Abstract
Background
Acute paraquat poisoning-induced multiple organ dysfunction syndrome (MODS) leads to the high mortality. This study aimed to investigate the clinical significance of microRNA-200b-3p (miR-200b-3p), an upstream inhibitor of high-mobility group box 1 (HMGB1), in acute paraquat poisoning patients for the prediction of MODS and survival.
Methods
This study enrolled 80 patients with MODS induced by paraquat and 94 healthy volunteers. The interaction between miR-200b-3p and HMGB1 was identified by luciferase reporter assay. miR-200b-3p levels were measured by quantitative real-time (QRT) PCR. High-mobility group box 1 levels were measured by enzyme-linked immune sorbent assay (ELISA). Receiver operating characteristic analysis was used to evaluate the diagnostic value of miR-200b-3p in screening MODS patients. The relationship between miR-200b-3p and the 28-day survival of MODS patients was evaluated by Kaplan–Meier curves and log-rank test. Cox regression analysis was used to assess the prognostic value of miR-200b-3p. Correlation between miR-200b-3p and HMGB1 was confirmed by Pearson’s correlation analysis.
Results
miR-200b-3p directly target HMGB1. miR-200b-3p, decreased in MODS patients, had high diagnostic value to screen MODS patients from healthy controls. Additionally, serum miR-200b-3p was decreased in non-survivors, and patients with low miR-200b-3p level had poor 28-day survival. Serum miR-200b-3p could independently predict the survival prognosis. Moreover, serum HMGB1 level was increased in MODS patients, and was negatively correlated with miR-200b-3p level.
Conclusion
Decreased miR-200b-3p may function as a biomarker for the diagnosis and survival prognosis of MODS patients, and miR-200b-3p may be involved in the progression of acute paraquat-induced MODS via regulating inflammatory responses by targeting HMGB1.
Introduction
Paraquat, a type of organic compound containing 1,1’-dimethyl-4, 4’-bipyridinium dichloride, is applied as an herbicide; In developing countries, intentional self-mutilation by oral paraquat has become a serious public health problem. 1 Paraquat poisoning is always fatal after ingestion and leads to the death of 50–90% of affected individuals; although comprehensive clinical therapy has been adopted, the mortality rate is still high. 2 Multiple organ dysfunction syndrome (MODS) caused by paraquat poisoning has been found to be a leading cause of death in poisoned individuals. 3 Thus, exploring effective approaches for indicating the diagnosis and survival prognosis of MODS patients caused by paraquat poisoning is urgently needed to solve clinically.
Aberrant inflammatory response plays a crucial role in the progression of MODS. 4 The early treatment of patients with paraquat poisoning is mainly anti-inflammatory treatment. 5 Currently, some aberrantly expressed genes and microRNAs (miRNAs) in the development of inflammatory response have been found to serve as therapeutic targets for patients with paraquat-induced MODS.6,7 For example, miR-27a may regulate inflammatory responses to play a role in the progression of paraquat poisoning caused MODS, which is modulate by interleukin (IL)-10. 6 miR-219-5p may participant in MODS process by regulating inflammatory responses via affecting toll-like receptor 4 (TLR4). 7
High-mobility group box 1 (HMGB1) is an early mediator of injury and inflammatory response, 8 which has been proved to play a role in the molecular mechanisms of paraquat toxicity.9–11 For example, Huang et al. have reported that HMGB1 is significantly increased in a concentration and time-dependent manner after exposure to paraquat, and is involved in paraquat-induced neuron death. 9 High-mobility group box 1-TLR4-IL-23-IL-17A axis plays an important role in the pathogenesis of paraquat-caused acute lung injury. 10 High-mobility group box 1 level in fatalities is higher than that in survivors, and can be used as an independent predictor of 30-day mortality in patients with acute paraquat poisoning, suggesting the crucial role of HMGB1 in the injury of poisoned patients. 11 In addition, it has been known that miRNAs can regulate HMGB1 in diseases, such as miR-140-5p 12 and miR-627. 13 A previous study has shown that HMGB1 is the target of miR-200b and is regulated by miR-200b. 14 We then performed bioinformatics prediction in our study and found that miR-200b-3p can bind to HMGB1. In addition, miR-200b-3p has been reported to regulate inflammatory response.15,16 Thus, we speculated that miR-200b-3p may regulate HMGB1 to play a role in MODS caused by acute paraquat poisoning.
