Abstract
MicroRNA (miRNA) has been reported to play important roles in regulating drug-induced liver injury. Ethyl acetate extract isolated from rhizoma Dioscoreae bulbifera (EF) has been reported to induce hepatotoxicity in our previous studies. This study aims to observe the altered liver miRNA profile and its related signalling pathway involved in EF-induced hepatotoxicity. Serum alanine/aspartate aminotransferase assay showed that EF (450 mg/kg)-induced hepatotoxicity in mice. Results of miRNA chip analysis showed that the expression of eight miRNAs was up-regulated and of other nine miRNAs was down-regulated in livers from EF-treated mice. Further, the altered expression of miR-200a-3p, miR-5132-5p and miR-5130 was validated using real-time polymerase chain reaction (PCR) assay. There were total seven predicted target genes of miR-200a-3p, miR-5132-5p and miR-5130. Only one kyoto encyclopedia genes and genomes pathway was annotated using those target genes, which is protein processing in endoplasmic reticulum (ER). Furthermore, liver expression of DnaJ subfamily A member 1, a key gene involved in protein processing in ER based on the altered miRNAs, was increased in EF-treated mice. In conclusion, the results demonstrated that EF altered the expression of liver miRNA profile and its related signalling pathway, which may be involved in EF-induced hepatotoxicity.
Keywords
Introduction
MicroRNAs (miRNAs) are endogenously expressed small non-coding RNA molecule, containing about 22 nucleotides. 1 A large number of studies have shown that miRNAs have crucial roles in triggering the degradation and translational repression of cellular target mRNA. 1 –3 Accumulating studies have already demonstrated the critical role of miRNAs in many human physiological and pathological processes, including cell proliferation, cell differentiation, apoptosis and cellular immune responses. 1 –4
Drug-induced liver injury (DILI) is a type of liver diseases caused by the drug itself or its metabolites. According to the report of World Health Organization, DILI has risen to the fifth cause of death in the world. 5 A prospective study of acute liver failure at 17 tertiary care centres in the United States demonstrated that approximately above 50% of acute liver failure in clinic was caused by drugs. 6,7 With the wide acceptance of traditional Chinese medicines in the world, hepatotoxicity induced by herbal and dietary supplements has been a rising cause for DILI. 6,8
The rhizome of Dioscorea bulbifera L. is mainly used to treat thyroid diseases and a variety of cancers including gastric cancer, liver cancer and breast cancer in clinic. 9 However, the toxicity induced by D. bulbifera has severely limited its clinical application in cancer treatment. It has been reported that the main toxic target organ of D. bulbifera is liver, and its exposure could lead to severe liver damage, including liver swelling, fatty degeneration and even death. 10 According to the previous studies in our research group, the ethyl acetate extract isolated from D. bulbifera (EF) and diosbulbin B, a main compound isolated from D. bulbifera, caused serious hepatotoxicity in experimental animals. 11 –13 Moreover, such hepatotoxicity is mainly oxidative stress related and inflammatory liver injury. 11 –14 However, the concrete mechanism engaged in the hepatotoxicity induced by D. bulbifera is still unclear. In addition, there is no report about the involvement of miRNA in regulating the hepatotoxicity induced by D. bulbifera.
In this study, the altered liver miRNA profile in mice treated with EF was observed, and some altered miRNAs were found and further validated using real-time polymerase chain reaction (PCR) assay. Moreover, the predicted target genes based on the differentially expressed miRNAs were annotated with kyoto encyclopedia genes and genomes (KEGG) pathways and the expression of critically involved target gene was further analysed.
