Abstract
Retinoblastoma (Rb) is the most common intraocular malignant tumor in infants. Here, we investigated the function and mechanism of cyclophosphamide (CTX) in the development of Rb. Real-time quantitative polymerase chain reaction (RT-qPCR) results showed that paired box protein 5 (Pax5) expression was down-regulated in Rb tissues and cell lines. Methylation-specific PCR (MSP) results showed that the methylation level of Pax5 was up-regulated in Rb. After treatment with CTX, the Pax5 expression in Rb cell lines was increased significantly. The methylation of Pax5 and the expression of DNA methyltransferases (DNMTs) were down-regulated in the CTX group. Cyclophosphamide inhibited cell proliferation, migration, and invasion, promoted cell apoptosis via the Notch1 pathway. DNA methyltransferase inhibitor SGI-1027 had synergistic effects with CTX. Paired box protein 5 siRNA was transfected into Y79 cells treated with CTX. The expression of DNMTs, Pax5, the Notch1 pathway and apoptosis marker protein was detected by Western blotting, and changes in cell behavior were detected, respectively. Results showed that knockdown of Pax5 reversed the effects of CTX. Moreover, the Notch1 activator Valproic acid (VPA) abolished the inhibitory effects of CTX on Rb development. Moreover, CTX inhibited tumor growth in nude mice. These findings demonstrated that CTX up-regulated Pax5 expression by down-regulating DNMTs expression, and then inhibited the Notch1 signaling pathway activation and Rb growth.
Introduction
Retinoblastoma (Rb) is a malignant tumor derived from photoreceptor precursor cells. It is common in children under 3 years old and has a family genetic predisposition. It is the most common intraocular malignant tumor in infants and young children, but rarely seen in adults. The clinical manifestations of Rb are complex, including conjunctival hyperemia, corneal edema, iris neovascularization, vitreous opacity, elevated intraocular pressure and strabismus.1,2 This disease is prone to intracranial and distant metastasis, and often endangers the life of children. Therefore, early detection, diagnosis and treatment are the key to improve the cure rate and reduce the mortality rate. 3 Retinoblastoma protein is an inhibitor of E2F, which can inhibit cell cycle progression through G1/S restriction point, and control the growth and differentiation of cells. 4 As the first cloned tumor suppressor gene, RB gene has been found to be abnormally expressed in a variety of tumors, including hematological tumors. 5 Human retinoblastoma susceptibility gene (RB1) was located on chromosome 13q14. Rb is caused by the biallelic mutation and inactivation of RB1. The expression of RB protein is decreased in Rb tissues, and low expression of RB1 promotes the occurrence of Rb.6,7
Paired box protein 5 (Pax5) is a member of Pax gene family. The Pax gene family consists of nine genes, encodes a group of Pax transcription factors that play an important role in cell development and disease formation. 8 It has been reported that Pax5 promoter methylation existed in a variety of tumors, and the decrease of Pax5 protein expression caused by methylation could significantly promote the occurrence and development of tumors.9,10 In recent years, it has been found that the loss of tumor suppressor genes caused by abnormal DNA methylation played an important role in the occurrence and development of Rb. 11 Previous studies have shown that Pax5 expression is down-regulated due to the promoter hypermethylation, which promotes the development of Rb. 12 Demethylation drugs can be used to inhibit DNA methyltransferases (DNMTs), reverse DNA methylation, and induce tumor cells to differentiate into normal cells. They have clinical anti-tumor activity and have been used in the treatment of many kinds of tumors.
Cyclophosphamide (CTX) is used as an antineoplastic agent to treat a variety of tumors, including Rb. 13 In colon cancer, endostatin/cyclophosphamide normalize tumor micrangium through inhibiting the Notch signaling pathway. 14 The Notch family is composed of four highly conserved transmembrane receptors, which participates in embryonic development and many physiological processes of normal cells. Notch 1 is a member of the Notch family and is related to many types of cancer. Notch 1 is closely connected with many signaling pathways that play a therapeutic role in tumorigenesis, and together affect cell apoptosis, proliferation, and chemosensitivity. 15 It was found that Notch1 expression was significantly up-regulated in Rb cells. 16 Many non-coding RNAs affected Rb progress by inhibiting the Notch signaling pathway, for example, miR-34b-5p inhibited proliferation, stemness, migration, and invasion of RB cells through inhibiting the Notch signaling. 17 LincRNA-ROR induced EMT in Rb by acting as a sponge of miR-32-5p to activate the Notch signaling pathway. 18
In this study, we measured the methylation level and expression of Pax5 in Rb tissues and cell lines, the effect of CTX on Pax5 expression, cell proliferation, apoptosis, migration, and invasion. The aim was to explore the role and the mechanism of CTX in Rb.
