Abstract
The organic compound di(2-ethylhexyl) phthalate (DEHP) is widely used as a plasticizer in many products. Exposure to DEHP has been reported to lead to adverse pregnancy outcomes by suppressing placenta growth and development. The aim of this study was to determine the gene expression profiles of rat placenta exposed to (DEHP) and identify genes crucial for the DEHP response. Three groups of Wistar rats were administered an intragastric dose of 1,000 mg/kg DEHP, 500 mg/kg DEHP, or corn oil, RNA was isolated from placenta tissue, and hybridization was performed. Gene expression profiles were analyzed by identifying functional enrichment, differentially expressed genes (DEGs), protein–protein interaction (PPI) networks and modules, and transcription factor (TF)-miRNA-target regulatory networks. We obtained 2,032 DEGs, including cytochrome P450, family 2, subfamily R, polypeptide 1 (CYP2R1), sterol O-acyltransferase 2 (SOAT2), and 24-dehydrocholesterol reductase (DHCR24) from the steroid biosynthesis pathway and somatostatin receptor 4 (SSTR4) and somatostatin receptor 2 (SSTR2) in the neuroactive ligand-receptor interaction pathway. The PPI network included 476 nodes, 2,682 interaction pairs, and three sub-network modules. Moreover, eight miRNAs, three TFs, and 176 regulatory pairs were obtained from the TF-miRNA-target regulatory network. CYP2R1, SOAT2, DHCR24, SSTR4, and SSTR2 may affect DEHP influence on rat placenta development.
Keywords
Introduction
The organic compound di(2-ethylhexyl) phthalate (DEHP; C6H4 (CO2C8H17)2) has been shown in epidemiological studies to adversely affect insulin sensitivity and glucose homeostasis. 1 DEHP is often used as a plasticizer in many products, such as blood bags, infusion tubing, and umbilical artery catheters 2 and is used in the manufacture of consumer products like toys, infant products, wall coverings, and cosmetics. 3 –5 The widespread use of DEHP makes it likely that people are frequently exposed to DEHP. Although DEHP is somewhat degradable, phthalates can cross the placenta lining and build up in the bodies of children, creating health problems. 6
Previous studies have suggested that exposure to DEHP leads to adverse pregnancy outcomes due to the suppressed placenta growth and development. 7 Fiandanese et al. demonstrated that maternal exposure to DEHP and polychlorinated biphenyls results in reproductive dysfunction of adult male mouse offspring. 8 In addition, Pocar et al. found that exposure of pregnant female mice to DEHP leads to altered reproductive health of the female offspring. 9 Several additional studies have also explored the effects of maternal DEHP exposure during pregnancy on offspring. 10 –12 For example, Yu et al. indicated that DEHP exposure could exacerbate the incidence of fetal intrauterine growth restriction by dysregulating placenta thyroid hormone receptors. 13 Zhang et al. suggested that DEHP as well as its metabolite, mono (2-ethyl-hexyl) phthalate, could inhibit the cell growth in mouse placenta and dysregulate the progesterone secretion. 14 Although numerous studies have presented the effects of DEHP on the placenta during pregnancy, the molecular mechanisms involved in this process remain unknown.
Transcriptome analysis allows the investigation of gene expression pattern changes response to diseases, pathogens, and environmental challenges. 15 Using transcriptome RNA sequencing analysis, Midic et al. found that exposure of primate trophoblast stem cells to environmental toxicants led to substantial expression changes of a large number of genes, which were implicated in cytokine signaling, DNA damage repair, cell movement function, and anti-viral response. 16 In the present study, RNA-sequencing analysis was performed to investigate changes in gene expression in rat placenta exposed to DEHP at the gestational stage. We applied bioinformatics analyses in order to screen the crucial genes and to elucidate potential molecular mechanisms. This study is expected to provide theoretical basis and research targets to explore effects of DEHP on the placenta during pregnancy.