Thus, the purpose of this study is to investigate the correlation of miR-200b-3p with HMGB1, and to explore the expression and clinical significance of miR-200b-3p in patients with MODS caused by acute paraquat poisoning. This study provides a new insight into the diagnosis and therapy for paraquat-induced MODS.
Material and methods
Patients and sample collection
All the procedures of this study were approved by the Ethics Committee of Qingdao Hospital of Traditional Chinese Medicine (Qingdao Hiser hospital) and were conducted in accordance with the Declaration of Helsinki. Each patient signed written informed consent when entering the emergency room (ER). The present study included 80 patients with acute paraquat poisoning who visited the ER of Qingdao Hospital of Traditional Chinese Medicine (Qingdao Hiser hospital) between 2013 and 2018. The patients took paraquat orally for 2–100 mL and arrived at the hospital within 24 h. Each patient was treated according to a standard comical treatment protocol. Patients were excluded from this study if they (1) had immune or immune-related diseases, such as diabetes mellitus, hypertension, renal failure, connective tissue disease or malignancy or (2) incomplete follow-up data. Venous blood was collected immediately after patient admission. In addition, 94 healthy volunteers who had healthy examination during the same period at this hospital were enrolled into the control group. The venous blood samples from all health controls were also collected. Serum was then isolated from blood by centrifugation and stored at −80°C for further use. The patients were followed up to death or 28 days.
Data collection
Baseline clinical characteristics of all patients were recorded within 24 h after admission, including: age, gender, time from paraquat ingestion to emergency department, white blood count (WBC), blood urea nitrogen (BUN), serum creatinine (Scr), alanine transaminase (ALT), aspartate transaminase (AST), arterial partial pressure of carbon dioxide (PaCO2), blood lactic acid (Lac), and cardiac troponin I (cTnI).
Cell culture and transfection
The 293 cell line was purchased from Shanghai Cell Bank of Chinese Academy of Science (Shanghai, China), and the cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS; Gibco, Grand Island, New York). miR-200b-3p mimic (5’-UAAUACUGCCUGGUAAUGAUGA-3’) and its negative control (mimic NC, 5’-UUCUCCGAACGUGUCACGU-3’) were purchased from GenePharma (Shanghai, China), which were used to regulate the expression of miR-200b-3p. miR-200b-3p mimic and mimic NC were transfected into 293 cells using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) following the protocols of manufacturer. Cells were collected after 24 h of transfection and then used for the following analysis.
Luciferase reporter assay
This study used starBase V3.0 (https://starbase.sysu.edu.cn) to predict that the 3’-UTR of HMGB1 includes complementary sequence of miR-200b-3p. The luciferase reporter assay was performed to verify whether there has a direct interaction between miR-200b-3p and HMGB1. High-mobility group box 1-mutant-type (MUT) 3’-UTR and HMGB1-wild-type (WT) 3’-UTR containing the binding site of miR-200b-3p were, respectively, inserted in the pGL-control vector (Promega, Madison, WI, USA). The combined vectors were then co-transfected with miR-200b-3p mimic and mimic NC into 293 cells using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA). After 48 h of transfection, relative luciferase activity was measured using a dual-luciferase reporter assay system (Promega). Firefly luciferase activity was normalized to Renilla luciferase activity.
