Abstract
Objective
Long non-coding RNA (lncRNA
Methods
The expression of HOXC-AS3 and TGF-β2 were detected by RT-qPCR. Overexpression assays were performed to explore the interaction between HOXC-AS3 and TGF-β2. A follow-up study was performed to explore the prognostic value of HOXC-AS3 for CRC. The direct interaction between HOXC-AS3 and miR-1269 was assessed with RNA-RNA pulldown assay. Transwell assays were performed to determine the role of HOXC-AS3 and TGF-β2 in regulating CRC cell invasion and migration.
Results
HOXC-AS3 was significantly downregulated in CRC tissues, while TGF-β2 was significantly upregulated in CRC tissues compared to that in adjacent non-cancer tissues of CRC patients. The follow-up study showed that low expression levels of HOXC-AS3 in CRC tissues were closely correlated with poor survival. Correlation analysis showed that HOXC-AS3 and TGF-β2 were inversely correlated across CRC tissues but not non-cancer tissues. Overexpression of HOXC-AS3 in the two cell lines resulted in downregulation of TGF-β2, while the expression of HOXC-AS3 was not affected by TGF-β2. Transwell migration and invasion assay showed that overexpression of TGF-β2 increased cell invasion and migration, while overexpression of HOXC-AS3 decreased cell migration and invasion. In addition, overexpression of HOXC-AS3 attenuated the effects of overexpression of TGF-β2. MiR-1269 increased the expression of TGF-β2. HOXC-AS3 directly interacted with miR-1269 in CRC cells.
Conclusions
Upregulation of HOXC-AS3 inhibited TGF-β2-induced colorectal cancer (CRC) cell migration and invasion possibly by sponging miR-1269.
Introduction
In both males and females, incidence of colorectal cancer (CRC) ranks the third place among all types of malignancy. 1 Previous studies have characterized a series of risk factors for CRC, such as red meat consumption and smoking. 2 With the historical changes in those factors, incidence of CRC has been decreasing over the past several decades. 2 However, CRCs are prone to tumor metastasis, and traditional therapies are limited due to the development of resistance.3,4 Therefore, treatment outcomes of CRCs are generally poor. At present, molecular mechanisms of the pathogenesis of CRC remain largely unknown,5, 6 which limits the development of novel therapeutic approaches for CRC.
The TGF-β signaling, as a key regulator of cell behaviors, has critical roles in cancer biology.7,8 During the initiation of cancer, TGF-β inhibits cancer cell proliferation to play a tumor suppressive role.7,8 With the development of cancer, TGF-β induces epithelial-mesenchymal transition to promote tumor metastasis.8,9 It has been well established that TGF-β can exert its roles by interacting with long (>200 nt) non-coding RNAs (lncRNAs), 10 which do not encode proteins but regulate gene expression to promote or inhibit cancer development. 11 HOXC-AS3 has been characterized as a key player in several types of cancer, such as lung cancer and ovarian cancer.12,13 In ovarian cancer, HOXC-AS3 regulates miR-96, 13 which can regulate the production of TGF-β. 14 Therefore, HOXC-AS3 may also interact with TGF-β. This study aimed to investigate the interaction between HOXC-AS3 and TGF-β in CRC.
Materials and methods
Patient information and a 5 year follow-up
This study selected 63 CRC patients (gender: 35 males and 28 females, 34–69 years old, mean age 48.3 ± 5.2 years old) from 135 CRC patients admitted to The Third Hospital of Hebei Medical University between January 2012 and January 2014. All patients were followed up for 5 years to record their survival conditions. Inclusion criteria: 1) newly diagnosed CRC patients; 2) patients were willing to and complete the 5 year follow-up. Exclusion criteria: 1) other clinical disorders were observed; 2) recurrent cases; 3) patients who were lost during the follow-up; 4) patients died of other causes during the follow-up. Based on clinical findings and AJCC staging criteria, the 63 patients included 8, 12, 23 and 20 cases at clinical stage of I-IV, respectively. There were 15 cases at grade 1, 28 cases at grade 2 and 20 cases at grade 3. All patients were educated with the experimental details and all of them signed the informed consent. This study was approved by the Ethics Committee of aforementioned hospital.