Materials and methods
Animals
The Ethics Committee and Animal Management Committee of Seventh Medical Center of Chinese PLA General Hospital approved all experiments. A total of 100 Wistar rats weighing approximately 200 g each were obtained from the Dalian Medical University Animal Center. All rats were raised at a constant environmental temperature of 21 ± 3°C.
Experimental protocols
Sixty female and 30 male rats were mated at a ratio of 2:1 and the vaginal plug of female rats checked the next morning. The female sexual cycle (determined by vaginal smears) was the same for all the female rats. Vaginal smears in early estrus are characterized by a large number of enlarged oval nuclear epithelial cells, with few white blood cells and keratinized epithelial cells. Estrus is characterized by the disappearance of nuclei of expanded epithelial cells, the aggregate of squamous exfoliation of keratinized epithelial cells, and few white blood cells. Late estrus is characterized by a majority of cells in the vaginal secretions being white blood cells, mixing with some keratinized epithelial cells. The resulting 43 pregnant rats were randomly divided into three groups: High (n = 17), Middle (n = 17), and Control (n = 9). After a 4-h fast, the rats in the High, Middle, and Control groups were administered 1,000 mg/kg DHEP (Sigma, St. Louis, MO, USA), 500 mg/kg DHEP, or corn oil, respectively, by gavage once per day for 7 days from day 7 to day 12 of pregnancy.
Microarray hybridization and analysis
On day 20 of pregnancy, we anesthetized the rats with intraperitoneal injection of 2% sodium pentobarbital (50 mg/kg) and then obtained the placenta.
Total RNA was extracted using TRIZOL reagent according to the manufacturer’s instructions. RNA integrity and concentration were then evaluated using a microplate reader (Infinite M100 PRO, TECAN). Total RNA were labeled using the Agilent Quick Amp Labeling Kit. mRNA was reverse transcribed to synthesize cDNA, followed by transcription to produce labeled cRNA. The labeled cRNA was hybridized to the Agilent SurePrint G3 gene expression 8 × 60 K microarray based on the manufacturer’s instructions. After washing, the arrays were scanned using the Agilent DNA microarray scanner. Data were extracted and normalized using the Agilent Feature Extraction Software (version 11.0.1.1) and raw data preprocessed using the Agilent GeneSpring GX v11.5.1. The Limma package was used to performed differential expression analysis. Differentially expressed genes (DEGs) were screened using pairwise comparison (High vs. Ctrl, Mid vs. Ctrl, High vs. Mid) with threshold of |log 2 FC| > 2.0 and P < 0.05. The DEGs in the three comparable group were then merged to performed time series analysis.
Short time-series expression miner (STEM) analysis
The time series analysis is a clustering analysis appropriate for longitudinal data and has been widely used in studies focusing on dynamical biological processes or patterns of change over time. 17 STEM 18 (version 3.11, http://www.cs.cmu.edu/∼jernst/stem/) was used here to analyze the gene expression pattern changes with DEHP dose. In STEM analysis, genes with consistent expression trend were clustered. The following parameters were used: ≤ 20 clusters, >0.7 correlation coefficient of gene expression in each cluster, and P-value <0.05. The genes in significant clusters were used in the following analysis.
Functional enrichment analysis for DEGs
To explore the involved functions of genes, Gene Ontology (GO) 19 and Kyoto Encyclopedia of Genes and Genomes (KEGG) 20 pathway enrichment analyses were conducted utilizing the online Database for Annotation, Visualization and Integrated Discovery (DAVID Version 6.8, https://david-d.ncifcrf.gov/). 21 The enriched GO terms and KEGG pathways with count ≥2 and P value <0.05 were considered significantly enriched.
