Abstract
Introduction
Arsenic contamination has always been a major worldwide environmental concern. More than 200 million people across the world are reported to be chronically exposed to arsenic concentrations at or above the WHO threshold. 1 Arsenic poisoning is usually manifested by characteristic skin lesions of melanosis and/or keratosis and skin cancer, as well as organ damage of liver, bladder and lung.2–5 Notably, arsenic poisoning is not easily reversible and there is no effective therapeutics. Even after blocking arsenic exposure for many years, patients are at risk for aggravation of skin and multi-organ damage.5–7 Therefore, there is an urgent need for the assessment of arsenic poisoning risk to serve the purpose of early identification and prevention.
Identification of reliable biomarkers is critical for the risk assessment of adverse health outcomes. 8 Because of their excellent performance in responding to environmental stimuli and in involvement in environmentally triggered phenotypes and diseases, epigenetic modifications have been considered as potential biomarkers for the assessment of adverse health risk induced by environmental pollutants. 9 DNA methylation, a focus of environmental epigenetics, has been demonstrated to be a key regulator of the toxicity mechanisms of environmental pollutants, and has been considered to be a viable and reliable marker for the identification of early health damage upon exposure to pollutants.10–12 Chronic arsenic exposure can lead to a diverse promoter methylation profile, which has been found to be involved in disease outcomes by inducing aberration expression in disease-specific genes.13,14 However, DNA methylation biomarkers that can be used in the assessment of arsenic poisoning risk have not yet been identified.
Immune imbalance is an early critical event in the initial development and progression of arsenic poisoning.15,16 The forkhead-family transcription factor FOXP3, which is specifically expressed in Treg cells, is critical for maintenance of immune homeostasis and tolerance. 17 Anomalous DNA methylation of FOXP3, which plays an important role in inhibiting FOXP3 expression and subsequent immunosuppressive dysfunction, has been considered to be a critical marker for identifying the risk of poor prognosis in immune-related diseases such as autoimmune diseases and tumors.18–20 Our previous studies demonstrated that the aberration of FOXP3 methylation could be sustained following continued exposure to arsenic and was involved in arsenic poisoning by triggering inflammation.21,22 These results suggest that FOXP3 methylation might be an early epigenetic biomarker of arsenic poisoning, but it is unknown whether FOXP3 methylation exhibits accuracy, sensitivity and reliability in prediction of arsenic poisoning risk.
In this study, a population of coal-borne arsenic poisoning was recruited to assess the value of FOXP3 methylation in the identification of arsenic poisoning risk. Firstly, we identified the mediation efficacy of FOXP3 methylation on the relationship between arsenic exposure and arsenic poisoning. Additionally, we constructed a nomogram model with FOXP3 methylation as an epigenetic predictor to predict the probability of individual arsenic poisoning and evaluated the model’s ability to discriminate risk. This study revealed the great potential of FOXP3 methylation for the identification of arsenic poisoning risk, and provided a new approach to the application of epigenetic markers in accurately risk assessment of adverse outcomes.
Methods
Investigation site and study population
This study was conducted at two investigation sites. One of the sites is a coal-borne arsenic poisoning area in China, Jiaole village of Xingren county, Guizhou province, where residents had earlier been exposed to excessive amounts of arsenic through contaminated food and air because of the usage of arsenic-rich coal for cooking, drying food and heating. The reference site is Shang Batian village (approximately 12 km from Jiaole village), which is an arsenic-free area but shared similar dietary habits and coal-burning lifestyle with Jiaole village. We invited professional clinicians to perform the physical check of subjects and the diagnosis of arsenic poisoning according to the Endemic Arsenic Poisoning Diagnostic Criteria (WS/T 211–2015). 129 participants were recruited for this study. The arsenic poisoning group included 88 arsenic poisoning residents from the Jiaole village with 28 mild, 32 intermediate and 28 severe arsenic poisoning. The 41 healthy villagers in the control group were recruited from the Shang Batian village. All participants were permanent residents of the local area. Exclusion criteria included participants with presence or history of occupational arsenic exposure and autoimmune diseases, or a recent history of infection or use of drugs that affect immune function, or genetic diseases. The Ethics Committee of the Guizhou Medical University approved this study (No 201403001), and all subjects gave informed consent to the study.