Materials and methods
Tissue samples collection
There were 20 Rb patients, including nine males and 11 females, aged 4 months to 10 years old. They were diagnosed by orbital computed tomography (CT) scan, magnetic resonance imaging (MRI), eye ultrasound and fundus examinations under general anesthesia. The exclusion criteria were as follows: patients with severe congenital deficiency, including immune deficiency; patients with a family history of mental illness cannot cooperate with follow-up. The specimen of Rb tissue and adjacent retinal tissue were collected in the pathology department of Xi’an Hospital of Traditional Chinese Medicine. At the same time, the aqueous humor of Rb patients and cataract patients was collected, the DNA was extracted to detect the methylation level of Pax5, and the methylation rate of Pax5 in Rb patients was analyzed. This study was approved by the Ethics Committee of Xi’an Hospital of Traditional Chinese Medicine (number: XHTCM-2020-005). Informed consent was signed by all participants.
Cell culture and treatment
Normal retinal pigment epithelial cells hTERT RPE-1 and Rb cell lines (HXO-RB44, SO-RB50, WERI-Rb1, Y79) were maintained in RPMI 1640 medium containing with 10% fetal bovine serum (FBS). The cells were cultured in a humid incubator with 5% CO2 at 37°C. Cyclophosphamide group was treated with 200 μΜ (CTX, Hengrui Pharmaceutical Co., Ltd, Lianyungang, Jiangsu, China) for 24 h, and the same volume of normal saline (NS) was used as the control. Then transfected the Pax5 siRNA into cells by using Lipofectamine 2000 (Invitrogen, Carlsbad, California, USA). After 6 h, the medium was changed with a fresh culture medium containing 10% FBS. Then the cells were incubated in the incubator for follow-up experiments. Real-time quantitative polymerase chain reaction (RT-qPCR), cell proliferation assay, cell apoptosis assay and Transwell assay were performed as described previously. 19
Cell proliferation assay
After 24 h of treatment, the cells were re-suspended and counted, then seeded in a 96-well plate with 3 × 103 cells/well, and each sample set three replicate wells. The cells were cultured in the incubator, 10 μL of CCK-8 solution was added into the sample wells at 24, 48 and 72 h. Then the cells were incubated in the dark for 2 h to make the CCK-8 reagent fully react with living cells. The absorbance of the samples at 450 nm was measured by multifunctional microplate reader and recorded.
Cell motility assay
1 × 106 cells/mL cell suspension in serum-free medium was inoculated into the upper chambers of the 8-μm pore size Transwell. The culture medium containing 10% FBS was added to the lower chamber used as a chemoattractant. After 24 h of incubation at 37°C, cells in the upper side of the insert were thoroughly wiped off with a cotton swab, cells that migrated to the lower layer of microporous membrane were washed, fixed with 5% glutaraldehyde at 4°C, and stained with 0.1% crystal violet for 10 min. The number of cells were counted under a microscope.
Transwell invasion assay
The cells were digested and suspended in serum-free medium to prepare 1 × 105 cells/mL cell suspension, then added into the upper chamber of Transwell containing Matrigel. The culture medium containing 20% FBS was added to the lower chamber. After cultured at 37°C for 48 h, the cells that invaded to the lower layer of microporous membrane were washed, fixed with 5% glutaraldehyde at 4°C, and stained with 0.1% crystal violet for 10 min. Then the number of cells were counted under a microscope.
Cell apoptosis assay
After 24 h of treatment, the cells were washed twice with pre-cooled PBS, and suspended with 200 μL of binding buffer. 5 μL of Annexin V-FITC reagent was added, and the cells were incubated in the dark for 15 min. Then 10 μL of propidium iodide (PI) solution (20 μg/mL) was added, and the cells were incubated in the dark for 5 min. Cell apoptosis was detected by flow cytometer within 1 h after incubation.
Real-time quantitative polymerase chain reaction
The RNA of tissues or cells were extracted by Trizol according to the instruction. After the concentration was detected by optical density at 260 nm, the first strand cDNA was synthesized by using PrimeScript™ RT Master Mix (Takara, Dalian, Liaoning, China). Real-time quantitative polymerase chain reaction was performed by using the cDNA as a template. The reaction procedure was as follows: 95°C for 30 s; 95°C for 5 s, 60°C for 30 s, 40 cycles. The melt curve was 65°C–95°C, and the temperature was increased 0.5°C every 5 s. The experimental results were calculated by the 2−ΔΔCt method.
