Abstract
Aim
Hsa_circ_0000285, a novel circular RNA, has been proven to extensively take part in the pathogenesis of numerous tumors. In hepatocellular carcinoma (HCC), very little is known about hsa_circ_0000285 until now. Hence, this research aims to determine hsa_circ_0000285’s functional role and underlying mechanisms in HCC.
Methods
The expressions of miR-582-3p, hsa_circ_000028, and cyclin B2 (CCNB2) among the HCC cells and tumor samples were determined by performing western blotting and qRT-PCR analyses. The impacts of hsa_circ_000028 on the proliferative and migratory abilities of HCC cells were examined through the execution of CCK-8 and wound-healing assays. Meanwhile, the expressions of the proteins Bcl-2 and Bax were detected via western blotting. Tumor xenograft models were established to examine how hsa_circ_000028 functions during the mediation of HCC tumor growth in vivo. RNA immunoprecipitation and luciferase reporter experiments were performed for the validation of the interactions of miR-582-3p, hsa_circ_000028, and CCNB2 with each other.
Results
Elevated hsa_circ_0000285 and CCNB2 expressions, and a decreased miR-582-3p expression were observed among the HCC cell lines and tumors. Hsa_circ_0000285 bound to miR-582-3p competitively to improve CCNB2 levels. Silencing of hsa_circ_0000285 promoted apoptosis and repressed proliferation and migration among HCC cells. Moreover, silencing hsa_circ_0000285 also impeded the growth of HCC tumors in vivo. Inhibiting hsa_circ_0000285 or CCNB2 reversed the miR-582-3p-knockdown-mediated promotion of malignant HCC cell phenotypes.
Conclusion
Our study has demonstrated that hsa_circ_0000285 fosters the development of malignant HCC cells phenotypes through the modulation of the miR-582-3p/CCNB2 axis. Thus, these results suggest that hsa_circ_0000285 is a prospective target for HCC treatment.
Introduction
Primary liver cancer, a major type of hepatic malignancy, originates from bile duct cells, hepatocytes, and a mixture of the two types of cells. 1 About 90% of the primary liver cancer cases have been accounted to hepatocellular carcinoma (HCC). Primary HCC, a grave threat to human health globally, ranked sixth among the most prevalent cancers and third among the chief causes of cancer mortality worldwide in 2020. 2 At present, the etiological factor of HCC is not understood fully. Some studies have shown that infections with hepatitis B and C virus are closely related to the occurrence of HCC. Meanwhile, other risk factors including the intake of aflatoxin, long-term heavy alcohol intake, non-alcoholic fatty liver, and other genetic factors also stimulate the pathogenesis of HCC.3,4 Although the treatments have been increasing in recent years, they have not fundamentally changed the high mortality rate of HCC. Early HCC lacks specific clinical symptoms. Furthermore, most patients suffering from HCC are already at the advanced stages by the time they are diagnosed, thus missing the best time for radical treatment. In addition, HCC’s 5-yr overall survival rate remains poor, with rates between 30% and 50%, due to frequent recurrence and intrahepatic metastasis. 5 Considering that, seeking new biomarkers is imperative for timely HCC diagnosis and treatment, and for the improvement of the patients’ quality of life.
Circular RNAs (circRNAs) have been observed to control gene expression, thus influencing the development of a variety of tumors, including HCC. 6 For instance, circUBE2J2 serves as a competing endogenous RNA (ceRNA) that regulates the KLF7/miR-370-5P axis, consequently repressing HCC cell migration and proliferation. 7 In another study, circMRPS35 is highly expressed in HCC specimens. 8 Another example is circHPS5. Its m6A modification regulates HMGA2 expression, which promotes the epithelial-mesenchymal transition (EMT) and cancer stem-cell-like phenotypes of HCC cells, thereby facilitating HCC development. 9 Hsa_circ_0000285 is a cancer-related circRNA that is widely involved in various cancers. For instance, the levels of hsa_circ_0000285 are reduced in the tissue and serum of bladder cancer and cisplatin-resistant bladder cancer patients. 10 Also, the levels of hsa_circ_0000285 are closely related to tumor growth and metastasis in bladder cancer. 10 In other studies, hsa_circ_0000285 accelerates osteosarcoma 11 and cervical cancer 12 progression by acting as an oncogene. Hsa_circ_0000285 has been documented to boost cell migration and proliferation in osteosarcoma by regulating miR-599/TGFB2 axis. 11 Inhibiting hsa_circ_0000285 suppresses the malignant development of cervical cancer through the modulation of the miR197-3p/ELK1 axis. 12 Thus, hsa_circ_0000285 has a dual function in the pathogenesis of different cancers. However, it is worth noting that there are no research showing hsa_circ_0000285’s biological function and mechanism in HCC.