RNA extraction and quantitative real-time PCR
Total RNA was extracted from serum samples and cells by TRIzol reagent (Invitrogen, Carlsbad, CA, USA). RNA purity and concentration were assessed by NanoDrop 2000 (Thermo Fisher Scientific, Inc.). The extracted RNA was reverse transcribed into cDNA using PrimeScript RT Reagent Kit (TaKaRa, Japan). The levels of miR-200b-3p were detected by quantitative real-time-PCR (qRT-PCR), which was performed using SYBR green I Master Mix kit (Invitrogen, Carlsbad, CA, USA) and a 7500 Real-Time polymerase chain reaction (PCR) System (Applied Biosystems, USA). U6 was used as an endogenous reference gene for the normalization of miR-200b-3p. The final relative levels were calculated using the 2−ΔΔCt method. 17
Enzyme-linked immune sorbent assay for HMGB1
High-mobility group box 1 levels in the serum of patients were measured by enzyme-linked immune sorbent assay (ELISA) kits (ArigoBio) according to the instructions of manufacturer. Microtiter plates have been pre-coated with HMGB1 specific antibodies. Then, each well was added with a primary antibody conjugated to Horseradish Peroxidase (HRP), which can bind to HMGB1, and then incubated at 37°C. After that, substrate solution (TMB) was added to each well after any unbound antibody–enzyme reagent was washed away. Finally, the reaction was stopped by adding acid, and the levels of HMGB1 were calculated by detecting the optical density (OD) value at 450 nm using microplate reader (Bio-Rad, Hercules, CA, USA). The results were expressed in ng/ml.
Statistical analysis
All statistical analyses were conducted using SPSS 22.0 software (IBM Corp.) and GraphPad Prism 7.0 software (GraphPad Software, Inc.). The analysis results of data were shown as mean ± SD. Differences in measurement data were compared using unpaired Student’s t-test or one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test, and the differences between categorical variables were compared by Chi-square test. The diagnostic ability of miR-200b-3p to screen MODS patients from healthy volunteers was evaluated by performing receiver operating characteristic (ROC) analysis. Kaplan–Meier curves and log-rank test were used to assess the association of miR-200b-3p with the survival of patients. The prognostic value of miR-200b-3p was confirmed by Cox regression analysis. Pearson’s correlation analysis was used to evaluate the correlation between miR-200b-3p and HMGB1. A difference with a p < .05 indicated statistical significance.
