Abstract
The objective of this study was to determine if the association between exposure to ambient air pollutants such as sulfur dioxide, nitrogen dioxde (NO2), nitrous oxide (NO), and PM10, and variation in lung function measures was modified by genotype. A validated questionnaire was administered to 71 African children to evaluate prevalence of respiratory symptoms. Atopy was evaluated by skin-prick testing and bihourly measures of lung function (spirometry) were collected. Gaseous air pollutant concentrations were monitored continuously. CD14 polymorphism was genotyped and plasma CD14 levels were measured. There was no statistically significant association between the CD14 (159) CT+TT polymorphism with any asthma-related phenotype. There was a significant association between lung function (forced expiratory volume in 1 second intraday variability) and NO2 and NO among participants carrying the CD14 CT/TT genotype for lags 1, 2, and the 5-day average. Similarly, statistically significant gene–pollutant interactions (p < 0.05) were found with NO and CD14 CT/TT at lag 2 and for the 5-day average. While there was no association with any respiratory phenotype (as determined by symptoms), the CD14 CT/TT genotype appeared to be protective to increased exposure to NO2 and NO.
Introduction
With adverse respiratory health increasing morbidity among children, 1 the effects of air pollution become increasingly important and it is biologically plausible that many pollutants modify innate and acquired immune responses. 2 Genetic factors may also modify the effect of these exposures and are critical determinants of lung inflammation and pulmonary function. 3 It is hypothesized that genetic differences that affect responsiveness to endotoxin underlie the variability in airway responsiveness. CD14 is a high-affinity receptor for endotoxin, 4 which acts by transferring endotoxin and other bacterial ligands from circulating lipopolysaccharide-binding protein to the toll-like receptor 4 (TLR4)/MD-2 signalling complex. Acute endotoxin-rich bioaerosol exposure is a known cause of acute lung disease and asthma exacerbation. Myeloid differentiation primary response gene 88 (MyD88)-dependent TLR signalling has been suggested as a potential mechanism for the inflammatory actions of pollutants. 2
The gene encoding CD14 is localized on chromosome 5q31.1,
5
which is a region that has been linked to asthma
6,7
and pulmonary function.
8
Polymorphisms in CD14 that influence the propensity to develop asthma and allergy have been described. A cytosine to thymine transition at base pair
To our knowledge only one study has to date been published on the effects of CD14 polymorphism on an asthmatic African population. This study found that the CD14 (-159) C-allele was significantly higher in the asthmatic group when compared with controls (p ≤ 0.005). 19 In our South African population of indigenous African children, the frequency of the CD14 (-159) polymorphism, soluble CD14 and the risk conferred on respiratory outcomes, was determined. Additionally, we evaluated if the association between exposure to ambient air pollutants such as sulfur dioxide (SO2), NO2, NO, and PM10, and variation in lung function measures was modified by genotype.
Methodology
Study population
A sample of 71 indigenous African children (between 9 and 11 years old) from 7 primary schools was recruited for this study, from the south and north communities of Durban, South Africa. At each of the 7 schools, two fourth grade classrooms were randomly assigned as classroom 1 and 2 and children were randomly selected from these classrooms. Details of the study sampling strategy have been presented elsewhere. 20 The south Durban region, recognized as one of the most highly industrialized and most heavily polluted areas in Southern Africa, 21 includes extensive residential areas. The University of KwaZulu-Natal and the Durban University of Technology’s Ethics Committees approved this study. Parental consent for genetic analyses was obtained for all children.