Questionnaire survey and sample collection
Information on demographic factors, lifestyle and residence history of the participants was obtained by a questionnaire survey. Morning urine samples from each subject were collected to measure the urinary arsenic concentration. Fasting venous blood samples were collected using EDTA-anticoagulant tubes, and then were used to isolate peripheral blood lymphocytes (PBLCs) using a lymphocyte separation medium (MP Biomedicals, California, American). The lymphocytes were used for genomic DNA extraction.
Detection of urinary arsenic concentration
The content of urinary arsenic was measured with inductively coupled plasma mass spectrometry (ICP-MS) (Thermo Fisher, Massachusetts, American) according to our previous method. 6 In this study, the urinary arsenic concentration was adjusted with urinary creatinine.
DNA methylation detection
The pyrosequencing was performed to measure the DNA methylation as described previously. 23 The salting out procedure was performed to isolate genomic DNA from the PBLCs. 500 ng DNA were taken for bisulfite modification according to the instructions of DNA methylation kit (Zymo Research, Orange, CA), which was aim to convert unmethylated cytosines to uracils. Establish PCR system to amplify the target region according to manufacturer’s instructions of PyroMark PCR Kit (Qiagen, Germany), and then the sequencing was performed in the PyroMark Q96 sequencer. The target sequences of the FOXP3 gene that we selected for pyrosequencing contain five CpG sites targeting from – 78 bp to +10 bp of the transcription start site. The primers are illustrated in the supplemental Table S1. Sentinel samples and negative controls were also set in each plate. We took the DNA methylation results into statistical analysis when the variations of sentinel samples between each plate were below 5%.
Statistical analysis
Statistical analysis was performed by using SPSS version 21.0 and R 4.1.2. Independent-sample t-test was used to analyze the differences of normal variables (age and FOXP3 methylation) between two groups, while Mann–Whitney U test and χ2 test were performed to explore the differences of non-normal variable (urinary arsenic) and categorical variables (sex, smoking status, and alcohol use), respectively. General linear model (univariate analysis) was performed to assess the differences of FOXP3 methylation between two groups with adjustment for age. The correlation between urinary arsenic and FOXP3 methylation was analyzed by multiple linear regression model. Binary and ordinal logistic regression models were used to explore the correlation of urinary arsenic and FOXP3 methylation with risk of arsenic poisoning and its severity, respectively. The regression models listed above were adjusted for age, sex, smoking status and alcohol use. The mediation model was created with the mediation R package to determine the mediation effect of FOXP3 methylation on the relationship between arsenic exposure and arsenic poisoning. To evaluate the application value of FOXP3 methylation in the risk prediction of arsenic poisoning, using the variables of age, sex, smoking status, alcohol use and FOXP3 methylation as input, multivariable logistic regression analysis was performed to identify the combination of predictors, and a model-based nomogram was created to assess the individual risk of arsenic poisoning. To assess the nomogram’s performance in discriminating arsenic poisoning, internal validation of the model was performed using the bootstrap method and the area under the curve (AUC) of individual arsenic poisoning risk predicted by nomogram was calculated by the receiver operating characteristics (ROC) curve analysis. A significant level α = 0.05 was used.
Results
The demographic data and urinary arsenic load of subjects
Demographic data and factor characteristics of subjects.

aIndependent-sample t-test.
bTwo-tailed χ2 test.
cMann-Whitney U tests.
dGeneral linear model (univariate analysis).
Next, the association between the urinary arsenic levels and the arsenic poisoning risk was analyzed. Compared with the controls, the levels of urinary arsenic in the arsenic poisoning group were significantly higher (p < 0.001) (Table 1). The results from logistic regression models showed that the levels of urinary arsenic were positively associated with the prevalence and the severity of arsenic poisoning (arsenic poisoning prevalence: OR = 1.397, p < 0.001; arsenic poisoning severity: OR = 1.108, p = 0.001) (Table S2).
