Abstract
Genotoxicants can be identified as aneugens and clastogens through a micronucleus (MN) assay. The current high-content screening-based MN assays usually discriminate an aneugen from a clastogen based on only one parameter, such as the MN size, intensity, or morphology, which yields low accuracies (70–84%) because each of these parameters may contribute to the results. Therefore, the development of an algorithm that can synthesize high-dimensionality data to attain comparative results is important. To improve the automation and accuracy of detection using the current parameter-based mode of action (MoA), the MN MoA signatures of 20 chemicals were systematically recruited in this study to develop an algorithm. The results of the algorithm showed very good agreement (93.58%) between the prediction and reality, indicating that the proposed algorithm is a validated analytical platform for the rapid and objective acquisition of genotoxic MoA messages.
Background
The currently used micronucleus (MN) mode of action (MoA) detection method is usually based on MN parameters such as the MN size, 1 fluorescence intensity, 2 and morphology. 3 However, these methods employ a single MN parameter. Because each of these parameters might contribute to the results, 1 –3 single parameter determination may yield lower accuracy than multiple parameter evaluation. Current studies usually set a threshold; for instance, Westerink et al. 4 set a threshold for the fraction area of 0.17 to determine the MoAs, but this value might not be applicable to data from other laboratories because different cultivation environments or cell lines might generate different fractional areas. 4,5 In addition, current methods cannot be automated with high-content screening (HCS or another automated form) to assess MoAs. 4 In most biomonitoring studies, large quantities of samples are prepared by different laboratories and in parallel. Thus, the development of an automated objective algorithm that can yield comparative results is important. Therefore, we proposed a Bayesian k-means algorithm that can maximize hidden variables, marginalize the random parameter irregularity in large data sets and, consequently, improve the precision of the results by accurately depicting the cell information.
k-Means clustering is a cluster analysis methodology that aims to partition n observations into k clusters such that each observation belongs to the cluster with the nearest mean. 6 In our study, we first tested various chemicals to obtain MN parameter data and then used the proposed k-means algorithm to partition the MN parameters into a defined set of discrete clusters. In this process, the value of k was determined using the testing data such that it maximized the similarity among the MN parameters within clusters and the dissimilarity among the clusters. The main advantage of k-means clustering is its ability to consider multiple parameters. The final result of the algorithm is a set of k clusters with similar MN MoAs. k-Means clustering was combined with the Bayesian method, which has become an efficient learning algorithm. 7
Materials and methods
Chemicals
Twelve clastogens and eight aneugens representing a broad range of genotoxic activities were selected from the literature (Table 1). All test chemicals were purchased from reliable sources and were of the highest commercially available grade. The reagents and chemical solution were prepared based on published methods 27 with slight modifications.
List of chemicals.

CAS: Chemical Abstracts Service.
Cell treatment
The human gastric cancer cell line MGC-803 was purchased from the Cell Culture Center, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences, and the School of Basic Medicine of Peking Union Medical College, Beijing, China.
Phenol red-free Dulbecco’s modified Eagle medium (DMEM; Lot 17205072 R; Corning cellgro®, Corning, New York, USA) was employed for routing culture, and cells were cultivated in complete culture medium at 37°C in a carbon dioxide (CO2) incubator under an atmosphere with 5% CO2 (Sanyo, Fukuoka-ken, Japan). The complete culture medium consisted of DMEM supplemented with 10% fetal calf serum (Gibco, California, USA) and penicillin/streptomycin (100 IU/mL–1; Gibco, California, USA).
To ensure adequate attachment, exponentially growing cells were plated at a density of 9000 cells/well in flat-bottomed tissue culture-treated 96-well plates (CellCarrier®; PerkinElmer, Hamburg, Germany) 12 h prior to treatment with the test chemicals.
At the time of exposure, the test chemicals were added from 102 stock solutions to obtain a solvent concentration of 1% (v/v). Logarithmically growing cells were exposed to four concentrations of each chemical. The cell exposure treatments were performed in duplicate for a duration corresponding to twice the normal cell doubling time (24 h).
After 24 h of treatment, the exposure culture medium was replaced with 100 μL of a fluorophore mixture containing Hoechst 33342 (1 μM, product ID: 4082 S; Cell Signaling Technology, Danvers, Massachusetts, USA) and 3,3′-dioctadecyloxacarbocyanine (5 μM, product ID: D4292; Sigma-Aldrich, St Louis, Massachusetts, USA) in DMEM. These two dyes were used for staining nuclei and mitochondria, respectively, and the cells were incubated for 20 min at 37°C. The fluorophore mixture was then discarded, and 20 μL of phosphate-buffered saline was added.
Quality controls
Duplicate wells of four concentrations of each chemical in three repeat assays were scored, and controls were run alongside each of the chemicals. Due to the large number of cases, 2,3 a robust, dose-dependent occurrence of MNs was considered appropriate for the assay. The MoA signatures of the chemicals that were identified as inducing significant elevations to 150% MNs were examined. Each chemical was inspected to ensure considerable induction of MNs (greater than or equal to 1.5-fold increase over the mean solvent control) at concentrations that did not distinctly cause excessive toxicity. First, the cytotoxicity was calculated using the following equation