Results
miR-200b-3p targets HMGB1
The complementary binding sequences of miR-200b-3p with HMGB1 predicted through starBase V3.0 platform were shown in Figure 1(a). Relative miR-200b-3p expression was found to be increased by miR-200b-3p mimic (Figure 1(b), p < .001). The results of luciferase reporter assay showed that miR-200b-3p overexpression markedly suppressed relative luciferase activity in HMGB1-WT group (p < .05), whereas, no change was observed in HMGB1-MUT group (Figure 1(c)). HMGB1 is a direct target gene of miR-200b-3p. (A) The complementary binding sequences between miR-200b-3p and HMGB1. (B) Expression of miR-200b-3p was upregulated by the miR-200b-3p mimic. (C) Relative luciferase activity in HMGB1-WT group was inhibited by overexpression of miR-200b-3p. *p < .05, ***p < .001 vs. Untransfected. HMGB1, high-mobility group box 1; NC, negative control; WT, wild type; MUT, mutant type.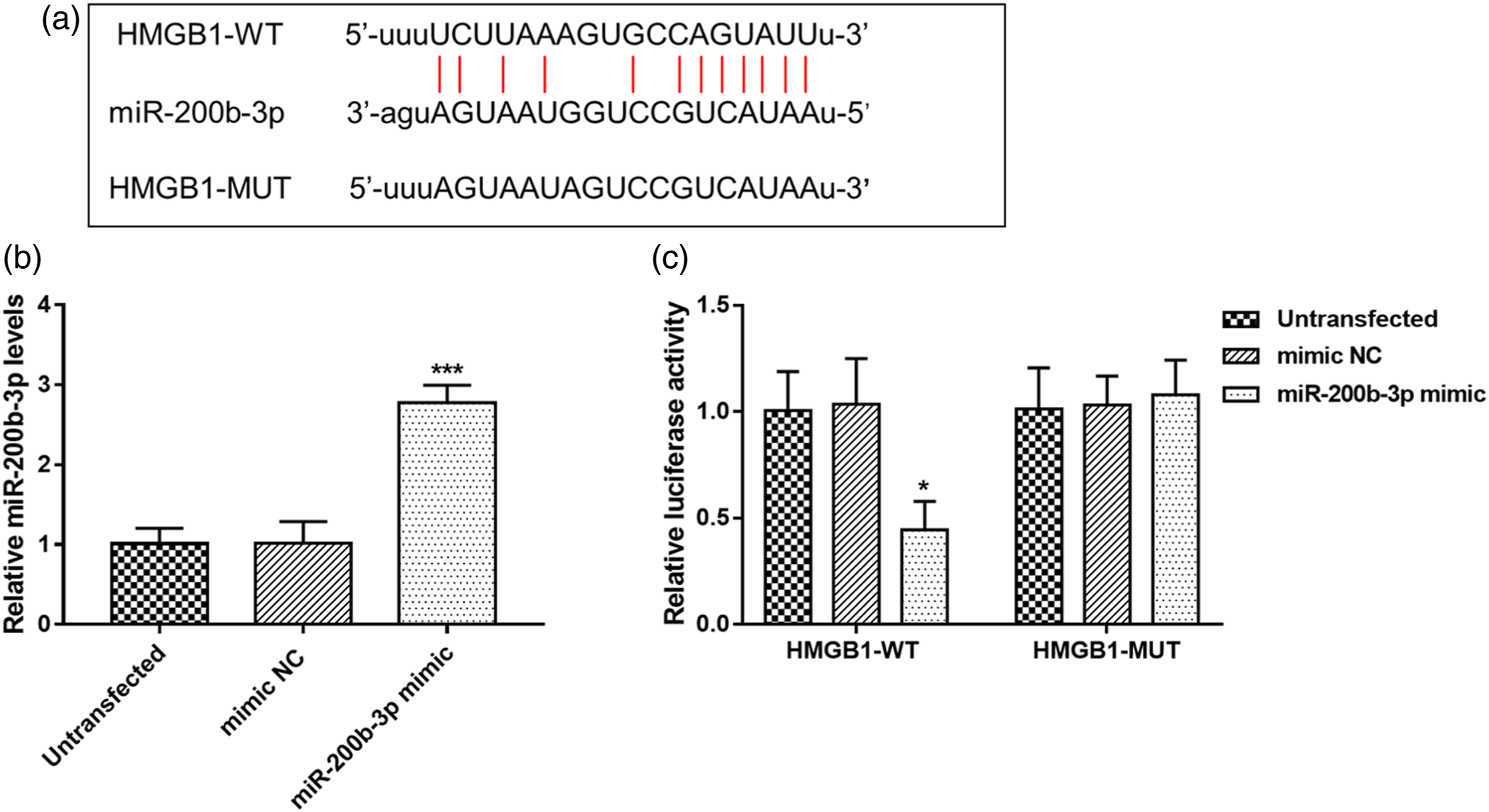
Serum miR-200b-3p expression in patients with MODS caused by paraquat poisoning
The expression of serum miR-200b-3p, measured by qRT-PCR method, was significantly decreased in MODS patients compared with that in healthy controls (Figure 2, p < .001). Expression of serum miR-200b-3p in patients with MODS caused by paraquat poisoning and healthy controls. ***p < .001 vs. Healthy controls. MODS, multiple organ dysfunction syndrome.
Association of miR-200b-3p with clinical characteristics of MODS patients induced by paraquat poisoning
Association of miR-200b-3p expression with the clinical characteristics of MODS patients induced by paraquat poisoning.

MODS, multiple organ dysfunction syndrome; WBC, white blood count; BUN, blood urea nitrogen; Scr, serum creatinine; ALT, alanine transaminase; AST, aspartate transaminase; PaCO2, partial pressure of carbon dioxide; Lac, arterial blood lactic acid; cTnI, cardiac troponin I.