PPI network and modules analyses
Protein–protein interactions (PPI) were predicted using STRING 22 (Version: 10.0, http://www.string-db.org/) database with highest confidence (PPI score = 0). The PPI network was then visualized using Cytoscape (Version 3.2.0, http://www.cytoscape.org/) software based on the retrieved PPIs. To analyze the topology property of the nodes in PPI network, the CytoNCA plug-in (Version 2.1.6, http://apps.cytoscape.org/apps/cytonca) 23 was used with the parameters set without weight. Afterward, hub nodes located in the PPI network were obtained according to the ranking of network connectivity. In addition, the functional modules were identified from the PPI network by using the Molecular Complex Detection (MCODE, Version 1.4.2, http://apps.cytoscape.org/apps/MCODE) plugin of Cytoscape. 24 The functional modules means the neighboring nodes interacted with each other as a function unit than other nodes.
TF -miRNA-target regulatory network
The microRNAs (miRNAs) and transcription factor (TFs) that target the genes in the PPI network were predicted using the WebGestalt GAST tool 11 (http://www.webgestalt.org/option.php) based on the Overrepresentation Enrichment Analysis (ORA) method. The species chosen was rnorvegicus and the reference gene sequences were Gene IDs from the selected platform agilent_wholegenome_4x44k_v1. A count number ≥2 was set as the enriched parameter. The significantly enriched miRNA-gene pairs were selected with P, and the significantly enriched TFs-gene pairs were selected using false discovery rates <0.05.
Statistical analysis
Statistical analyses were performed using the SPSS 19.0 software (SPSS Inc., Chicago, IL, USA). Measurement data were compared and analyzed by one-way analysis of variance (ANOVA), followed by Tukey’s post hoc test, and the count data were evaluated by the χ2 test.
Results
Effect of DEHP on maternal and fetal rats
Weight changes of pregnant rats are presented in Table 1. No significant differences were observed in weight changes of pregnant rats at 0 and 9 days of pregnancy (P > 0.05). At 18 days of pregnancy, the weight changes of pregnant rats was significantly reduced in the Middle and High groups compared to the Control group (P < 0.05). In addition, live fetuses in Middle and High groups were significantly lower than those of the Control group (Table 2, P < 0.05). Additionally, fetus malformation dramatically increased in the Middle and High groups compared to the Control group (Table 2, P < 0.05).
Comparison of weight change in pregnant rats.

Note: At 18 day of pregnancy, the weight changes of pregnant rats was significantly reduced in the Middle and High groups compared to the Control group. *P < 0.05, compared with the control group.
Comparisons of the general characteristics of fetal rats.

Note: Live fetuses of fetal rats in Middle and High groups were significantly lower than those of the Control group. Fetus malformation dramatically increased in the Middle and High groups compared to the Control group. *P < 0.05, compared with the control group.
RNA integrity
Three distinct bands were observed in electrophoresis results, and the A260/280 ratios of total RNA ranged 1.8-2.0. Total RNA integrity of all three groups was thus acceptable and could be used for this study. The workflow of this study is shown in Supplemental Figure 1.
DEG identification
Compared to the Control group, we identified 951 DEGs (424 up-regulated and 527 down-regulated) in the High group and 3787 DEGs (1951 up-regulated and 1836 down-regulated) in the mild group. Additionally, 3456 DEGs (1565 up-regulated and 1891 down-regulated) were identified between the High and Mild group, and a combined set of 2032 DEGs was used for subsequent analysis.
Expression profile analysis
Three significantly enriched clusters (of classes) (Figure 1) were obtained, comprising 896, 672, and, 464 genes in Class 10, Class 5, and Class 1, respectively. Expression patterns of genes in each class showed different concentration-dependent variation. Genes in all three significantly enriched classes were used for subsequent analysis.

Gene expression clustering analyzed by short time-series expression miner (STEM). In STEM analysis, the genes with consistent expression trend were grouped into one cluster. A total of 16 clusters (squares) were obtained, and three clusters (colored squares) were significant. The number in the upper left represents the serial number of the cluster, the black line in the middle the overall trend of genes expression in this cluster, and the number on the lower left portion represent the P value of gene expression similarity in this cluster.
