Abstract
Background
Bladder cancer (BCa) is a common genitourinary malignancy with higher incidence in males. Long intergenic non-protein coding RNA 265 (LINC00265) is identified as an oncogene in many malignancies, while its role in BCa development remains unknown.
Purpose
To explore the functions and mechanism of LINC00265 in BCa
Research Design
Reverse transcription quantitative polymerase chain reaction was performed to examine LINC00265 expression in BCa cells. Cell counting kit-8 assays, colony formation assays, TdT-mediated dUTP Nick-End Labeling assays, and Transwell assays were conducted to examine BCa cell viability, proliferation, apoptosis, and migration. Luciferase reporter assays and RNA immunoprecipitation assays were carried out to explore the binding capacity between miR-4677-3p and messenger RNA fibroblast growth factor 6 (FGF6) (or LINC00265). Xenograft tumor model was established to explore the role of LINC00265 in vivo.
Results
LINC00265 was highly expressed in BCa cells. LINC00265 knockdown inhibited xenograft tumor growth and BCa cell viability, proliferation and migration while enhancing cell apoptosis. Moreover, LINC00265 interacted with miR-4677-3p to upregulate the expression of FGF6. FGF6 overexpression reversed the suppressive effect of LINC00265 knockdown on malignant phenotypes of BCa cells.
Conclusions
LINC00265 promotes the viability, proliferation, and migration of BCa cells by binding with miR-4677-3p to upregulate FGF6 expression.
Introduction
Bladder cancer (BCa) is one of the top 10 common malignancies worldwide and accounts for approximately 170,000 cases of death annually. 1 Male sex, advanced age, and cigarette smoking are risk factors for BCa development. 1 Visible hematuria is the most common symptom of BCa, followed by microscopic hematuria, irritative voiding, or a tumor discovered on imaging. 2 According to the Global Cancer Observatory (https://gco.iarc.fr/), there were 549,393 new BCa cases worldwide in 2018 and 424,082 of them were males. Moreover, BCa was responsible for 199,922 deaths globally in 2018. 3 Despite the great progress achieved in BCa surgery, chemotherapy, and radiotherapy, the death rate of BCa is still more than 33%. 3 Hence, it is significant to find more biomarkers for the treatment of BCa.
Recently, increasing evidence has revealed that long noncoding RNAs (lncRNAs) can become diagnostic and therapeutic biomarkers for BCa.4,5 Long noncoding RNAs are transcripts with over 200 nucleotides in length, which is an important subclass of noncoding RNAs (ncRNAs). 6 According to the Human Genome Project, 98% human genomes are ncRNAs. 7 Long noncoding RNA plays an essential role in regulating biological processes, especially in oncogenesis. 8 Many lncRNAs, such as lncRNA small nucleolar RNA host gene 3 (SNHG3), 9 lncRNA thymopoietin antisense RNA 1 (TMPO-AS1), 10 and lncRNA DEAD/H-box helicase 11 antisense RNA 1 (DDX11-AS1) 11 were reported to affect BCa development by promoting cell proliferation, migration, or invasion. Long intergenic non-protein coding RNA 265 (LINC00265) is a newly discovered lncRNA that has been reported to play roles in osteosarcoma, colorectal, lung, and gastric cancer.12–15 For example, LINC00265 targets the epidermal growth factor receptor to induce colorectal cancer cell proliferation and migration. 16 LIN00265 promotes cell invasion and angiogenesis by targeting miR-382-5p to upregulate the expression of spermidine/spermine N1-acetyltransferase 1 and VAV guanine nucleotide exchange factor 3. 12 Since the function and mechanism of LINC00265 in BCa have not been explored yet, we herein focused on the role of LINC00265 in BCa.
Mechanistically, the lncRNA-mediated competing endogenous RNA (ceRNA) network is widely investigated and has been validated in tumor development.17,18 Following the ceRNA mechanism, lncRNAs competitively bind with microRNAs (miRNAs) to prevent the degradation of target genes of miRNAs.6,19,20 These target genes are messenger RNAs (mRNAs) whose 3’-untranslated region (3’-UTR) is directly targeted by miRNAs. Micro RNAs target mRNAs to enhance the degradation or restrict the translation of mRNAs post-transcriptionally. However, mRNA overexpression can reverse the suppressive effect of miRNAs on them due to the regulation of lncRNAs. 20 The lncRNA–miRNA–mRNA network has been extensively explored in BCa. For example, lncRNA SNHG3 acts as a ceRNA to increase the expression of GINS complex subunit 2 by interacting with miR-515-5p, thereby facilitating BCa cell proliferation and metastasis. 9 Long noncoding RNA cancer susceptibility 9 inhibits BCa cell apoptosis by interacting with miR-497-5p to upregulate the expression of chromobox 2 under the ceRNA network. 21 The ceRNA role of LINC00265 in other cancer types has also been reported. For example, LINC00265 facilitates the production of glycolysis and lactate in colorectal cancer by binding with miR-216b-5p as a ceRNA to regulate the expression of tripartite motif-containing 44. 22 LINC00265 serves as a ceRNA to upregulate the expression of chromobox 4 by interacting with miR-144-3p in gastric cancer. 15 Hence, we hypothesized that LINC00265 might serve as a ceRNA to regulate the expression of downstream genes and thus affect BCa development.
In conclusion, this study investigated the function and mechanism of LINC00265 in BCa, which might provide a promising biomarker for BCa prognosis and therapeutic treatment.
Materials and methods
Cell culture and transfection
Human uroepithelial cell line SV-HUC-1 were purchased from Kunming Cell Bank (KCB, Kunming, China), and BCa cell lines (1A6 and 5637) were purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). The above cells were cultured in Dulbecco’s modified eagle medium (DMEM; Thermo Fisher Scientific, Waltham, MA, USA) added with 10% fetal bovine serum (FBS; Hyclone, Logan, UT) and 1% penicillin and streptomycin (Invitrogen, Carlsbad, CA, USA). These cells were incubated in a humidified atmosphere at 37°C with 5% CO2. For cell transfection, short hairpin RNAs (shRNAs) target LINC00265 (sh-LINC00265#1/2) and the negative control (sh-NC), miR-4677-3p mimics and NC mimics, pcDNA3.1/FGF6 and control pcDNA3.1 were respectively transfected into 1A6 and 5637 cells with Lipofectamine 2000 (Invitrogen). All plasmids and vectors were purchased from Beyotime (Shanghai, China). The transfection efficiency was examined by RT-qPCR after 48 h.