Materials and methods
Chemical compounds and reagents
The rhizome of D. bulbifera was collected in Qingyang, Anhui Province and authenticated by Prof. Shou-Jin Liu (Anhui College of Traditional Chinese Medicine, Anhui, China). The voucher specimens were deposited in the herbarium of Institute of Chinese Materia Medica, Shanghai University of Traditional Chinese Medicine. The preparation of EF has already been reported in our previous published articles, and the content of diosbulbin B in EF was 13.72%. 14,15
The analytic kits for alanine aminotransferase (ALT) and aspartate aminotransferase (AST) assay were purchased from Nanjing Jiancheng Bioengineering Institute (Nanjing, China). Trizol reagent was purchased from Life Technology (Carlsbad, California, USA). MiRNeasy mini kit was obtained from Qiagen (Hilden, German). miRCURY LNATM array (version18.0) was purchased from Exiqon (Vedbaek, Denmark). miScript PCR Starter kit was obtained from Qiagen (Hilden, German). PrimeScriptRT Master Mix and SYBR Premix Ex TaqTM were purchased from Takara (Shiga, Japan). Antibody for immunoblotting anti-DnaJ subfamily A member 1 (Dnaja1) was purchased from Biobasic Inc (Shanghai, China).
Experimental animals
Specific pathogen-free male imprinting control region mice (weight: 20 ± 2 g) were purchased from Shanghai Experimental Animal Center of Chinese Academy of Sciences (Shanghai, China). The mice were fed with a standard laboratory diet and given free access to tap water, living in a controlled room temperature (22 ± 1°C) and humidity (65 ± 5%) with a 12-h light/12-h dark cycle. All animal experiments were performed according to the protocols approved by the Experimental Animal Ethical Committee of Shanghai University of Traditional Chinese Medicine.
Treatment of animals
The mice were randomly divided into four groups as following: (1) vehicle control group (n = 9), (2) EF (300 mg/kg body weight) group (n = 9), (3) EF (375 mg/kg body weight) group (n = 9) and (4) EF (450 mg/kg body weight) group (n = 9). Mice were orally administrated with different doses of EF or vehicle for once, and the doses were selected with minor modification according to our previous study. 14 Mice were killed at 24 h after EF administration, and blood and livers from each group were collected.
Serum biomarkers for liver injury
The blood samples obtained from mice of all groups were allowed to coagulate for 2 h. Serum was then isolated following centrifugation at 840 × g for 15 min. Serum ALT and AST activities were measured with kits purchased from Nanjing Jiancheng Bioengineering Institute (Nanjing, China) according to the manufacturer’s instructions.
miRNA microarray assay and data analysis
Total RNA from liver tissues of control and EF (450 mg/kg)-treated groups were harvested using Trizol and miRNeasy mini kit according to the manufacturer’s instructions. After having passed RNA quantity measurement using the NanoDrop 1000, the samples were labelled using the miRCURYTM array power labeling kit and hybridized on the miRCURY™ array (version 18.0). Following the washing steps, the slides were scanned using the Axon GenePix 4000B microarray scanner. Scanned images were then imported into GenePix Pro version 6.0 software (Axon) for grid alignment and data extraction.
Expressed data were normalized using the median normalization. After normalization, differentially expressed miRNAs were identified through fold change filtering. Only those miRNAs with the fold difference >2.0 were considered significant, and hierarchical clustering was performed to show different miRNA expression profiling among control and EF (450 mg/kg)-treated groups using MEV software (version 4.6, TIGR).
RNA isolation and real-time reverse transcriptase -PCR analysis
Liver total RNA was isolated using Trizol reagent according to the manufacturer’s instruction. The RNA content was determined and complementary DNA was synthesized using reverse transcriptase kit according to the instruction.
The primers for miRNAs were obtained from Sangon Biotech. (Shanghai, China), and the primer sequences used in this study are shown as follows: mmu-miR-1912-5p forward primer (5′-GTGCTCATTGCATGGGCTGT-3′), mmu-miR-34a-5p forward primer (5′ -GTGGCAGTGTCTTAGCTGGT-3′), mmu-miR-200a-3p forward primer (5′-GTAACACTGTCTGGTAACGATG-3′), mmu-miR-302a-3p forward primer (5′-TAAGTGCTTCCATGTTTTGGTGA-3′), mmu-miR-5129-5p forward primer (5′-ATGTGGGGGCATTGGTATTTTC-3′), mmu-miR-5132-5p forward primer (5′- GCGTGGGGTGGTGGACT-3′), mmu-miR-5130 forward primer (5′-CTGGAGCGCGCGGGC-3′), mmu-miR-652-5p forward primer (5′-CAACCCTAGGAGGGGGTG-3′) and RNU6-2 forward primer (5′-AACGCTTCACGAATTTGCGT-3′). Real-time PCR was performed using miScript PCR Starter kit according to the manufacturer’s instruction. The relative expression level of each miRNA was normalized to the level of RNU6-2.