Functional and pathway enrichment analyses of DEGs
KEGG pathway enrichment analysis results for Class 10, Class 5, Class 1, and all DEGs are presented in Table 3. Somatostatin receptor 4 (SSTR4) and somatostatin receptor 2 (SSTR2) were significantly enriched in the neuroactive ligand-receptor interaction pathway. Cytochrome P450, family 2, subfamily R, polypeptide 1 (CYP2R1), sterol O-acyltransferase 2 (SOAT2), and 24-dehydrocholesterol reductase (DHCR24) were enriched in the steroid biosynthesis pathway. GO analysis also indicated striking enrichment of genes involved in G-protein coupled receptor signaling, chemical stimulus detection, the plasma membrane, and olfactory receptor activity (Figure 2).
KEGG pathway enrichment results for the DEGs in Class 10, Class 5, Class 1, and union.

Note: Class 10, Class 5 and Class 1 are the three significant clusters in STEM clustering analysis. Union represents the union of the Class 10, Class 5 and Class 1. Count represents the number of genes in this pathway.

The significantly enriched Gene ontology (GO) annotations terms. The significantly enriched biological processes terms (A), cellular component terms (B) and molecular function terms (C). Vertical coordinates represent the enriched gene count, and horizontal axis represent the significantly enriched (top 10) GO terms; black lines represent the P value.
PPI network and module analyses
There were 2,682 interaction pairs and 476 nodes in the PPI network (Figure 3). Moreover, three sub-network modules (modules a, b and c) were identified using MCODE in the Cytoscape plugin. The sub-network module a (score = 61) included 1,830 interaction pairs and 61 nodes, sub-network module b (score = 15) contained 15 nodes and 105 interaction pairs, and sub-network module c included 26 nodes and 183 interaction pairs (score = 14.56).
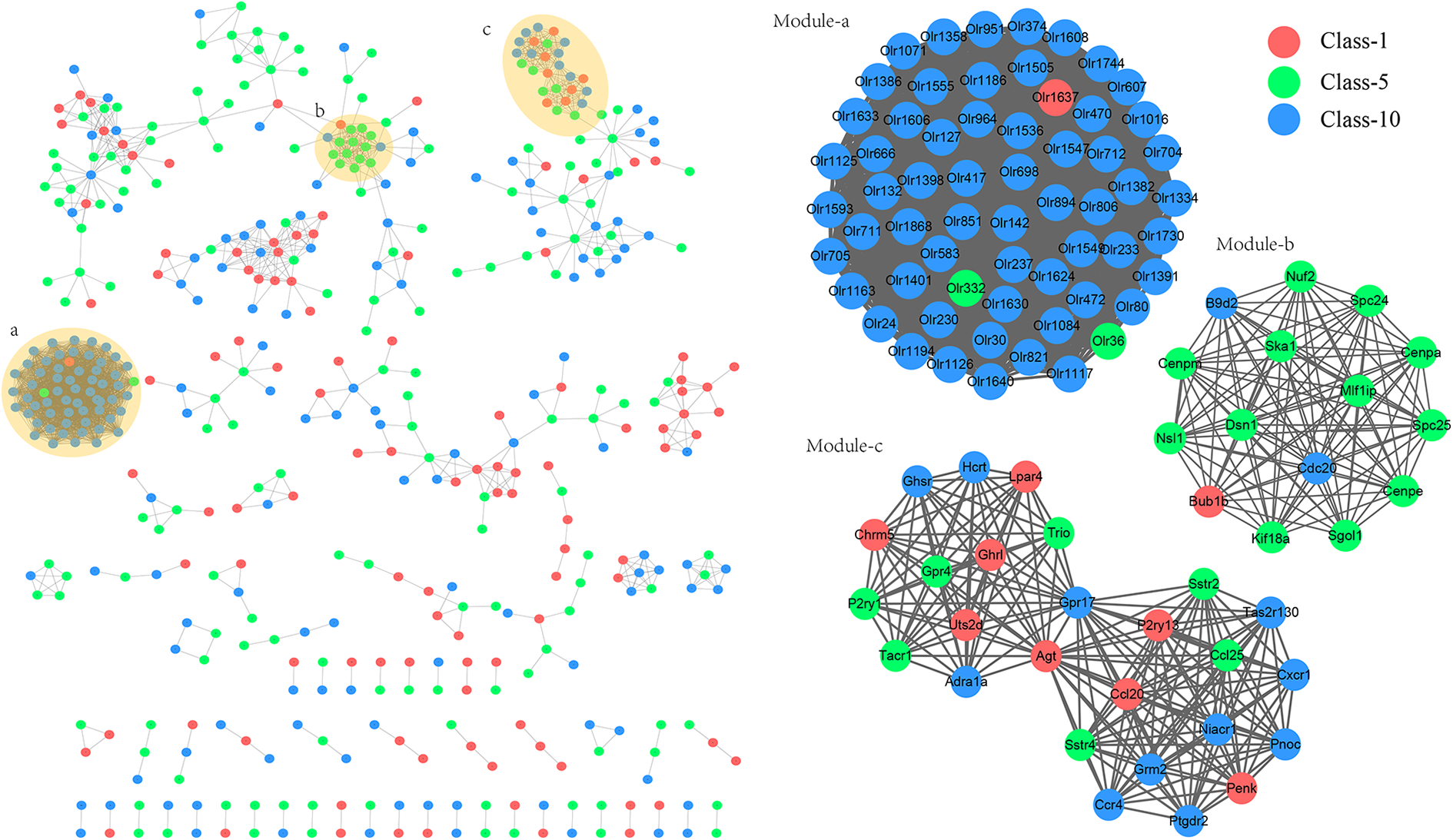