Cell-counting kit-8 (CCK-8) assay
After indicated transfection, the cells were plated in 96-well plates for 24 h, 48 h, and 72 h, respectively. During each interval, the cell-counting kit-8 kit (Dojindo, Kumamoto, Japan) was used to assess cell viability according to the manufacturer’s protocol. After added with colorant, cells were cultured for another 2 h, and then the absorbing result was examined by a microplate reader (Reagen, Shenzhen, China) at a wavelength of 450 nm. All experiments were conducted three times.
Colony formation assay
The assay was conducted to examine the proliferative capability of BCa cells. After transfection, 1A6 and 5637 cells were incubated in 6-well plates at a density of 4 × 103 cells per well for 2 weeks. Next, 4% paraformaldehyde fix solution was added to these plates to fix colonies, and 0.1% crystal violet (Beyotime) was added for staining. Colonies were then counted manually through a microscope (Olympus). All experiments were repeated in triplicate.
TdT-mediated dUTP nick-end labeling (TUNEL) assay
TdT-mediated dUTP nick-end labeling assays were performed to assess BCa cell apoptosis. The indicated cells were washed with phosphate-buffered saline three times. Next, the BCa cells were fixed with 4% paraformaldehyde (Sangon Biotechnology, Shanghai, China), and then permeabilized with 0.1% triton X-100 (Sigma-Aldrich, St. Louis, MO, USA) and 0.1 mol/l sodium citrate buffer (Caiyou Bio Technologies, Shanghai, China). Afterward, the In Situ Cell Death Detection kit (Vazyme) was utilized to mark apoptotic cells, and the nucleus were stained with 4’,6-diamidino-2-phenylindole (DAPI, CWBIO). Digital images were captured with a microscope (Olympus, Tokyo, Japan). All experiments were conducted in thrice.
Western blot analysis
Total protein samples were extracted by a radio-immunoprecipitation assay (RIPA) lysis buffer (Beyotime). Next, equal amounts of protein samples (100 µg) were fractionated by SDS-PAGE and then transferred to PVDF membranes (Millipore, Bedford, MA, USA). After blocked with 5% skim milk dissolved with PBS for 2 h at room temperature, the membranes were incubated with primary antibodies (Abcam, Cambridge, UK) of anti-FGF6 (ab89549; 1:1000), anti-matrix metallopeptidase 2 (MMP2) (ab92536; 1:1000), anti-matrix metallopeptidase 9 (MMP9) (ab76003; 1:1000), anti-PCNA (ab92552; 1:1000), anti-Cyclin D1 (ab16663; 1:150), anti-Bcl-2 (ab182858; 1:2000), anti-Bax (ab32503; 1:1000), anti-cleaved caspase 3 (ab32042; 1:500), anti-cleaved caspase 9 (ab2324; 1:1000), and anti-glyceraldehyde-3-phosphate dehydrogenase (GAPDH; ab8245; 1:2000) at 4°C for 12 h. GAPDH was set as a loading control. Afterward, the prepared membranes were washed with Tris Buffered saline Tween (TBST) and incubated with secondary antibodies (Abcam) for 2 h at room temperature. Finally, an enhanced chemiluminescence kit (Pierce, Waltham, MA, USA) was used to access the protein signals. The band intensity in each group was measured by Odyssey v1.2 software (LI-COR Biosciences, Lincoln, NE, USA) and normalized to GAPDH.
Transwell assay
A transwell assay was applied to detect BCa cell migration. The cells after transfection were resuspended in DMEM and seeded to the upper chamber, while DMEM containing 10% FBS was added to the lower chamber. After 48 h, a cotton swab was used to clear the non-migrated cells, while the migrated cells were fixed with methanol and stained with crystal violet. Five fields were randomly selected, and cells under these fields were counted under a microscope (Olympus). All experiments were repeated three times.
RNA immunoprecipitation (RIP) assay
For the RIP assay, Magna RIP RNA-binding protein immunoprecipitation kits (Millipore, Billerica, MA, USA) were used following the manufacturer’s protocol. Briefly, RIPA lysis buffer was added with 50-μL magnetic beads. Afterward, the cell lysates were incubated with the Ago2 antibody (CST, Danvers, MA, USA) or rabbit IgG and protein A/G magnetic beads at 4°C for 12 h while being rotated. After washing, the immunoprecipitated RNAs were purified by an RNeasy MinElute Cleanup Kit (Qiagen, Dusseldorf, Germany). The enrichment of LINC00265, miR-4677-3p, and FGF6 was evaluated by RT-qPCR.
Reverse transcription quantitative polymerase chain reaction (RT-qPCR) analysis
Total RNA was extracted using TRIzol Reagent (Invitrogen). Next, the Reverse Transcription Kit (Takara, Carlsbad, CA, USA) was applied to reverse-transcribe RNA into cDNA. Subsequently, RT-qPCR was conducted utilizing the SYBR Green real-time PCR kit (Takara, Carlsbad, CA, USA) on a CFX96 system (Bio-Rad, Hercules, California, USA). The RNA expression was calculated using the 2-ΔΔCt method. GAPDH was regarded as an internal control for LINC00265 and FGF6, while U6 was set as an endogenous control for miRNAs. The primer sequences used in RT-qPCR are as follows:
LINC00265:
Forward: 5’-AGTGCTATCACAGCTCACTG-3’
Reverse: 5’-AGAGGGAGACCCTACAGTC-3’
miR-4677-3p:
Forward: 5’-TCTGTGAGACCAAAGAACTACTGC-3’
Reverse: 5’-CTCTACAGCTATATTGCCAGCCAC-3’
FGF6:
Forward: 5’-GAAAGTGGCTATTTGGTGGG-3’
Reverse: 5’-ATTTCCAGCAGGCTGTAGG-3’
GAPDH:
Forward: 5’-CCTCCTGTTCGACAGTCAG-3’
Reverse: 5’-CATACGACTGCAAAGACCC-3’
U6:
Forward: 5’-CTTTGGCAGCACATATACCA-3’
Reverse: 5’-CTCATTCAGAGGCCATGCT-3’
Luciferase activity reporter assay
The wild-type (Wt) sequence of the LINC00265-containing binding site with miR-4677-3p was subcloned into pmirGLO vectors (Promega, Madison, WI, USA) to construct pmirGLO-LINC00265-Wt. A site-direct mutagenesis kit (Takara, Carlsbad, CA, USA) was used to mutate miR-4677-3p binding sites to construct mutant luciferase vectors of pmirGLO-LINC00265-Mut. Next, the above vectors were respectively co-transfected with miR-4677-3p mimics or NC mimics into 5637 and 1A6 cells utilizing Lipofectamine 2000 (Invitrogen, Shanghai, China). After 48 h of transfection, a dual-luciferase kit (Promega) was used to assess the relative luciferase activity.