Methylation-specific PCR
Total DNA was extracted according to the DNA extraction kit (Trans, Beijing, China). After testing the purity of genomic DNA, Methylation-Gold Kit (ZYMO, Beijing, China) was used to modify the genomic DNA with sodium bisulfite. The specific primers of methylated (M) and unmethylated (U) Pax5 gene were designed by Invitrogen (Waltham, Massachusetts, USA). The fragments were amplified by polymerase chain reaction (PCR), and detected by DNA agarose gel electrophoresis. The primer sequences used in the experiment were as follows: (M) F: 5’-TTT TTT TAA AAG TAT TTG TTT GGT C-3’, (M) R: 5’-GCG ACC CTC TAC GCT ATA CG-3’; (U) F: 5’-TTT TTA AAA GTA TTT GTT TGG TTG A-3’, (U) R: 5’-AAT CAC AAC CCT CTA CAC TAT ACA C-3’.
Quantitative MSP
The target DNA was purified by gel extraction kit, A260 and A280 were measured by an ultraviolet spectrophotometer, and the copy number concentration of the target DNA was calculated. The DNA was multiple diluted and used as a standard for fluorescence quantitative MSP. The reaction system without a template was used as a negative control, and pure water was used as a blank control. The PCR reaction conditions were 95°C for 30 s; 95°C for 5 s, 60°C for 30 s, 40 cycles. The methylation level was expressed by the methylation index (MI), calculated from the absolute copy number, MI = [methylated copy number/(methylated copy number + unmethylated copy number)]/100.
Western blotting
Cells were collected by centrifugation, and washed with pre-cooled phosphate-buffered saline (PBS). Then the cells were lysed with radioimmunoprecipitation assay lysis buffer containing protease inhibitor on ice for 30 min. After high-speed centrifugation, the supernatant protein lysate was collected. The protein concentration was detected by the bicinchoninic acid (BCA) kit (Beyotime, Shanghai, China). Then the same amount of protein was subjected to 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The target protein bands were transferred to polyvinylidene fluoride membranes. The membranes were blocked with 5% skim milk for 2 h at room temperature, then incubated with the corresponding antibody at 4°C overnight. After washing, the membranes were incubated with HRP-labeled IgG antibody at room temperature for 1 h. The enhanced chemiluminescence (ECL) kit was used to visualize the protein bands. The antibodies used in the experiment were as follows: anti-Pax5 (1:3000, ab109443, Abcam, Cambridge, London, UK), anti-DNMT1 (1:1000, ab188453, Abcam, Cambridge, London, UK), anti-DNMT3b (1:2000, ab79822, Abcam, Cambridge, London, UK), anti-Bcl-2 (1:1000, ab182858, Abcam, Cambridge, London, UK), anti-cleaved caspase 3 (1:500, ab49822, Abcam, Cambridge, London, UK), anti-caspase 3 (1:500, ab13847, Abcam, Cambridge, London, UK), anti-Notch1 (1:1000, ab52627, Abcam, Cambridge, London, UK), anti-Hes1 (1:500, ab108937, Abcam, Cambridge, London, UK), anti-Jagged1 (1:500, ab7771, Abcam, Cambridge, London, UK) and goat anti rabbit IgG (1:5000, ab6721, Abcam, Cambridge, London, UK).
Xenograft tumor mouse model
The animal experiments were approved by the Animal Experimental Ethical Committee of Xi’an Hospital of Traditional Chinese Medicine. The right eyeball of Rb patient was excised under general anesthesia. Then the eyeball was cut open immediately under aseptic conditions, the tumor tissue was removed and placed in RPMI-1640 culture medium (containing 20% FBS). After being cut with ophthalmic scissors, the tumor tissue was implanted subcutaneously in 18 male BALB/c nude mice (6 weeks old, weight 20–25 g). The mice were randomly divided into three groups with six in each group. Mice in the CTX group were injected intraperitoneally with 20 mg/kg CTX dissolved in 1 mL NS once every 2 days for 4 weeks. Mice in the NS group were injected intraperitoneally with 1 mL of NS. Mice in the control group were fed normally without any treatment. During 4 weeks of observation, the tumor volume was recorded every week. After euthanasia, the tumor was weighed and the expression of Pax5, apoptosis, and pathway marker proteins were detected by Western blotting.
Statistical analysis
The data were analyzed by SPSS 20.0 statistical software and expressed as mean ± standard error of mean (SEM). The data of two groups were compared by independent sample t test, and the data between multiple groups were compared by one-way ANOVA analysis. The difference was statistically significant in the case of p < 0.05. All experiments were repeated three times independently, and the average value was taken as the experimental results.
