Abstract
The malignancy of non–small cell lung cancer (NSCLC) largely results from its invasive manner. Secreted frizzled-related proteins (SFRPs) have been recently found to suppress the invasiveness of some cancers. On the other hand, the methylation of SFRPs increases protein degradation to reduce the activity of SFRPs, resulting in increased tumor cell invasion and cancer metastasis. However, the role of SFRPs in the invasion of NSCLC has not been reported. Here we analyzed the regulation of SFRPs in NSCLC cells and its effects on cell invasion. We found that SFRP2 mRNA was significantly decreased and methylation of SFRP2 gene was significantly increased in NSCLC tissue, compared to the paired adjacent nontumor tissue. Moreover, SFRP2 expression was significantly decreased in NSCLC cell lines. In NSCLC cell lines, the SFRP2 expression would be restored by the demethylation of SFRP2 gene with 5′-aza-deoxycytidine in NSCLC cell lines, at the levels of both mRNA and protein. Thus, the cell invasion would be suppressed. Furthermore, the demethylation of SFRP2 gene appeared to inhibit Zinc Finger E-Box Binding Homeobox 1 (ZEB1) and matrix metallopeptidase 9 (MMP9), two key factors that enhance NSCLC cell invasion. Thus, SFRP2 may inhibit NSCLC invasion by suppressing ZEB1 and MMP9, while its methylation promotes NSCLC invasion.
Keywords
Introduction
Non–small cell lung cancer (NSCLC) is the most common malignant lung tumor in humans, 1 –3 and its poor prognosis largely results from local invasion and distal metastasis. 4 –15
Recently, with the rapid development of chemotherapy, the prognosis of patients with nonmetastatic NSCLC has been substantially improved. However, the survival rate for patients with metastatic NSCLC remains low. Hence, strategies that target the NSCLC invasion, migration, and metastases are very critical for NSCLC therapy.
The family of secreted frizzled-related proteins (SFRPs) has been recently found to suppress the invasiveness of some cancers. 16 –21 On the other hand, the methylation of SFRPs increases protein degradation to reduce the activity of SFRPs, resulting in the increased tumor cell invasion and cancer metastasis. Till now, five human SFRPs have been detected. 22 –25
SFRPs have been shown to have a variety of functions, for example, inhibition of activation of canonical Wnt signaling. SFRPs are found to downregulate in several cancers, which are often indicators of poor prognosis. 22 –25 Indeed, in recent years, accumulating evidence suggests SFRPs as tumor suppressors, due to their silence in cancer via promoter hypermethylation. 26 –28 However, whether the methylation of SFRP2, a member of SFRPs, may be involved in the pathogenesis of NSCLC remains unknown.
Here we investigated the expression levels of SFRP2 in NSCLC specimens and NSCLC cell lines. The demethylation of SFRP2 gene was induced by drugs in NSCLC cell lines, then the effects of demethylation on SFRP2 levels and cell invasiveness were analyzed.
Materials and methods
Patient tissue specimens
Thirty NSCLC patients were included in the current study. The specimens from these patients were histologically diagnosed at The Fourth People’s Hospital of Ji’nan from 2011 to 2015. The NSCLC tissue versus nontumor tissue (NT) was determined by gross observation and confirmed by histology. For the use of these clinical materials for research purposes, prior patient’s consents and approval from the Institutional Research Ethics Committee were obtained.
NSCLC cell lines and primary osteoblasts
The malignant NSCLC cell lines A549, Calu-3, SK-LU1, and a primary lung epithelial cell (NL20), were obtained from the American Type Culture Collection (ATCC, Rockville, Maryland, USA) and cultured at 37°C with 5% CO2. The culture media were Roswell Park Memorial Institute medium (RPMI) 1640 medium with 15% fetal bovine serum (R&D systems, Minneapolis, Minnesota, USA). The cells were regularly analyzed for the absence of mycoplasma contamination.