The primes for Dnaja1 and actin were obtained from Sangon Biotech (Shanghai, China), and the primer sequences used in this study are shown as follows: Dnaja1 forward primer (5′-AGGCCGAGGTGGTAAGAAAG-3′) and reverse primer (5′-GCACTCCATGCACACTGACT-3′); actin forward primer (5′-TTCGTTGCCGGTCCACACCC-3′) and reverse primer (5′-GCTTTGCACATGCCGGAGCC-3′). Real-time PCR was performed using SYBR® Premix Ex TaqTM kit. Relative expression of target genes was normalized to actin, analysed by the 2−ΔΔCt method and given as ratio compared with the control.
Target prediction and pathway analysis
The target genes of altered miRNAs were predicted by the overlapped data from Mirbase prediction software (http://www.ebi.ac.uk), Miranda prediction software (http://www.microrna.org/microrna/home.do) and TargetScan prediction software (http://www.targetscan.org/). The predicted target genes were annotated with KEGG pathways using the DAVID gene annotation tool (http://david.abcc.ncifcrf.gov). The enrichment p-value of the PathwayID was used according to EASE method. 16
Protein extraction and Western-blot analysis
After treatment, liver tissues were homogenized in ice-cold lysis buffer containing 50 mM Tris (pH 7.5), 150 mM sodium chloride, 1 mM ethylenediaminetetraacetic acid, 20 mM sodium fluoride, 0.5% NP-40, 10% glycerol, 1 mM phenylmethylsulfonyl fluoride, 10 μg/ml aprotinin, 10 μg/ml leupeptin and 10 μg/ml pepstatin A. After centrifugation, protein concentration of the resulting supernatant was determined and normalized to equal amount of protein of each sample. Protein samples were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis and electrophoretically transferred onto PVDF membrane, and then the membranes were probed with appropriate combination of primary and secondary antibodies. Proteins in the membranes were visualized by enhanced chemiluminescence kits. The protein bands were quantified by the average ratios of integral optic density following normalization to the expression of internal control β-actin, and the results were further normalized to control.
Statistical analysis
Data were expressed as means ± standard error of the mean. The significance of differences between two groups was evaluated by Student’s t-test and between multiple groups was evaluated by one-way analysis of variance with LSD post hoc test. The value of p < 0.001 for miRNA, mRNA and protein expression and p < 0.01 for ALT and AST activity was considered as statistically significant.
Results
EF-induced hepatotoxicity in mice
As shown in Figure 1, EF (450 mg/kg) markedly elevated the activities of serum ALT (p < 0.01) and AST (p < 0.01) at 24 h after mice was orally administrated with EF. The increased percentage of ALT and AST induced by EF (450 mg/kg) is 976.3% and 447.3%, respectively. EF (300 mg/kg) had no obvious effect on the activities of serum ALT and AST. It seems that EF (375 mg/kg) had the tendency to increase the activities of serum ALT and AST, but such tendency had no significance (p > 0.05).

EF elevated serum ALT (a) and AST (b) activities in ICR mice. Data is shown as means ± SEM (n = 9), **p < 0.01 compared with control. ICR: imprinting control region; ALT: alanine aminotransferase; AST: aspartate aminotransferase; SEM: standard error mean.
Effects of EF on liver miRNA expression profile
The miRNA array was further used to analyse the effects of EF on liver miRNA expression profile. The results of hierarchical clustering analysis showed that there were total 17 miRNAs differentially expressed in livers from control and EF (450 mg/kg)-treated groups (Figure 2). Among these altered 17 miRNAs, eight miRNAs were up-regulated (Table 1) and nine miRNAs were down-regulated (Table 2) in EF-treated mice.