Diagnostic potential of miR-200b-3p in patients with MODS induced by paraquat poisoning
The results of Figure 3 demonstrated that miR-200b-3p had high diagnostic potential in screening MODS patients from healthy controls with an area under the ROC curve (AUC) of 0.896. At the cutoff value of 0.640, the sensitivity and specificity were 81.25% and 85.11%, respectively. miR-200b-3p had high diagnostic value in differentiating MODS patients from healthy controls. AUC, area under the ROC curve; ROC, receiver operating characteristic; MODS, multiple organ dysfunction syndrome.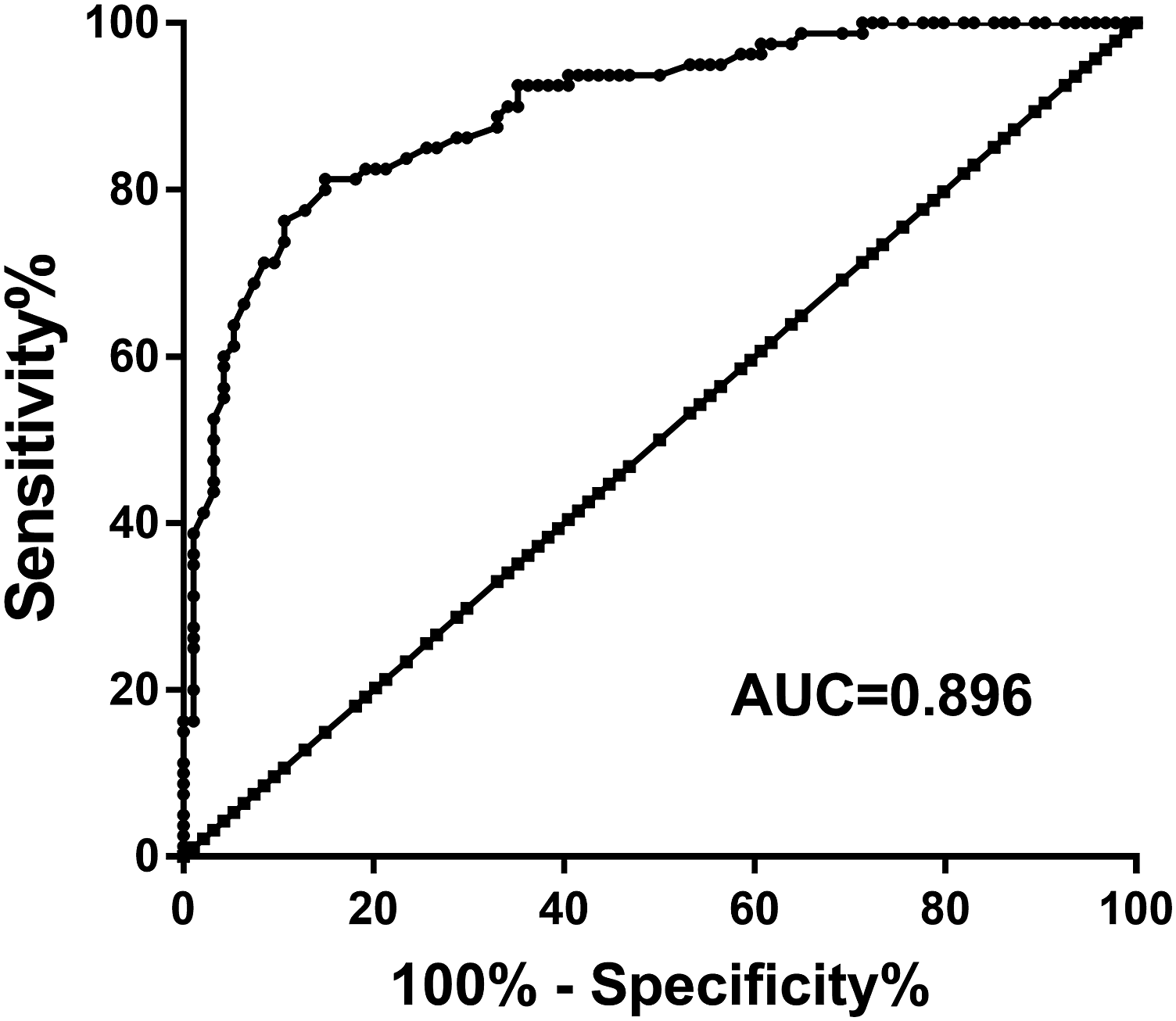
Predictive value of miR-200b-3p for the 28-day survival prognosis of MODS patients
As presented in Figure 4(a), the non-survivors had significantly lower miR-200b-3p expression than the survivors (p < .001). The Kaplan–Meier survival curves were then established to explore the association between miR-200b-3p expression and percent survival of patients. The Figure 4(b) revealed that low miR-200b-3p expression was associated with poor survival in MODS patients (log-rank p < 0.001). Cox analysis results are shown in Table 2. The results of univariate Cox analysis demonstrated that BUN, Scr, PaCO2, Lac and miR-200b-3p were significant factors influencing the survival of MODS patients. Significant baseline characteristics from univariate analysis were included in multivariate analysis. Multivariate analysis results demonstrated that miR-200b-3p [hazard ratio (HR) = 3.665, 95% confidence interval (CI) = 1.826–7.357, p < .001] was independently associated with the survival prognosis of MODS patients. Association between miR-200b-3p expression and 28-day survival prognosis of MODS patients. (A) Expression of miR-200b-3p in survivors and non-survivors. (B) Patients with low miR-200b-3p expression had poor survival prognosis (log-rank p < .001). ***p < .001 vs. Survivors. MODS, multiple organ dysfunction syndrome. Cox regression analysis for MODS patients induced by paraquat poisoning. MODS, multiple organ dysfunction syndrome; HR, hazard ratio; CI, confidence interval; WBC, white blood count; BUN, blood urea nitrogen; Scr, serum creatinine; ALT, alanine transaminase; AST, aspartate transaminase; PaCO2, partial pressure of carbon dioxide; Lac, arterial blood lactic acid; cTnI, cardiac troponin I.

Correlation of miR-200b-3p with HMGB1 in MODS patients
The level of serum HMGB1, measured by ELISA method, was significantly increased in MODS patients compared to that in healthy subjects (Figure 5(a), p < .001). Additionally, a negative correlation between HMGB1 levels and miR-200b-3p levels in patients with MODS induced by paraquat poisoning was found (Figure 5(b), r = −0.647, p < .001). Correlation between miR-200b-3p and HMGB1 in MODS patients. (A) Expression of serum HMGB1 levels in patients with MODS and healthy controls. (B) miR-200b-3p level was negatively correlated with serum HMGB1 levels in MODS patients (r = −0.647, p < .001). ***p < .001 vs. Healthy controls. HMGB1, high-mobility group box 1; MODS, multiple organ dysfunction syndrome.