Results
Pax5 was down-regulated and the methylation level was up-regulated in Rb tissues and cells
To explore the effects of Pax5 in the progression of Rb, the expression of Pax5 in Rb tissues and cells were detected by RT-qPCR. The results showed that the Pax5 expression in Rb tissues was down-regulated fivefolds than adjacent tissues (Figure 1(a)). The methylation rate of Pax5 gene in aqueous humor of Rb patients was much higher than that of cataract patients (Figure 1(b)). Paired box protein 5 expression was down-regulated about threefolds in Rb cell lines (HXO-RB44, SO-RB50, WERI-Rb1, and Y79) compared to normal retinal pigment epithelial cells hTERT RPE-1 (Figure 1(c)). The methylation level of Pax5 gene in Rb cell lines was much higher than normal retinal pigment epithelial cells (Figure 1(d)). Paired box protein 5 was down-regulated and the methylation rate was up-regulated in Rb tissues and cell lines. (a) The expression of Pax5 in Rb tissues and adjacent normal tissues was detected by RT-qPCR (n = 20). (b) The methylation rate of Pax5 gene in aqueous humor of Rb patients was much higher than that of cataract patients (n = 20). (c) The expression of Pax5 in normal retinal pigment epithelial cells and Rb cell lines was detected by RT-qPCR (n = 3). (d) The methylation level of Pax5 gene in normal retinal pigment epithelial cells and Rb cell lines was detected by MSP (n = 3). Data were represented as mean ± SEM, * p < 0.05; ** p < 0.01. Pax5: paired box protein 5; Rb: retinoblastoma; SEM: standard error of the mean; RT-qPCR: real-time quantitative polymerase chain reaction; MSP: methylation-specific PCR.
CTX up-regulated Pax5 expression by inhibiting DNMTs
To investigate the influences of CTX on Pax5 methylation, after treatment with 200 μΜ CTX for 24 h, the expression of Pax5 was detected by Western blotting, and we found that Pax5 expression in the CTX group was up-regulated threefolds than the control group (Figure 2(a)). In order to further study its mechanism, the Y79 cell line was selected for later experiments. The methylation level of Pax5 was detected by MSP, and the results showed that CTX down-regulated the methylation level of Pax5 (Figure 2(b)). The expression of DNMTs in Y79 cell line was detected by Western blotting. The results showed that DNMT1 was down-regulated fivefolds and DNMT3b expression was down-regulated threefolds in the CTX group than the control group (Figure 2(c)). Cyclophosphamide up-regulated Pax5 expression by inhibiting DNMTs activity and Pax5 promoter methylation. After treatment of cells with 200 μΜ CTX for 24 h, (a) the expression of Pax5 in different cell lines was detected by Western blotting. (b) The methylation level of Pax5 was detected by MSP. (c) The expression of DNMTs was detected by Western blotting. Data were represented as mean ± SEM (n = 3). * p < 0.05. CTX: cyclophosphamide; DNMTs: DNA methyltransferases; Pax5: paired box protein 5; SEM: standard error of the mean; MSP: methylation-specific PCR.
CTX inhibited cell proliferation, migration and invasion, and promoted cell apoptosis via the Notch1 pathway
In order to study the effects of CTX on the behavior of Rb cells, cell proliferation, apoptosis, migration, and invasion were detected by CCK-8, flow cytometry and Transwell assay, respectively. The expression of apoptosis marker proteins, such as Bcl-2 and cleaved caspase-3 was detected by Western blotting. The results showed that CTX inhibited cell proliferation, migration and invasion, and promoted cell apoptosis (Figure 3(a) to (e)). Previous studies have shown that Pax5 repressed the activation of Notch1 pathway to promote B lymphopoiesis and block T cell development.
