Abstract
Background:
Epirubicin is a potent chemotherapeutic agent for the treatment of breast cancer. However, it may lead to cardiotoxicity and cardiomyopathy, and no reliable biomarker was available for the early prediction of epirubicin-induced cardiomyopathy.
Methods:
Global gene expression changes of peripheral blood cells were studied using high-throughput RNA sequencing in three pair-matched breast cancer patients (patients who developed symptomatic cardiomyopathy paired with patients who did not) before and after the full session of epirubicin-based chemotherapy. Functional analysis was conducted using gene ontology and pathway enrichment analysis.
Results:
We identified 13 significantly differentially expressed genes between patients who developed symptomatic epirubicin-induced cardiomyopathy and their paired control who did not. Among them, the upregulated Bcl-associated X protein was related to “apoptosis,” while the downregulated 5′-aminolevulinate synthase 2 (ALAS2) was related to both “glycine, serine, and threonine metabolism” and “porphyrin and chlorophyll metabolism” in pathway enrichment analysis.
Conclusions:
ALAS2 and the metabolic pathways which were involved may play an important role in the development of epirubicin-induced cardiomyopathy. ALAS2 may be useful as an early biomarker for epirubicin-induced cardiotoxicity detection.
Introduction
Anthracycline is a group of antitumor antibiotics, of which epirubicin is a very potent chemotherapeutic agent for the treatment of myeloid and lymphatic leukemia, non-Hodgkin’s lymphoma, and solid tumors such as breast cancer, lung cancer, and thyroid cancer. 1 However, because of its high affinity with myocardial tissue, its adverse effects of cardiotoxicity is common and may lead to irreversible myocardial injury, and eventually results in dilated cardiomyopathy and congestive heart failure. 2 At present, clinical parameters for cardiac toxicity monitoring, such as creatine kinase isoenzyme, 3 troponin, 4 and fatty acid-binding protein, 5 can sensitively detect anthracycline-induced myocardial injury. However, patients with abnormal changes in these biomarkers may have already experienced severe irreversible cardiotoxicity and being progressing into heart failure state. 6 Therefore, it is crucial to find timely, effective and reliable analytical biomarkers for the early prediction and evaluation of chemotherapy-induced cardiotoxicity to effectively prevent and monitor the side effects of anthracycline treatment in patients under chemotherapy treatment.
High-throughput RNA sequencing (RNA-seq) provides a unique opportunity to investigate the presence and quantity of RNA in a biological sample under specific conditions to detect subtle changes in gene expression level. 7,8 Through RNA-seq technology, previous in vitro study revealed that after 24 h exposure to doxorubicin, the transcriptomic response of induced pluripotent stem cell-derived cardiomyocytes is substantial. 9 To explore the possible genetic difference in vivo may help us to better elucidate the underlying molecular mechanisms of those patients at high risk of developing irreversible anthracycline-induced cardiomyopathy.
In the present study, we used RNA-Seq to comprehensively identify the differentially expressed genes (DEGs) related to symptomatic heart failure in breast cancer patients receiving full course epirubicin chemotherapy, which may help to identify potential biomarkers predict future anthracycline-induced cardiomyopathy risk.
Materials and methods
Study population
The study used a matched-pair case–control design. We initially enrolled 363 women consecutively treated for breast cancer with epirubicin-based chemotherapy at our hospital between May 2016 and May 2017. All patients were treated with a combination of epirubicin: 100 mg/m2 and cyclophosphamide: 600 mg/m2 at about 3-week intervals for eight cycles and have a normal cardiac function, with no heart failure symptoms and left ventricular ejection fraction (LVEF) ≥50%, at baseline. During a 6-month follow-up from their last session of chemotherapy, three patients developed cardiomyopathy and were identified as cardiotoxicity subjects. Among the same cohort, we identified three breast cancer patients without cardiotoxicity manifestation during follow-up, matching with three cardiotoxicity subjects separately forage (within ±5 years), type of surgery, pathologic stage (tumor stage and nodal stage), estrogen receptor status, progesterone receptor status, and human epidermal growth factor receptor 2 status to serve as control. All cardiotoxicity subjects and controls were contacted after identification to obtain informed consent and to provide a blood sample for RNA extraction if not already available. Cardiotoxic cardiomyopathy was defined as New York Heart Association functional class III and IV with a significant reduction in LVEF to below 50% after treatment during 6 months period of follow-up.