The response of FOXP3 methylation to arsenic exposure and arsenic poisoning risk
We quantified FOXP3 methylation in PBLCs of subjects using pyrosequencing. The result of liner regression analysis showed that the levels of FOXP3 methylation rose with the increasing concentration of urinary arsenic (β = 0.375, p < 0.001) (Figure 1(a)). We further explored the correlation of FOXP3 methylation with arsenic poisoning. As shown in Table 1, compared to the controls, the higher levels of FOXP3 methylation were observed in the arsenic poisoning group (p < 0.001). Additionally, the increased FOXP3 methylation was found to enhance the risk of arsenic poisoning (OR = 1.215, p < 0.001) and its progression (OR = 1.243, p < 0.001) (Figure 1(b)). Taken together, these results suggested that the FOXP3 hypermethylation might be an epigenetic marker which enabled sense arsenic-exposure insults and assess arsenic poisoning risk. The association of FOXP3 methylation with arsenic exposure and arsenic poisoning. (a) The association between arsenic exposure and FOXP3 methylation analyzed by a multiple linear regression model. The fitted dose-response trend line and the 95% CI were represented with the red solid line and the area between the red dotted lines, respectively. The β-coefficient (95% CI) was from the model to indicate the trends of alterations between arsenic exposure load and FOXP3 methylation. **p < 0.001. (b) Forest graph showed the effects of FOXP3 methylation on the risk of arsenic poisoning and its progression. The OR (95% CI) and p-value were from the logistic regression models. All models adjusted for age, sex, smoking status and alcohol use.
The mediation effect of FOXP3 methylation on the relationship between arsenic exposure and arsenic poisoning risk
To further clarify the relationship between arsenic exposure, FOXP3 methylation, and arsenic poisoning, we performed a mediation analysis to uncover the mediation effect of FOXP3 methylation on the relationship between arsenic exposure and arsenic poisoning risk. As shown in Figure 2, we found that the raised levels of arsenic exposure increased the risk of arsenic poisoning through direct effect (βc = 0.0052, 95% CI: 0.0001–0.010) and mediation effect of FOXP3 hypermethylation (β0 = 0.0020, 95% CI: 0.0002–0.010). The mediation effect accounted for 24.3% (95% CI: 9.56% - 42.00%). In other words, approximately 24.3% of the effect of the arsenic exposure on the arsenic poisoning risk was mediated by the FOXP3 hypermethylation, and the remaining 75.7% was induced by other perturbations due to the arsenic exposure. These results suggested that the FOXP3 hypermethylation mediated by the arsenic exposure might be one of the crucial pathways for the increased risk of arsenic poisoning. The mediation effect model of FOXP3 methylation on the relationship between arsenic exposure and arsenic poisoning. In this mediation analysis model, FOXP3 methylation was included as the mediator, arsenic exposure (the concentration of urinary arsenic) was the exposure variable, and arsenic poisoning was the outcome variable. With bootstrap analysis (1000 bootstrapped samples), the total, direct, mediation effects and their β-coefficient (95% CI) were estimated, adjusting for age, sex, smoking status and alcohol use. (a) The total effect represents the effect estimate of the exposure (urinary arsenic) on the outcome (arsenic poisoning) leaving the potential mediator (FOXP3 methylation) out of the model. Without considering FOXP3 methylation, arsenic exposure load was positively associated with the prevalence of arsenic poisoning. (b) The relationship between arsenic exposure, FOXP3 methylation, and arsenic poisoning. FOXP3 methylation was positively correlated with arsenic exposure and arsenic poisoning. Including FOXP3 methylation as a potential mediator in the model not only revealed a positive direct effect of arsenic exposure on the risk of arsenic poisoning (proportion of direct effect = 75.70%), but also found that arsenic exposure could increase the risk of arsenic poisoning through rising the FOXP3 methylation (proportion of mediation effect = 24.30%). **p < 0.001.