20
The decitabine inhibited the Notch1 signaling pathway by up-regulating the expression of Pax5, thereby inhibiting the growth of acute T lymphocytic leukemia tumor cells. We further detected the expression of Notch1 and its downstream proteins (Hes1 and Jagged1) by Western blotting and found that CTX inhibited the activation of Notch1 pathway (Figure 3(f)). Cyclophosphamide inhibited cell proliferation, migration and invasion, and promoted cell apoptosis. After treatment of Y79 cells with 200 μΜ CTX for 24 h, the cell proliferation (a), migration (b), invasion (c), and apoptosis (d) were detected by CCK-8, Transwell and flow cytometry assay, respectively. The expression of apoptosis marker proteins, such as Bcl-2 and cleaved caspase-3 (e), the Notch1 pathway and its downstream proteins Hes1 and Jagged1 (f) was detected by Western blotting. Data were represented as mean ± SEM (n = 3). * p < 0.05; ** p < 0.01. CTX: cyclophosphamide; SEM: standard error of the mean; MSP: methylation-specific PCR.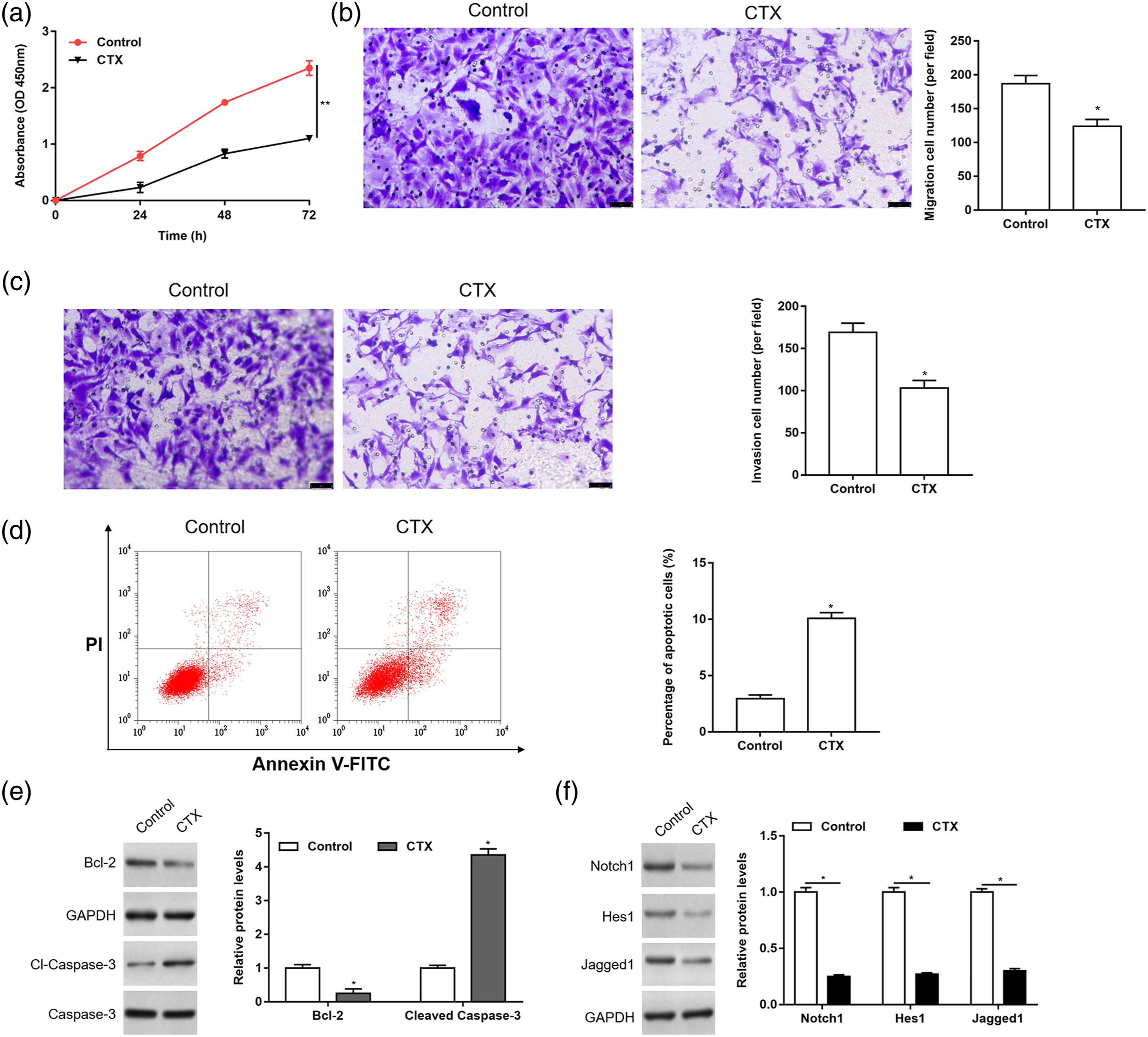
DNMT inhibitor and CTX had synergistic effects on the proliferation, migration, invasion, and apoptosis of Rb cells
In order to further test that CTX inhibited the proliferation, migration, and invasion of Rb cells by affecting the DNA methylation of Pax5 gene, Rb cells were treated with CTX alone or together with DNMT inhibitor SGI-1027 (10 μM), the expression of DNMTs, Pax5 and the Notch1 pathway marker proteins, the methylation level of Pax5 gene, and the changes of cell behavior were further detected. It was found that SGI-1027 down-regulated DNMTs expression (Figure 4(a)) and the methylation level of Pax5 gene (Figure 4(b)), up-regulated Pax5 expression (Figure 4(c)), and inhibited the Notch1 pathway activity (Figure 4(d)). SGI-1027 inhibited cell proliferation (Figure 4(e)), migration (Figure 4(f)) and invasion (Figure 4(g)), and promoted cell apoptosis (Figure 4(h) and (i)). And CTX treatment had a synergistic effect on regulating Pax5 expression (Figure 4(a) to (d)), jointly inhibiting the proliferation, migration, and invasion of Rb cells (Figure 4(e) to (g)), and promoting cell apoptosis (Figure 4(h) and (i)). DNA methyltransferase inhibitor and CTX had synergistic effects on the proliferation, migration, invasion, and apoptosis of Rb cells. Y79 cells were treated with CTX alone or together with DNMT inhibitor SGI-1027 (10 μM), (a) the expression of DNMTs was detected by Western blotting. (b) The methylation level of Pax5 was detected by MSP. The expression of Pax5 (c), Notch1 pathway marker proteins (d), and apoptosis marker proteins (i) was detected by Western blotting. Cell proliferation (e), migration (f), invasion (g), and apoptosis (h) were detected by CCK-8, Transwell, and flow cytometry assay, respectively. Data were represented as mean ± SEM (n = 3). * p < 0.05; ** p < 0.01. Rb: retinoblastoma; CTX: cyclophosphamide; DNMTs: DNA methyltransferases; MSP: methylation-specific PCR; SEM: standard error of the mean.
