Abstract
Di(2-ethylhexyl) phthalate (DEHP) is one of the most common organic pollutants and is added to various plastic products as a plasticizer. DEHP oxidative metabolite content in the human body is associated with DNA damage in sperm and decreased testosterone levels in blood. In this study, a DEHP-induced sperm DNA damage mouse model was replicated and improved, and the transcriptomic and proteomic characteristics of the model were observed. Male mice in the two groups were exposed to DEHP 1 g/kg/d or the same amount of normal saline for 60 days, and the sperm DNA fragmentation index (DFI) was detected by a sperm chromatin structure assay (SCSA). The mRNA and protein expression profiles of the testis were detected by RNA-seq and data-independent acquisition (DIA). The sperm DFI of the DEHP group was significantly increased. Compared with the control group, 111 differentially expressed genes (DEGs) and 2147 differentially expressed proteins (DEPs), such as Lamb2, Ahnak, Tkt, Dnah8 and Tbl2, were found in the DEHP group. These genes were mainly enriched in metabolic pathways, pathways in cancer and the PI3K-Akt signaling pathway. Our results showed that DEHP 1 g/kg/d can induce sperm DNA damage in a male mouse model after 60 days of intragastric administration. The reproductive toxicity of DEHP may be related to metabolic pathways in cancer and the PI3K-Akt signaling pathway.
Keywords
Introduction
The quality and concentration of male sperm have declined rapidly over the past 50 years. 1 It is suspected that this is closely related to the changes in our way of life in recent decades. Di(2-ethylhexyl) phthalate (DEHP) is one of the most widely used phthalates. 2 DEHP is used as a plasticizer added to plastic products, and humans may be exposed to low doses of DEHP for long periods of time through contact with various plastic products, such as plastic food packaging, intravenous fluids, cosmetics, and personal care products. 2
In recent years, studies on reproductive toxicity caused by DEHP have increased gradually. The concentration of DEHP metabolites in male urine was associated with higher infertility treatment failure. 3 Some studies have found that DEHP can cause acute and profound damage to animal sperm, thus posing a significant risk to male fertility. 4 It has been found that DEHP can cause sperm DNA damage in rats exposed to DEHP. 5 However, in-depth research on this model has not been performed. This study intends to replicate and improve a DEHP-induced sperm DNA damage mouse model and to detect the molecular biological characteristics of the model through transcriptomics and proteomics.
Method
Animals and Materials
A total of 16 male SPF ICR (CD1) mice (10 weeks old, body weight 30–40 g) were provided by Henan Skbex Biotechnology Co, Ltd (Anyang, China; license number: SCXK(YU)2020-0005). The mice were numbered and then randomly grouped based on a random number table. All animal experiments and operations were approved by the Ethics Committee of Henan Province Hospital of Traditional Chinese Medicine (approval no. PZ-HNSZYY-2020-018). The animals were raised in the individually ventilated cage (IVC) system of the central laboratory of Henan Hospital of Traditional Chinese Medicine (license no. SYXK (Henan) 2011-0002). In the IVC system, the temperature was maintained at 21–25°C, with a humidity of 38%–60%, a ventilation rate of 22 times/h, and a 12 h light-dark cycle. The mice had free access to standard animal feed and filtered water. The adaptive feeding lasted for 1 week prior to the formal experiment.
Di-2-ethylhexyl phthalate (DEHP) (cat. no.: D201154, lot no.: MKCK4506) was purchased from Sigma–Aldrich (Shanghai) Trading Co, Ltd, Shanghai, China.
Experimental design
After the adaptation period, the mice were stratified at random into two groups (six mice in each group). The control group was given normal saline, and the DEHP group was given DEHP 1 g/kg/d once daily for 60 days (equivalent to two cycles of spermatogenesis and maturation in mice) by intragastric administration on the basis of the literature. 6 After the 60-days modeling, the mice were weighed and anesthetized using sodium pentobarbital. The testis and epididymis were removed via laparotomy and then weighed. The tail of the epididymis was cut and dissected. The sperm were fully released after full shock in 1 mL normal saline for 1–3 min until the solution showed obvious turbidity. The sperm were stored at −80°C before the detection of the sperm DNA fragmentation index (DFI). Both testes were frozen in liquid nitrogen and kept at −80°C until the transcriptomic and proteomic tests.
Sperm chromatin structure assay
The sperm chromatin structure assay (SCSA) was conducted as documented previously with modification 7 at the Reproductive Laboratory of Henan Province Hospital of Traditional Chinese Medicine.