Symptom and demographic data
A questionnaire was administered to all caregivers, which included components addressing demographic information, and respiratory and other relevant symptoms, using standardized validated questions from sources such as the British Medical Research Council and the American Thoracic Society. Characterization of asthma severity (any asthma and persistent asthma) was previously reported. 20 A child was considered to have asthma of any severity if any of the following were true: three or more non-exercise-related symptoms (e.g. cough, wheeze, chest tightness); exercise-induced wheeze or cough reported at a frequency of three or more times during the previous year; doctor-diagnosed asthma, reactive airway disease and/or asthmatic bronchitis; or doctor-prescribed medication taken in the previous year. A child was considered to have ‘persistent asthma’ if, first, the child met the diagnostic criteria for asthma of any severity, and second, if any of the following were true: any daytime symptoms (cough, wheeze, exercise-induced cough and wheeze, and chest tightness) was reported as being present more than two times per week; sleep disturbances due to cough, wheeze, shortness of breath, or chest tightness reported more than two times per month; and/or daily use of doctor-prescribed medication.
Baseline spirometry was performed by experienced technologists using the American Thoracic Society criteria utilizing the Jaeger Flowscreen (ATS, 1995). Spirometers were calibrated at least twice a day with a 3 L syringe. Bronchial airway hyperreactivity (BHR) was categorized as marked (PC20, dose of methacholine causing a 20% fall in baseline forced expiratory volume in 1 second (FEV1), ≤ 4 mg/mL), probable (4 mg/mL < PC20 ≤ 8 mg/mL), borderline/possible (8 mg/mL < PC20 ≤ 16 mg/mL), and no hyperreactivity (PC20 ≥ 16 mg/mL). Marked, probable, and possible BHR were categorized as ‘any evidence of bronchial hyperreactivity’. Atopy was defined as a positive response to one or more of the following antigens: mixed cockroach; mixed dust mite; mould mix (Aspergillus, Cladosporium and Penicillium); cat; dog; mouse; rat; and grass, by skin-prick testing, with histamine as a positive control and saline as a negative control. A greater than 3-mm difference in mean diameter between allergen and control wheal was considered positive.
Bihourly measures of pulmonary function
A bihourly measure of lung function was collected at the schools for 5 days per week over 3-week periods in each of the four seasons using digital hand-held devices: the AirWatch® (iMetrikus, Carlsbad, California, USA) brand airway monitor. All schools were studied simultaneously in the same calendar periods. On each of the five schooldays during the week, participants were asked to perform a session of three consecutive forced expiratory manoeuvres every one-and-a-half to two hours (four times per 5.5-h school day: at approximately 08h00, 09h45, 11h30, and 13h20). The highest FEV1 from a valid expiratory manoeuvre for each of the four daily sessions was used in data analyses. An expiratory manoeuvre was considered valid if the FEV1 result (1) was between 30% and 120% of each child’s personal best as defined by that child’s highest recorded FEV1 during baseline spirometry (performed by experienced technicians using the American Thoracic Society criteria); and (2) came from expiratory manoeuvre that was recorded by the Airwatch device as ‘error-free’.
Environmental monitoring of ambient pollutants
During each of the 4 intensive 3-week phases, gaseous air pollutant concentrations were continuously monitored: NO2 and NO were sampled at 7 monitoring sites using continuous gas-phase chemiluminescence detection; and SO2 was monitored continuously at 16 sites, including all 7 schools, using ultraviolet fluorescence spectrometry. PM10 was monitored gravimetrically at 12 sites: at each school using 24-h integrated measurements, and at 5 additional sites using a tapered element oscillating microbalance. Each pollutant was sampled using standard reference methods and protocols. Details of these measurements and the quality assurance program are reported elsewhere. 20
Genotyping and analysis of plasma CD14 concentration
All genotyping assays were conducted by a researcher who was blind to child ID and disease status. Genomic DNA was extracted using a PUREGENE DNA isolation kit (Gentra, Minneapolis, Minnesota, USA). The CD14 genotypes were determined by Taqman Assays (Applied Biosystem, Foster City, California, USA) using the Assay-by-Design (Taqman® SNP Genotyping Assays) mix. The fluorescence of polymerase chain reaction products was detected by the ABI Prism ®7000 sequence detection system (Applied Biosystems, Foster City, California, USA). Positive and negative controls were included in the analyses. Plasma CD14 levels were measured using the Quantikine Human sCD14 Immunoassay™ (R & D SYSTEMS, Minneapolis, Minnesota, USA). Reaction color intensity was measured using the Bio-Tek μQuant enzyme-linked immunosorbent asssay plate reader.