Subcellular fractionation assay
A PARIS Kit was applied to fractionate nuclear and cytoplasmic RNAs of 1A6 and 5637 cells. RT-qPCR analysis was performed to probe the levels of LINC00265, GAPDH, and U6 RNA in the cytoplasmic and nuclear fractions. GAPDH was a cytoplasmic control, while U6 was a control for nuclear parts.
Xenograft nude mice model
Twelve male BALB/c nude mice (6-week-old) were purchased from Vital River Laboratory Animal Technology (Beijing, China). These mice were randomly divided into two groups (n = 6/group). After transduction of lentivirus (LV)-sh-NC or LV-sh-LINC00265#1 into 1A6 cells, mice were subcutaneously injected with 0.1 mL of cell suspension containing 3 × 106 1A6 cells. After 3 days, tumor volume was recorded every 3 days according to the following formula: volume = length × width2 × 0.5. On day 15, mice were euthanized and then tumors were weighed.
Statistical analysis
Statistical Product and Service Solutions (SPSS) 23.0 (SPSS Inc, USA) software was used for statistical evaluation. 23 The data are shown as the mean ± standard deviation (SD). Student’s t test and analysis of variance (ANOVA) followed by Tukey’s post hoc test were employed to compare the difference between groups or among multiple groups. The value of p less than 0.05 was considered as statistically significant.
Results
LINC00265 knockdown inhibits BCa cell proliferation and migration and promotes cell apoptosis
First, the expression of LINC00265 in BCa cell lines and uroepithelial cell line was measured by RT-qPCR. We discovered that LINC00265 expression was significantly upregulated in 5637 and 1A6 cell lines compared with that in the SV-HUC-1 cell line (Figure 1(a)). Next, we silenced LINC00265 expression by transfection of sh-LINC00265#1/2 into BCa cells to conduct the following loss-of-function assays. As shown in RT-qPCR, LINC00265 expression was efficiently silenced by sh-LINC00265#1/2 (Figure 1(b)). The CCK-8 assay illustrated that LINC00265 knockdown significantly decreased the viability of 1A6 and 5637 cells compared with that of SV-HUC-1 cells (Figures 1(c) and (d)). Furthermore, silencing LINC00265 decreased the number of cell colonies according to the colony formation assay, suggesting that LINC00265 depletion suppressed BCa cell proliferation (Figure 1(e)). As was shown in the TUNEL assay, the apoptosis of BCa cells was enhanced due to LINC00265 deficiency (Figure 1(f)). Protein levels of proliferation markers (PCNA and cyclin D1) were decreased due to LINC00265 depletion according to the result of Western blot analysis (Figure 1(g)). Additionally, protein levels of Bax, cleaved caspase 3, and cleaved caspase 9 were upregulated, while the Bcl-2 level was reduced by silencing LINC00265 (Figure 1(h)). Moreover, Western blot analysis was performed to measure the effect of LINC00265 knockdown on protein levels of migration-related factors (MMP2 and MMP9) in BCa cells, implying that LINC00265 knockdown decreased the protein levels of MMP2 and MMP9 in 1A6 and 5637 cells (Figure 1(i)). Additionally, the transwell assay suggested that the number of migrated cells was decreased by LINC00265 deficiency (Figure 1(j)). Overall, the above results indicated that LINC00265 knockdown promotes BCa cell apoptosis and inhibits cell viability, proliferation, and migration. LINC00265 knockdown inhibits cell viability, proliferation, and migration and promotes cell apoptosis in BCa. (A) LINC00265 expression was upregulated in BCa cell lines (1A6 and 5637) compared with that in the uroepithelial