Briefly, the sperm suspension was thawed, transferred into 800 μL of cold TNE buffer, and passed through a 100 μm nylon strainer. The filtered sperm suspension was diluted with the same buffer to a sperm concentration of 2–3 × 106/ml. The sperm suspension (100 μL) was introduced into a FACS tube containing acid detergent solution (200 μL). Then, spermatozoa were stained with acridine orange (600 μL). Each sample was analyzed and gated out by flow cytometry (BD Accuri™ C5) for 5000 sperm. The ratio of single-stranded (red) to double-stranded (green) fluorescence (%DFI) was computed with BD Accuri™ C6 plus software (BD).
Transcriptomic analysis
RNA extraction, library preparation and sequencing
After 60 days of intervention, six mice were randomly selected from the control and DEHP groups (three from each group) for transcriptomic analysis. According to the kit instructions, total RNA was isolated with TRIzol reagent (Invitrogen, United States). The quality of the isolated RNA was examined using an Agilent 2100 Bioanalyzer Platform (Agilent Technologies, Inc, United States), and RNase-free agarose gel electrophoresis was used for confirmation. The eukaryotic mRNA was aggregated with oligo (dT) beads, and the aggregated prokaryotic mRNA was eliminated with the Ribo-Zero™ Magnetic Kit (Epicentre, United States). The aggregated mRNAs were also split into short fragments by fragmentation buffer and converted into first-strand cDNA using arbitrary primers. Generation of second-strand cDNA was performed using DNA polymerase I, RNase H, and dNTPs along with the buffer. The cDNA fragments were purified using the QIAQuick PCR isolation kit (Qiagen, Netherlands), the ends were repaired, the bases were introduced, and ligation of cDNA fragments was performed with Illumina sequencing adapters. The linked products were sequenced by an Illumina NovaSeq 6000.
Analysis of differentially expressed genes
Differential expression analysis of genes (DEGs) between groups was performed using DESeq2 8 software (two samples were analyzed with edgeR). Genes harboring an FDR (false discovery rate) ≤0.05 and an absolute fold change ≥2 were considered DEGs. Then, Gene Ontology (GO) and KEGG pathway enrichment assessments were performed for the DEGs. GO terms or pathways with p ≤0.05 and Q-value ≤0.05 signified remarkably abundant GO terms or cascades by FDR-corrected hypothesis testing.
Proteomic analysis
DIA proteome analysis
Samples were placed in lysis buffer (2% SDS, 1 mg/mL protease inhibitor, 7
Analysis of differentially expressed proteins
As described above, GO and KEGG pathway analyses were carried out on the DEPs (differentially expressed proteins) between the two groups.
Correlation analysis of the transcriptome and proteome
Analyses of GO and KEGG pathway associations
The genes/proteins and differential genes/proteins detected in the transcriptomic and proteomic analyses were counted separately. Correlation analysis of GO terms and KEGG pathway information in the transcriptome and proteome was performed to compare the two groups.
Four-quadrant map analysis
To screen out genes and proteins with the same expression trends, we used R (v3.5.1) software to draw four-quadrant maps based on the changes in gene and protein expression in the transcriptome and the proteome. Quantitative and enrichment assessments were performed for genes and proteins in each region of the four-quadrant map. We focused on the first and third quadrants, where genes and proteins showed the same expression trends.
Results
DEHP-induced sperm DNA damage in mice
The sperm DFI of DEHP-exposed mice was markedly higher than that of control mice (4.15% vs. 29.23%, p = 0.0022; Figure 1(a)). According to the DFI results, three mice were selected at random from the control and DEHP groups for transcriptomic and proteomic analyses. Sperm chromatin structure assay (SCSA) results. SCSA of sperm cells collected from the cauda epididymis (caudal sperm) in mice after exposure to DEHP. (a) Histogram of flow cytometry results. Data are presented as the mean ± SE. In a t test, asterisks show significant differences between the DEHP and control groups (**p < 0.01). (b) Venn diagrams of the transcriptome and proteome. (c) Venn diagrams of the relationships between all genes, proteins, and differentially expressed genes and proteins.
Transcriptomic and proteomic analyses of DEGs and DEPs
A total of 19,690 genes and 15,266 proteins were analyzed by transcriptome and proteome analyses between the control and DEHP groups. A total of 8301 genes and 15,121 proteins were detected in both omics analyses (Figure 1(b)). Venn diagrams were constructed to show the relationships between all genes, proteins, and differentially expressed genes and proteins (Figure 1(c)).