Discussion
Previous evidence has suggested the critical role of HMGB1 in the process of paraquat poisoning.9,10 In addition, miRNAs are known regulators of important biological processes in animals and plants, which play crucial roles in the processes of various diseases. 18 Our preliminary bioinformatics study revealed that miR-200b-3p could directly target HMGB1. Then our study results demonstrated that HMGB1 was inhibited by miR-200b-3p overexpression. Therefore, we speculated that miR-200b-3p had a great influence on paraquat caused MODS. Further study found the lower miR-200b-3p expression in MODS patients than that in healthy participants. Besides, we found the significant association of miR-200b-3p with WBC, BUN, Scr, PaCO2, and Lac. BUN and Scr are critical indexes reflecting kidney function, and PaCO2 and Lac are critical indicators reflecting lung and respiratory function. Therefore, miR-200b-3p may be involved in the process of lung and kidney injury in MODS patients. miR-200b-3p has been reported to reduce inflammatory responses in pulmonary emphysema 19 and suppress renal fibrosis in mouse, 20 which support the results of our study. Notably, it has been reported that miR-200b-3p can be related to the progression of hepatic fibrosis 21 and modulate cardiac fibrosis in rat. 22 Thus, we conclude that miR-200b-3p may be involved in the progression of MODS patients.
miRNAs used as biomarkers for organ damage have been reported.23,24 Notably, miR-219-5p 7 and miR-27a 6 can be used as biomarkers for the diagnosis and survival prognosis of MODS patients induced by paraquat. In this study, we further investigated the clinical value of miR-200b-3p in patients with MODS caused by paraquat poisoning. ROC analysis results indicated the high diagnostic value of serum miR-200b-3p in discriminating between MODS patients and healthy controls. Serum miR-200b-3p, which was lowly expressed in non-survivors, was related to the 28-day survival of patients. Besides, miR-200b-3p was found to serve as an independent prognostic factor for MODS patients. It is worth noting that miR-200b-3p can be used as a biomarker for other diseases, such as oral squamous cell carcinoma 25 and breast cancer. 26 Therefore, we conclude that serum miR-200b-3p expression may be an effective clinical biomarker for diagnosing MODS and predicting the 28-day survival prognosis of paraquat-induced MODS patients.