Echocardiography
Blood samples were immediately processed into plasma by centrifugation and were kept at −80°C until further analysis. Concentrations of cardiac troponin i samples, collected at the two time points, were measured on an Elecsys 1010 platform (Hoffman-LaRoche Ltd, Basel, Switzerland). The analytical sensitivity of the cTni immunoassays was 0.010 ng/mL.
Echocardiography investigation was performed before the first doses of epirubicin treatment and was repeated 6 months later after the last drug administration, using GE ViVidE 7 ultrasound machine (GE Healthcare, USA) with a 3.5-MHz transducer. LVEF was measured by Simpson’s method in the two-dimensional echocardiographic apical four-chamber view.
Plasma isolation
Blood samples were collected into ethylenediaminetetraacetic acid blood collection tubes immediately before the first doses of epirubicin treatment and 48 h after the last drug administration. Centrifugation was performed at 16,000 × g for 10 min. The upper plasma was then aspirated and transferred to a new sterilized centrifuge tube. After another centrifugation at 16,000 × g for 5 min, trace cells and impurities were removed. Plasma was then stored in diethyl pyrocarbonate-treated sterile centrifuge tubes at −20°C for future RNA analysis.
RNA extraction, library construction, and sequencing
RNA was extracted from 1 mL of plasma using the TRIzol reagent (Invitrogen, Carlsbad, California, USA) and purified with RNeasy mini kit (Qiagen) according to the manufacturer’s protocol. Nanodrop spectrophotometer (IMPLEN GmbH, Munich, Germany) was used to measure the purity and concentration of sample RNA. cDNA libraries were constructed using conventional BGISEQ-500 library construction protocol. 10 Briefly, (1) Oligo (dT) magnetic beads are used to select mRNA with polyA tail, then obtain the target RNA after purification. (2) The target RNA was fragmented and reverse transcripted to double-strand cDNA (dscDNA) by N6 random primer. (3) End repair the dscDNA with phosphate at 5′ end and stickiness “A” at 3′ end, then ligate an adaptor with stickiness “T” at 3′ end to the dscDNA. (4) Two specific primers are used to amplify the ligation product. (5) Denature the polymerase chain reaction (PCR) product by heat and the single-strand DNA is cyclized by splint oligo and DNA ligase. (6) The library was validated with Agilent 2100 Bioanalyzer (Agilent DNA 1000 Reagents) to assess fragment size and concentration. Then, we performed sequencing on the prepared library with the BGISEQ-500 sequencing system (BGI, Wuhan, China) for high-throughput RNA measurements.
Analysis of sequencing data
Primary sequencing data, called as raw reads, were subjected to quality control (QC) by using the FastQC v0.11.5 software to determine if a resequencing step is needed. After QC, raw data were filtered to remove the reads containing adapter, reads in which unknown bases are more than 10%, and low quality reads (the percentage of low-quality bases is over 50% in a read, we define the low-quality base to be the base whose sequencing quality is no more than 5), to generate the clean data and aligned to the reference sequences. For sequence mapping, we use Bowtie2 11 to map clean reads to reference genes and use Hierarchical Indexing for Spliced Alignment of Transcripts 12 to reference genome. The NOISeq 13 method was used to screen DEGs between two groups.
Enrichment analysis of differentially expressed genes
Annotation analysis of Gene Ontology (GO) is performed for screened DEGs. Firstly, we mapped all DEGs to GO terms in the database (http://www.geneontology.org/), calculating gene numbers for every term, then used the hypergeometric test to find significantly enriched GO terms in the input list of DEGs, based on “GO Term Finder” (http://www.yeastgenome.org/help/analyze/go-term-finder). We performed pathway enrichment analysis of DEGs based on the Kyoto Encyclopaedia of Genes and Genomes (KEGG, http://www.kegg.jp/kegg/pathway.html) database 14 to further identify significantly enriched metabolic pathways or signal transduction pathways in DEGs comparing with the whole genome background.