A total of 111 DEGs were screened, including four downregulated and 107 upregulated genes (Figure 2(a) to (c)), and a total of 2147 DEPs were screened, including 785 downregulated and 1362 upregulated proteins (Figure 2(d) to (f)). Analysis of DEGs and DEPs between the two groups. (a)–(c) Bar charts, volcano charts and heatmaps of DEGs. (d)–(f) Bar charts, volcano charts and heatmaps of DEPs. The blue bar/dots/stripes represent the downregulated genes/proteins, and the red bar/dots/stripes represent the upregulated genes/proteins.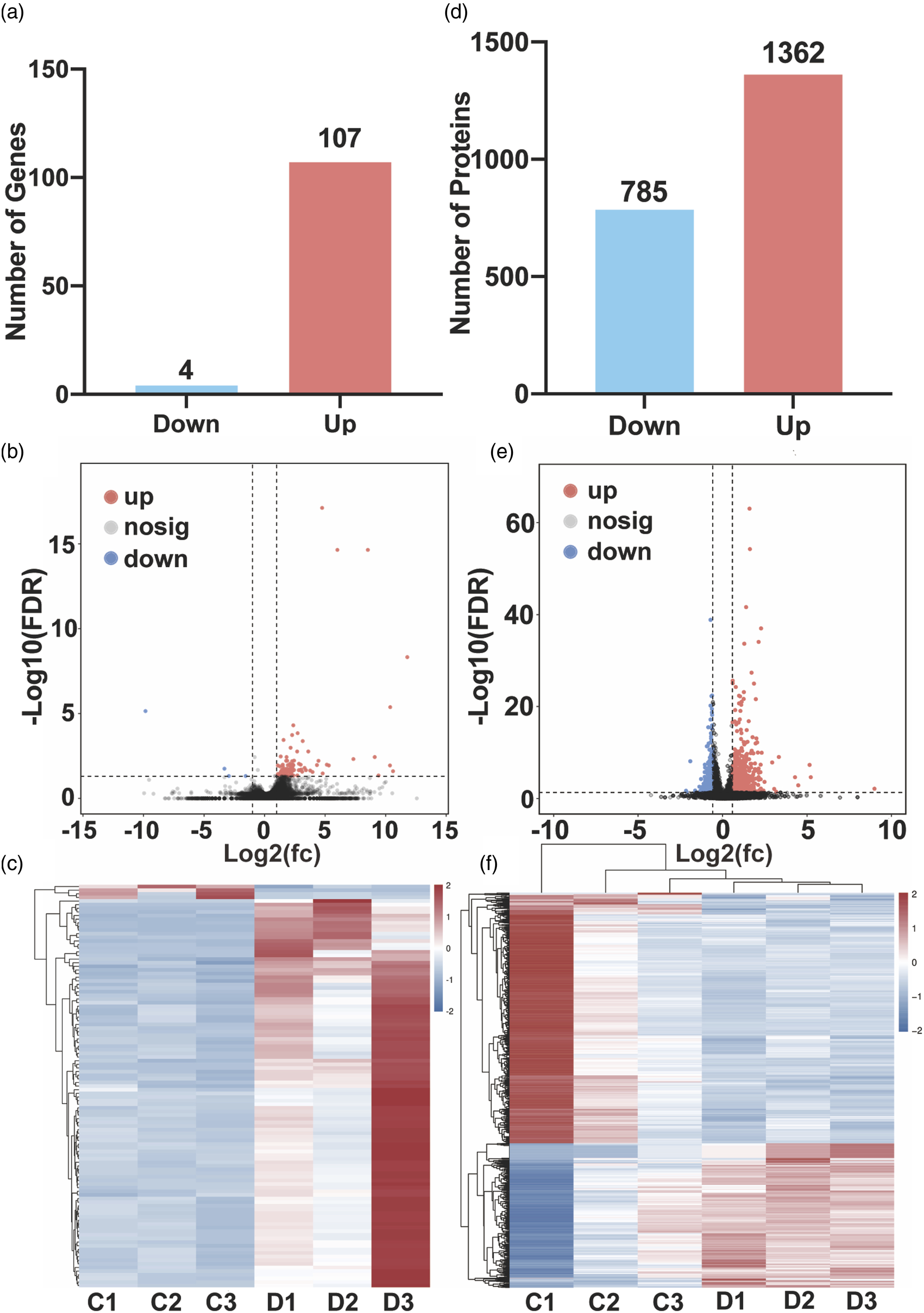
GO and KEGG analyses of the transcriptome and proteome
Both omics analyses revealed high similarity among the three categories in GO analysis. In the biological process (BP) category, the top five GO terms with the largest numbers of genes and proteins were cellular process (GO: 0009987), metabolic process (GO: 0008152), biological regulation (GO: 0065007), organic substance metabolic process (GO: 0071704) and regulation of biological process (GO: 0050789). In the cellular component (CC) category, the top four GO terms with the largest numbers of genes and proteins were cell (GO: 0005623), cell part (GO: 0044464), intracellular part (GO: 0044424) and intracellular (GO: 00056622). In the molecular function (MF) category, the top three GO terms with the largest numbers of genes and proteins were binding (GO: 0005488), protein binding (GO: 0005515) and catalytic activity (GO: 0003824) (Figure 3(a) to (c)). Histogram of GO/KEGG enrichment and classification of DEGs and DEPs. (a)–(c) The top 30 GO terms with the lowest Q-value and p value of the transcriptome and proteome between the two groups (a: biological process, b: cellular component, c: molecular function). (d): The top 30 KEGG pathways with the lowest Q-value and p value of the transcriptome and proteome between the two groups. The abscissa indicates the gene/protein number, and the ordinate indicates the GO terms/KEGG pathway names.