cell line (SV-HUC-1), as shown in RT-qPCR analysis. (B) LINC00265 expression was successfully knocked down by transfection of sh-lINC00265#1/2 into BCa cells, according to RT-qPCR. (C)–(D) The CCK-8 assay was performed to measure the effect of LINC00265 knockdown on BCa cell viability. (e) The proliferation of BCa cells transfected with sh-LINC00265#1/2 was assessed by the colony formation assay. (F) The apoptosis of BCa cells with the above transfection was probed by the TUNEL assay. (G)–(I) Western blot analysis was performed to detect protein levels of proliferation markers (PCNA and cyclin D1), apoptosis markers (Bcl-2, Bax, cleaved caspase 3, and cleaved caspase 9) and migration-related factors (MMP2 and MMP9) in BCa cells with the above transfection. (J) The effect of LINC00265 knockdown on BCa cell migration was measured by the transwell assay. **p < 0.01, ***p < 0.001. Note: BCa: bladder cancer; TUNEL: TdT-mediated dUTP nick-end labeling.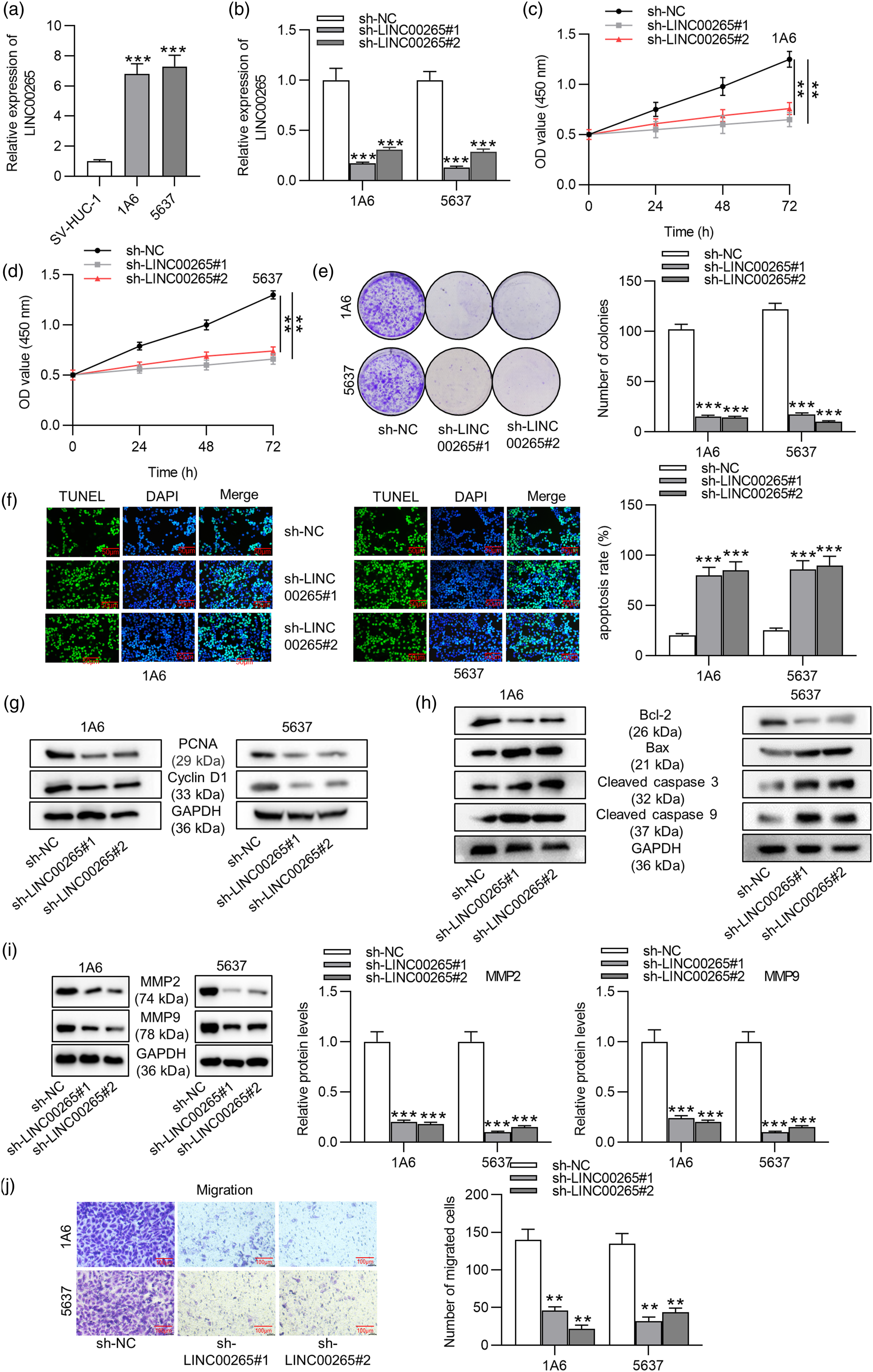
LINC00265 interacts with miR-4677-3p in BCa cells
To verify the hypothesis that LINC00265 might serve as a ceRNA in BCa, we first predicted the location of LINC00265 in cells using the lncLocator database (http://www.csbio.sjtu.edu.cn/bioinf/lncLocator/).
24
The data suggested that LINC00265 is mainly located in cytosol (Figure 2(a)). Next, we conducted a subcellular fractionation assay to determine the main location of LINC00265. The results showed that LINC00265 was primarily located in the cytoplasm of BCa cells, indicating that LINC00265 functions at the post-transcriptional level (Figure 2(b)). Subsequently, we explored the downstream genes of LINC00265 using bioinformatics tools. MicroRNA let-7d-5p, let-7b-5p and miR-4677-3p were predicted by starBase (http://starbase.sysu.edu.cn/) to be the possible downstream miRNAs of LINC00265 under the condition of 10 cancer types.
25
Afterward, RT-qPCR analysis was performed to detect the expression levels of the three miRNAs after transfection with sh-LINC00265#1/2, revealing that let-7d-5p and miR-4677-3p were significantly upregulated due to LINC00265 deficiency, while the expression of let-7b-5p failed to be affected (Figures 2(c) and 2(d)). To further identify the downstream miRNAs of LINC00265 in BCa, the relative expression levels of miR-4677-3p and let-7d-5p in 1A6 and 5637 cells were examined. The results of RT-qPCR analysis exhibited that miR-4677-3p expression was downregulated in BCa cells, while no significant changes in let-7d-5p expression were detected compared with their expression levels in control groups (Figure 2(e)). Therefore, we identified miR-4677-3p for further study. The luciferase reporter assay was conducted to explore the interaction between LINC00265 and miR-4677-3p. The binding site between miR-4677-3p and LINC00265 was predicted from starBase (Figure 2(f)). In addition, miR-4677-3p was overexpressed by miR-4677-3p mimics in BCa cells for the following luciferase reporter assay. The overexpressing efficiency was measured by RT-qPCR analysis (Figure 2(g)). As shown in the luciferase reporter assay, miR-4677-3p mimics decreased the luciferase activity of LINC00265-Wt, while they failed to significantly decrease that of LINC00265-Mut compared with the control groups (Figure 2). LINC00265 interacts with miR-4677-3p in BCa cells. (A) The subcellular location of LINC00265 was predicted by the lncLocator database. (B) The subcellular fractionation assay was conducted to determine the distribution of LINC00265 in BCa