MicroRNAs (miRNAs), which are twenty-two nucleotides in length, control the expressions of their target genes at post-transcriptional levels. It is widely known that miRNAs have functions that are essential to biological processes as well as to various human diseases. 13 For instance, a previous report has confirmed that miR-3682 promotes the malignant phenotypes of HCC cells through the inactivation of the AMPK pathway. 14 In another study, miR-125b-5p induces epithelial-mesenchymal transition, stemness, and sorafenib resistance among HCC cells, thereby suggesting that miR-125b-5p is a potential biomarker for the poor prognosis of HCC. 15 Our bioinformatics analysis revealed that miR-582-3p was sponged by hsa_circ_0000285 and targeted Cyclin B2 (CCNB2). Notably, accumulating studies have discovered that the invasion and migration of cancer cells are hampered by miR-582-3p, consequently impairing the malignant phenotypes of prostate cancer and gastric cancer.16,17 CCNB2, from the cyclin family, has been discovered to have a high expression in various malignancies, such as breast cancer and lung adenocarcinoma.18,19 The upregulation of CCNB2 accelerates tumorigenesis and is a prospective biomarker for the poor prognosis of non-small cell lung cancer (NSCLC), lung adenocarcinoma, and HCC.20–22 Additionally, an earlier study has identified CCNB2 as the miR-582-3p downstream target in acute myeloid leukemia. 23 Nonetheless, whether miR-582-3p/CCNB2 can affect HCC progression has yet to be reported.
In this work, we aim to evaluate the effect of the hsa_circ_0000285/miR-582-3p/CCNB2 axis on HCC progression by confirmatory experiments in both HCC cell lines and a xenograft tumour model. We for the first time hypothesize that HCC pathogenesis may be accelerated by hsa_circ_0000285 as it regulates the miR-582-3p/CCNB2 axis. Our findings will aid in the identification of a potential biomarker for predicting HCC progression and a potential therapeutic target for its therapy.
Materials and methods
Participants
This study recruited 36 HCC patients, all of whom signed and provided written informed consent. Thirty-six pairs of tumor tissues and their corresponding normal adjacent tissues (NATs) were acquired from HCC patients that underwent hepatectomy. HCC patients with the following condition were included: (i) the patients were diagnosed with HCC at first time in our hospital; (ii) the patients signed the informed consent. The following patients' samples were excluded: (i) those who received preoperative radiotherapy and chemotherapy; (ii) those whose tissue lacked the typical histomorphologic features of HCC, and (iii) those who had a history of another organ malignancy. All protocols had been approved by the Medical Ethics Committee of School of Clinical Medicine & The First Affiliated Hospital of Chengdu Medical College (Approval number: 2021CYFYIRB-BA-Feb11) and were executed in strict adherence to the Declaration of Helsinki.
Cell culture
The HCC cell lines Hep 3B, Huh7, and SNU-182; and the human normal liver cell line THLE-2 were acquired from the Chinese Academy of Sciences (Shanghai, China). Dulbecco’s Modified Eagle Medium (DMEM; Gibco, Grand Island, NY, USA) was utilized in cultivating the cell lines. The medium used also had 10% fetal bovine serum (FBS; Gibco) and 1% penicillin/streptomycin (Solarbio, Beijing, China). The cell lines were maintained at 37°C in an incubator that had 5% CO2.
Cell transfection
Small interfering RNAs (siRNAs) which targeted hsa_circ_0000285 (si-circ_0000285) and CCNB2 (si-CCNB2) were synthesized for the knockdown of hsa_circ_0000285 and CCNB2. The scrambled siRNA (si-NC) was used as control. The miR-582-3p inhibitor was utilized for miR-582-3p silencing, with inhibitor-NC as control. GeneChem (Shanghai, China) synthesized all the plasmids used. Using a Lipofectamine 2000 Transfection Reagent (Invitrogen, Carlsbad, CA, USA), the plasmids were individually transfected into the Huh7 and Hep 3B cells. After a 48-hr incubation, the cells were gathered, and quantitative real-time PCR (qRT-PCR) was performed to assess transfection efficiency.
CCK-8 assay
Cell Counting Kit-8 (CCK-8; Beyotime, Shanghai, China) was utilized to assess the proliferative capacity of the cells. Huh7 and Hep 3B cells (2000 cells/100 μL) were plated on culture plates containing 96-well and cultivated for 0, 24, 48, and 72 h. The cells were then maintained for another hour at 37°C with 10 µL of a 5 mg/mL CCK-8 reagent. Finally, a microplate reader (Thermo Fisher Scientific, Waltham, MA, USA) was operated to test the absorbance in each well at a wavelength of 450 nm. 24
Wound-healing assay
Huh7 and Hep 3B cells (4 × 105 cells/well) were inoculated on 6-well plates and then cultivated at 37°C to reach 100% confluency. Afterward, the monolayer of cells was scraped with a sterilized 200 µL pipette tip and then maintained in a serum-free DMEM for 24 h. The wounds were observed through an optical microscope (Olympus, Tokyo, Japan). 24 Cell migration rate = (width at 0 h - width at 24 h)/width at 0 h.
Flow cytometry assay
The Annexin V-FITC Apoptosis Detection Kit (Beyotime) was used for flow cytometry. 25 Transfected cells were collected after 48 h of culture and a total of 5 × 104 cells were gathered for detection. These cells were transferred into 190 μL of flow cytometry binding buffer, followed by labelling them with 5 μL of Annexin V-FITC and 5 μL propidium iodide at room temperature for 15 min. Finally, the apoptotic cells were distinguished with the help of a flow cytometer (FACScan™, BD Biosciences, Franklin Lakes, NJ, USA).