Protein–protein interaction (PPI) networks and functional modules. Red nodes, green nodes, and blue nodes represent the genes in cluster 1, cluster 5 and cluster 10 in STEM analysis, respectively. Module-a, module-b, and module-c represent the identified functional modules with score >10 from PPI network.
KEGG pathway enrichment results for modules are in Table 4. Genes in the sub-networks module a and module b were enriched in olfactory transduction and cell cycle pathways. The genes in sub-network module c were enriched in the pathway of neuroactive ligand-receptor interactions.
KEGG pathway enrichment results for genes in modules.

Note: module-a, module-b, and module-c are the significant modules identified from protein–protein interaction network. The enrichment analysis are performed for the genes in those modules. Count represents the number of genes in this pathway.
TF-miRNA-target regulatory network
Eight miRNAs, three TFs, and 176 regulatory pairs were obtained, including 30 genes in class 10, 35 genes in class 5, and 26 genes in class 1. The regulatory network of TF-miRNA-target was established by Cytoscape (Figure 4).

Transcription factor (TF)-microRNA-target regulatory network. Red nodes, green nodes, and blue nodes represent the genes in cluster 1, cluster 5 and cluster 10 in STEM analysis, respectively. Rhombus represent transcription factors and triangles represent microRNAs. Lines represent the interaction between two nodes, and arrow represent the direction of gene regulation.
Discussion
The hormone-like activity of DEHP makes it an endocrine disruptor. A previous study has shown that the effects of DEHP demonstrate endocrine toxicity. 25 In the present study, we speculated that DEHP critically impacts placenta development due to its endocrine toxicity and modifications of genetic material associated with the placenta.