cells. (C)–(D) The starBase website was used to predict the downstream miRNAs of LINC00265. The effects of LINC00265 knockdown on expression levels of the three candidate miRNAs in BCa cells were measured by RT-qPCR analysis. (E) RT-qPCR analysis was performed to assess the expression levels of miR-4677-3p and let-7d-5p in BCa cells to further identify the downstream miRNA of LINC00265. (F) starBase was utilized to predict the binding site between miR-4677-3p and LINC00265. (G) RT-qPCR analysis was performed to measure the overexpression efficiency of miR-4677-3p. (H) The binding between LINC00265 and miR-4677-3p was verified by the luciferase reporter assay in BCa cells transfected with NC mimics and miR-4677-3p mimics. **p < 0.01, ***p < 0.001. Note: BCa: bladder cancer; miRNA: microRNAs.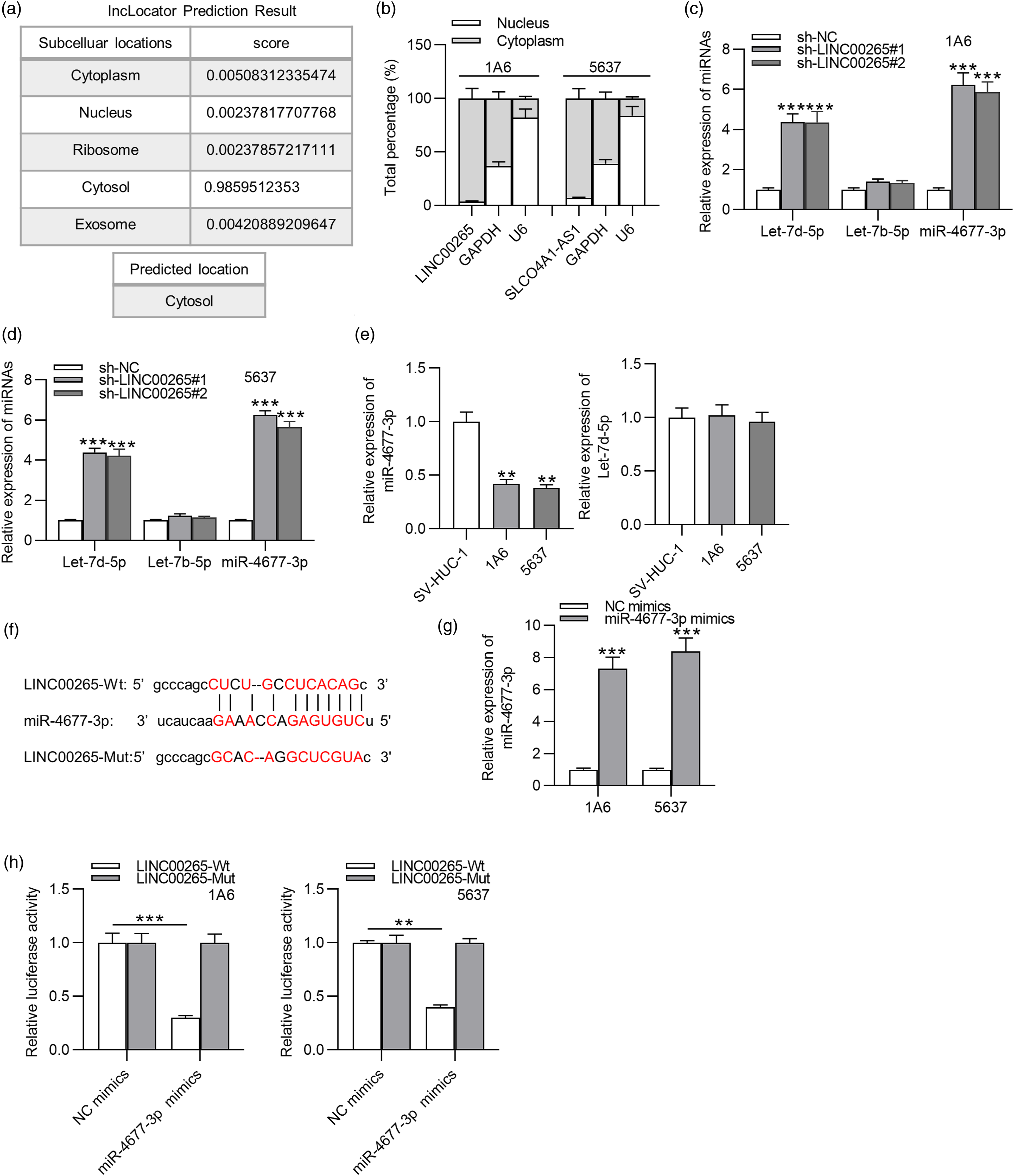
MiR-4677-3p directly targets FGF6 in BCa cells
The bioinformatic tool, TargetScan (http://www.targetscan.org/), was used to identify the target genes of miR-4677-3p.
26
The expression of top 10 potential target genes in BCa cells with or without transfection of sh-LINC00265#1 was examined by RT-qPCR analysis. The results showed that only FGF6 expression was both upregulated in BCa cells and downregulated by LINC00261 knockdown (Figure 3(a)). RT-qPCR and Western blot analyses were performed to assess the effect of miR-4677-3p overexpression on FGF6 levels, implying that miR-4677-3p overexpression decreased FGF6 expression at both mRNA and protein levels (Figures 3(b) and (c)). Additionally, the effect of LINC00265 knockdown on FGF6 expression was also measured. Relative mRNA expression and protein levels of FGF6 were downregulated by silencing LINC00265 (Figure 3(d)). Subsequently, the binding between miR-4677-3p and FGF6 was explored by the luciferase reporter assay. The potential binding area between LINC00265 and FGF6 was predicted from TargetScan (Figure 3(e)). The luciferase reporter assay demonstrated that miR-4677-3p overexpression reduced the luciferase activity of FGF6-Wt, while that of FGF6-Mut was not significantly altered in 1A6 and 5637 cells (Figure 3(f)). According to the RIP assay, LINC00265, miR-4677-3p, and FGF6 were markedly enriched in Ago2 precipitations compared with those in the IgG control group (Figure 3(g)). The above results indicated that FGF6 was directly targeted by miR-4677-3p, and LINC00265 upregulated FGF6 expression by binding with miR-4677-3p in BCa cells. LINC00265 upregulates FGF6 expression by interacting with miR-4677-3p. (A) The bioinformatics tool TargetScan was applied to predict potential target genes of miR-4677-3p. The mRNA FGF6 was identified since it not only exhibited high expression in BCa cells but was also downregulated by LINC00265 knockdown, as shown in the Venn diagram. (B)–(C) RT-qPCR and Western blot analyses were carried out to test the influence of miR-4677-3p overexpression on mRNA expression and protein levels of FGF6 in BCa cells. (D) The effects of LINC00265 knockdown on mRNA expression and protein levels of FGF6 were evaluated by RT-qPCR and Western blot analyses, respectively. (E) The binding site between miR-4677-3p and FGF6 was predicted by starBase. (F) The luciferase reporter assay was performed to investigate the binding between miR-4677-3p and FGF6 in BCa cells. (G) The RIP assay was performed to explore the relationship among LINC00265, miR-4677-3p, and FGF6. **p < 0.01, ***p < 0.001. Note: FGF6: fibroblast growth factor 6; BCa: bladder cancer; mRNA: messenger RNA; RIP: RNA immunoprecipitation.