Xenograft mouse model
Male BALB/c nude mice that are 4 to 6 weeks old (Shanghai Laboratory Animal Center, Shanghai, China) were maintained in a specific-pathogen-free (SPF) housing. The mice were randomly assigned to the sh-circ_0000285 group or the sh-NC group; each group had five mice. Lentiviruses harboring short hairpin RNA (shRNA) that target circ_0000285 (sh-circ_0000285) and its negative control (sh-NC) were generated by GeneChem. Sh-NC or sh-circ_0000285 were stably transfected into Huh7 cells. With a density of 1 × 106 cells/200 μL, the transfected cells were administered subcutaneously into the right armpit of each mouse. The volumes of the tumors were monitored every week with the aid of a vernier caliper. Tumor volume (cm3) = ab2/2; where a is the long diameter (cm), and b is the short diameter (cm). After 5 weeks, 10% chloral hydrate was used to anesthetize the mice before euthanizing them by means of CO2 inhalation. The tumors were gathered, and their weights were recorded afterwards. 26 All procedures implemented had been approved by the Ethics Committee of The First Affiliated Hospital of Chengdu Medical College (Approval number: 20210019).
Luciferase reporter assay
GeneChem constructed the pmir-Glo vectors containing hsa_circ_0000285 (WT-hsa_circ_000028/Mut-hsa_circ_0000285) and CCNB2 3′ UTR (WT-CCNB2/Mut-CCNB2) mutant (Mut) or wild-type (WT) sequences that could bind to miR-582-3p. The mimic-NC and miR-582-3p mimic were also obtained from GeneChem. With the aid of a Lipofectamine 2000 Transfection Reagent, a combination of a vector and either the mimic-NC or miR-582-3p mimic was introduced into the Huh7 and Hep 3B cells. Following 48-h incubation, a dual-luciferase reporter assay system (Promega, Madison, WI, USA) was utilized to assess the luciferase activities in the transfected cells. 26
Isolation of total RNA from the cytoplasm and nucleus
Following the product protocol, total RNA from nuclear and cytoplasmic fractions were isolated from Hep 3B, Huh7, and tissue samples with the help of a PARIS™ Kit (Thermo Fisher Scientific, Waltham, MA, USA). The total RNAs (5 μg) from Huh7 and Hep 3B cells were treated with 1 μL ribonuclease R (RNase R; Geneseed, Guangzhou, China) for 15 min. These RNAs were then used for qRT-PCR analysis. 27
qRT-PCR
Primer sequences.

Western blotting
The cells and tissue specimens were subjected to RIPA Lysis Buffer (Beyotime) treatment for the isolation of total proteins. 29 The protein concentrations were assessed with the aid of a BCA Protein Assay Kit (Beyotime). Electrophoresis of the total protein was accomplished on an SDS-PAGE gel before the separated proteins were transplanted onto PVDF membranes. Skimmed milk (5%) was added then the membranes were maintained at room temperature for 2 hr to seal them. They were then kept at 4°C overnight with the primary antibodies anti-Bax (1:1000 dilution; #ab182733), anti-Bcl-2 (1:2000 dilution; #ab182858), anti-CCNB2 (1:1000 dilution; #ab185622), and GAPDH antibody (1:2500 dilution; #ab9485). Subsequently, goat anti-rabbit HRP-IgG antibody (1:2000 dilution; #ab6721) was supplemented and then the membranes were maintained for 1 hr at room temperature. All antibodies used were sourced from Abcam (Cambridge, MA, USA). The immunoreactive bands were developed with the aid of an enhanced chemiluminescence (ECL) reagent (Engreen Biosystem, Beijing, China). Finally, the Gel Imaging System and Image J software were utilized to obtain the images and analyze the data, respectively.
RNA immunoprecipitation (RIP) assay
As previously reported, 30 a Magna RIP RNA Binding Protein Immunoprecipitation Kit (EMD Millipore, Billerica, MA USA) was utilized for the argonaute 2 (AGO2) RIP experiment. RIPA Lysis Buffer was utilized to treat the cells. The Protein A/G Magnetic Beads were exposed to anti-IgG or anti-AGO2 antibodies at 4°C for 6 h to coat them. Subsequently, the cell lysates and the beads-antibody complex were maintained together at 4°C for an hour. After subjecting them to proteinase K treatment, the captured RNAs were eluted from the beads–antibody–RNA complex. The miR-582-3p and hsa_circ_0000285 expressions were then quantified via qRT-PCR.
Statistical analysis
SPSS 22.0 statistical software (IBM, Armonk, NY, USA) was employed for the statistical analysis. The assays were each performed thrice, and the figures were reported as the mean ± standard deviation (SD). To analyze the statistical differences between two groups and among many groups, a student’s t test and one-way ANOVA with Tukey’s post-test, respectively, were employed. The relationships among CCNB2, miR-582-3p, and hsa_circ_0000285 were ascertained by means of Pearson’s correlation coefficient. A p < .05 is indicative of a statistically significant result.