Tissue collection
All patients were subjected to biopsy. During biopsy, CRC tissues (primary tumor) and adjacent non-tumor tissues (within 2 cm around tumors) were collected from each patient. All tissues were subjected to histopathological analysis to confirm that correct tissues were collected.
CRC cells and transient transfections
HOXC-AS3 and TGF-β2 expression vectors were constructed using pcDNA3.1 vector (Sangon, Shanghai, China) as the backbone. Mimic of miR-1269 and negative control (NC) miRNA as well as HOXC-AS3 siRNA and NC siRNA were obtained from Invitrogen (Shanghai, China). CR4 and RKO cells were harvested at confluence of 70–80%, followed by transfection of 10 mM HOXC-AS3 or TGF-β2 expression vector, 10 mM pcDNA3.1 vector (NC), 50 mM miR-1269 mimic, 50 mM NC miRNA (NC), 50 Mm HOXC-AS3 siRNA, or 50 mM NC siRNA into 106 cells through Lipofectamine 2000 reagent (Thermo Fisher Scientific)-mediated transient transfections. Control (C) cells were cells without transfections. The interval between transfections and subsequent experiments was 24 h.
RNA extraction and qPCR
Tissues collected from CRC patients were ground in liquid nitrogen, followed by mixing 0.01 g tissue powder or 106 CR4 or RKO cells with 0.5 mL RiboZol (VWR, USA) to extract total RNAs. All RNA samples were digested with DNase I to remove genomic DNA. PrimeScript RT Reagent Kit (Takara, Japan) was used to perform reverse transcriptions to synthesize cDNAs. Using cDNA as template, QuantiFast SYBR Green PCR Kit (Qiagen, Shanghai, China) was used to prepare all qPCR reaction mixtures. The expression levels of HOXC-AS3 and TGF-β2 were determined with 18S rRNA as the endogenous control. All qPCR reactions were repeated 3 times and data were normalized using 2−ΔΔCq method.
RNA-RNA pulldown assay
MEGAscript™ T7 Transcription Kit (Thermo Fisher Scientific) was used to prepare in vitro transcripts of both HOXC-AS3 and a NC RNA. These two RNAs were labeled with Biotin using Pierce™ RNA 3’ End Biotinylation Kit (Thermo Fisher Scientific). These two labeled RNAs were named Bio-NC and Bio- HOXC-AS3, respectively, and were transfected into cells. Cell lysis was performed at 48 h after transfections, and RNA-RNA pulldown was performed using magnetic beads. RNA was purified using RiboZol, followed by RT-qPCR to determine the expression levels of miR-1269.
Western blot
CR4 and RKO cells were harvested at 24 h post-transfections and were mixed with 0.5 mL RIPA solution to extract total proteins. After denaturing in boiled water for 5 min, proteins were separated by molecular weights through electrophoresis using 12% SDS-PAGE gel. After that, proteins were transferred to PVDF membranes and blocking was performed in 5% non-fat milk at room temperature for 2 h. PVDF membranes were then blotted using primary antibodies of rabbit GAPDH (1:800, ab181602, Abcam) and TGF-β2 (1:800, ab36495, Abcam) at 4°C for at least 12 h. Following that, membranes were further incubated with secondary antibody of HRP Goat Anti-Rabbit (IgG) (1:1,100, ab6721, Abcam) at room temperature for 2 h. Signals were developed using ECL (Sangon, Shanghai, China). Gray values were measured and processed using Image J 1.48 software.