LINC00265 promotes BCa cell viability, proliferation, and migration by upregulating FGF6 expression
Rescue experiments were conducted to further explore whether LINC00265 modulates malignant phenotypes of BCa cells by upregulating FGF6 expression. First, FGF6 was successfully overexpressed by transfection of pcDNA3.1/FGF6 into 1A6 and 5637 cells, as shown by RT-qPCR analysis (Figure 4(a)). Since sh-LINC00265#1 displayed better knockdown efficiency than sh-LINC00265#2, we identified sh-LINC00265#1 for the following rescue assays. The CCK-8 assay revealed that FGF6 overexpression rescued the inhibitory effect of LINC00265 knockdown on BCa cell viability (Figure 4(b)). The colony formation assay demonstrated that overexpressing FGF6 reversed the suppressive effect on BCa cell proliferation induced by LINC00265 knockdown (Figure 4(c)). In addition, FGF6 overexpression reversed the promotion of cell apoptosis mediated by LINC00265 knockdown, as shown in the TUNEL assay (Figure 4(d)). Western blot analysis exhibited that protein levels of proliferation markers (PCNA and cyclin D1) were reduced by LINC00265 deficiency and then reversed by overexpressing FGF6 (Figure 4(e)). In addition, FGF6 overexpression reversed the elevation of Bax, cleaved caspase 3, and cleaved caspase 9 protein levels and rescued the downregulation of the Bcl-2 protein level induced by LINC00265 knockdown (Figure 4(f)). Moreover, overexpressing FGF6 rescued the decrease in MMP2 and MMP9 protein levels mediated by LINC00265 knockdown (Figure 4(g)). According to the results of the transwell assay, FGF6 overexpression rescued the suppressive effect of LINC00265 depletion on BCa cell migration (Figure 4(h)). Thus, we concluded that LINC00265 facilitates BCa cell viability, proliferation, and migration by upregulating FGF6 expression. LINC00265 modulates malignant phenotypes of BCa cells by upregulating FGF6 expression. (A) The overexpression efficiency of FGF6 in BCa cells was detected by RT-qPCR analysis. (B)–(C) The suppressive effect of LINC00265 depletion on BCa cell viability and proliferation was rescued by FGF6 overexpression, as shown by CCK-8 and colony formation assays. (D) The apoptosis of BCa cells promoted by LINC00265 silencing was reversed by FGF6 overexpression, according to the TUNEL assay. (E)–(G) Western blot analysis was performed to detect the protein levels of proliferation markers (PCNA and cyclin D1), apoptosis markers (Bcl-2, Bax, cleaved caspase 3, and cleaved caspase 9) and migration-related key factors (MMP2 and MMP9) in BCa cells transfected with sh-NC, sh-LINC00265#1, and sh-LINC00265#1 + pcDNA3.1/FGF6. (H) The transwell assay was carried out to measure the migratory capacity of BCa cells with the above transfection. *p < 0.05, **p < 0.01, ***p < 0.001. Note: BCa: bladder cancer; FGF6: fibroblast growth factor 6; TUNEL: TdT-mediated dUTP nick-end labeling.
Silencing LINC00265 inhibits xenograft tumor growth in vivo
After the establishment of xenograft nude mice model, tumor volume was measured every day, and tumors of LV-sh-NC and LV-sh-LINC00265#1 were weighed after mice were euthanized on day 15. We discovered that LINC00265 depletion suppressed tumor growth and decreased tumor weight (Figures 5(a) and (b)). The expression levels of LINC00265, miR-4677-3p and FGF6 in tumor tissues were examined by RT-qPCR. LINC00265 and FGF6 expression was downregulated, while miR-4677-3p expression was increased in the LV-sh-LINC00265#1 group (Figure 5(c)). The results verified that LINC00265 upregulates FGF6 expression by