Results
An upregulation of hsa_circ_0000285 was observed among HCC cells and tissue samples
To establish the biological role of hsa_circ_0000285 in HCC, we first inspected its levels of expression in specimens from HCC patients. The qRT-PCR outcomes revealed higher hsa_circ_0000285 levels in the tumor tissues than in the NATs (Figure 1(a)). We also discovered that, unlike in the THLE-2 cells, hsa_circ_0000285 was upregulated among the three HCC cell lines (Figure 1(b)). Hep 3B and Huh7 cells demonstrated the highest hsa_circ_0000285 levels among the cell lines used, so they were chosen for further investigation. Nuclear and cytoplasmic RNAs were extracted from Hep 3B and Huh7 cells, and their hsa_circ_0000285 levels were measured. As illustrated in Figure 1(c), hsa_circ_0000285 levels were higher within the cytoplasm than within the nuclei of the Hep 3B and Huh7. The total RNAs obtained from Hep 3B and Huh7 cells were treated with RNase R. We observed that RNase R treatment did not affect the levels of hsa_circ_0000285 within the cells but reduced the expression of its corresponding linear gene, HIPK3 (Figure 1(d)). Hsa_circ_0000285 was aberrantly upregulated among the HCC cells and tumor samples. This indicates that hsa_circ_0000285 possibly exerts an oncogenic function in HCC pathogenesis. Hsa_circ_0000285 was upregulated in the HCC cells and tumor samples. (A) Relative hsa_circ_0000285 expressions in HCC tumors and NATs were detected via qRT-PCR. (B) Hsa_circ_0000285 levels among the normal hepatocyte THLE-2 and HCC cell lines Huh7, Hep 3B, and SNU-182 as assessed via qRT-PCR. (C) qRT-PCR analysis of hsa_circ_0000285 within the cytoplasm and nuclei of Huh7 and Hep 3B (D) qRT-PCR analysis of hsa_circ_0000285 and HIPK3 levels in the total RNA of Hep 3B and Huh7 cells after subjecting them to RNase R treatment. *p < .05, **p < .001 vs. THLE-2 or Control.
Hsa_circ_0000285 knockdown hampered migration and proliferation among HCC cells, boosted their apoptosis, and repressed the growth of HCC tumors.
Hsa_circ_0000285’s function in HCC was explored through its silencing among Hep 3B and Huh7 cells. Following transfection with si-circ_0000285, hsa_circ_0000285 was dramatically reduced in Hep 3B and Huh7 cells (Figure 2(a)). The CCK-8 and wound-healing experiments revealed that knocking-down hsa_circ_0000285 diminished the proliferative (Figure 2(b)) and migratory capacities (Figure 2(c)) of Hep 3B and Huh7 cells. We also examined how knocking-down hsa_circ_0000285 could influence the apoptosis-related proteins Bcl-2 and Bax through western blotting. The results demonstrated that the hsa_circ_0000285 deficiency in Hep 3B and Huh7 cells enhanced Bax protein levels while reducing that of Bcl-2 (Figure 2(d)). Additionally, we also detected the apoptotic rate by flow cytometry, which was found to be high in Huh7 and Hep 3B cells transfected with si-circ_0000285 (Figure 2(e)). Furthermore, tumor-bearing mouse models were established to verify hsa_circ_0000285’s influence on HCC in vivo. The knockdown of hsa_circ_0000285 reduced the tumor volume and weight among the mice (Figure 2(f)). All these findings demonstrate that hsa_circ_0000285 knockdown can repress HCC progression. Knockdown of hsa_circ_0000285 stimulated apoptosis, inhibited the proliferation and migration of HCC cells, and suppressed the growth of HCC tumors. Huh7 and Hep 3B were inserted with either a si-NC or a si-circ_0000285. (A) Relative hsa_circ_0000285 expressions detected via qRT-PCR. (B, C) CCK-8 (B) and wound-healing (C) experiments for cell proliferation and migration. (D) Western blotting analysis of the protein expressions of Bcl-2 and Bax. (E) Evaluation of apoptosis rate via flow cytometry experiment. (F) The volumes and weights of tumors from sh-NC- or sh-circ_0000285-infected Huh7-tumor-bearing nude mice. **p < .001 vs. sh-NC or si-NC group.
MiR-582-3p was a hsa_circ_0000285 target gene
To elucidate the biological function of hsa_circ_0000285 in the progression of HCC, its downstream target gene was identified. CircInteractome database was employed to identify the binding site of miR-582-3p and hsa_circ_0000285 (Figure 3(a)). Subsequently, RIP and luciferase reporter experiments were conducted to substantiate hsa_circ_0000285 and miR-582-3p′s relationship. A poor luciferase activity was observed among the Huh7 and Hep 3B cells carrying the combination of WT-hsa_circ_000028 and miR-582-3p mimic. Meanwhile, no notable changes in luciferase activities were seen among the Huh7 and Hep 3B transfected with MUT-hsa_circ_000028 (Figure 3(b)). Furthermore, the RIP analysis demonstrated that miR-582-3p and hsa_circ_0000285 expressions were considerably elevated in the Anti-Ago2 group, unlike in the Anti-IgG group (Figure 3(c)). These results confirm the interaction of hsa_circ_0000285 with miR-582-3p in Hep 3B and Huh7 cells. Furthermore, we observed decreased levels of miR-582-3p in the tumor tissues of HCC patients (Figure 3(d)). Consistently, a downregulation of miR-582-3p was also detected in Hep 3B and Huh7 (Figure 3(e)). Pearson’s correlation analysis uncovered a negative association existing between the expressions of miR-582-3p and hsa_circ_0000285 in HCC patient specimens (Figure 3(f)). Altogether, our findings substantiate that miR-582-3p is a hsa_circ_0000285 target gene, and that it is downregulated in the HCC cells and tissue samples. MiR-582-3p interacted with hsa_circ_0000285 and was downregulated in HCC cells and tumor samples. (A) CircInteractome identified the prospective miR-582-3p and hsa_circ_0000285 binding sites. (B, C) MiR-582-3p/hsa_circ_0000285 interaction was validated via the luciferase reporter (B) and RNA immunoprecipitation (C) assays. (D) Relative miR-582-3p expressions, analyzed via qRT-PCR, in the tumor tissues and NATs of HCC patients. (E) Relative miR-582-3p expressions among the HCC cell lines, Huh7 and Hep 3B, and normal hepatocyte THLE-2 was measured through qRT-PCR. (F) Correlational analysis of miR-582-3p and hsa_circ_0000285 expressions in HCC patient specimens revealed a negative association.