Cell migration and invasion analysis
CR4 and RKO cells were counted, and 5 × 104 cells were mixed with 1 mL Eagle’s Minimum Essential Medium (serum-free) to prepare single cell suspensions. Transwell upper chamber was filled with 0.1 mL cell suspension and Transwell lower chamber was filled with Eagle’s Minimum Essential Medium (20% FBS). To mimic in vivo cell invasion, Transwell upper chamber was coated with Matrigel (356234, Millipore, USA) at 37°C for 8 h. Invasion and migration were allowed for 12 h under conditions of 37°C and 5% CO2. The membranes were then cleaned using a cotton swab and stained with 1% crystal violet (Sangon) at room temperature for 20 min. Invasion and migration cells of five randomly selected visual fields were counted under a light microscope. The value of C group was set to “100”. All other groups were normalized to C group.
Statistical analysis
Mean values of three biological replicates of each experiment were calculated and used for all data analyses. Differences between two types of tissue (CRC vs non-tumor) or among multiple cell transfection groups were analyzed by performing paired t test and one-way ANOVA followed by Tukey test, respectively. Correlations were analyzed by linear regression. For survival analysis, 63 CRC patients were divided into low (
Results
The expression of HOXC-AS3 and TGF-β2 were altered in CRC
Expression levels of HOXC-AS3 and TGF-β2 in CRC and non-tumor tissues were measured by performing qPCR. Expression data were compared between two types of tissue by paired t test. It showed that HOXC-AS3 was significantly downregulated in CRC tissues (Figure 1(a)), while TGF-β2 