targeting miR-4677-3p. Moreover, protein levels of factors associated with proliferation, apoptosis, and migration were detected by Western blot analyses in tumor cells. PCNA and cyclin D1 levels were decreased by LINC00265 silencing (Figure 5(d)). The Bcl-2 protein level was reduced while Bax, cleaved caspase 3, and cleaved caspase 9 levels were upregulated due to LINC00265 deficiency (Figure 5(e)). MMP2 and MMP9 protein levels were decreased by LINC00265 silencing (Figure 5(f)). The results of Western blot analyses were consistent with those of in vitro experiments. Overall, LINC00265 promotes xenograft tumor growth in vivo. Silencing LINC00265 inhibits xenograft tumor growth in vivo. (A) The volume of tumors in LV-sh-NC and LV-sh-LINC00265#1 groups was measured every 3 days. (B) Tumors in two groups were weighed after mice were euthanized. (C) LINC00265, miR-4677-3p, and FGF6 expression in tumor tissues were measured by RT-qPCR. (D)–(E) Protein levels of proliferation markers (PCNA and cyclin D1), apoptosis markers (Bcl-2, Bax, cleaved caspase 3, and cleaved caspase 9) and migration markers (MMP2 and MMP9) in tumor cells were probed by Western blot analyses. **p < 0.01, *** p < 0.001. Note: FGF6: fibroblast growth factor 6.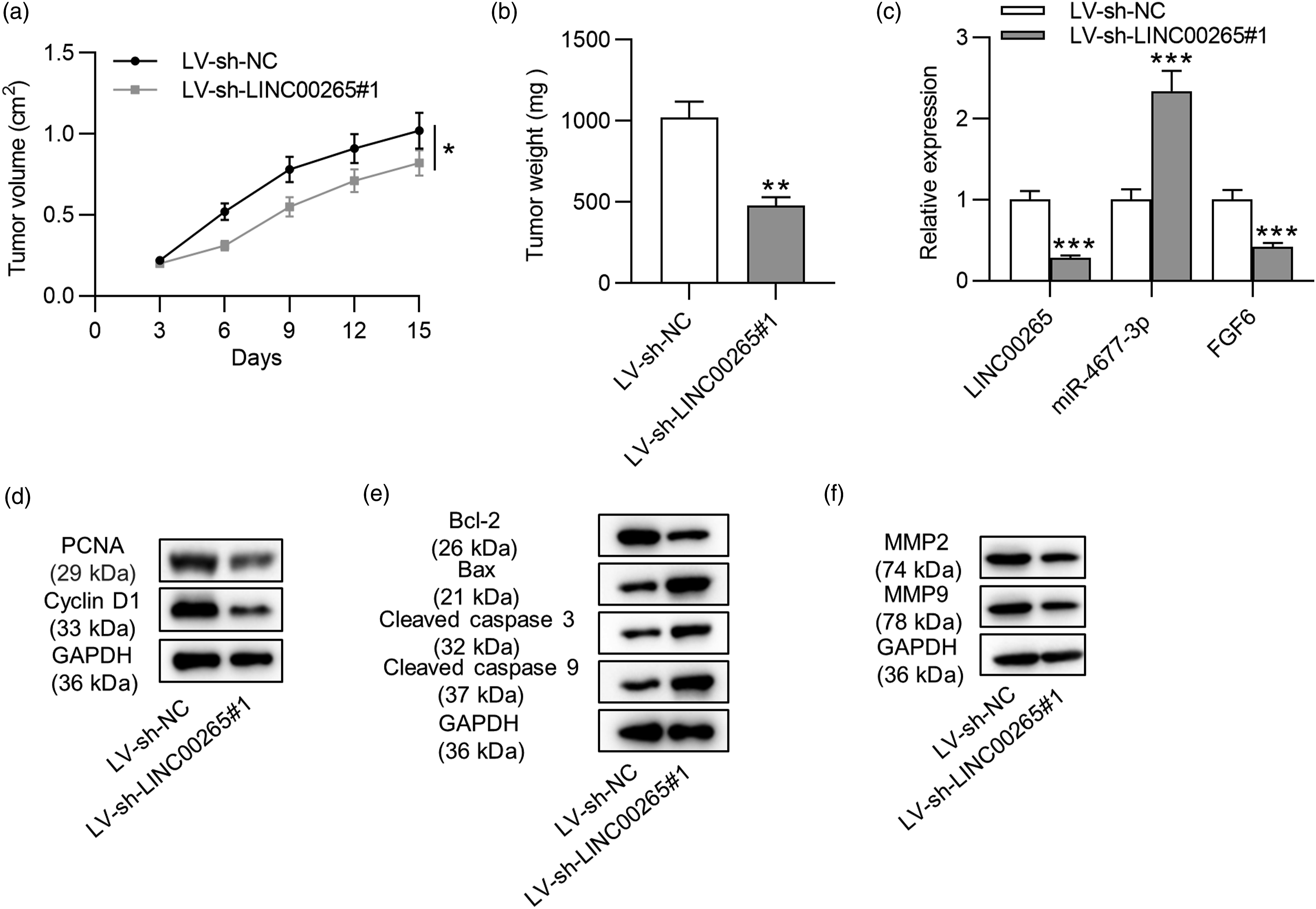
Discussion
Increasing reports suggest that lncRNAs are essential regulators involved in biological processes including cell proliferation, apoptosis, invasion, and stem cell maintenance.
27
Mechanistically, lncRNAs regulate gene expression by binding with miRNAs, which is widely known as the ceRNA hypothesis.
28
LINC00265 has been reported to be an oncogene in many types of cancer, including osteosarcoma, colorectal, lung, and gastric cancer.12–16 Moreover, the ceRNA role of LINC00265 has been verified in lung adenocarcinoma.
14
However, the role of LINC00265 in BCa has not been reported yet. In the present study, the functions and regulatory mechanism of LINC00265 in BCa were explored. We found that LINC00265 exhibits high expression in BCa cells. LINC00265 knockdown inhibits the viability, proliferation, and migration of BCa cells and promotes cell apoptosis, as shown by loss-of-function experiments. Moreover, silencing LINC00265 downregulates the protein levels of migration-related factors (MMP2 and MMP9) as well as levels of proliferation-associated protein (PCNA and cyclin D1). Additionally, the Bcl-2 protein level is decreased, while levels of Bax, cleaved caspase 3, and cleaved caspase 9 proteins are increased after LINC00265 depletion. Furthermore, in vivo experiments implied that LINC00265 deficiency suppresses xenograft tumor growth (Figure 6). Experimental design for in vitro and in vivo assays. (A) For in vitro assays, cells were transfected for 48 h. Next, RT-qPCR and Western blot analyses were conducted to examine gene expression levels, and subcellular fractionation assays were performed to determine the distribution of LINC00265. CCK-8 assays were performed to detect cell viability, which lasted for 3 days. Transwell assays were designed for cell migration detection, which lasted for 2 days. Colony formation assays were conducted to examine cell proliferation, which lasted for 14 days. After 2 days of cotransfection of luciferase reporters and plasmids, luciferase reporter assays were conducted. (B) For in vivo assays, tumor volume was recorded 3 days after subcutaneously injection of lentivirus (LV) into mice (n = 6/group). Volume record was performed every 3 days. On day 15, mice were euthanized and then tumors were weighed. RT-qPCR and Western blot were performed to examine gene expression in tumor tissues. Protein levels of proliferation, apoptosis, and migration markers were examined in tumor cells by Western blot.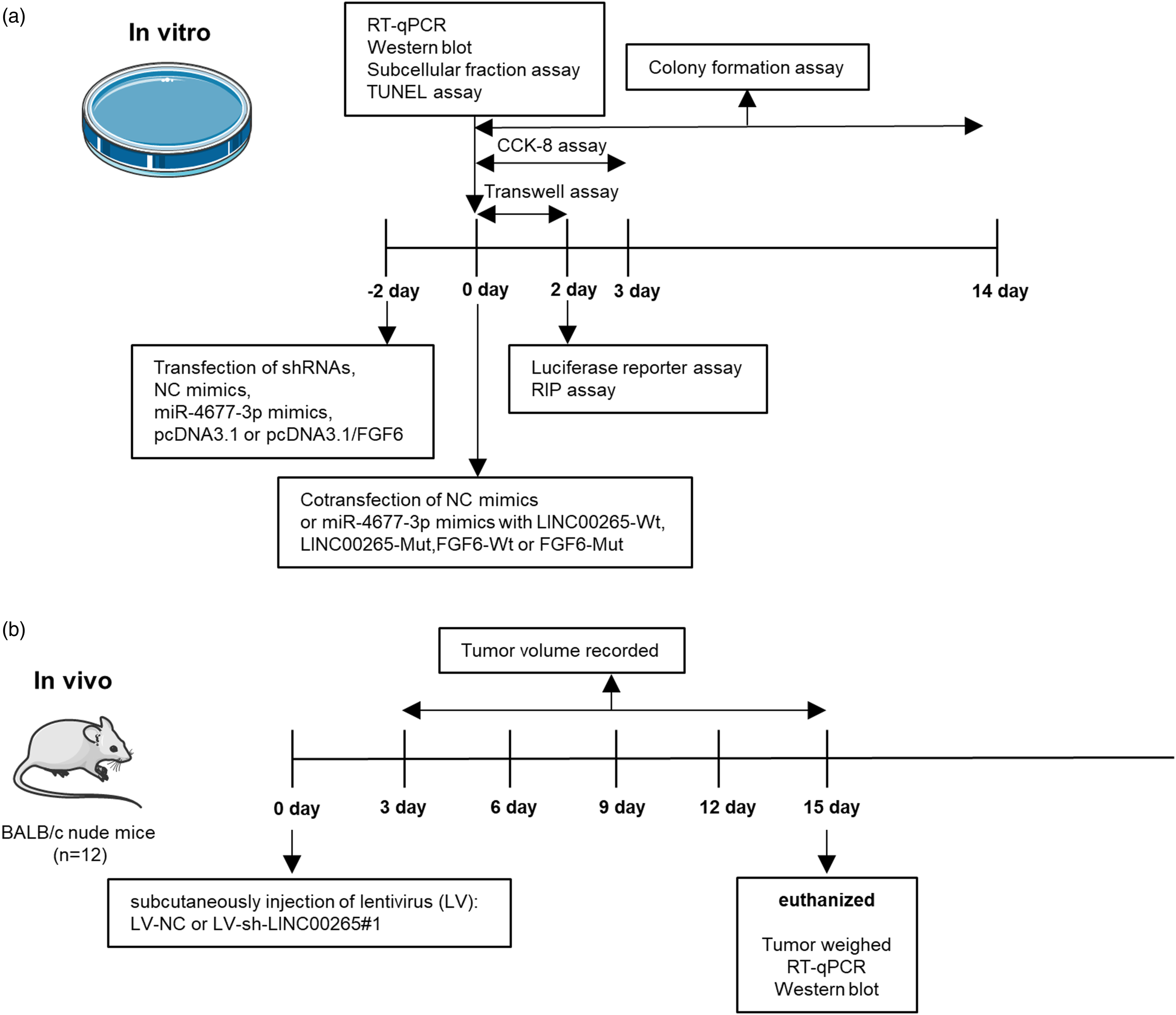