The hsa_circ_0000285 knockdown inhibited the development of HCC through miR-582-3p targeting
To scrutinize whether hsa_circ_0000285/miR-582-3p can affect HCC, a combination of si-circ_0000285 and miR-582-3p inhibitor was introduced into the Huh7 and Hep 3B cells. The qRT-PCR results revealed that knocking-down hsa_circ_0000285 improved miR-582-3p levels and reversed the miR-582-3p downregulation, which was caused by the miR-582-3p inhibitor (Figure 4(a)). The impacts of silencing hsa_circ_0000285 and/or knocking-down miR-582-3p on the proliferative and migratory capacities of HCC cells were assessed by carrying out CCK-8 and wound-healing experiments respectively. As shown in Figures 4(b) and 4(c), hsa_circ_0000285 silencing repressed the proliferative and migratory abilities of Huh7 and Hep 3B, whereas the miR-582-3p deficiency engendered the opposite of that outcome. The miR-582-3p inhibitor-mediated stimulation of proliferation and migration was abrogated by the hsa_circ_0000285 knockdown. Western blotting analysis revealed that hsa_circ_0000285 silencing in Huh7 and Hep 3B elevated the protein levels of Bax and decreased that of Bcl-2. Following the transfection of Hep 3B and Huh7 cells with miR-582-3p inhibitor, the levels of Bcl-2 protein increased while that of Bax decreased. However, this outcome was abrogated by hsa_circ_0000285 deficiency (Figure 4(d)). As expected, the flow cytometry experiment revealed that hsa_circ_0000285 knockdown increased the rate of apoptosis, but this could be mitigated by inhibiting miR-582-3p. Furthermore, miR-582-3p inhibition reduced apoptosis rates in Hep 3B and Huh7 cell lines transfected with si-circ 0000285 (Figure 4(e)). Altogether, these findings show that knocking-down hsa_circ_0000285 inhibits the development of HCC through miR-582-3p targeting. Hsa_circ_0000285 knockdown inhibited the proliferation and migration of Huh7 and Hep 3B cells and promoted their apoptosis of by targeting miR-582-3p. Huh7 and Hep 3B cells were co-transfected with si-circ_0000285, si-NC, inhibitor-NC, or/and miR-582-3p inhibitor. (A) Relative miR-582-3p expressions were determined via qRT-PCR analysis. (B, C) CCK-8 (B) and wound-healing (C) assays for the assessment of the proliferation and migration of cells. (D) Relative levels of Bcl-2 and Bax proteins as assessed via western blotting. (E) Apoptosis rate was evaluated by flow cytometry experiment. *p < .05, **p < .001 vs. si-NC group; +p < .05, ++p < .001 vs. inhibitor-NC group; #p < .05, ##p < .001 vs. si-circ+inhibitor group.