Pax5 siRNA reversed the effects of CTX on the proliferation, migration, invasion, and apoptosis of Rb cells
After treatment with CTX, Pax5 siRNA was transfected into Rb cells to knock down the expression of Pax5, the expression of DNMTs, Pax5 and the Notch1 pathway marker proteins, and the methylation level of Pax5 gene were detected 24 h later. It was found that DNMTs expression was down-regulated fivefolds, the methylation level of Pax5 gene was down-regulated fourfolds, and Pax5 expression was up-regulated threefolds in the CTX treatment group compared with the control group. Paired box protein 5 siRNA reversed the up-regulation of Pax5 expression by CTX, but did not influence the expression of DNMTs and the methylation level of Pax5 gene (Figure 5(a) to (c)). Cyclophosphamide inhibited the activity of Notch1 pathway, but Pax5 siRNA reversed the inhibitory effects (Figure 5(d)). Further examination of cell behavior revealed that Pax5 siRNA reversed the inhibitory effects of CTX on the proliferation, migration and invasion, and the promotional effects on apoptosis of Rb cells (Figure 5(e) to (i)). Pax5 siRNA reversed the effects of CTX on the proliferation, migration, invasion and apoptosis of Rb cells. After treatment of Y79 cells with CTX for 24 h, Pax5 siRNA was transfected into the cells, (a) the expression of DNMTs was detected by Western blotting. (b) The methylation level of Pax5 was detected by MSP. The expression of Pax5 (c) and Notch1 pathway marker proteins (d) was detected by Western blotting. Cell proliferation (e), migration (f), invasion (g), and apoptosis (h) were detected by CCK-8, Transwell, and flow cytometry assay, respectively. (i) The expression of apoptosis marker proteins was detected by Western blotting. Data were represented as mean ± SEM (n = 3). * p < 0.05; ** p < 0.01. Rb: retinoblastoma; CTX: cyclophosphamide; DNMTs: DNA methyltransferases; MSP: methylation-specific PCR; SEM: standard error of the mean.