The combined transcriptomic and proteomic KEGG pathway analyses showed that the DEGs and DEPs between the control and DEHP groups were consistent in metabolic pathways (ko01100), pathways in cancer (ko05200), PI3K-Akt signaling pathway (ko04151), Salmonella infection (ko05132), and human cytomegalovirus infection (ko05163) (Figure 3(d)).
Four-quadrant map screening of consistent DEGs and DEPs
Comparisons of the transcriptomes and proteomes between the control and DEHP groups were made (Figure 4). In the four-quadrant map, genes with differentially expressed mRNA patterns consistent with the corresponding proteins were distributed in the first and third quadrants. The upregulated mRNAs and the corresponding proteins were distributed in the first quadrant, while the downregulated mRNAs and corresponding proteins were distributed in the third quadrant. In the first quadrant, 788 upregulated genes and proteins in the DEHP group were screened out, and the top 10 are listed in Table 1, including Lamb2, Ahnak, Tkt, Aldh1l1 and Lamc1. In the third quadrant, 37 downregulated genes and proteins in the DEHP group were screened out, and the top 10 are listed in Table 2, including Dnah8, Tbl2, Sil1, Sh3gl3 and Crat. Four-quadrant diagrams. The horizontal axis is the multiple of transcriptome difference (log2), and the vertical axis is the multiple of protein difference (log2). Each dot represents a gene/protein: red dots represent DEGs and DEPs with consistent trends (both upregulated or downregulated); yellow dots represent DEGs and DEPs with opposite trends; green dots represent differentially expressed genes but nondifferentially expressed proteins; blue dots represent differentially expressed proteins but nondifferentially expressed genes; and black dots represent neither differentially expressed genes nor differentially expressed proteins. Top 10 upregulated DEGs and DEPs (DEHP vs control). Top 10 downregulated DEGs and DEPs (DEHP vs control).


Discussion
Worldwide, more than 15% of couples of reproductive age suffer from infertility, and the incidence is increasing year by year; infertility caused by male factors accounts for approximately 50%. 9 The quality of men’s sperm has declined significantly over the last 80 years. Taking sperm concentration as an example, in 1940, the sperm concentration of healthy men was 113 × 106/ml. 10 By 1980, it had fallen to 60×106/ml, 11 and in the last decade, that number fell to 15 × 106/ml,12,13 one-seventh of what it was in 1940.
Sperm DNA integrity plays a critical role in pregnancy. Compared with the traditional semen analysis method, sperm DFI can evaluate male fertility in a more objective way. Many patients with normal semen analysis parameters have high sperm DFI. 14 Increased SDF is detrimental to embryo development and pregnancy outcomes and may lead to pregnancy miscarriage, recurrent spontaneous abortions, and low live birth rates. 15 It has been shown that a DFI>27% will significantly lower conception and pregnancy rates, and normal pregnancies will not be achieved. 16 In couples who experienced unexplained recurrent miscarriages, the male partners had a higher DFI rate than male partners in normal couples. 17 Therefore, sperm DFI is more likely to determine the outcome of pregnancy.