where N samples is the number of cells in the chemical treated wells and N solvent control is the number of cells in the solvent control wells. 28 The chromosome damaging effect was determined in terms of the MN frequency (MN%) and calculated using the following equation:

where N cell with MN is the number of cells with MNs and N T is the total number of selected cells, that is, the number of cells = selected nuclei (all cells with the exception of necrotic/apoptotic cells). 28
HCS scoring
An Operetta™ HCS instrument with Harmony software version 3.1 (PerkinElmer) and a 20_NA objective was initially employed to image and analyze the stained cells. The image analysis was performed based on a published method 4 with slight modifications. First, a module for the detection of normal cells was employed. Using this module, the intensity, area, and roundness of the nuclei were then analyzed to exclude necrotic/apoptotic cells; and the membrane area was subsequently analyzed to exclude false MNs. Second, a module for detecting MNs in normal cells was employed, and a machine learning algorithm was utilized to identify MNs. MN data (e.g. fraction of nuclear intensity, roundness, and area) were collected to construct our MoA analysis algorithm. An example of the analysis of MGC-803 cell images is shown in Figure 1, which displays the main steps in the process and the main steps of the process used for MN localization.

Main steps of HCS scoring. (a) Identification of nuclei; (b) selection of nuclei; (c) identification of the cytoplasm; (d) identification of the MN; (e) feature extraction MN localization. MN: micronucleus; HSC: high-content screening.
Seventeen fields per well were selected for the analysis. 29 The control wells contained 300–500 cells in each field (the cell number in the dosing wells was smaller due to cytotoxicity but was not less than half of that in the control well to avoid MN caused by cytotoxicity). 30
The MoA-related signatures obtained using Harmony software version 3.1 are listed in Table 2.
MN parameters.

MN: micronucleus.
Algorithm
The main concept of the algorithm is shown in Figure 2.

(a) The plot points represent the MNs, and the boxes represent the 96-well plates. The two wells on the left were exposed to a clastogen, whereas the others were exposed to an aneugen. (b) All of the MN parameters values were input into the algorithm. (c) k-Means clustering pre-clustered the MN parameter values. (d) Three central points were generated after the k-means clustering of all of the wells. (e) The statistical characteristics of the clusters of aneugens and clastogens were calculated. (f) A well containing an unknown sample was analyzed. (g) The MNs were clustered to the three central points shown in Figure 2(d) based on their parameter values. (h) The characteristics of the clusters shown in Figure 2(g) were statistically analyzed and then compared to clusters of known aneugens and clastogens to calculate (with a Bayesian formula) the aneugen probability of the interested chemical. MN: micronucleus.
A version of the k-means algorithm was implemented based on a description by Maggioli et al. 31 in Python version 2.7. The data were randomly divided into a training set and a testing set (at a ratio of 9:1) to avoid overfitting. 32 The algorithms were repeated 20 times for each setting (one MN parameter or more), each time with a different combination of the MN data sets for training and testing. The algorithm was executed as follows. First, Bayesian k-means clustering was performed using a single MN parameter. Then, to evaluate whether the multiple parameter model would improve the accuracy, an iterative forward-stepping approach was used.
We defined accuracy (precision rate) using the following equations:
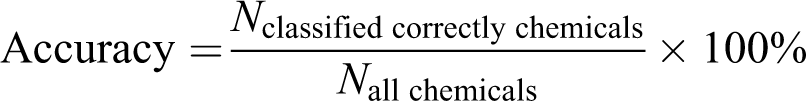
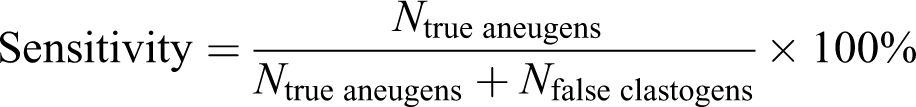

For example, as shown in Figure 3, one parameter (e.g. intensity) data of the MN in a 96-well plate were incorporated into the algorithm, and the aneugen probability of the well (predictor variables) was calculated using our method. The chemical in a well was classified as an aneugen by the algorithm if the aneugen probability was greater than the threshold of 50%. The aneugen probabilities of all wells of a chemical were predicted. If more than 50% of the wells of a chemical were classified as an aneugen, the chemical was predicted to be an aneugen.