Cell invasion assay
The cell invasion assay was performed with commercial kit (5 μm pore size; Cell Biolabs, San Diego, California, USA). The cells were seeded on the upward side of the chamber and the Dulbecco’s Modified Eagle’s Medium (DMEM) containing serum was added in the downward side of the chamber. After incubating at 37°C for 24 h, the cells could be observed to migrate through the pores in the membrane. The cells in the downward side of the chamber would be removed and collected. After lysis, the cells would be determined with CyQuant GR dye (Invitrogen, Hongkong, China) and read with a microplate reader (FluoStar, Fluorescence channel: 485/520 nm). The data was processed with the software of microplate reader.
Quantitative PCR (RT-qPCR)
Total RNA was extracted with RNeasy kit (Qiagen, Hilden, Germany). For complementary DNA (cDNA) synthesis, cDNA was randomly primed from 2 μg of total RNA with the Omniscript reverse transcription kit Applied Biosystems (ABI). RT-qPCR was subsequently performed in triplicate using the QuantiTect SyBr green PCR system (Qiagen). All primers were purchased from Qiagen. Data were collected and analyzed using 2–△△Ct method. Values of genes were first normalized against α-tubulin and then compared to the experimental controls.
Western blot
Total protein was extracted from cell lysates and homogenized in radioimmunoprecipitation assay buffer (RIPA lysis buffer; Sigma-Aldrich, San Diego, California, USA) on ice. The supernatants were collected after centrifugation at 12,000 × g at 4°C for 20 min and the total protein concentration was quantified with the bicinchoninic acid assay (BCA assay; Bio-rad, China). The total protein solutions were heat-degenerated and separated on Sodium Dodecyl Sulfate (SDS) gels. After separation, the proteins were electro-transferred on a Polyvinylidene fluoride (PVDF) membrane. The PVDF membrane was blocked with 5% skim milk. Then, the first antibodies against target proteins were added and incubated. The first antibodies were rabbit anti-SFRP2 (Abcam, Cambridge, Massachusetts, USA) and anti-α-tubulin (Cell Signaling, San Jose, California, USA). α-tubulin was used as protein loading controls. The PVDF membrane was incubated again with HRP labeled second antibody (Jackson ImmunoResearch Labs, West Grove, Pennsylvania, USA). The enhanced chemiluminescent substrate was added and read with autoradiograms. Images shown in the figures were representative from five individuals. Densitometry of Western blots was quantified with NIH ImageJ software (Bethesda, Massachusetts, USA). The protein levels were first normalized to α-tubulin and then normalized to experimental controls.
Methylation-specific polymerase chain reaction analysis
Methylation-specific primers for SFRP2 were designed. Forward primer: 5′-GGGTCGGAGTTTTTCGGAGTTGCGC-3′. Reverse primer: 5′-CCGCTCTCTTCGCTAAATACGACTCG-3′. The length of the PCR product was 138 bp. For the controlled non-methylation-PCR, forward primer: 5′-TTTTGGGTTGGAGTTTTTTGGAGTTGTGT-3′ and reverse primer: 5′-AACCCACTCTCTTCACTAAATACAACTCA-3′. The length of the PCR product was 145 bp. The PCR conditions were as follows: 5 min at 95°C; 45 s at 94°C for 35 cycles; 30 s at 50°C; 45 s at 72°C; and final 10 min at 72°C. The 138-bp methylation-specific polymerase chain reaction (MSP) product was isolated using electrophoresis in a 1.5% agarose gel and observed with an UV gel imaging system (ImageQuant 350; GE Healthcare Co., Little Chalfont, UK).
Bisulfite sequencing analysis of promoter methylation
The methylated CpGs (shorthand for 5′ --C --phosphate --G --3′, that is, cytosine and guanine separated by only one phosphate; phosphate links any two nucleosides together in DNA) were avoided in the design of bisulfite sequencing (BS) primers. The DNA samples were sulfonated with sodium bisulfite. For the BS product, the forward primer: 5′-TTGTTTGTAAGGTAATTATTAG-3′. Outside reverse primer: 5′-ATTTTCTTAACCTTTTTTATAC-3′ and inside reverse primer: 5′-AAACAAAAAAAAAACCAAAC-3′. The length of the product was about 200 bp. The PCR amplification conditions were as follows: 5 min at 95°C; 45 s at 94°C for 35 cycles, 30 s at 52°C; 30 s at 72°C; and final 10 min at 72°C. The 200-bp amplification product was isolated with electrophoresis in a 1.5% agarose gel and visualized under UV light. The PCR product was sequenced (Beijing Genomics Institute, Beijing, China). The sequences of amplification products from the samples were compared with sequences of completely sulfonated promoter target (JellyFish 1.3; Field Scientific, Madison, Wisconsin, USA). The methylation level was calculated as (mC/C – G) × 100%.