The efficacy of FOXP3 methylation in the risk prediction of arsenic poisoning
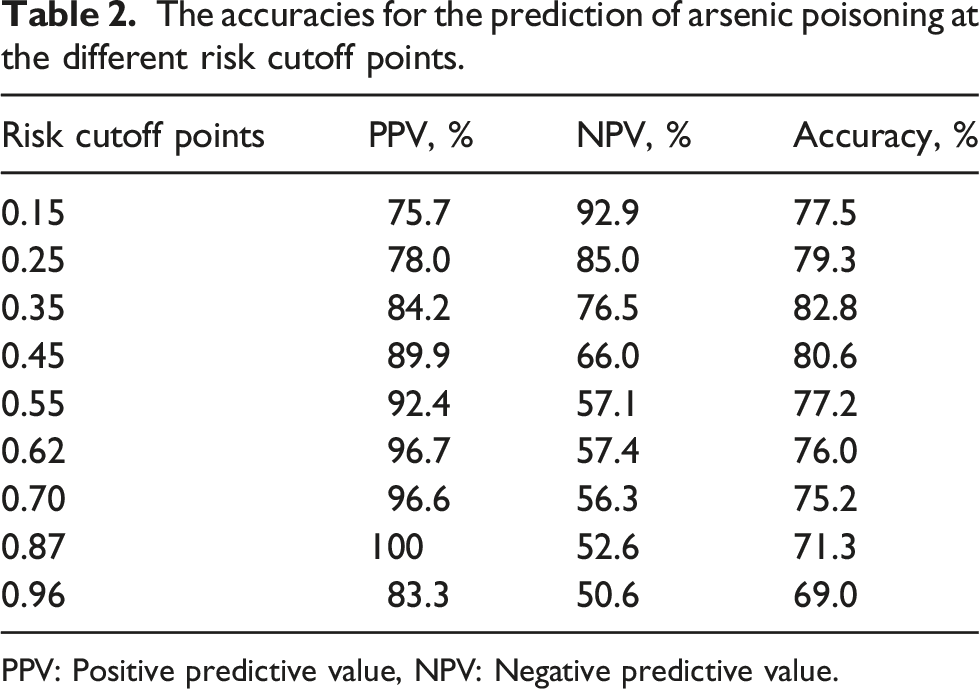
The results listed above suggested that the alteration of FOXP3 methylation was a critical factor in the effect of arsenic poisoning risk. In order to evaluate the power of FOXP3 methylation on discriminating the arsenic poisoning risk, the powerful combination of predictors, FOXP3 methylation and age, was identified by multivariable logistic regression analysis (Table S3). Then, the new model for the assessment of arsenic poisoning risk was established with the combination of these two predictors and depicted in the nomogram (Figure 3(a)). This model was able to quantify the probability of individual arsenic poisoning through the accumulating points from age and FOXP3 methylation levels. The results of the Harrell concordance index (C-index) of the nomogram model showed that there was 89.7% chance that the predicted arsenic poisoning risk was consistent with the actual. The calibration curves of the internal validation performed by the bootstrap re-sampling method also showed that the arsenic poisoning probabilities predicted by the nomogram agreed well with the actual probabilities (Figure 3(b)). Notably, we established the ROC curves using the risk of individual arsenic poisoning predicted by the nomogram model (Figure 4(a)). The nomogram had a powerful prognostic ability with an area under the curve (AUC) of 0.897 (0.845–0.949). The prediction accuracies of the nomogram at the different risk cutoff points were listed in Table 2. The positive predictive value (PPV) rose as the risk scores increased, but the negative predictive value (NPV) decreased. By using the cutoff point 0.35 with the best accuracy, the nomogram yielded accuracy of 82.2%, PPV of 84.2%, NPV of 76.5% (Figure 4(b)). These results indicated a good performance of FOXP3 methylation in the identification of arsenic poisoning risk. Nomogram for predicting the risk of arsenic poisoning. (a) The predictors (FOXP3 methylation and age) were used to create the nomogram to predict the individual probability of arsenic poisoning. The actual levels of predictors were transformed into “points” value according to the ruler of their axis. Then