration of western blotting of PUM2 and SOX2 in HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. *p < 0.05, **p < 0.01, ***p < 0.001 (n = 3, one-way ANOVA for analysis).
Cell proliferation, ability of invasion and DNA damage were then evaluated in the HTR-8a cells that contained exogenous exosomes. Our results showed that HG-exosomes increased DNA damage (Figure 3(B) and (D)) and reduced invasiveness (Figures 3(C) and (D)) and proliferate activity (Figure 3(E)) of HTR-8a cells, illustrated by γ-H2AX pS139 immuno-fluorescence, trans-well assay and CCK8 assay, respectively. While incorporation of GW4869, the exosome inhibitor, ameliorated the adverse effects of HG-exosomes in cell proliferation, invasiveness, and DNA damage (Figures 3(B)–3(E)). mRNA and protein levels of PUM1, PUM2 and SOX2 in HTR-8a cells were further detected. The results revealed that compared to NG-HUVEC-Exo, HG-HUVEC-Exo increased PUM2 expression and decreased SOX2 expression in HTR-8a cells, and GW4869 almost completely rescued SOX2 perturbation induced by HG-HUVEC-Exo (Figures 3(F) and (G)). Although HG-HUVEC-Exo induced the increase of PUM1 mRNA in HTR 8a cells, the increase degree of PUM1 mRNA was lower than the increase degree of PUM2. Moreover, HG-HUVEC-Exo did not induce notable increase of PUM1 protein level in HTR 8a cells. Thus, we choice PUM2 for further study. Collectively, these data indicate that endothelial cell-derived HG-exosomes impair cell proliferation and invasiveness and trigger DNA damage of trophoblasts.
As indicated earlier by RNA-sequencing data, exosomes from HG-HUVECs illustrated alteration in modulation of RNA stability. Therefore, we wondered whether altered expression of SOX2 in HTR-8a cells resulted from RNA instability. Stability of RNA from cells treated with Actinomycin D for 0–12 h was evaluated by SOX2-targeting PCR, and we found that mRNA of SOX2 degraded faster in HG-exosome treated cells than that in NG-exosome treated cells (Figure 4(A)). HTR-8a cells treated by HG-HUVEC-Exo showed instability of SOX2 mRNA and increased interaction between PUM2 protein and SOX2 mRNA. (A) Evaluation of mRNA stability of SOX2 in HTR-8a cells treated by exosomes harvested from HUVECs subjected to conditions as indicated. Evaluation of 18S is included as the internal control. (B) Quantitation of SOX2 mRNA immuno-precipitated by PUM2 antibody in NG-HUVEC-Exo or HG-HUVEC-Exo treated HTR8a cells. IgG precipitation was performed to evaluate non-specific interaction. ***p < 0.001 (n = 3, one-way ANOVA for analysis).
Considering that studies have proved that PUM1/2 directly interacts with mRNA and that SOX2 is a downstream target of PUM2,16,17,22, we certified the interaction between PUM2 protein and SOX2 mRNA by immunoprecipitation. We applied PUM2 antibody to acquired complex of immune-precipitates in the HTR-8a cells treated with NG-HUVEC-Exo and HG-HUVEC-Exo. The immune-precipitates were subjected to qRT-PCR for analysis of SOX2 mRNA. Our results showed a higher SOX2 mRNA level in precipitated complex from HG-HUVEC-Exo treated HTR-8a cells as compared to NG-HUVEC-Exo treated HTR-8a cells (Figure 4(B)), suggesting that higher levels of PUM2 were carried into HTR-8a cells by HG-HUVEC exosomes and PUM2 protein strongly interacted with SOX2 mRNA, possibly affecting the stability of SOX2 mRNA.
Cell damage of HG-HUVEC-Exo treated trophoblasts was mediated by instability of SOX2 mRNA
To further establish the mechanisms underlying the abnormal phenotypes of HG-HUVEC-Exo treated cells, we explored the role of SOX2 mRNA stability in the process. SOX2-knockdown HTR-8a cells (oligo-nucleotide sequence of si-SOX2: 5′-CCCGCATGTACAACATGATGG-3′) were treated with NG-HUVEC-Exo, and SOX2-overexpressed (SOX2-OE) HTR-8a cells were treated with HG-HUVEC-Exo. We found that HTR-8a controls treated by HG-HUVEC-Exo demonstrated significantly promoted cell damage (Figures 5(A)–(C)) and reduced invasiveness (Figures 5(B) and (C)) and cell proliferation (Figure 5(D)) than HTR-8a controls treated by NG-HUVEC-Exo, while SOX2-OE in HTR-8a cells rescued the impaired phenotypes (Figures 5(A)–(D)). Protein and mRNA levels of PUM2 and SOX2 of cells from different groups were then detected to confirm successful knockdown or overexpression of SOX2 (Figures 5(E) and (F)). These results imply that the unfavorable cell phenotypes of trophoblasts induced by HG-HUVEC-Exo are mediated by instability of SOX2 mRNA. Overexpression of SOX2 rescued cell damage induced by HG-HUVEC-Exo. (A) Immunofluorescence of γ-H2AX pS139 (green) in HTR-8a cells treated by osmotic mannitol control, NG-HUVEC-Exo (NG-control) or HG-HUVEC-Exo (HG-control), SOX2-knockdown HTR-8a cells treated by NG-HUVEC-Exo (NG-si-SOX2) or SOX2-overexpressed HTR-8a cells treated by HG-HUVEC-Exo (HG-SOX2 OE). Nucleus was counterstained by DAPI (blue). (B) Illustration of trans-welled cells of HTR-8a cell from groups as indicated in a). (C) Statistical analysis of cell counts of HTR-8a cells in (A) (upper panel) and (B) (lower panel). (D) Illustration of proliferate rate of HTR-8a cells from groups as indicated. (E) Statistical analysis ofmRNA levels of PUM2 and SOX2 in HTR-8a cells from groups as indicated. (F) Representative illustration of western blotting of PUM2 and SOX2 in HTR-8a cells from groups as indicated. **p < 0.01, ***p < 0.001 (n = 3, one-way ANOVA for analysis).