Statistics
All statistical analyses were carried out using the GraphPad Prism 6.0 statistical software (GraphPad Software, Inc., La Jolla, California, USA). All values in cell and animal studies are depicted as mean ± standard deviation and are considered significant if p < 0.05. All data were statistically analyzed using one-way ANOVA with a Bonferroni correction, followed by Fisher’s exact test for comparison of two groups. For analyzing clinic samples, power analysis was performed.
Results
SFRP2 levels decrease and SFRP2 gene methylation increases in NSCLC specimens
We examined the levels of SFRP2 in NSCLC specimens. The SFRP2 mRNA was observed to be significantly decreased in malignant NSCLC tumors as compared, in contrast to the paired adjacent NT, shown by individual levels (Figure 1(a)), and by mean ± SD (Figure 1(b)). Moreover, SFRP2 protein levels were significantly decreased in NSCLC tissue, compared to NT (Figure 1(c)). Next, we checked SFRP2 gene methylation in NSCLC specimens and found that the methylation of SFRP2 gene was significantly increased in NSCLC, compared to NT (Figure 1(d)). Hence, SFRP2 levels decrease and SFRP2 gene methylation increases in NSCLC specimens.

SFRP2 levels decrease and SFRP2 gene methylation increases in NSCLC specimens. (a and b) SFRP2 mRNA levels were analyzed in NSCLC tissue as compared to the paired adjacent NT, shown by individual levels (a), and by mean ± SD (b). (c) SFRP2 protein levels were analyzed in NSCLC tissue versus NT by Western blot. (d) The quantification of SFRP2 gene methylation by BS in specimens. *p < 0.05. N = 30. SFRP2: secreted frizzled-related protein 2; NSCLC: non–small cell lung cancer; NT: nontumor tissue; SD: standard deviation; BS: bisulfite sequencing.
Demethylation of SFRP2 gene is induced in NSCLC cells
Next, we examined the levels of SFRP2 and gene methylation in NSCLC cell lines, SK-LU1, Calu-3 and A549, using NL20, a normal human epithelial cell as a control. We found that all the NSCLC cell lines had decreased SFRP2 levels, compared to NL20 cells (Figure 2(a)), consistent with clinical findings. Then, we used 5′-aza-deoxycytidine (5-aza-dCyd) to treat these NSCLC cell lines. We found that demethylation of SFRP2 gene was efficiently induced by 5-aza-dCyd in NSCLC cells, shown by representative MSP chain reaction gels (Figure 2(b)), and by quantification (Figure 2(c)). Thus, demethylation of SFRP2 gene is induced by 5-aza-dCyd in NSCLC cells.

Demethylation of SFRP2 gene is induced by 5-aza-dCyd in NSCLC cells. (a) SFRP2 mRNA levels were analyzed in NSCLC cell lines, SK-LU1, Calu-3 and A549, and normal human lung epithelial cells (NL20). (b and c) 5′-aza-deoxycytidine (5-aza-dCyd) was used to treat NSCLC cell lines, SK-LU1, Calu-3 and A549, and NL20. The MSP was done, shown by quantification (a), and by representative MSP gels (b). *p < 0.05. N = 5. SFRP2: secreted frizzled-related protein 2; NSCLC: non–small cell lung cancer; MSP: methylation-specific polymerase chain reaction.
Demethylation of SFRP2 gene increases SFRP2 expression in NSCLC cells
We found that demethylation of SFRP2 gene by 5-aza-dCyd in NSCLC cell lines SK-LU1, Calu-3, and A549 significantly increased SFRP2, at both mRNA (Figure 3(a)) and protein (Figure 3(b)) levels. Thus, demethylation of SFRP2 gene increases SFRP2 expression in NSCLC cells.