the points of each predictor were summed to obtain a “total points”. Finally, the individual probability of arsenic poisoning was got from the axis of “arsenic poisoning risk” corresponding to the value of “total points”. (b) The calibration curve was drawn for the internal validation of arsenic poisoning probability predicted by the nomogram, which was performed using bootstrap method with 1000 replications to assess the consistency between predictive and actual probability of arsenic poisoning. The x-axis and y-axis represented the predicted probability and the actual probability of arsenic poisoning, respectively. The green and the blue dashed lines represented the perfect prediction and the actual performance of the nomogram model, respectively. The bias of calibration curves was shown by the solid pink line. The discriminative ability of the nomogram in arsenic poisoning risk. (a) The ROC curve was used to derive the area under the curve (AUC) and cut-off values of the individual arsenic poisoning risk predicted from the nomogram, depicted in the figure, demonstrated good discriminative ability of the model. (b) The nomogram scores were obtained by subtracting the cut-off value 0.35 from the nomogram-predicted individual risk of arsenic poisoning, and shown in the figure. If a score was greater than zero, the subject was predicted to be arsenic poisoning by the nomogram, and if not, to be non-arsenic poisoning. While, the actual arsenic poisoning and non-arsenic poisoning subjects were displayed with blue and pink bars, respectively. When the cut-off value of individual arsenic poisoning risk is 0.35, the nomogram yielded positive predictive value (PPV) of 84.2%, negative predictive value (NPV) of 76.5%, accuracy of 82.2%. The accuracies for the prediction of arsenic poisoning at the different risk cutoff points. PPV: Positive predictive value, NPV: Negative predictive value.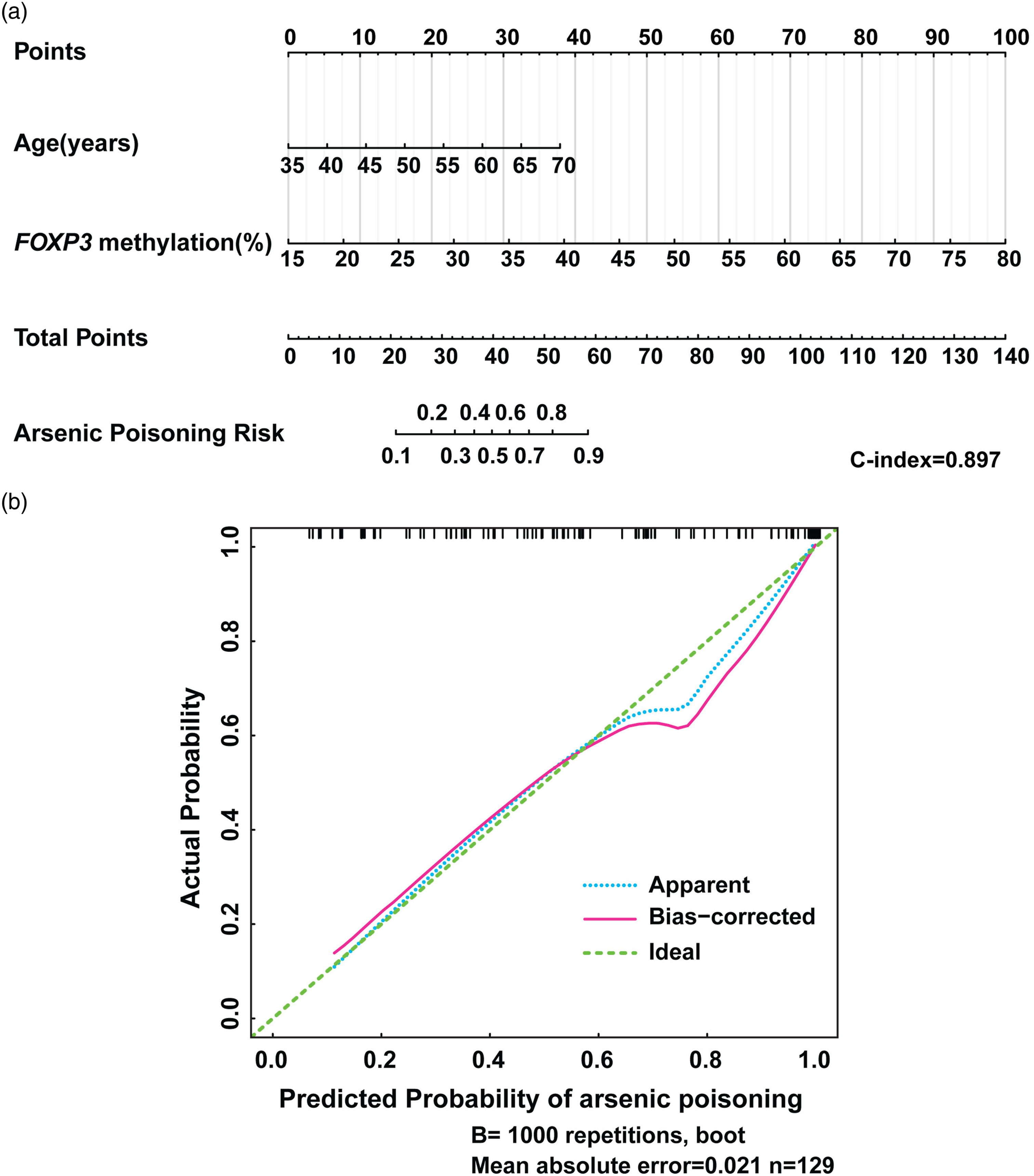

Discussion