We found the direct interaction of miR-200b-3p with HMGB1 by bioinformatics analysis, which was consistent with previous study. 14 Then, serum HMGB1 level was found to be increased in MODS patients caused by paraquat poisoning, and was negatively correlated with the miR-200b-3p. The above results indicated that miR-200b-3p may be involved in the progression of MODS patients by targeting HMGB1. Dysregulation of inflammatory response is well known to play a crucial role in the process of paraquat-induced MODS. 27 Besides, HMGB1, as a nuclear protein present in all cell types, can promote inflammation. 28 For instance, a study by Wang et al. has indicated that nesfatin-1 can relieve acute lung injury by decreasing inflammation through regulating HMGB1, suggesting a mediator role of HMGB1 in inflammation related acute lung injury. 29 Besides, grape seed proanthocyanidin extract (GSPE) can mitigate renal fibrosis through suppressing the complement component 3 (C3)/HMGB1/TGF-β1 pathway. 30 In addition, miR-200b-3p can modulate inflammatory responses in various diseases.15,16 Therefore, miR-200b-3p may play a role in paraquat caused MODS via regulating HMGB1-related inflammatory responses.
The potential targets of miR-200b-3p were predicted using the miRWalk database (left side of Figure 6). Then we co-screened TargetScan, miRDB, and miRTarBase databases, and obtained 10 target genes (right side of Figure 6). By searching the literature on the relationship between these 10 target genes and organ injury, we found that RND3 and DNMT3A were related to liver injury,31,32 RAB21 was related to kidney injury by regulating neutrophil polarization,
33
ETS1 was related to lung, intestinal, and heart injury,34–36 and ETS1 interacted with HMGB1 in heart injury.
36
The prediction of above target genes is of great significance to further explore the mechanism of miR-200b-3p on organ damage after paraquat poisoning in the future, and also provide reference materials for related studies by other researchers. It should be noted that ETS1 is found to be related to multiple organ injury, especially lung cell injury, which is of great significance to explore the most common lung injury caused by paraquat poisoning. The potential targets of miR-200b-3p were predicted by miRWalk database (figure left), and the resulting ten target genes were jointly screened by TargetScan, miRDB and miRTarBase databases (figure right).
However, this study had some limitations. At first, the sample size was small, and a large sample is needed in future studies. Second, although we deduced that miR-200b-3p might regulate inflammatory responses by targeting HMGB1, the precise mechanisms of miR-200b-3p involving in the progression of paraquat poisoning were not deeply investigated in the present study. Further studies are necessary to provide more evidence for the role and mechanisms of miR-200b-3p, and our predicted functional targets of miR-200b-3p might provide some references.
In conclusion, the results of this study indicate that aberrant miR-200b-3p may be a new diagnostic and prognostic biomarker for MODS patients induced by paraquat poisoning. Additionally, miR-200b-3p may be involved in the pathogenesis of MODS by regulating HMGB1-related inflammatory responses. This study will provide a new target for the diagnosis and treatment of paraquat-induced MODS patients.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval and consent to participate
The experimental procedures were all in accordance with the guideline of the Ethics Committee of Qingdao Hospital of Traditional Chinese Medicine (Qingdao Hiser hospital) and has approved by the Ethics Committee of Qingdao Hospital of Traditional Chinese Medicine (Qingdao Hiser hospital). This study complies with the Declaration of Helsinki.
A signed written informed consent was obtained from each patient.
Consent for publication
Written informed consent for publication was obtained from each participant.
Availability of data and materials
The data used and analyzed can be obtained from the corresponding author under a reasonable request.