Taken together, we established a model depicting the regulatory machinery of hyperglycemia on trophoblasts (Figure 6). We illustrate that hyperglycemia induces exosome secretion from endothelial cells, and the exosomes carry highly-expressed PUM2 proteins into trophoblasts, subsequently leading to abnormal placentation via PUM2-mediated regression (PMR). We conclude that transcriptional factor SOX2 is a major target of PMR in trophoblasts and instability of SOX2 is crucial for impaired phenotypes of trophoblasts. Graphic illustration of the regulatory machinery of hyperglycemia in trophoblasts. Upon stimulation of high-leveled glucose, the vascular endothelial cells secreted exosomes that contains high level of PUM2 mRNA. After exosomes enter into trophoblasts, PUM2 mRNA is translated to PUM2 protein. PUM2 protein impair the SOX2 mRNA stability, resulting in the reduction of SOX2. SOX2 deficiency is associated to the dysregulation of trophoblasts.
Discussion
Though evidences have shown an established relationship between hyperglycemia and PE,2,12,13 however, the underlying pathogenesis remain elusive. The subject of how hyperglycemia affects trophoblasts and contributes to PE needs further investigation. Vessel function is extensively disturbed in conditions of hyperglycemia including diabetes, and endothelial dysfunction have been considered as a major contributor. However, is endothelial dysfunction the cause or the effect of abnormal placentation in GDM? More experimental evidence is needed to answer this question. As indicated by our results, the exosomes secreted by high glucose-induced endothelial cells impaired the normal function of trophoblasts, which implicated the possibility that endothelial dysfunction in GDM may be the cause of abnormal placentation. While reduced proliferative activity and invasiveness of trophoblasts are closely related to abnormal placentation, down-regulated proliferation and invasiveness of HG-HUVEC-Exo treated trophoblasts shown by our results also supported the idea that hyperglycemia induces abnormal placentation.
As shown by GO analysis, the DEGs we obtained from sequencing of exosomes demonstrated enrichment in multiple biologic processes related to RNA modulation and regulation of oxidative stress, which indicated that high-leveled glucose might induce an oxidative state of endothelial cells and subsequent alteration in expression of related genes and proteins.
SOX2 is a transcription factor with its target site extensively distributed in many genes, many of which are also important transcription factors. 30 According to our data, SOX2 is prone to degrade in trophoblasts under the condition of HG. Multiple studies have reported that PUM2 is critically involved in mediating repression of RNA and protein expression.16,17,22 Therefore, we hypothesized that instability of SOX2 might be mediated by high-leveled PUM2 in HG-induced exosomes, and our results confirmed that there is a direct interaction between PUM2 and mRNA of SOX2 as expected. Our data also showed that SOX2 is required for maintenance of normal state of trophoblasts under the condition of hyperglycemia. Therefore, we speculate that suppressed expression of SOX2 may ultimately trigger oxidative stress and cell damage in trophoblasts through extensive transcription regulation. In addition, in normal conditions of trophoblast development, SOX2 has been shown to mediate self-renewal of trophoblast stem cells as a critical downstream target of fibroblast growth factor (FGF) signaling,31,32 which also supports our result that SOX2 is repressed in dysregulated trophoblasts.
The function of exosomes in intercellular communication have been addressed ever since the identification of exosomes. Placenta is a crucial bridge connecting the maternal side and the fetal part, thus the role of exosomes in placenta have been intensively investigated. For instance, a study investigated the effect of high glucose induced exosomes on endothelial function. 14 Another study also showed that placenta-related exosomes detected in serum inhibited the expression of eNOS in endothelial cells, subsequently affecting development of trophoblasts and progression of PE. 33 These evidences inspired us with the idea that high-leveled glucose in the vessel may regulate trophoblasts by releasing exosomes because there is no direct contact between high-leveled glucose and trophoblasts. Collective results of our experiments supported the model that hyperglycemia induces release of endothelial exosomes to trigger abnormal placentation and trophoblast dysregulation. However, all the experiments in this study were conducted in cultured cells, further evidence in animal models or patient-derived samples may further consolidate our conclusions.
Conclusions
We uncover that the endothelial cell-secreted exosomes induced by high-leveled glucose drive dysregulation of trophoblasts and abnormal placentation through PUM2-mediated repression of SOX2, providing novel insight into abnormal placentation during PE development and potential therapeutic targets for GDM-associated PE.
Footnotes
Authors contributions
Aibing Zhu, Suwan Qi, Wenjuan Li and Dashu Chen conceived and designed the study. Xiaomin Zheng, Jianjuan Xu performed experiments. Aibing Zhu collected the data. Suwan Qi and Yaling Feng interpreted the data. Wenjuan Li and Dashu Chen analyzed the data and wrote the manuscript. All authors reviewed the results and approved the final version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Nanjing Medical University Science and Technology Development Fund (NMUB20210316), Jiangsu Provincial Maternal and Child Health Research Project grant number F202135.
Consent for publication
Written informed consent for publication was obtained from all participants.
Data availability
The data used to support the findings of this study are available from the corresponding author upon request.