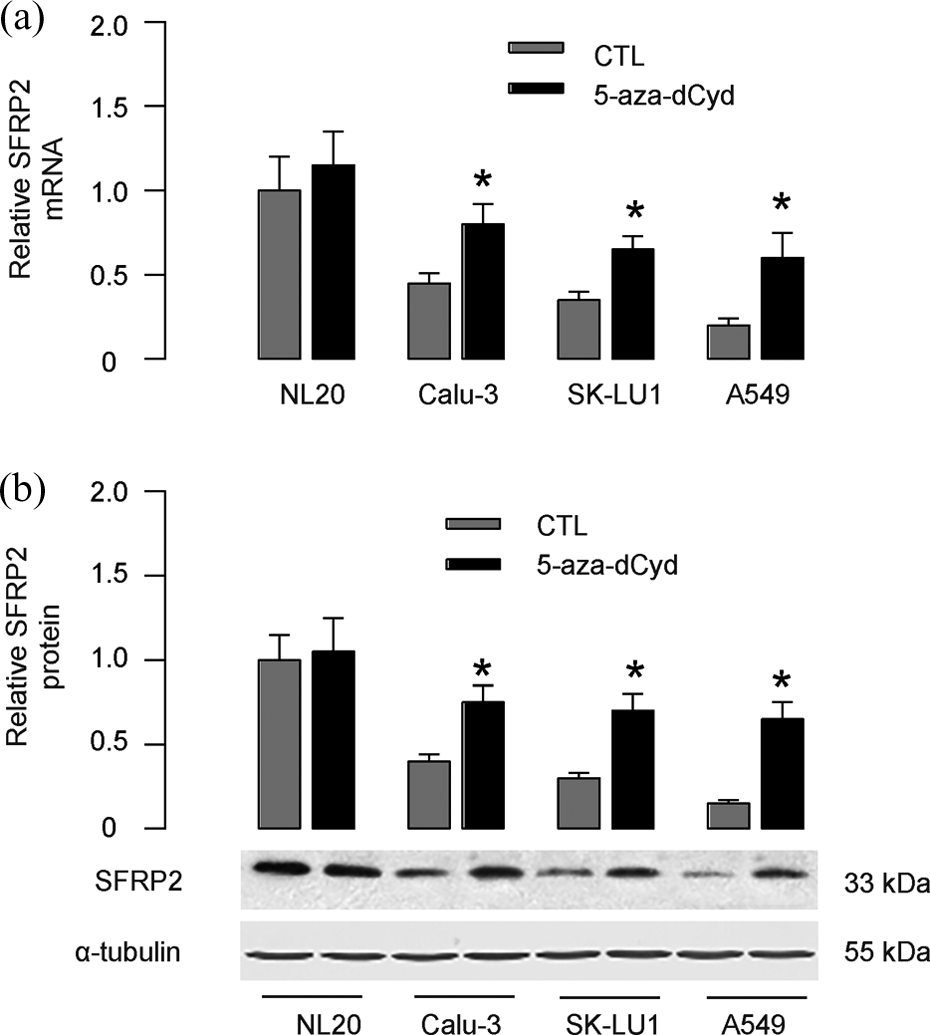
Demethylation of SFRP2 gene increases SFRP2 in NSCLC cells. (a and b) 5′-aza-deoxycytidine (5-aza-dCyd) was used to treat NSCLC cell lines, SK-LU1, Calu-3 and A549, and the NL20. The SFRP2 levels were analyzed by RT- qPCR (a) and Western blot (b). *p < 0.05. N = 5. SFRP2: secreted frizzled-related protein 2; NSCLC: non–small cell lung cancer.
Demethylation of SFRP2 gene suppresses NSCLC cell invasion
Next, the effects of demethylation of SFRP2 on the invasiveness of NSCLC cells were detected with cell invasion assay. We found that demethylation of SFRP2 gene by 5-aza-dCyd in NSCLC cell lines SK-LU1, Calu-3, and A549 significantly suppressed cell invasion, shown by quantification (Figure 4(a)), and by representative images (Figure 4(b)). Thus, SFRP2 gene demethylation suppresses NSCLC cell invasion.