Hierarchical clustering analysis of altered miRNA expression in livers from control or EF-treated mice. Hierarchical clustering is performed with miRNAs of 2 fold-changed. Rows, miRNA; column, control (Cont) group and EF-treated group. For each miRNA, red colour indicates miRNA with high expression, whereas green colour indicates miRNA with low expression. miRNA: microRNA.
The up-regulated miRNAs in EF-treated group.

miRNA: microRNA.
The down-regulated miRNAs in EF-treated group.
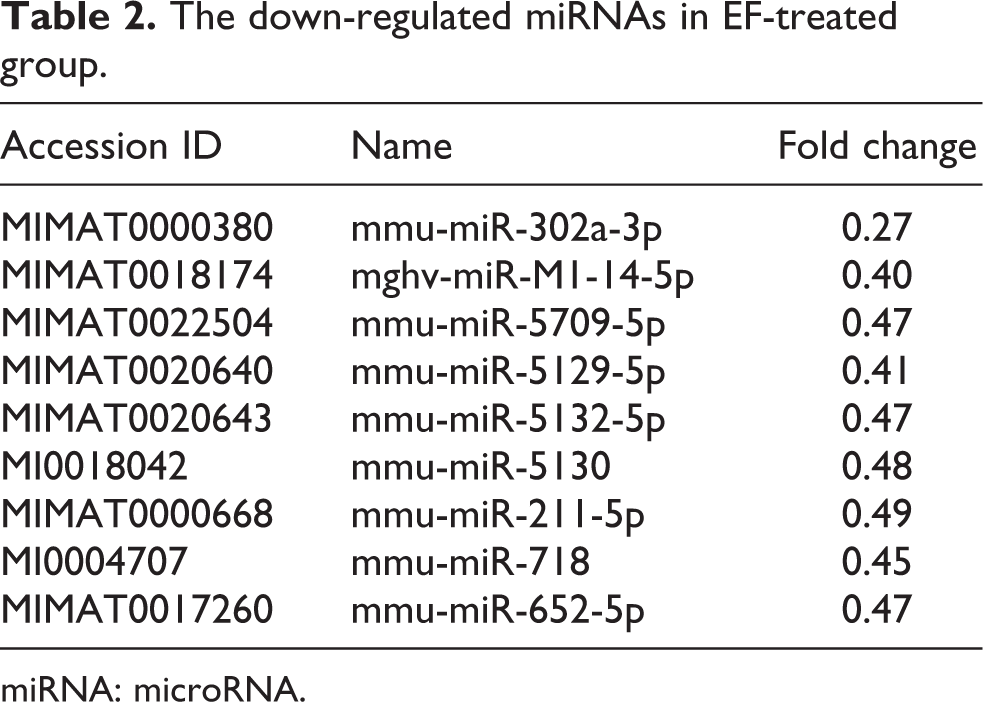

miRNA: microRNA.
Validation of differentially expressed miRNA
Next, we chose eight miRNAs from the above 17 miRNAs, which had higher expression in liver (>100), to further validate its expression in control and EF-treated groups using real-time PCR assay. The results (Figure 3) showed that the expression of miR-200a-3p was increased in EF-treated mice, but the expression of miR-5132-5p and miR-5130 was decreased in EF-treated mice. Thus, we can see that among these above eight miRNAs, the alteration of three miRNAs (miR-200a-3p, miR-5132-5p and miR-5130) was the same as the results of miRNA array.