Quantitative reverse transcriptase PCR validation
We verified the 5′-aminolevulinate synthase 2 (ALAS2) expression by quantitative reverse transcriptase PCR (RT-qPCR) after the screening of the DEGs. Reverse transcription for cDNA synthesis was performed as per the manufacturer’s instructions using an RT Reagent Kit (Thermo Scientific) with ABI PRISM 7000HT Sequence Detection System (Applied Biosystems, Foster, California, USA). After normalization to glyceraldehyde 3-phosphate dehydrogenase, Δcycle threshold (ΔCt) method (ΔCt = CTRNA − CTGAPDH) was used to determine the relative ALAS2 RNA expression levels.
Statistical analysis
The NOISeq method was used to assess the DEGs in the two groups using the following criteria as default: fold-change ≥ 2.0 and diverge probability ≥ 0.8. An associated p value <0.05 for the GO term enrichment among DEGs and KEGG enrichment analysis was considered indicative of a statistically significant difference. An independent sample t-test was used to analyze the RT-qPCR validation results.
Results
Clinical characteristics of subjects and controls
In this study, we collected peripheral blood samples from three breast cancer patients who developed epirubicin-based chemotherapy-induced symptomatic cardiomyopathy and three paired patients without epirubicin-induced cardiotoxicity. Table 1 presents the clinical characteristics of enrolled patients according to the matched criteria.
Characteristics of subjects and matching controls.

ER: estrogen receptor; PR: progesterone receptor; HER2: human epidermal growth factor receptor 2; LVEF: left ventricular ejection fraction.
RNA-seq, reads mapping, and gene expression
As a result of high-throughput RNA-seq, we averagely obtained 23,781,383 raw sequencing reads and then 23,580,736 clean reads after filtering low quality. Table 2 briefly summarizes information on sequencing data for each sample.
Summary of sequencing data for each sample.a

a Clean data rate (%) = clean reads number/raw reads number.
After filtering, clean reads were mapped to reference using HISAT/Bowtie2 tool. The average mapping ratio with the reference gene was 75.68% and Table 3 listed the separate mapping rate for each sample. The average genome mapping ratio is 92.35% as given in Table 4. We counted the number of identified expressed genes for each sample as shown in Figure 1.

Number of identified genes of different samples.
Alignment statistics of reads align to the reference gene.a

a Total mapped reads (%) = unique match (%) + multi-position match (%).
Alignment statistics of reads align to the reference genome.a

a Total mapped reads (%) = unique match (%) + multi-position match (%).
Gene expression profile associated with epirubicin-induced cardiotoxicity
Comparing the epirubicin-induced cardiotoxicity patients’ group to the control patients’ group, 12 genes were significantly differentially expressed (fold change >1.5, false discovery rate < 0.1), among which nine were upregulated while three were downregulated (Figure 2). Detailed information for the 12 genes is listed in Table 5.

Heat map diagram of the expression patterns of DEGs in subjects developed epirubicin-induced cardiomyophathy and controls who did not. DEG: differentially expressed gene.
Twelve DEGs between the two groups.

DEG: differentially expressed gene; G0S2: G0/G1 switch gene 2; BAX: Bcl-2-associated X protein; SNORD: small nucleolar RNAs, C/D box; ALAS2: 5′-aminolevulinate synthase 2; IL: interleukin.
GO and pathway enrichment analysis of DEGs
Annotation analysis of GO is performed for further investigation of the biological processes, cellular component, and molecular function that were possibly involved in these DEGs. A total of seven significantly enriched GO terms were obtained, including the immune system process, hemopoiesis, hematopoietic or lymphoid organ development, cellular metal ion homeostasis, immune system development, and response to stress. Downregulated gene ALAS2 and upregulated gene Bcl-associated X protein (BAX) were found to be involved in all the seven process categories, while upregulated IL32 was only found related to “immune system process” and “response to stress.”
Pathway enrichment analysis was implemented to assess the activated biological pathways during epirubicin-induced myocardial toxicity. As a result, these DEGs were significantly enriched in 20 KEGG pathways for upregulated and downregulated genes. Among them, the three pathways with the most significant p value include glycine, serine, and threonine metabolism (p = 0.005065, ko00260), apoptosis (p = 0.005267, ko04215), and porphyrin and chlorophyll metabolism (p = 0.006177, ko00860). Among the upregulated DEGs, BAX was related to “apoptosis,” while the downregulated DEGs, ALAS2, was related to both “glycine, serine, and threonine metabolism” and “porphyrin and chlorophyll metabolism” (Table 6).