MiR-582-3p had an interaction with 3′UTR of CCNB2
Using the starBase database, the miR-582-3p and CCNB2 3′ UTR binding sites were revealed (Figure 5(a)). As displayed in Figure 5(b), the combined transfection of WT-CCNB2 and miR-582-3p mimic reduced the luciferase activities among Huh7 and Hep 3B. Meanwhile, no significant changes in luciferase activities were observed among Huh7 and Hep 3B cells carrying MUT-CCNB2. This indicates that hsa_circ_0000285 has interacted with CCNB2 3′ UTR. CCNB2 was obviously upregulated in the tumor samples from HCC patients unlike in the NATs (Figure 5(c)). Consistently, CCNB2 levels were higher in Hep 3B and Huh7 than in THLE2 (Figure 5(d)). Through Pearson’s correlation coefficient, it was revealed that CCNB2 expression was inversely associated to that of miR-582-3p in HCC patient specimens (Figure 5(e)). Thus, miR-582-3p may affect HCC development via targeting CCNB2. MiR-582-3p and CCNB2 3′ UTR interacted. (A) StarBase predicted the potential miR-582-3p/CCNB2 binding sites. (B) CCNB2 and miR-582-3p interactions within Huh7 and Hep 3B cells, as verified through the luciferase reporter assay. (C) The relative expressions of CCNB2 in the tumor tissues and NATs of HCC patient was ascertained by means of qRT-PCR analysis. (D) Relative expressions of CCNB2 among the HCC cell lines, Hep 3B and Huh7, and the normal hepatocyte THLE-2 were assessed through qRT-PCR. (E) The correlational analysis between the miR-582-3p and CCNB2 expressions in HCC patient specimens revealed a negative association.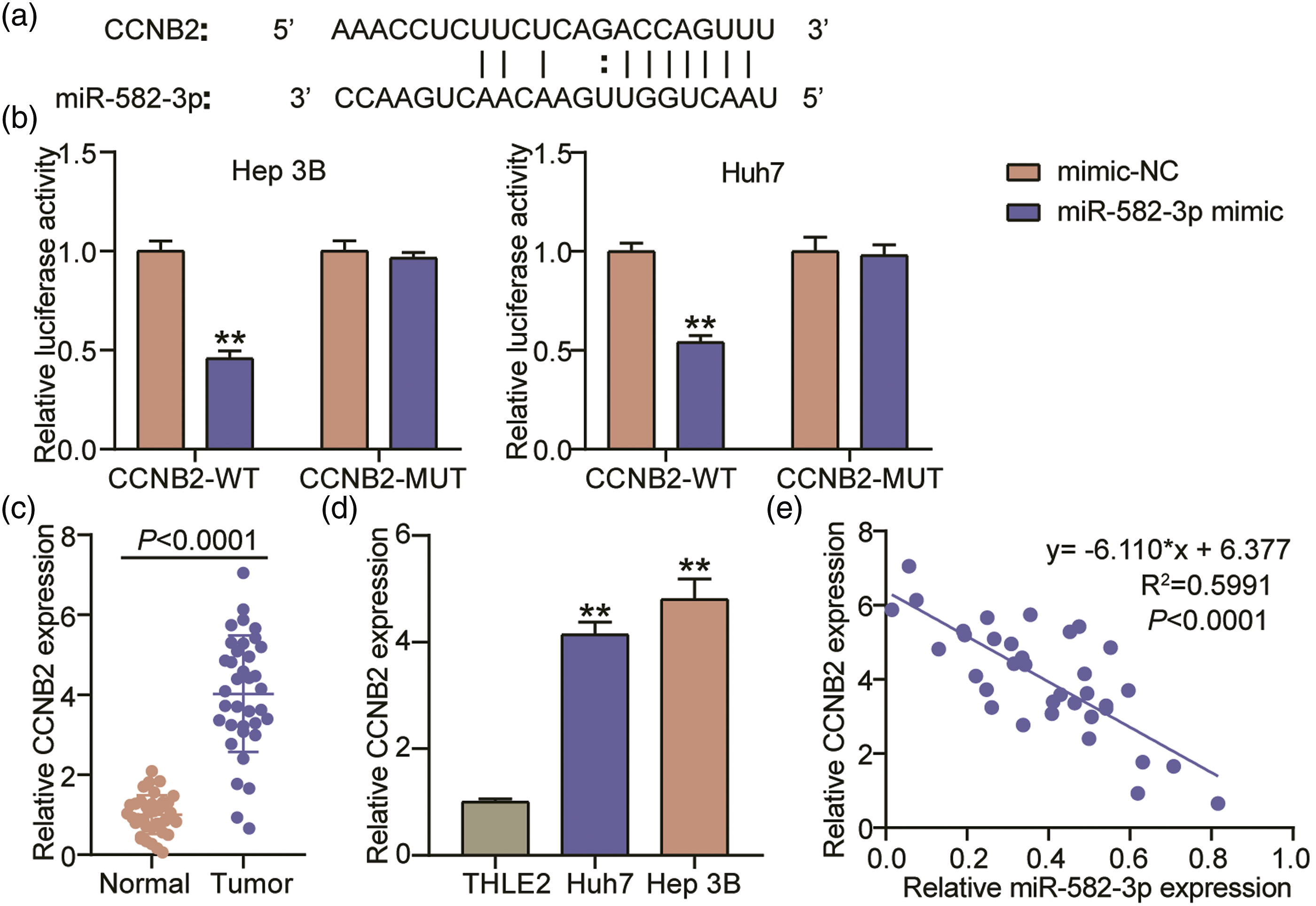
MiR-582-3p knockdown boosted HCC progression by targeting CCNB2
Finally, we validated how miR-582-3p/CCNB2 functions in HCC. Western blotting revealed the considerable reduction in CCNB2 protein levels among Huh7 and Hep 3B cells in the presence of si-CCNB2. Meanwhile, elevated levels of CCNB2 were observed among the miR-582-3p-silenced Huh7 and Hep 3B cells. Nonetheless, the miR-582-3p inhibitor-mediated stimulation of CCNB2 expression was suppressed by knocking-down CCNB2 (Figure 6(a)). CCNB2 silencing deteriorated the proliferative and migratory abilities