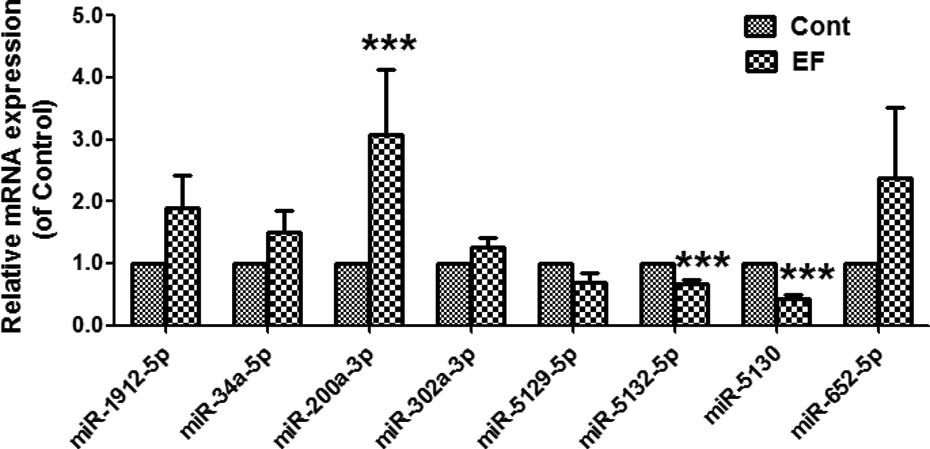
The expression of altered miRNAs from miRNA array was detected by real-time PCR assay. Data is shown as means ± SEM (n = 5), ***p < 0.001 compared with control. miRNA: microRNA; PCR: polymerase chain reaction; SEM: standard error mean.
miRNA target prediction and pathway annotation
Further, the target genes of these above three altered miRNAs (miR-200a-3p, miR-5132-5p and miR-5130) were predicted by the overlapped data from Mirbase, Miranda and TargetScan prediction softwares. As shown in Figure 4, there are total 4178 target genes of miR-200a-3p, 24 target genes of miR-5130 and 5 target genes of miR-5132-5p, respectively. However, there was no same target genes of miR-200a-3p, miR-5132-5p and miR-5130 (Figure 4(b)). We can see from Figure 4(b), there were five same target genes of miR-200a-3p and miR-5130 and two same target genes of miR-200a-3p and miR-5132-5p. The list of the above seven same target genes was shown in Table 3.

The predicted target genes of altered miRNAs. (a) Signal network. Blue box nodes represent miRNA, and red cycle nodes represent mRNA. Edges show the inhibitory activity of miRNA on mRNA. (b) Target genes of miRNAs (miR-200a-3p, miR-5132-5p and miR-5130) were predicted by the overlapped data from Mirbase, Miranda and TargetScan prediction softwares. miRNA: microRNA.
List of the same target genes of differentially expressed miRNAs.
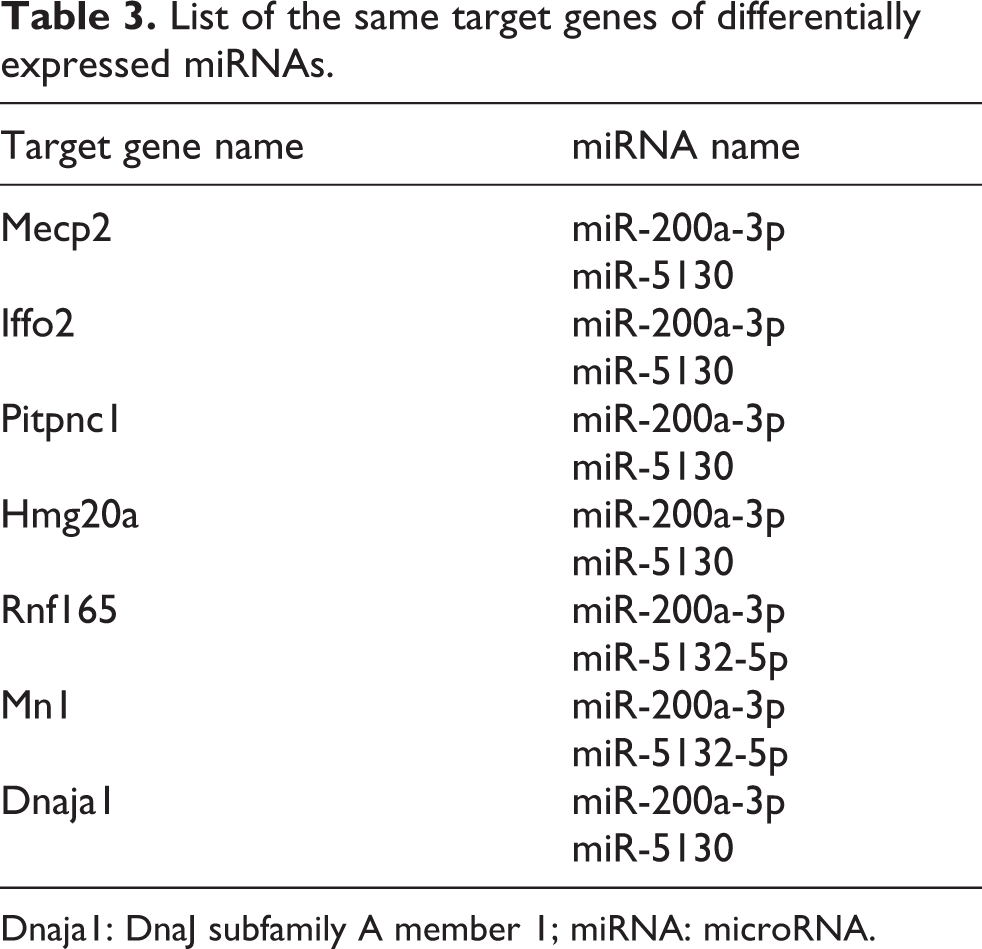

Dnaja1: DnaJ subfamily A member 1; miRNA: microRNA.