was significantly upregulated (Figure 1(b)) in CRC tissues compared to that in adjacent non-tumor tissues of CRC patients ( The expression of HOXC-AS3 and TGF-β2 were altered in CRC. Expression levels of HOXC-AS3 (a) and TGF-β2 (b) in non-tumor and CRC tissues were measured by performing qPCR. Expression data were compared between two types of tissue by performing paired t test. HOXC-AS3 (c) and TGF-β2 mRNA (d) expression in both CRC cell lines of CR4 and RKO and normal HCnEpC was also determined by qPCR. qPCR reactions were performed in triplicate manner and data were presented as mean values, (*, 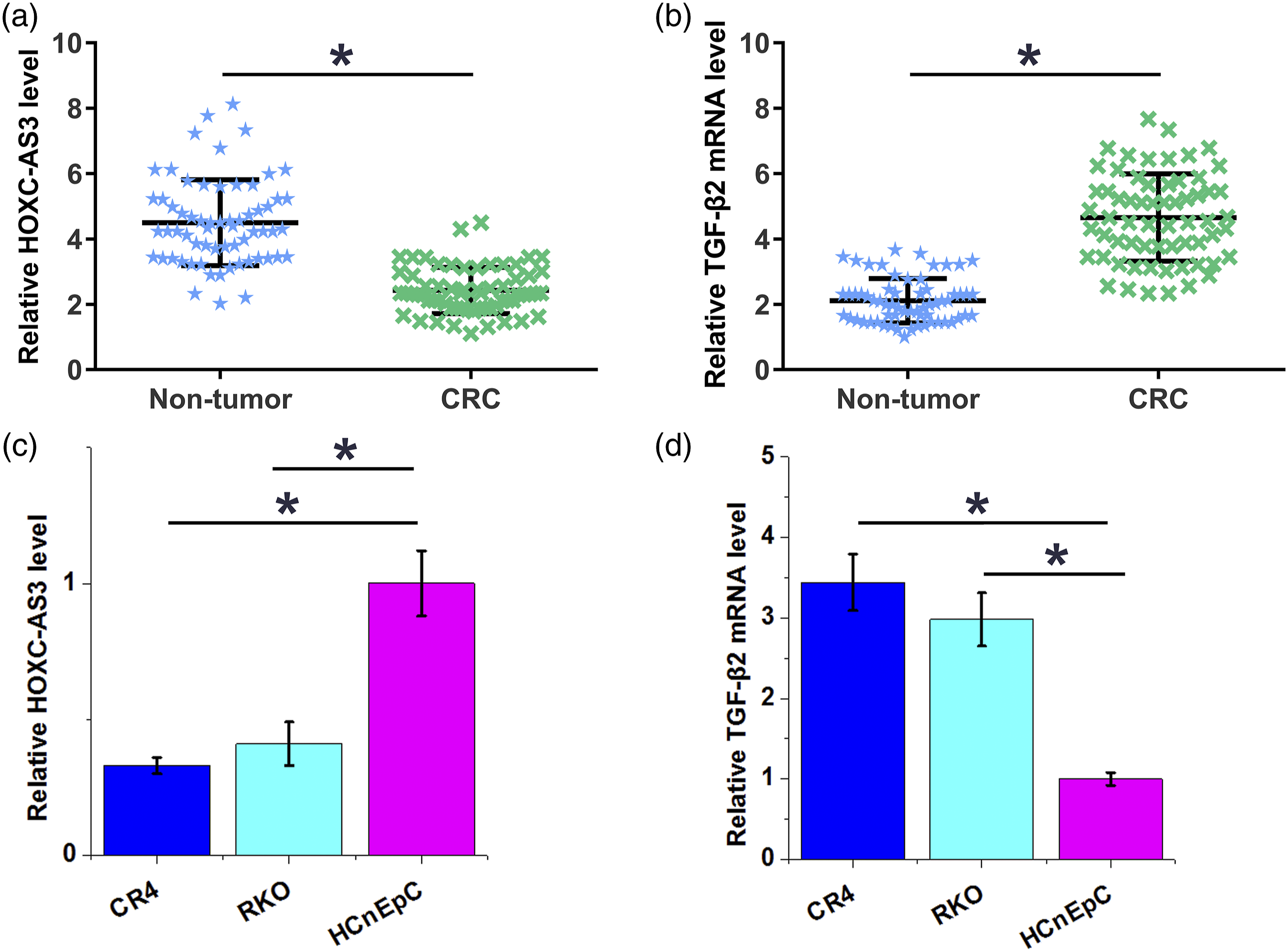
Low expression levels of HOXC-AS3 predicted poor survival
Chi-squared test revealed no obvious association between the expression levels of HOXC-AS3 in CRC tissues and patients’ age, gender, cancer stage and cancer grade (all, Low expression levels of HOXC-AS3 predicted poor survival. Survival curves of low (
The expression of HOXC-AS3 and TGF-β2 were inversely correlated in CRC tissues
Correlations between the expression of HOXC-AS3 and TGF-β2 were analyzed by performing linear regression. It showed that the expression of HOXC-AS3 was inversely and significantly correlated with the expression of TGF-β2 in CRC tissues (Figure 3(a)). In non-tumor tissues, the expression of HOXC-AS3 and TGF-β2 were not correlated (Figure 3(b)). An inverse correlation was observed between the expression of HOXC-AS3 and TGF-β2 in CRC tissue. The correlations between HOXC-AS3 and TGF-β2 in CRC tissues (a) and non-tumor tissues (b) were analyzed by performing linear regression.
HOXC-AS3 downregulated TGF-β2 in CRC cells