of the Huh7 and Hep 3B cells. Furthermore, miR-582-3p inhibition among the Hep 3B and Huh7 cells accelerated migration and proliferation, thus reversing the effect of CCNB2 deficiency (Figures 6(b) and 6(c)). The influence of miR-582-3p/CCNB2 on apoptosis was examined via western blotting. Knocking-down CCNB2 in Huh7 and Hep 3B enhanced the protein expression of Bax while repressing that of Bcl-2. Introducing the miR-582-3p inhibitor caused a downregulation of Bax and an upregulation of Bcl-2. Nonetheless, these changes effected by the miR-582-3p inhibitor was extinguished by knocking-down CCNB2 (Figure 6(d)). Cellular apoptosis was accelerated by CCNB2 knockdown, suppressed by miR-582-3p inhibition, and partially slowed by miR-582-3p inhibition during CCNB2 knockdown (Figure 6(e)). Therefore, miR-582-3p inhibition boosted the proliferation and migration of HCC cells and suppressed their apoptosis by regulating CCNB2 expression. MiR-582-3p knockdown promoted HCC progression by targeting CCNB2. Huh7 and Hep 3B were transfected with si-CCNB2, si-NC, inhibitor-NC, or/and miR-582-3p inhibitor. (A) Western blotting analysis of CCNB2 expression in the Huh7 and Hep 3B. (B, C) CCK-8 (B) and wound-healing (C) experiments for cell proliferation and migration. (D) Relative levels of Bcl-2 and Bax proteins as estimated via western blotting. (E) Evaluation of apoptosis rate by flow cytometry experiment. **p < .001 vs. si-NC group; +p < .05, ++p < .001 vs. inhibitor-NC group; ##p < .001 vs. si-CCNB2+inhibitor group.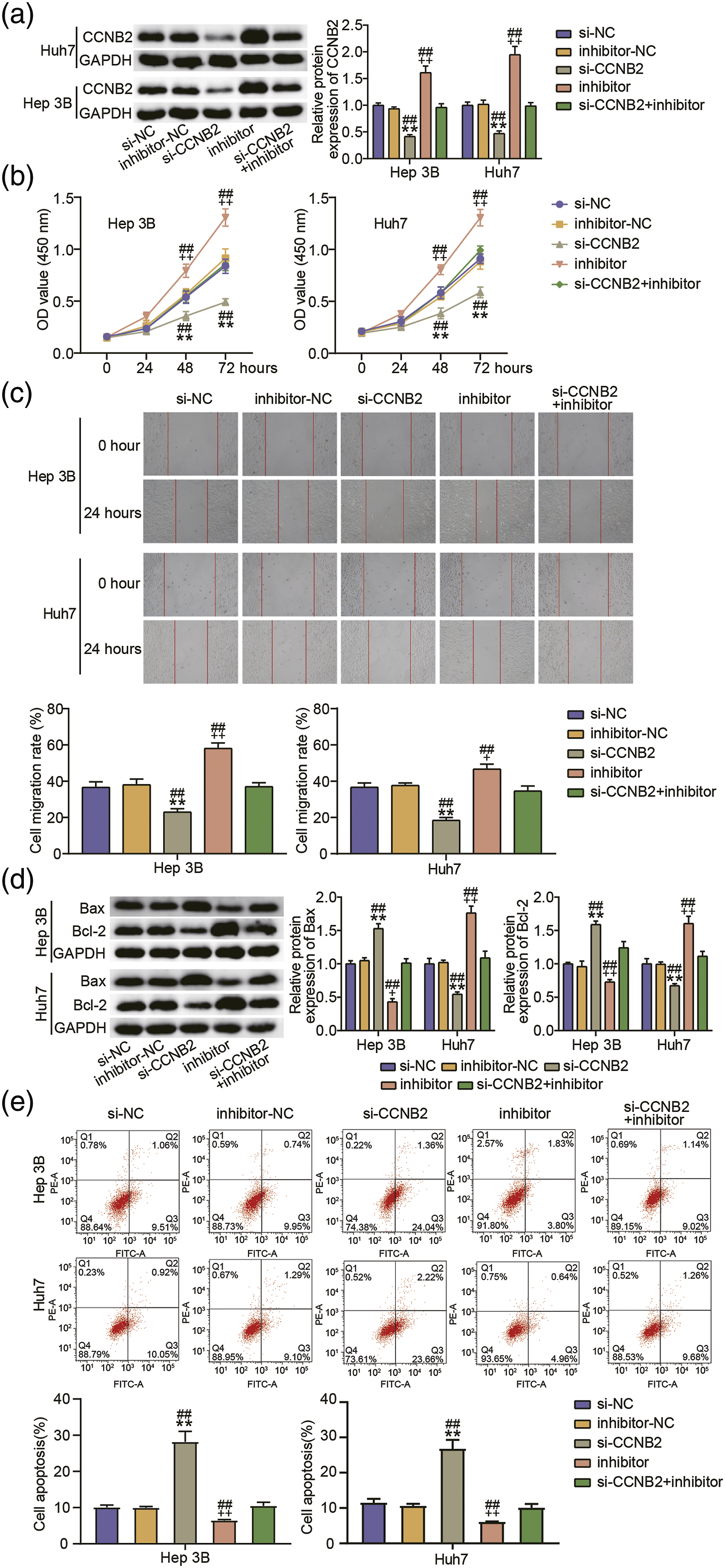
Discussion
HCC is a malignant tumor which gravely imperils human health. In this work, we found an upregulation of hsa_circ_0000285 and CCNB2, and a downregulation of miR-582-3p in HCC cell lines and tumor specimens. Hsa_circ_0000285 enhanced the expression of CCNB2 by serving as a miR-582-3p sponge. Furthermore, hsa_circ_0000285 deficiency repressed the malignant phenotypes of HCC. The reduced proliferative and migratory abilities of HCC cells, as well as the increased apoptosis rate induced by hsa_circ_0000285/CCNB2 knockdown, were partially reversible by miR-582-3p inhibition.