KEGG pathway enrichments were further performed by mapping these above seven predicted target genes. As shown in Figure 5(a), only one KEGG pathway (p < 0.05) was annotated, which is protein processing in ER. Also, the KEGG pathway analysis pointed out that Dnaja1 (also named heat shock protein 40 (HSP40), orange word) was the most important target gene of this pathway based on the differentially expressed miRNAs (Figure 5(b)). In addition, we can see that Dnaja1 was the same target gene of both miR-200a-3p and miR-5130 (Table 3). Further, the mRNA and protein expression of Dnaja1 in liver was analysed. The results showed (Figure 6(a)) that EF (450 mg/kg) increased liver mRNA expression of Dnaja1, and Dnaja1 mRNA expression is about 2.4 times of the normal group. In addition, EF (450 mg/kg) also increased hepatic Dnaja1 protein expression, but EF (300, 375 mg/kg) had no effect on Dnaja1 expression (Figure 6(b) and (c)).

KEGG pathway analysis. (a) Pathway analysis based on miRNA (miR-200a-3p, miR-5132-5p and miR-5130) targeted genes. The vertical axis is the pathway category, and the horizontal axis is the enrichment of pathways. (b) The KEGG pathway of protein processing in ER. Dnaja1 was marked orange. KEGG: kyoto encyclopedia genes and genomes; miRNA: microRNA; ER: endoplasmic reticulum; Dnaja1: DnaJ subfamily A member 1.

The mRNA and protein expression of Dnaja1 in livers from control and EF-treated groups. (a) Dnaja1 mRNA expression was detected by real-time PCR assay (n = 6). (b) Hepatic Dnaja1 expression was detected by immunoblotting and β-actin was used as a loading control. Results represent three independent experiments. (c) The quantitative densitometric analysis of Dnaja1 (n = 3). Data is shown as means ± SEM. ***p < 0.001 compared with control. Dnaja1: DnaJ subfamily A member 1; SEM: standard error mean; PCR: polymerase chain reaction.