Pathways enriched for the deferentially expressed genes corresponding to the proteins of the protein–protein interaction network.
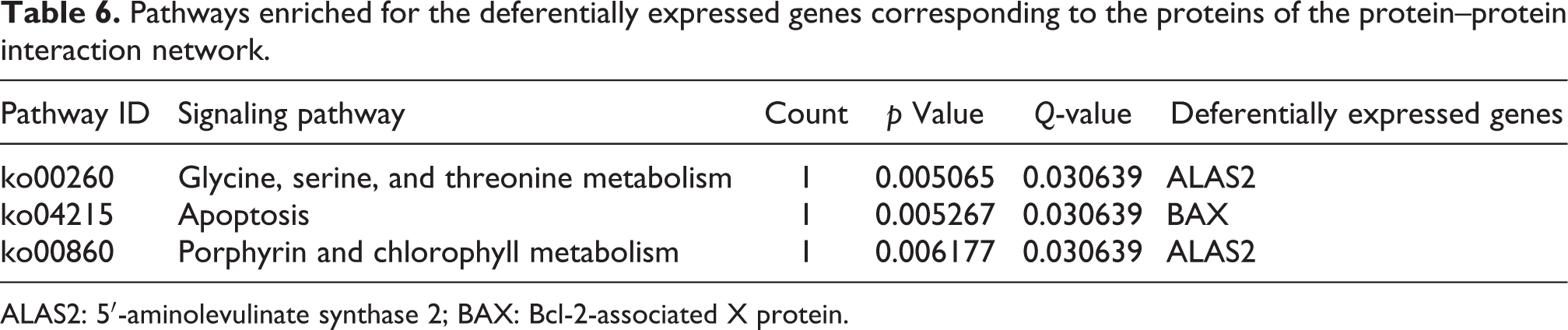
ALAS2: 5′-aminolevulinate synthase 2; BAX: Bcl-2-associated X protein.
RT-qPCR validation for ALAS2 expression
To validate the high-throughput RNA-seq result of the different ALAS2 expression, RT-qPCR was conducted in all six patients’ blood samples. RT-qPCR results showed that ALAS2 was downregulated in three patients with epirubicin-induced cardiomyopathy, compared with paired patients without developing epirubicin-induced cardiotoxicity, in the trend matched with those acquired in the RNA-seq analysis (Figure 3).
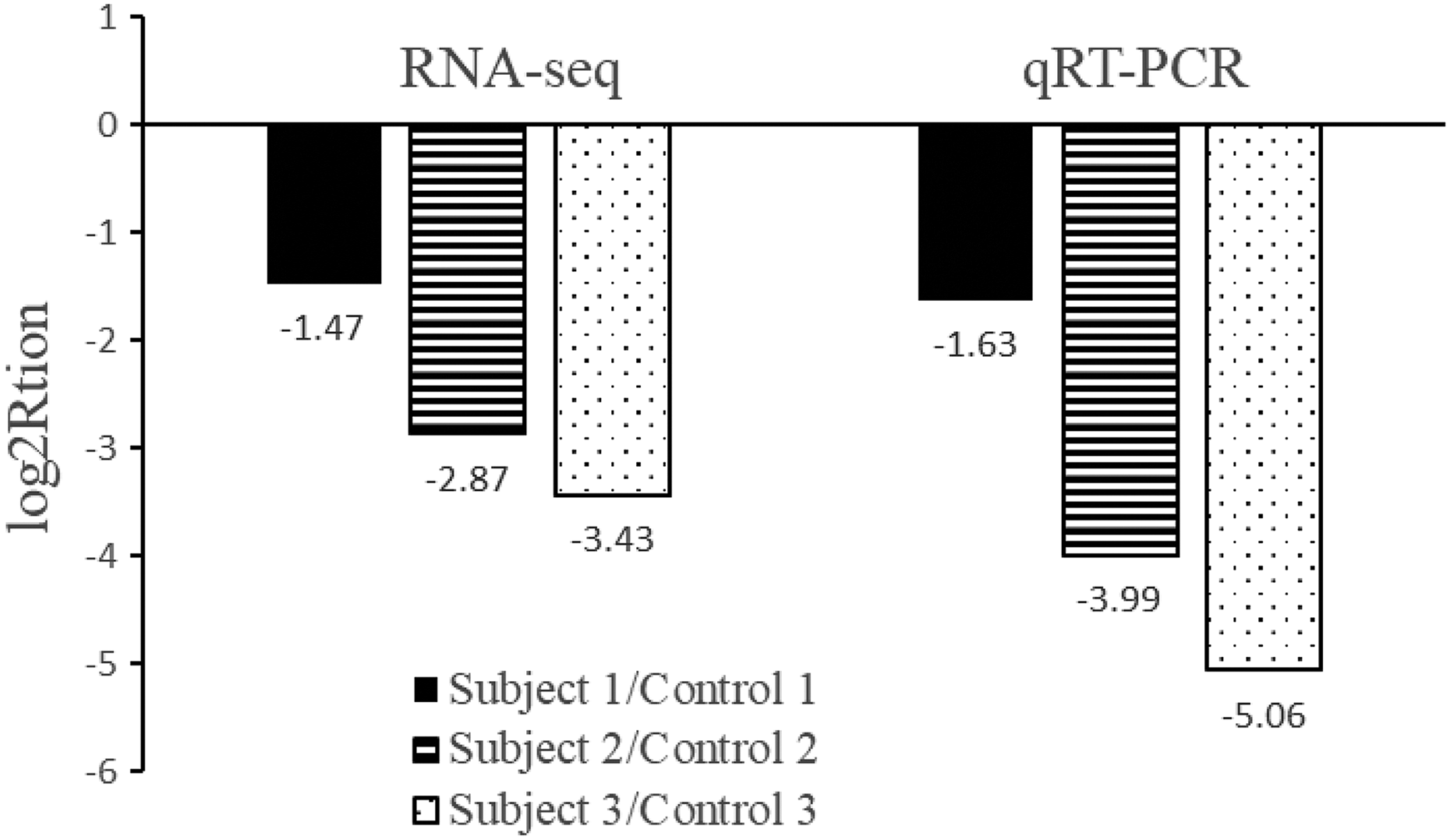
RT-qPCR validation of ALAS2 expression for patients with or without epirubicin-induced cardiomyopathy. The height of the columns in the chart represents the log-transformed fold change in ALAS2 expression across the subject and control in each matched pair. RT-qPCR: quantitative reverse transcriptase polymerase chain reaction; ALAS2: 5′-aminolevulinate synthase 2.
Discussion
In the present study, a total of 12 DEGs were identified between the epirubicin-induced myocardial toxicity and control samples. Of these, nine genes were significantly upregulated and three were downregulated. In ingenuity pathway analysis for the DEGs, downregulated ALAS2 exhibited the highest degree and therefore was identified to be highly involved in the occurrence and propagation processes of anthracycline-induced cardiotoxicity.
Anthracyclines, including epirubicin, doxorubicin, and aclacinomycin, are widely used in the treatment of hematological malignancies and solid tumors such as acute leukemia, lymphoma, breast cancer, gastric cancer, soft tissue sarcomas, and ovarian cancer. However, cardiotoxicity is the most serious toxic side effect of anthracyclines, and it lacks effective preventive and therapeutic measures. 