Mechanistically, the lncRNA-mediated ceRNA network, that is, the lncRNA–miRNA–mRNA network is extensively investigated in various cancers including BCa.7,29 For example, the lncRNA plasmacytoma variant translocation 1 gene promotes cellular processes of BCa cells by upregulating the expression of b-cells lymphoma-2–associated transcription factor 1 via interaction with miR-194-5p. 30 LncARSR promotes cell proliferation and metastasis by upregulation of SOX4 via miR-129-5p in BCa following the ceRNA mechanism. 31 In the study, LINC00265 is mainly located in cytoplasm, which indicates that LINC00265 regulates gene expression at the post-transcriptional level. Therefore, we explored the downstream genes of LINC00265 under the ceRNA network. According to bioinformatics analysis and our experiments, LINC00265 binds with miR-4677-3p in BCa cells. In previous reports, miR-4677-3p was reported to be targeted by LINC02418 to release the expression of kinetochore scaffold 1 (KNL1), thus facilitating the proliferation and motility of lung adenocarcinoma cells. 32 miR-4677-3p is bound by LINC00313 to upregulate the expression of cyclin-dependent kinase 6 (CDK6), thus promoting cell migration and epithelial–mesenchymal transition in cervical cancer. 33 miR-4677-3p was reported to interact with LINC02418, and LINC02418 increases the expression of the sec61 gamma subunit (SEC61G) via miR-4677-3p to modulate cell proliferation, invasion, and apoptosis in non–small cell lung cancer. 34 In the current study, miR-4677-3p exhibits low expression in BCa cells. The expression of miR-4677-3p is negatively correlated with LINC00265 expression in BCa cells. LINC00265 interacts with miR-4677-3p to upregulate the expression of FGF6.
Fibroblast growth factor 6 was found to be the target gene of miR-4677-3p in BCa cells. The mRNA expression and protein levels of FGF6 are positively regulated by LINC00265 and negatively modulated by miR-4677-3p in BCa cells. As shown by rescue assays, FGF6 overexpression rescues the suppressive effect on BCa cell viability, proliferation, and migration and reverses the enhancement of cell apoptosis mediated by LINC00265 knockdown. The results indicate that LINC00265 promotes malignant phenotypes of BCa cells by upregulating FGF6 expression. Previously, FGF6 was found to be aberrantly expressed in bladder and other cancer types, such as gastric cancer, colorectal cancer, and cervical carcinoma, indicating its contribution to cancer development. 35 FGF6 downregulation is associated with iron-metabolism dysfunction in cancer cells. 36
In conclusion, LINC00265 promotes cell viability, proliferation, and migration and inhibits cell apoptosis in BCa by serving as a ceRNA for miR-4677-3p to upregulate the expression of FGF6. This study might provide a novel biomarker for BCa diagnosis and treatment.
However, the potential downstream signaling pathway of FGF6 and the possible upstream factor regulating LINC00265 were not explored in this study. More experiments will be designed in our future studies to explore other possible genes involved in the LINC00265/miR-4677-3p/FGF6 axis in BCa progression.
Footnotes
Acknowledgment
We appreciate all participants for their contribution to the study.
Declaration of conflicting interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