Results of aneugen probabilities obtained using a single parameter with the Bayesian k-means algorithm. A series of probabilities is plotted for each chemical, and each plot point represents a well in a 96-well plate. The parameters (such as intensity, specified at the top of the figures) were incorporated into the algorithm, and then, based on the intensity data corresponding to the selected MNs in this 96-well plate, the aneugen probability of a well was calculated. The plot points on the top half (greater than 50%) refer to the wells that were exposed to an aneugen according to the Bayesian k-means clustering results, whereas the plot points on the bottom half refer to the wells that were exposed to a clastogen. MN: micronucleus.
Results
Parameter settings for Bayesian k-means clustering
To obtain effective MN MoA data for constructing the algorithm, 20 MN-positive chemicals were tested. The data were then used in the algorithm. In this section, we present the results obtained using a single parameter to discriminate aneugens and clastogens, as in previous studies 1 –3 (Figure 3), and compare these results with the results obtained using multiple parameters. The predictor variables (aneugen probabilities) were graphed for each of the tested chemicals (graphs are presented in Figures 3 and 4, and data are listed in Table 3).

Aneugen probabilities obtained using the multi-parameter Bayesian k-means algorithm. A series of probabilities is plotted for each chemical, and each plot point represents a well in a 96-well plate. The parameters (such as roundness, specified at the top of the figures) were incorporated into the algorithm, and then, based on the roundness data for the selected MNs in this 96-well plate, the aneugen probablity of a well was calculated. The plot points in the top half (greater than 50%) refer to the wells that were exposed to an aneugen according to the Bayesian k-means clustering, whereas the plot points in the bottom half refer to the wells that were exposed to a clastogen. MN: micronucleus.
Predictive accuracy for individual compounds when the algorithm was subjected to quantitative variables.

aPredictive accuracy for individual compounds =
In the best-case scenario, the aneugen probability of an aneugen should be higher than the threshold of 50% (aneugen plot point allocated to the area above the 50% line), whereas the clastogen dots should be allocated to the opposite area. Figure 3 and Table 4 indicate that the accuracy and detection rate of the single parameter method are not sufficient to discriminate aneugens from clastogens. Thus, we employed additional parameters to improve the accuracy of the prediction method. As determined using our method, a higher degree of predictive accuracy can be obtained (Figure 4) from the inclusion of additional cell information through the application of additional parameters (compared with Figure 3, most of the aneugens exhibited an aneugen probability above the 50% line, which means that the chemicals were classified correctly by our method), which improved the accuracy and detection rate compared with those obtained with a single parameter. All of the subset of six parameters were used for the clustering algorithm to attain the best performance (data not shown), and the final results were surprisingly accurate. First, we tested genotoxic chemicals to obtain the MN data and randomly divided the data into a training set and a testing set at a ratio of 9:1 to train and test the algorithm. The optimal performance, namely a precision rate of 93.69%, was obtained with the inclusion of the following: distance, compactness, and roundness (Figure 4 and Table 4).
Accuracy obtained with one parameter (documented method) and multiple parameters (our method).a