Arsenic contamination is widespread in the environment. Arsenic poisoning, an irreversible disease characterized by progressive lesions of skin and multiple organs, is a major health threat to the residents of arsenic-contamination areas. Identification of the reliable epigenetic markers is considered to be an effective strategy for the early recognition of arsenic poisoning risk and the prevention of the initial development and aggravation of arsenic poisoning. In this study, we revealed that the FOXP3 methylation was an epigenetic biomarker which could both respond to arsenic exposure load and mediate an increased risk of arsenic poisoning. We further confirmed that the nomogram model with FOXP3 methylation as a core predictor was reliable in the discrimination of arsenic poisoning risk. These results provided a promising epigenetic marker for accurate assessment of the arsenic poisoning risk.
The inflammation is a crucial mechanism, as well as the early biological event of arsenic induced non-tumor diseases such as progressive skin and multiple organ damage, 5 atherosclerotic vascular diseases, 24 and cardiovascular diseases. 25 Our previous studies demonstrated that arsenic exposure could induce FOXP3 hypermethylation, which was involved in arsenic poisoning by triggering inflammation.21,22 On this basis, this study, using mediation analysis, further revealed that FOXP3 methylation was an epigenetic modification that could both respond to arsenic exposure load and significantly mediate an increased risk of arsenic poisoning. The premise that the epigenetic modifications can be used as biomarkers for the assessment of adverse health risks induced by environmental pollutants is their excellent performance both in response to environmental stimuli and in involvement in environmentally triggered phenotypes and diseases. 9 Thus, we suggest that FOXP3 methylation has the potential to be a biomarker for assessment of arsenic poisoning. Additionally, the literature revealed that the aberrant DNA methylation of FOXP3 was a sensitive marker of environmental pollutants such as black carbon, 26 PM2.5 27 , tobacco smoke, 28 and asbestos fibers, 29 and was involved in lung dysfunction or asthma caused by these pollutants through the disruption of immunosuppressive function and homeostasis of pro/anti-inflammatory. These findings added to the evidences that aberrant FOXP3 methylation may be a universal marker for the risk assessment of environmental exposure-induced inflammation-based health threats.