The covalently closed loop structure of circRNAs allows them to stably exist within the cytoplasm or nucleus and enables them to be resistant to RNase R treatment.
31
There is a dearth of understanding about the involvement of hsa_circ_0000285 in HCC. Herein, we observed elevated hsa_circ_0000285 levels among the HCC cell lines and tumor specimens. Interestingly, previous research has suggested that the expression of hsa_circ_0000285 varies in different cancer types. For example, it was reported to be downregulated in bladder cancer tissues 10
Accumulating research have confirmed that miR-582-3p exerts anti-tumor influence in HCC.34–36 Elevated miR-582-3p levels in HCC cells has been documented to suppress migration, invasion, and proliferation. 34 Additionally, functional studies have shown that circRNAs that sponge miRNAs participate in tumorigenesis. 37 For example, miR-582-3p is a key molecule in the circ_HIPK3/miR-582-3p/DLX2 axis and hinders HCC progression. 35 Circ_104075 enhances YAP expression via miR-582-3p sponging, thereby accelerating the progression of HCC. 36 Consistently, our present work documented that miR-582-3p levels were reduced among HCC cells and tumor samples. More specifically, miR-582-3p served as hsa_circ_0000285’s downstream target, and the miR-582-3p silencing-mediated promotion of HCC progression was abrogated by hsa_circ_0000285 knockdown. Thus, hsa_circ_0000285 boosted HCC development via targeting miR-582-3p.
CCNB2 plays a crucial role in regulating cell cycle. Previously, an upregulation of CCNB2 was observed in HCC tumor tissues, and its knockdown was found to inhibit HCC progression. 22 Several research have demonstrated that CCNB2 may become a potential prognostic biomarker for HCC.38,39 Consistently, our data also uncovered that HCC tumor tissues and cells exhibited a significant upregulation of CCNB2. Moreover, an earlier research has reported that CCNB2 is miR-582-3p′s downstream target, and miR-582-3p represses cell cycle in acute myeloid leukemia through the inhibition of CCNB2 expression. 23 We reconfirmed that miR-582-3p negatively regulated CCNB2 by interacting with 3′ UTR of CCNB2, and that the miR-582-3p/CCNB2 axis participated in regulating HCC progression. The CCNB2 knockdown abrogated miR-582-3p inhibitor’s effects, namely the stimulation of migration and proliferation of HCC cells and inhibition of apoptosis.
Certainly, there are some limitations and shortcomings in this work. Only the impact of hsa_circ_0000285 on tumor formation was verified through in vivo experiments. For future works, the function of miR-582-3p/CCNB2 in HCC shall be verified in vivo. More specifically, a tumor xenograft mouse model shall be established by transplanting HCC cell lines carrying a miR-582-3p inhibitor and/or a CCNB2 knockdown vector. Furthermore, the influence of CCNB2 and miR-582-3p on tumor formation shall be observed. Additionally, more hsa_circ_0000285/miR-582-3p/CCNB2-axis-related malignant phenotypes of HCC shall be investigated further. As a cell cycle regulator, CCNB2 has been documented participate in cancer progression by modulating immune infiltration, cell cycle, and distant metastasis.20,21,38 We will further determine whether hsa_circ_0000285/miR-582-3p/CCNB2 axis can regulate those processes to affect HCC development.
In summary, this work has demonstrated that hsa_circ_0000285 promotes the malignant HCC phenotypes by means of controlling the miR-582-3p/CCNB2 axis. The current study’s findings suggest that hsa_circ_0000285 is a promising target for HCC diagnosis and treatment. CircRNAs have emerged as promising therapeutic targets due to their stability and tissue- or cell type-specific expression. As a result, several tools for targeting circRNAs have been developed, such as nanoparticle- and exosome-mediated delivery of circRNA-based therapeutics, synthetic circRNAs, conditional and CRISPR-Cas9-mediated circRNA knockout or knockdown, CRISPR/Cas9-mediated circRNA knockout, and CRISPR/Cas13-mediated circRNA knockdown. 40 Since hsa_circ_0000285 was found to be oncogenic in the current study, we hypothesise that siRNA targeting the junction sequence could downregulate it and thus prevent HCC progression.
Footnotes
Author’s contributions
JH conducted the experiments and data analysis. HCZ conceived and designed the study. YD and ZMY acquired the data. JH performed the data analysis and interpretation. All authors have read and approved this manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical approval
All protocols had been approved by the Medical Ethics Committee of School of Clinical Medicine & The First Affiliated Hospital of Chengdu Medical College (Approval number: 2021CYFYIRB-BA-Feb11). The processing of clinical tissue samples had been in strict compliance with the ethical standards of the Declaration of Helsinki. All patients signed a written informed consent. All animal experiments had been conducted in strict adherence to the ARRIVE guidelines and were carried out in accordance with National Research Council’s Guide for the Care and Use of Laboratory Animals.
Informed consent
All patients signed a written informed consent. All participants provided consent for publication.