HOXC-AS3 and TGF-β2 expression vectors were transfected into CR4 and RKO cells. Compared to NC and C two controls, the expression levels of HOXC-AS3 and TGF-β2 were significantly increased at 24 h post-transfections (Figure 4(a), HOXC-AS3 downregulated TGF-β2 in CRC cells. HOXC-AS3 and TGF-β2 expression vectors were transfected into CR4 and RKO cells. Overexpression of HOXC-AS3 and TGF-β2 was confirmed by performing qPCR (a). The effects of overexpression of HOXC-AS3 on the expression of TGF-β2 was analyzed by performing qPCR and western blot (B). The effects of TGF-β2 on the expression of HOXC-AS3 were analyzed by performing qPCR. All experiments were repeated 3 times and data were presented as mean values.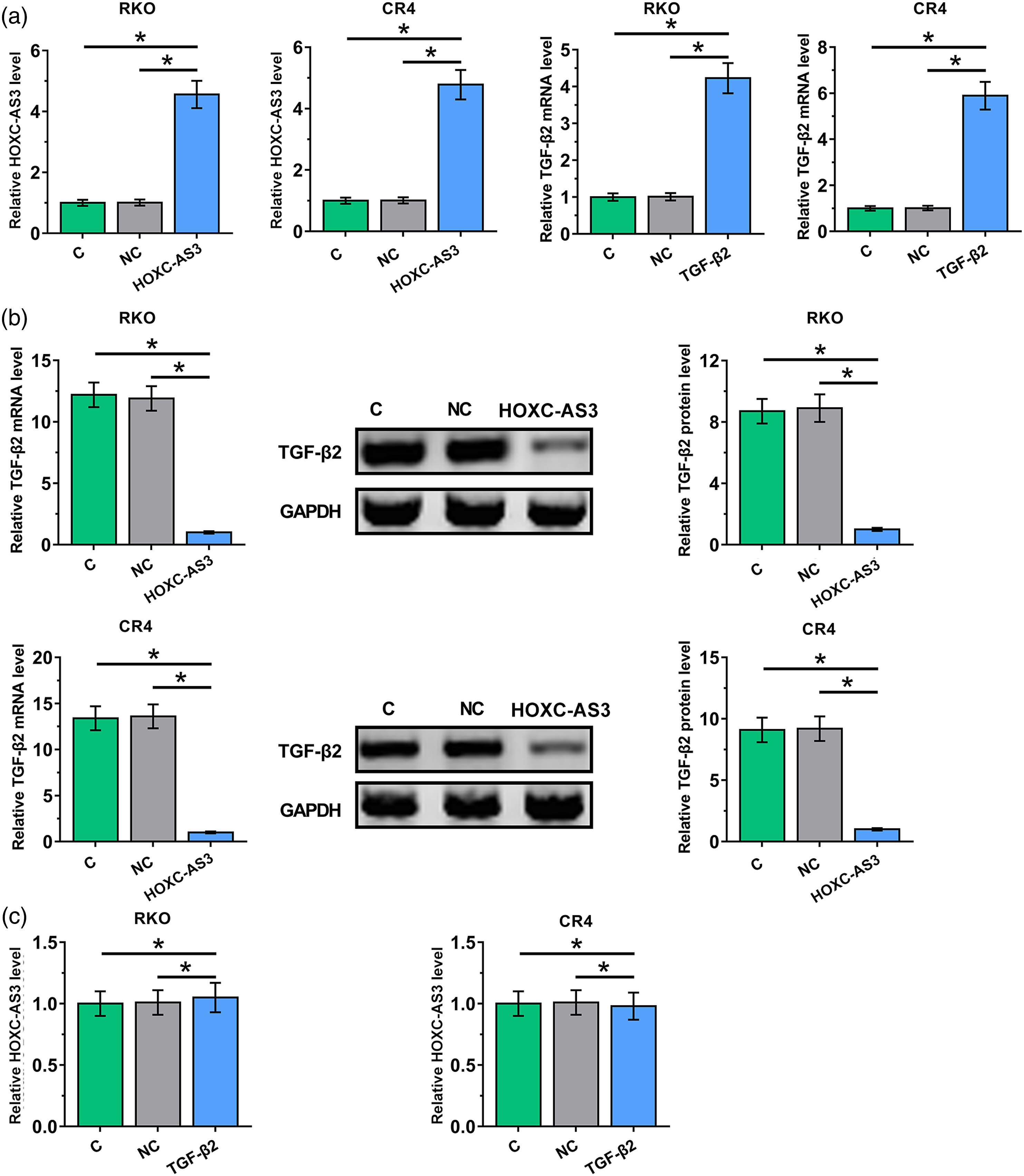
HOXC-AS3 suppressed CRC cell invasion and migration through TGF-β2
Compared to NC and C groups, Transwell migration and invasion assay showed that overexpression of TGF-β2 resulted in promoted, while overexpression of HOXC-AS3 in the two cell lines resulted in inhibited invasion (Figure 5(a)) and migration (Figure 5(b)) of CRC cells. In addition, overexpression of HOXC-AS3 attenuated the effects of overexpression of TGF-β2 ( HOXC-AS3 suppressed CRC cell invasion and migration through TGF-β2. The effects of HOXC-AS3 and TGF-β2 overexpression on CRC cell invasion (a) and migration (b) were analyzed by performing Transwell invasion and migration assays. All experiments were repeated 3 times and mean values were presented (*, 
HOXC-AS3 sponges miR-1269 to downregulate TGF-β2 in CR4 and RKO cells
miR-1269 was reported to upregulate TGF-βs by forming a positive regulatory loop with them.