15 Clinical studies and practical observations have shown that the anthracycline-induced cardiotoxicity tends to be progressive and irreversible and may develop after the initial administration. 16 Therefore, screening high-risk patients, early monitoring, and prevention of anthracycline-induced cardiotoxicity is particularly important. Currently, there are limitations in the clinical methods for the prevention and treatment of cardiac toxicity of anthracyclines. Regarding diagnosis, troponin abnormalities, clinical symptoms, and echocardiographic abnormalities are the main indicators for the diagnosis of cardiac toxicity in the present clinical practice, but sensitivity and specificity are relatively poor. 17 When these indicators are abnormal, patients may have been with severe cardiac toxicity or progressing to irreversible heart failure. 18
Unlike the mechanism of its antitumor activity, the main mechanism of anthracycline-induced cardiotoxicity is the results of iron-mediated reactive oxygen species and the myocardial oxidative stress. 19 Anthracyclines may chelate iron ions and then trigger the production of oxygen radicals, especially the hydroxyl radicals, leading to lipid peroxidation and myocardial mitochondrial DNA damage in myocardial cells. 20 Heme is the main form of iron ion in the body, and its synthesis is accomplished by an eight-step enzyme-catalyzed reaction. 5-Aminolevulinic acid synthase (ALAS) catalyzes the formation of 5-aminolevulinic acid from succinyl-coenzyme A and glycine, which is the first step in the synthesis of hemoglobin. 21 In the present study, we found that ALAS2 gene expression was significantly downregulated in the patients who developed myocardial toxicity.