Our results show that CYP2R1, SOAT2, and DHCR24 were differentially expressed in the placenta of rats exposed to DEHP, compared with control, and were significantly enriched in steroids synthesis. CYP2R1 is a member of the cytochrome P450 (CYP) superfamily of enzymes, which has been reported to mediate steroid biosynthesis. 26,27 Steroid biosynthesis is a complicated process involving multiple enzymes as well as cofactors, and CYP enzymes are a major kind of steroidogenic enzymes. 28 Notably, studies had reported that the disturbance of CYPs system homeostasis was implicated in DEHP induced apoptosis injury, 29 immunosuppression, 30 and nephrotoxicity. 31 DHCR24 (24-dehydrocholesterol reductase) catalyzes the reduction of the delta-24 double bond of sterol intermediates during cholesterol biosynthesis. 32 Several studies have indicated that DHCR24 plays a crucial role in cholesterol biosynthesis by modulating lipid raft formation to facilitate signal transduction and trafficking. 33 –35 SOATs are able to catalyze intracellular cholesterol ester (CE) synthesis, and include ACAT-1 and ACAT-2 two forms, which function in maintaining the cellular cholesterol homeostasis. 36,37 As a kind of endocrine-disrupting compound, DEHP was reported to target the reproductive tract through the up-regulation of steroid hormone receptors, which are involved in the regulation of reproductive development by acting as transcriptional factors. 38 Zhou et al. indicated that mono-(2-ethylhexyl) phthalate, the active metabolite of DEHP, could stimulate the steroid synthesis and progesterone synthesis in Leydig cells by activating hormone-sensitive lipase and steroidogenic acute regulatory. 39 Although there are no studies that report the associations between effect in rat placenta of gestational DEHP exposure and steroid biosynthesis, the results of our study suggest that gestational DEHP exposure affect placenta development probably through mediating steroid biosynthesis by the regulation of CYP2R1, SOAT2, and DHCR24.
It is well known that because DEHP can cross the placenta and enter fetal circulation it can affect neurodevelopment and cause teratogenic anomalies by disrupting fetal brain development. 40,41 We thus speculated that the genes associated with neurodevelopment might underlie the toxicity of DEHP in placenta. Here, our results showed that SSTR4 and SSTR2 from the neuroactive ligand-receptor interaction pathway are upregulated by DEHP treatment. Somatostatin is known to restrain the release of many hormones. 42 In particular, G protein-coupled receptors likely mediate the biological effects of somatostatin. 43 SSTR4 and SSTR2 are members of the GPCR superfamily, and SSTR4 is most prevalent in fetal and adult brains and lungs, while SSTR2 is most prevalent in the kidneys and cerebrum. 43 Overall, these results suggest that SSTR4 and SSTR2 are involved in DEHP-triggered placenta toxicity through the neuroactive ligand-receptor interaction pathway.
It has been demonstrated that DEHP is highly toxic, and DEHP exposure can result in embryo mortality. 44 Our data show that the numbers of live fetuses in the Middle and High groups were obviously lower than those in the Control group. As well, fetus malformation dramatically increased in the Middle and High groups compared with that in the Control group. Additionally, the number of dead fetuses in the High group was higher than that in the Middle group, indicating the DEHP effect on placenta toxicity is dose-dependent.
There are some limitations in this study. The present predicted results were not verified by the laboratory data; for example, the expressions of key DEGs in tissue or serum should be confirmed by quantitative PCR and relevant testing kits. Further study should confirm whether steroid biosynthesis changes in serum. More research is required to confirm the involved functions of these genes in the toxic process of DEHP in placenta. Meanwhile, the regulatory relationships between DEGs and TFs will be verified.
Conclusion
In summary, our results suggest that CYP2R1, SOAT2, and DHCR24 of the steroid biosynthesis pathway and SSTR4 and SSTR2 of the neuroactive ligand-receptor interaction pathway may be critical for DEHP-induced placentatoxicity. In particular, these genes appear be involved in rat placenta development after exposure to different doses of DEHP.
Supplemental material
Supplemental_Figure_1 - Gene expression in rat placenta after exposure to di(2-ethylhexyl) phthalate
Supplemental_Figure_1 for Gene expression in rat placenta after exposure to di(2-ethylhexyl) phthalate by Wan Xu, Hongyan Wu and Lixin Shang in Human & Experimental Toxicology
Footnotes
Data availability statement
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Ethical approval
The Ethics Committee and Animal Management Committee of Seventh Medical Center of Chinese PLA General Hospital approved all experiments.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Eleventh Five-Year Programs for Science and Technology Development of Military [Program No. 06G019].
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