aThe percentages represent the average probabilities after a reasonable number of iterations that the 20 chemicals were correctly classified.
Algorithm progress
Some clastogen signatures are similar to those of aneugens. Many algorithms, including the k-means algorithm, cannot correctly distinguish aneugens and clastogens. A highly divided k cluster corresponds to highly accurate clustering. However, we found that the accuracy with k = 6 is lower than the accuracy with k = 5 (data not shown). We hypothesize that a high number of clusters will result in the generation of some small clusters and that the MNs of these small clusters do not sufficiently represent the features of this cluster with sufficient accuracy. To improve the shortcomings of this classification process, k-means clustering was combined with the Bayesian classifier.
The percentages in Figure 5 represent the calculated probability that a chemical is an aneugen. According to the Bayes definition, aneugens have aneugen probabilities that exceed 50%. Figure 5 shows that the hybrid classifier exhibited a higher accuracy and detection rate because the aneugen-exposed plot point that was misclassified as having been exposed to clastogens (the aneugen probabilities of most aneugen plot point were lower than 50%, left image) using k-means clustering was correctly reclassified as aneugens (the aneugen probabilities of most aneugen plot point were higher than 50%, right image) using Bayesian k-means clustering. These results are attributed to the k-means clustering technique, which was utilized as a preclassification tool. The clustering technique used a preclassification component to group similar data into respective classes, which helped the proposed hybrid learning approach produce better results than a single classifier. The hybrid approach also enables the reclassification of misclassified data during the first stage, which improves the accuracy and detection rate.
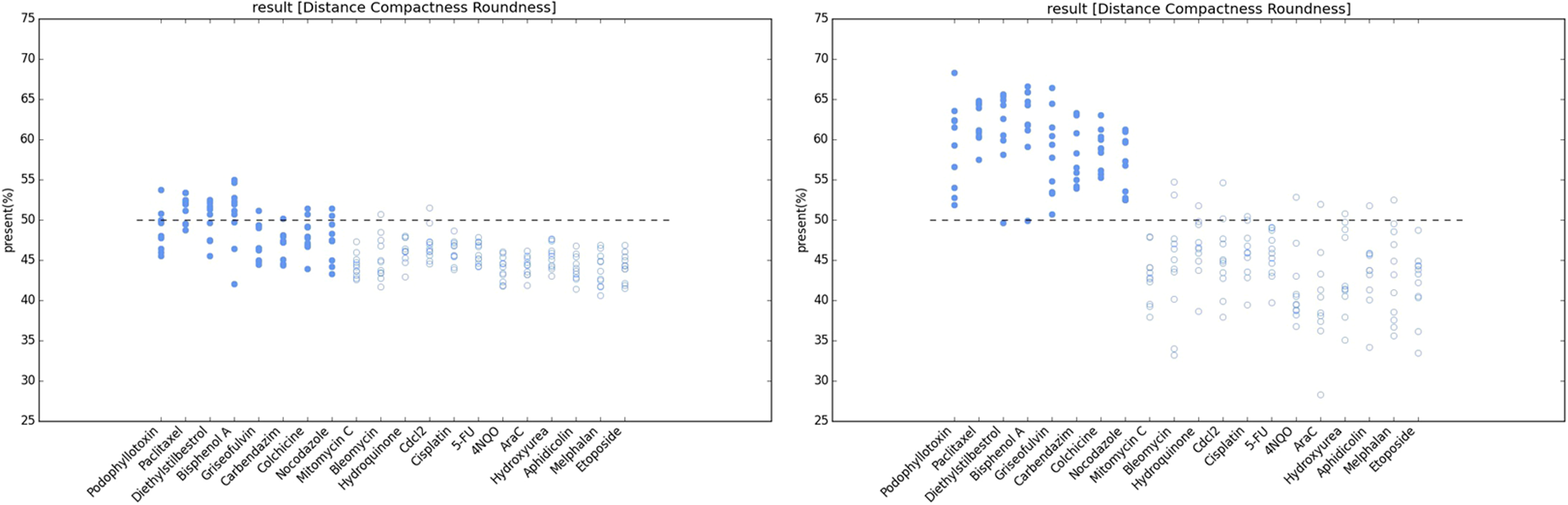
Results of k-means (left image) and Bayesian k-means (right image) clustering. A series of probabilities is plotted for each chemical, and each plot point represents a well in a 96-well plate. The parameters roundness, distance and compactness were incorporated in the algorithm, and then, based on data of the three parameters for the selected MNs in this 96-well plate, the aneugen probability of a well was calculated. The plot points in the top half (greater than 50%) refer to the wells that were exposed to an aneugen according to the algorithm, whereas the plot points in the bottom half refer to the wells that were exposed to a clastogen. MN: micronucleus.
Discussion
Several advantages of an automated MoA detection algorithm were demonstrated in our study. First, using the Bayesian k-means algorithm to process the data, which can reveal more complex correlations between the parameters and the contribution degree of various patterns, the MoA distinction obtains competitive results compared with current methods, such as those using a single parameter, namely, area, intensity, or morphology (Table 4).
Second, this approach enables the application of this automated technique not only to HCS but also to other experimental approaches (e.g. flow cytometry) that can provide sufficient MN characteristics.
Third, early high-throughput screening should constantly strive toward maximum sensitivity and minimum cost. This method is not only much cheaper than other MN MoA distinction methods, such as multiple parameter models using Western blotting, 33 but also reduces manual work.
Thus, the proposed approach is of significant utility. While this approach appears optimal based on the results from the current data set, it is important to recognize that the analyses considered a single cell line, one treatment schedule, and only 20 chemicals. Each laboratory should parametrize their own historical data and employ this algorithm as a starting point to determine their laboratory- and cell line-specific MoA signatures.
Footnotes
Acknowledgements
This research was financially supported by National Natural Science Foundation of China(21437006), Frontier Science Key Program of the Chinese Academy of Sciences (QYZDY-SSW-DQC004) and National Natural Science Foundation of China (51290283).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