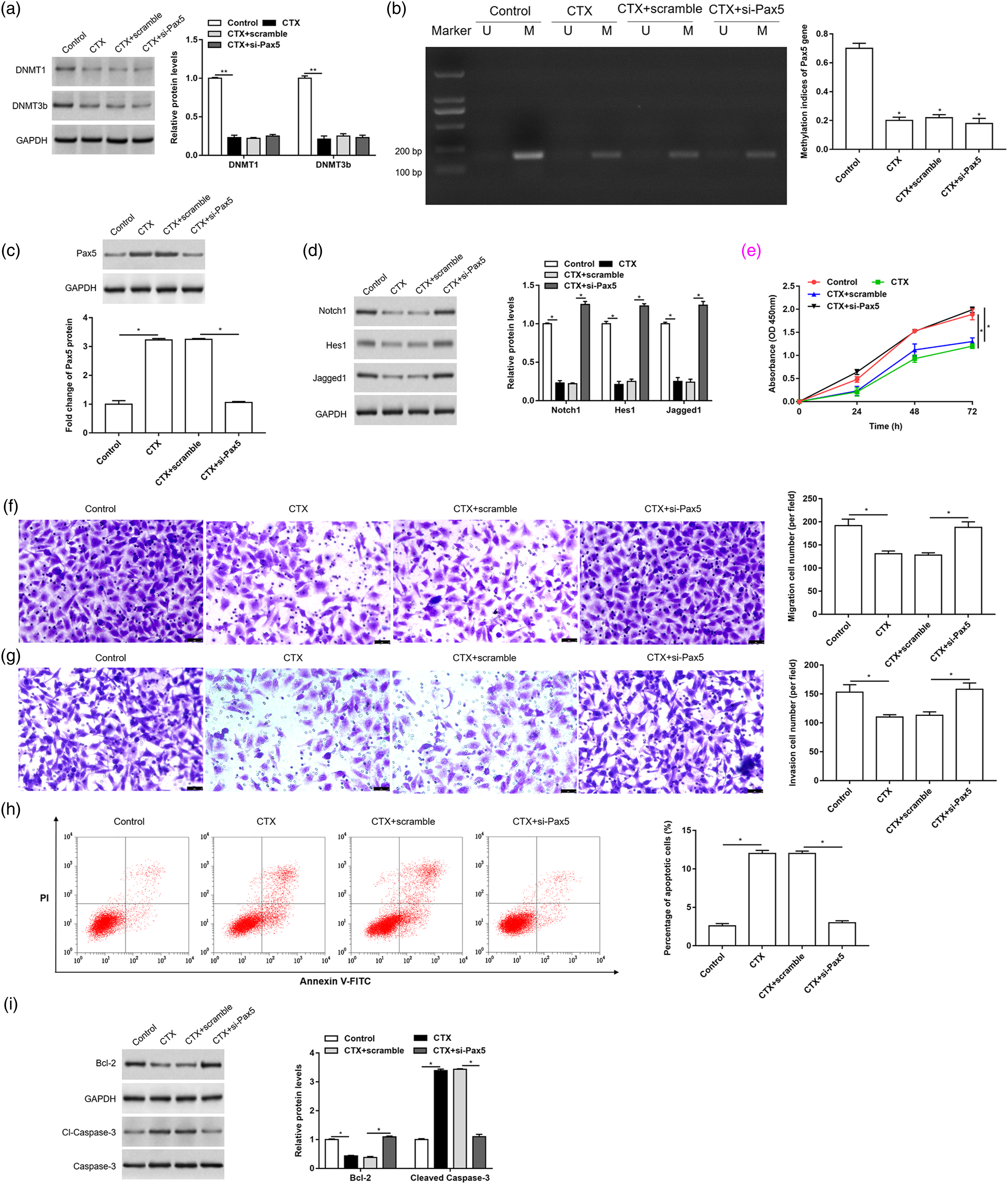
Notch1 activator reversed the effects of CTX on the proliferation, migration, invasion and apoptosis of Rb cells
In order to further test that CTX inhibited tumor growth through the Notch1 pathway, the cells were treated with the Notch1 activator (Valproic acid, VPA) after CTX treatment, and the changes in cell behavior were further detected. It was found that CTX up-regulated the expression of Pax5 and inhibited the activity of Notch1 signaling pathway (Figure 6(a) to (d)). Cyclophosphamide inhibited cell proliferation, migration, invasion, and promoted cell apoptosis. While VPA could eliminate the inhibitory effects of CTX on cell proliferation, migration and invasion (Figure 6(e) to (g)), and the promotional effects on cell apoptosis (Figure 6(h) and (i)). Notch1 activator reversed the effects of CTX on the proliferation, migration, invasion and apoptosis of Rb cells. After treatment of Y79 cells with CTX for 24 h, the cells were treated with Notch1 activator (Valproic acid, VPA). The expression of Pax5 (a and b) and Notch1 pathway marker proteins (c and d) was detected by Western blotting. Cell proliferation (e), migration (f), invasion (g), and apoptosis (h) were detected by CCK-8, Transwell, and flow cytometry assay, respectively. (i) The expression of apoptosis marker proteins was detected by Western blotting. Data were represented as mean ± SEM (n = 3). * p < 0.05. CTX: cyclophosphamide; SEM: standard error of the mean.