Discussion
miRNAs belong to one class of small non-coding RNAs, and it plays crucial regulatory roles in animal and plants by targeting mRNAs for cleavage or translation repression. 1 –3 Increasing evidences demonstrated that aberrant miRNA expression was implicated in a variety of diseases such as cancer (including breast cancer, gastric cancer, liver cancer, prostate cancer, colon cancer, renal cancer and lung cancer), cardiovascular diseases (such as atherosclerosis, cardiac hypertrophy and myocardial infarction), inflammation and neurodegeneration. 17 Recently, miRNAs such as miR-34a, miR-561, miR-122 and miR-33 were reported to play an important role in regulating the hepatotoxicity induced by aflatoxin B1, acetaminophen, carbon tetrachloride, thioacetamide and statin. 18 –21
In this study, we found that EF-induced hepatotoxicity in mice (Figure 1), and such results are consistent with our previous reported studies. 11,12,14 Further, liver miRNA expression profile was observed. The results showed that eight miRNAs were up-regulated (Table 1) and nine miRNAs were down-regulated (Table 2) in livers from EF-treated mice (Figure 2). Moreover, the increased expression of miR-200a-3p and the decreased expression of miR-5132-5p and miR-5130 in EF-treated mice livers were further validated by real-time PCR assay (Figure 3). Previous studies have shown the down-regulation of miR-200a-3p in some types of cancer and it’s potential involvement in the initiation and progression of cancer, and those results pointed out that miR-200a-3p might be a tumour suppressor by inhibiting cancer cell proliferation and migration. 22 –24 Also, a previous report demonstrated that miR-200a-3p was involved in the promotion of oxidative stress-mediated liver cell death. 25 In addition, miR-200a-3p (originally named miR-200a) was reported to be involved in transforming growth factor-β1-induced hepatic stellate cell activation. 26 Although there is still no direct evidence indicated that miR-200a-3p was involved in DILI, those above reported studies imply the possibility of miR-200a-3p in regulating DILI. Thus, the increased expression of miR-200a-3p may be involved in regulating the liver injury induced by EF. There is still no report about the study on miR-5132-5p and miR-5130, and they both have few predicted target genes (Figure 4). This study is the first report about the potential involvement of miR-5132-5p and miR-5130 in DILI.
Protein processing in ER was the only 1 KEGG pathway (p < 0.05) that was annotated for miR-200a-3p, miR-5132-5p and miR-5130 (Figure 5(a)). ER is responsible for protein folding, assembling and post-translational modifications. 27 Only the properly folded proteins are transported from ER to the Golgi apparatus, whereas the unfolded or misfolded proteins are targeted for ER-associated degradation via the ubiquitin-proteasome system. 28 The accumulated aggregation of misfolded or unassembled proteins in ER will lead to ER stress and thus seriously affect cell homeostasis. 27,28 It is reported that ER stress is closely related with oxidative stress and plays important roles in a variety of human disorders, including neurodegenerative diseases, diabetes mellitus, inflammation and liver diseases. 29 Increasing studies demonstrated that ER stress was involved in hepatotoxicity induced by some drugs including perfluoroalkyl acids, 30 zidovudine, 31 efavirenz 32 and acetaminophen. 33 However, the concrete contribution of ER stress to hepatotoxicity is still controversial. 34 This study indicates the potential involvement of ER stress in EF-induced liver injury, but it also needs further deep investigation to elucidate the concrete engaged mechanism.
The DnaJ proteins, also known as HSP40, generally act as cochaperones to the molecular chaperone Dnak (Hsp70), which are critically involved in ER-associated degradation and contribute to the relief of ER stress. 35 Dnaja1 is an important member of DnaJ proteins, and it has been reported that Dnaja1 associated with Dnak (HSP70) promoted tumorigenesis by inhibiting cell death. 36 In this study, we found that Dnaja1 was the only one predicted target gene of protein processing in ER signalling pathway based on the altered miRNAs (Figure 5(b)), and the expression of Dnaja1 was increased in EF-treated mice livers (Figure 6). Dnaja1 is the predicted target gene of both miR-200a-3p and miR-5130 (Table 3), but the expression of Dnaja1 was increased in EF-treated mice livers, which may be mainly due to the down-regulated expression of miR-5130. These above results imply that the increased expression of Dnaja1 may contribute to the elevated body self-defence against EF-induced hepatotoxicity by inhibiting hepatocytes death, and its concrete function needs future deep investigation.
In conclusion, this study demonstrated the altered expression of miR-200a-3p, miR-5132-5p and miR-5130 in mice livers during EF-induced hepatotoxicity, indicating the potential involvement of these miRNAs in EF-induced liver injury for the first time. Further, protein processing in ER signalling pathway based on those above altered miRNAs and the predicted target gene Dnaja1 was found to be critically involved in EF-induced liver injury.
Footnotes
Acknowledgements
The authors would like to thank Yanfen Geng and Hongxia Shen (KangChen Bio-tech Inc., Shanghai) for their kind assistant in miRNA array analysis.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was financially supported by State major science and technology special projects during the 12th five year plan (2015ZX09501004-002-002 and 2012ZX09505001-002).