DEHP exposure was associated with changes in sex hormone levels, sperm motility, sperm concentration, sperm DNA integrity and sperm apoptosis.18–20 Some reports have evaluated DEHP-caused sperm DFI. Experimental studies have found that DFI in F1 male rats was significantly increased when pregnant rats were given DEHP at 5 and 500 mg/kg/d from Day 0 of gestation to parturition, and the increase in DFI in the 500 mg/kg/d group continued from F1 to F3. 21 Although one study reported that DEHP caused DNA damage in rat sperm, the results of this study showed a DFI value of 6.4% at a DEHP exposure dose of 1 mg/kg compared with the control group (2.1%). 5 Although the DEHP group was statistically significant compared with the control group, our study aimed to investigate transcriptomic and proteomic changes after DEHP exposure. Therefore, a larger DFI gap between the two groups is needed. It is necessary to add DEHP dosage on the basis of this model, but considering the cost of modeling, it may be more economical to replace it with a mouse model. Therefore, mice were used in this study. We replicated the DEHP-induced DNA damage mouse model and used the SCSA assay, which is more accurate than in the literature. 6
Oxidative stress and mitochondrial dysfunction in germ cells are thought to be responsible for the phthalate-induced disruption of spermatogenesis in rodents. 21 Adolescent high-fat diets exacerbate reproductive dysfunction associated with prenatal low-dose DEHP exposure in male adult offspring. 22
Reproductive toxicity caused by DEHP is hereditary. Prenatal exposure of mice to 300 mg/kg DEHP reduced spermatogenesis in male offspring and affected sperm DNA methylation in gene promoters, which are thought to be involved in sperm chemotaxis and posttranscriptional regulatory mechanisms. 23 Similar studies have also found that DEHP damages the reproductive function of male rats, affects the expression of DNA methyltransferase, and leads to a change in the genomic imprinted methylation pattern that is passed on to the next generation, which unbalances the key role of imprinted genes in the male reproductive system and eventually leads to cryptorchidism in offspring. 24
Although our study is the first to characterize the transcriptome and proteome of a DEHP-induced mouse model of DNA damage, it has been found that DEHP may affect semen quality by causing dysregulation of seminal plasma polyunsaturated fatty acids (PUFAs) and acylcarnitine metabolism. 26 This is consistent with the results obtained by our analysis; the main enrichment pathway of DEHP after exposure was the metabolic pathway. A high-fat diet can increase the degree of testicular riboflavin and biotin metabolism induced by DEHP. 23
Another category enriched in our results is pathways in cancer. The risk of cancer from exposure to DEHP in ambient air is below the acceptable limit (10−6), but in indoor and occupational settings, the risk of cancer is serious even for short periods of exposure (2 years). 27 Additional data from animal models and studies of humans exposed to environmental DEHP suggest that multiple molecular signals and pathways in several cell types in the liver, rather than a single molecular event, contribute to cancer in rats and mice. 28 Exposure to DEHP may promote cell proliferation by activating progesterone receptor signals, which may increase the risk of breast cancer. 29 Endometriosis, endometrial cancer and abnormal ovulation may occur after exposure to DEHP. 30
The PI3K/Akt pathway, which is a classic pathway related to cell proliferation and apoptosis, was also enriched in our results. DEHP can cause testicular damage and decreased sperm quality in F0-F3 generation mice. Activation of the PI3K/Akt/mTOR signaling pathway can promote the proliferation and differentiation of testicular cells in F1 and F2 generation mice and protect testicular cells from apoptosis. 31 After DEHP exposure, the body may activate autophagy by inhibiting the PI3K-Akt-mTOR signaling pathway to protect the testis from DEHP-induced reproductive damage. 32 Similar results were also observed in mouse cardiomyocytes. Transcriptome sequencing showed that DEHP decreased NRG1 expression and downregulated target genes related to the ErbB2/ErBB4-PI3K/AKT signaling pathway. 33
Conclusion
DEHP may be one of the important pollution factors causing sperm DNA damage. Intragastric administration of DEHP (1 g/kg/d) to male mice for 60 days produced a sperm DNA damage model. DEHP may damage sperm DNA through the metabolic, cancer and PI3K-Akt signaling pathways. 25
Footnotes
Acknowledgments
We appreciate the efforts of all the researchers whose articles were included in this study.
Author contributions
Methodology, Project administration and Writing-Original draft: Chenming Zhang and Zulong Wang. Formal analysis: Rubing Chen. Data curation: Shiqi Wang. Validation: Hao Zhang. Resources: Sicheng Ma. Visualization: Zhong Hua. Conceptualization, Funding acquisition, Writing- Reviewing and Editing: Zulong Wang. The manuscript was read and approved by all authors.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (No. 82174377, 81603632), Natural Science Foundation of Henan Province (No. 222300420485), Science and Technology Program of Henan Province in 2019 (No. 192102310159), the Special Scientific Research Project of National TCM Clinical Research Base (No. 2021JDZX2056, No. 2021JDZX2068) , Innovative Training program for College students in Henan Province (No. 202210471022, No. 202210471057) and Key Scientific Research Project of Colleges and Universities in Henan Province (No. 23A360005).