The potential use of epigenetic signatures as biomarkers is receiving increasing attention. Lines of evidences revealed the potential of aberration FOXP3 methylation for biomarker applications, such as identifying the severity of autoimmune diseases, 19 predicting acute rejection after organ transplantation, 30 and assessing tumor prognosis and efficacy of therapies. 20 However, the application of epigenetic markers has limitations in terms of the compatibility of influencing factors and the accurate prediction of individual risk. The nomogram has been considered as a reliable tool to accurately quantify the individual risk through a comprehensive assessment of the contribution of influencing factors to adverse outcomes. 31 More recently, the nomograms have been used to create the accurate risk prediction models of diabetes, tumor metastasis, and early recurrence of cancer. The combination of variables identified by these nomogram models has been recognized as an important reference for disease prevention and treatment strategies.32–34 In this study, we identified that FOXP3 methylation and age as the powerful predictors of arsenic poisoning risk. Based on this, the nomogram model was performed to integrate the contribution of these two predictors to the probability of individual arsenic poisoning to obtain an accurate risk score. The results suggested that the nomogram model might be an effective strategy in realizing the application of epigenetic markers in accurately quantifying the risk of adverse outcomes.
Additionally, the complexity of FOXP3 methylation patterns in response to the different adverse outcomes is a concern. Hypermethylated FOXP3 causes inflammation, autoimmune disease, and asthma by suppressing Treg cell-mediated immunosuppression,35–37 while FOXP3 hypomethylation provides an immunosuppressive microenvironment for tumor escape and organ transplantation tolerance.30,38 The pattern of FOXP3 methylation may be variable in the different patterns of environmental stress. Our previous results suggested that a reduction in FOXP3 expression was significantly associated with the inflammatory injury induced by chronic arsenic exposure through inactivating the immunosuppressive function. 39 However, sodium arsenite has also been reported to increase FOXP3 expression in the spleen and thymus in acute or sub-acute animal models, thus inducing immunosuppression.40,41 Similar results have been found for the regulation of benzene toxicity. Immunosuppression induced by increased FOXP3 expression has been suggested to be associated with hematopoietic toxicity induced by sub-acute exposure to benzene, 42 while the inactivation of immunosuppressive function induced by inhibition of FOXP3 gene function has been reported to be associated with the increased sensitivity to disturbances in autonomic nervous regulation in chronic benzene-exposure. 43 Since DNA methylation is a critical regulator of FOXP3 expression, it is reasonable to speculate that the different patterns of FOXP3 expression in response to environmental stimulation could be induced by changes in DNA methylation patterns. Identifying the patterns of FOXP3 methylation in response to environmental pollutants or pathological outcomes is necessary to ensure the biological plausibility of this epigenetic biomarker in the application of the risk assessment and to target it for the appropriate prevention strategies.
There are some limitations to this study. Firstly, while this study revealed the potential of FOXP3 methylation in the identification of arsenic poisoning risk, cohort studies and more extensive sample sizes are needed to further validate the ability of FOXP3 methylation to predict the risk of arsenic poisoning and its progression. In addition, future research should consider to incorporating multiple DNA methylation markers that trigger arsenic poisoning to improve the model.
Conclusion
This study identified that the FOXP3 hypermethylation in human peripheral blood lymphocytes played a critical mediating role in the increased risk of arsenic poisoning. Moreover, we revealed that the nomogram model with FOXP3 methylation as a core predictor could accurately assess the risk of arsenic poisoning. These results provided a reliable epigenetic biomarker for the identification of arsenic poisoning risk and a novel approach to the application of epigenetic biomarkers in the accurate prediction of environmental health risks.
Supplemental Material
Supplemental Material - The hypermethylation of FOXP3 gene as an epigenetic marker for the identification of arsenic poisoning risk
Supplemental Material for The hypermethylation of FOXP3 gene as an epigenetic marker for the identification of arsenic poisoning risk by Lu Ma, Xiaolin Fang and Aihua Zhang in Human & Experimental Toxicology
Footnotes
Author contributions
AZ supervised the research project, organized population study, formulated the research question and reviewed the manuscript. LM participated in study of investigation, collated, and analyzed the data and responsible for writing the manuscript. XF participated in measurement of indexes including the FOXP3 methylation and urine arsenic concentration. All authors read and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundations of China, Grant Number: 81430077, U1812403.
Ethical approval
This study was approved by the Ethics Committee of the Guizhou Medical University (No 201403001). The informed consent was obtained from each participant before conducting the study.
Data availability
The data supporting the conclusion of this work are included with the manuscript and its supplemental materials. The datasets generated or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