Demethylation of SFRP2 gene inhibits NSCLC cell invasion. (a and b) 5′-aza-deoxycytidine (5-aza-dCyd) was used to treat NSCLC cell lines, SK-LU1, Calu-3, and A549. The cell invasion was analyzed in a transwell cell invasion assay, shown by quantification (a), and by representative images (b). *p < 0.05. N = 5. SFRP2: secreted frizzled-related protein 2; NSCLC: non–small cell lung cancer.
Demethylation of SFRP2 gene decreases ZEB1 and MMP9 in NSCLC cells
Finally, we aimed to analyze the effects of SFRP2 gene demethylation on the factors that regulate NSCLC cell invasiveness. We found that demethylation of SFRP2 gene decreased the levels of ZEB1 and matrix metallopeptidase 9 (MMP9) by Western blot (Figure 5). Together, our data suggest that SFRP2 may thus function as a suppressor for NSCLC invasion, and the methylation of SFRP2 gene may promote the invasiveness of NSCLC through ZEB1 and MMP9.

Demethylation of SFRP2 gene decreases ZEB1 and MMP9 in NSCLC cells. Demethylation of SFRP2 gene decreased the levels of ZEB1 and MMP9 by Western blot in NSCLC cell lines, SK-LU1, Calu-3, and A549. Representative Western blots were shown. N = 5. NSCLC: non–small cell lung cancer; SFRP2: secreted frizzled-related protein 2; MMP9: matrix metallopeptidase 9.
Discussion
Previous studies have mainly studied the role of SFRP2 in colorectal carcinoma, showing that extensive methylation of the SFRP2 promoter was present primarily in colorectal carcinoma. 29 –35 However, a role of SFRP2 gene methylation in the tumorigenesis of NSCLC is unknown. Here, we investigated the expression levels of SFRP2 in NSCLC specimens. Since the sample size is not big, the power of the investigation was evaluated before further statistical analysis. We found that the levels of both SFRP2 mRNA and protein were significantly decreased, and this downregulation of SFRP2 levels may result from the promoter methylation.
Since the SFRP2 expression was also significantly decreased in all examined NSCLC cell lines, SK-LU1, Calu-3, and A549, but not in the normal lung epithelial cells NL20, these data suggest that SFRP2 gene promoter may exist in a relative low methylation state in normal lung tissue, while significantly methylated when the cells undergo phenotypic changes toward malignant phenotype. Thus, DNA methylation appears to be a regulation mechanism on SFRP2 function, which contributes to the tumorigenesis and development of NSCLC. Therefore, induction of demethylation of the SFRP2 promoter may prevent the development and progression of NSCLC.
Based on these clinical findings, we were prompted to analyze the effects of demethylation of SFRP2 on NSCLC cell invasiveness. The demethylation of SFRP2 gene by 5-aza-dCyd in NSCLC cell lines restored SFRP2, at both mRNA and protein levels, which led to suppression of cell invasion. Furthermore, the demethylation of SFRP2 gene appeared to inhibit expression of ZEB1 and MMP9. The transcription factor ZEB1 possesses zinc-finger clusters in both the N-terminal and C-terminal regions and a homeodomain in the central region. ZEB1 is a key promoter of epithelial–mesenchymal transition, possibly through its specific binding to the promoter of E-cadherin to repress its expression. 36 MMP9 is an important matrix proteinase that degrades a basement membrane protein collagen type IV and has been shown to be involved in the metastatic spread of various tumors, including NSCLC. 37 –40 Here, we specifically defined these two factors as targets of SFRP2, and our data suggest that SFRP2 may function as a suppressor for NSCLC invasion, and the methylation of SFRP2 gene may promote the invasiveness of NSCLC through ZEB1 and MMP9.
The present study also had limitations. The molecular mechanisms were analyzed in cultured NSCLC cells, but not primary NSCLC tissue. The studies using animal model may further provide evidence for the conclusions drawn here. In addition, the precise molecular mechanisms underlying the control of SFRP2 gene promoter methylation still require further investigation.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