15
Therefore, miR-1269 was overexpressed in both CR4 and RKO cells to explore the role of miR-1269 in the regulation of TGF-β2. It was observed that overexpression of miR-1269 increased the expression levels of TGF-β2 in both cell lines (Figure 6(a), HOXC-AS3 sponges miR-1269 downregulate TGF-β2 in CR4 and RKO cells. MiR-1269 was overexpressed in both CR4 and RKO cells, in which the expression of TGF-β2 mRNA was detected by performing RT-qPCR (A). IntaRNA 2.0 was applied to predict the interaction between HOXC-AS3 and miR-1269 (B). The direct interaction between them was confirmed by RNA-RNA pulldown assay (*, 
Discussion
In this study, we mainly investigated the role of HOXC-AS3 in CRC. We observed that HOXC-AS3 was downregulated in CRC and regulated CRC cell invasion and migration. The roles of HOXC-AS3 in this process are likely mediated through the interactions with TGF-β2.
The involvement of HOXC-AS3 has been reported in several types of cancer.12,17 It was reported that overexpression of HOXC-AS3 promotes lung cancer by modulating the FUS/FOXM1 axis, 12 and upregulation of HOXC-AS3 sponges miR-3922-5p to promote tumor metastasis in breast cancer. 17 These studies showed the potential oncogenic role of HOXC-AS3 in cancer biology. However, HOXC-AS3 is downregulated in gastric cancer, and it regulates the miR-15b-5p/E2F3 axis to suppress cancer cell proliferation and metastasis. 18 These studies suggested the different functions of HOXC-AS3 in different types of cancer. In this study, we observed downregulation of HOXC-AS3 in CRC and inhibited invasion and migration of CRC cells after overexpression of HOXC-AS3. Our data suggested the tumor suppressive role of HOXC-AS3 in CRC. In addition, we showed that low expression levels of HOXC-AS3 were closely correlated with poor survival of CRC patients, indicating that measurement of the expression levels of HOXC-AS3 before therapies may assist the prognosis assignment of CRC patients. The upregulation of HOXC-AS3 was recently reported in ovarian cancer and glioblastoma.13,19 A recent study also reported that HOXC-AS3 was slightly upregulated in CRC. 20 The opposite observations are likely due to the use of different cell lines and patients involved in the present studies.
In this study, we showed that HOXC-AS3 can inhibit the expression of TGF-β2 to suppress the invasion and migration of CRC cells. It is known that miR-1269 can form a positive regulatory feedback with TGF-β, 15 which was shown to directly interact with HOXC-AS3 in this study. Therefore, miR-1269 may mediate the interaction between HOXC-AS3 and TGF-βs. It is known that lncRNAs may serve as the endogenous competing miRNAs to regulate the expression of downstream targets. 21 We therefore speculated that HOXC-AS3 could sponge miR-1269 to upregulate TGF-β2.
Conclusion
In conclusion, HOXC-AS3 is downregulated in CRC and inhibits TGF-β2-induced CRC cell migration and invasion by sponging miR-1269.
Supplemental Material
Supplemental Material - LncRNA HOXC-AS3 overexpression inhibits TGF-β2-induced colorectal cancer cell migration and invasion by sponging miR-1269
Supplemental Material for LncRNA HOXC-AS3 overexpression inhibits TGF-β2-induced colorectal cancer cell migration and invasion by sponging miR-1269 by Tong-tong Zhang, Hai-peng Chen, Su-yang Yu, and Shi-peng Zhao in Human & Experimental Toxicology
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Natural Science Foundation of Liaoning Province, China grant number (20180550769).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