CTX inhibited tumor growth in nude mice
The nude mouse subcutaneous tissue xenograft model was established to further verify the function of CTX in vivo. The tumor volume was monitored and recorded every week (Figure 7(a)). The mice were euthanized 4 weeks later, and the tumor weight was measured (Figure 7(b)). The expression of DNMTs, Pax5, the Notch1 pathway marker proteins and apoptosis marker proteins in tumor tissues was detected by Western blotting. The methylation level of Pax5 gene was detected by MSP. It was found that DNMTs expression was down-regulated threefolds to inhibit the methylation of Pax5 in the CTX group (Figure 7(c) and (d)). Paired box protein 5 was up-regulated threefolds and further inhibited tumor growth by inhibiting the activity of Notch1 signaling pathway in vivo (Figure 7(e) to (g)). CTX inhibited tumor growth in nude mice. The nude mouse subcutaneous tissue xenograft model was established and injected intraperitoneally with 20 mg/kg CTX for 4 weeks (a) The tumor volume was monitored every week (n = 6). The mice were anesthetized 4 weeks later, and the tumor weight (b) was measured (n = 6). (c) The expression of DNMTs in tumor tissues was detected by Western blotting. (d) The methylation level of Pax5 was detected by MSP. The expression of Pax5 (e), the Notch1 pathway marker proteins (f), and apoptosis marker proteins (g) in tumor tissues was detected by Western blotting. Data were represented as mean ± SEM (n = 3). * p < 0.05. CTX: cyclophosphamide; DNMTs: DNA methyltransferases; MSP: methylation-specific PCR; SEM: standard error of the mean.
Discussion
At present, there are no effective measures to prevent the occurrence of Rb. For families with a history of Rb, genetic testing and counseling can reduce the birth rate of children patients. The early fundus screening and intervention of newborns are carried out to improve the prognosis. 21 The previous treatments of Rb were eyeball removal and radiotherapy, which have been gradually replaced by chemotherapy in recent years.22,23 CTX has been used in the treatment of many tumors, such as prostate cancer and breast cancer.24,25 Studies have shown that CTX can be used in the treatment of Rb.26,27 Therefore, it is important to explore the role and mechanism of CTX in Rb progress.
Pax5 exhibits different properties of promoting or suppressing cancer in different tumors.28,29 It was found that Pax5 induced the upregulation of FOXP4-AS1/FOXP4 axis to promote the tumorigenesis of prostate cancer. 30 On the other hand, as a tumor suppressor gene, Pax5 played an important role in inhibiting cell proliferation and invasion, inducing cell apoptosis in gastric cancer. Paired box protein 5 methylation was related to the prognosis of gastric cancer patients. 31 The expression of Pax5 in non-small cell lung cancer (NSCLC) was down-regulated due to the hypermethylation in Pax5 promoter region. The demethylation could enhance the Pax5 expression and inhibit the cell invasion and migration of NSCLC. 32 Livide et al. found that the promoter of Pax5 gene was methylation in Rb. 12 Our results showed that the Pax5 expression was down-regulated in Rb tissues and cell lines, and the methylation level was up-regulated. Cyclophosphamide inhibited the methylation of Pax5 and up-regulated the expression of Pax5.
DNA methylation plays an important role in cancer biology, the down-regulation of certain genes due to methylation further affects the occurrence and development of tumors.33–35 Methylation can chemically modify some proteins and nucleic acids to form methylated products. In biological system, methylation can regulate the expression of genes and the function of proteins. The process of DNA methylation is usually completed under the catalysis of DNMTs. DNA methyltransferase include DNMT1, DNMT2, DNMT3a and DNMT3b. As one of the key mechanisms of tumor-related gene inactivation, abnormal methylation in gene promoter affects almost all important biological functions, including DNA damage repair, cell cycle regulation, cell apoptosis and cell adhesion. 36 The results of this study showed that CTX inhibited the methylation of Pax5 by down-regulating the expression of DNMTs. The up-regulated Pax5 further inhibited the proliferation, migration and invasion of Rb cells, and promoted cell apoptosis.
Paired box protein 5 promoted B lymphopoiesis and blocked T cell development by repressing the activation of Notch1 pathway. 20 Decitabine inhibited the Notch1 pathway by up-regulating the expression of Pax5, thereby inhibiting the growth of acute T lymphocytic leukemia tumor cells. The Notch1 pathway plays an important role in tumor growth, 37 Jagged1 is a ligand of Notch, hairy and enhancer of split homolog-1 (Hes1) is a downstream target gene of Notch. Many studies have shown that as an oncogene, Hes1 was not only abnormally expressed in a variety of solid tumors, but also related to the proliferation and metastasis of malignant tumor cells.38-40 The results of further treatment of cells with the Notch1 pathway activator showed that CTX inhibited tumor growth by inhibiting the Notch1 signaling pathway in Rb. Intraperitoneal injection of CTX could inhibit the growth of tissue xenograft tumor in nude mice.
In conclusion, these findings demonstrated that CTX inhibits the methylation of Pax5 gene by inhibiting the expression of DNMTs, and up-regulates the expression of Pax5 gene. Furthermore, Pax5 inhibits the growth of Rb by inhibiting the activity of Notch1 signaling pathway.
Footnotes
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