This result was not consistent with the report by Todorova et al. 22 In their research, ALAS2 gene expression was found to be 1.48 log2-fold upregulated in breast cancer patients after the first cycle of anthracycline-based chemotherapy. However, there was no significant difference in ALAS2 gene expression between women who developed abnormal LVEF after a full course of chemotherapy and women with normal LVEF in their study.
The following possibilities could explain this inconsistency: first, in the present study, we examined the gene expression after a full course of anthracycline-based chemotherapy. There may be a complex time-dependent pharmacodynamic change, but not necessarily in a linear manner, of ALAS2 gene expression related to the total cumulative dose of epirubicin in the body. Second, the patients with downregulated expression of ALAS2 in our study exhibited obvious symptoms of heart failure together with evidence of myocardial damage, including high-sensitivity troponin i elevation and LVEF reduction below 50% after chemotherapy treatment, while Todorova et al.’s study enrolled patients only with a presymptomatic or subclinical decrease in LVEF evaluation. 22 There may be an inflection point or even turning point of ALAS2 gene expression related to the different levels of epirubicin-induced myocardial injury.
The dichotomous pattern of biomarkers related to clinical prognosis has been reported in the previous studies. Secemsky et al. found that in patients infected with human immunodeficiency virus and developing cardiac dysfunction, soluble suppression of tumorigenicity 2 (ST2) associated with all-cause mortality in a nonlinear pattern with an inflection point occurring near 50 ng/mL. 23 O’Donoghue et al. also found that in patients with acute myocardial infarction, multi biomarkers, such as NT-pro B-type natriuretic peptide, troponin T, ST2, galectin-3, and growth-differentiation factor-15 were related with cardiovascular death or HF in a dichotomous manner. 24
The inconsistency of the present study with the previous research may serve as an important clue for the pivotal role of ALAS2 in the mechanisms of epirubicin-induced myocardial toxicity development. The ALAS-heme pathway activation might cause mild to moderate damage through oxidative stress. On the other hand, severe cardiomyocyte damage may suppress the ALAS-heme pathway by direct cell injury and mitochondrial dysfunction or indirect upregulation of protective feedback factors that inhibit the ALAS2 gene expression.
In the present study, we also found that BAX gene expression was significantly upregulated in patients who developed clinical myocardial toxicity compared with patients without epirubicin-induced cardiotoxicity. BAX has a pro-apoptotic effect, which can reduce the threshold of apoptosis and regulate apoptosis through directly affecting mitochondrial function and membrane integrity. 25 By changing the mitochondrial permeability via the formation of the BAX channel, BAX may affect the induction of cell death through the production of oxygen free radicals, the pumping of calcium ions into the cytoplasm, and the release of mitochondrial proteins into the cytoplasm to activate caspases. 26 In consistent with our findings, Chaudhari et al. reported that the upregulation of BAX was identified in the human-induced pluripotent stem cell-derived cardiomyocytes incubated with doxorubicin. 27 However, because BAX can be produced by diverse cell types and exert its function on a variety of cell function pathways, there may be some limitation for BAX to be used as a predictor of epirubicin-induced cardiotoxicity.
In conclusion, the results of the present study showed that ALAS2 was related to the epirubicin-induced myocardial toxicity, and for the first time indicate that “glycine, serine, and threonine metabolism” and “porphyrin and chlorophyll metabolism” pathway may be associated with the development of symptomatic cardiomyopathy after anthracycline-based chemotherapy. Further mechanism research is needed to better explore the potential biomarker value of ALAS2 in breast cancer patients at a high risk of drug-induced cardiotoxicity.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Beijing Municipal Natural Science Foundation (Grant No. 7162089).