Statistical analysis
CD 14 was dichotomized into two groups, based on the absence or presence of the polymorphic allele (wild-type homozygous (CD14 -159 CC) and heterozygote and homozygous variant (CD14 -159 CT+TT)). Differences in genotype frequency between cases (symptoms for any asthma or persistent asthma; atopic or increased BHR) and non-cases (no asthma symptoms, non-atopic, no BHR), were assessed by the χ2 test. Associations of genotype with either category of asthma, atopy, and BHR were examined using multivariate logistic regression models, utilizing gender and exposure to environmental tobacco smoke as covariates. Linear regression models were fitted using generalized estimating equations (GEEs, assuming normal distribution with identity link) using PROC GENMOD for SAS to accommodate the correlation structure arising from repeated measurements on the same individual.
Within-day variability for FEV1 was defined as 100 × (the maximum best FEV1 − minimum best FEV1)/maximum best FEV1, where the ‘best FEV1’ is the highest valid value for the specific time of day (08h00, 09h45 and 11h30, 13h20), thus providing a single summary lung function measurement per child, per day. We used a dominant coding model to assess the genotype effects in the statistical models. Covariates used in the GEE models included race, school, caregiver smoking, caregiver education, household income, and season. The gene–environment interaction was assessed for associations of exposure to SO2, NO, NO2, and PM10 with FEV1 intraday variability, using the CD14 (-159) polymorphism as the effect modifier. Daily exposure estimates were based on the child’s school: PM10 and SO2 used school-based measurements; and NO and NO2 used the spatial average across either north Durban (three schools) or south Durban (four schools), as not all schools had NO x monitors. Multiple imputation procedures (repeated five times for each exposure parameter) were used to obtain a complete data set. Lag effects were modelled to account for both acute and prior exposure effects, and included lags of 1–2 days as well as the 5-day average. The percentage change in within-day variability in FEV1 was estimated for an increase of one inter-quartile range in each pollutant (NO2: 8.19 ppb, NO: 29.7 ppb, PM10: 29.4 µg m−3 and SO2: 9.8 ppb). An adverse effect in lung function would be denoted by an increase in the estimate for intraday variability in FEV1. SAS (version 9.1) and STATA (version 9, College Station, Texas, USA) was used for analyses.
Results
Demographic, phenotypic, and genotypic characteristics are presented in Table 1. The average age of the children in this study was 10 years; almost 50% of them reported passive exposure to tobacco smoke. Almost 28% of the population were atopic and 35% reported having any category of asthma. Approximately 63% of the study population carried the wild-type CD14 genotype, while the polymorphic heterozygote (CD14 TT) was relatively rare (9%). The observed genotype frequencies did not show a significant departure from Hardy Weinberg equilibrium (p = 0.10). Table 2 summarizes 24-h average pollutant levels in Durban. SO2 levels varied widely across the study area, and the highest concentrations occurred in South Durban, reflecting the location of emitting industries. Table 3 shows the frequencies of the CD14 (159) C/T polymorphism in relation to persistent asthma, any grade of asthma, atopy, and BHR. There was no statistically significant association between the CD14 (159) CT+TT polymorphism with any asthma-related phenotype such as atopy, any asthma, persistent asthma, and bronchial hyper responsiveness. Plasma sCD14 (soluble CD14) levels were not significantly different between the CD14 (159) CC and the CD14 (159) CT+TT genotypes (8.8 ng/mL ± 4.1 ng/mL vs. 7.8 ng/mL ± 4.7 ng/mL; p = 0.47). Similarly, soluble CD14 protein levels did not vary significantly between case and non-case for asthma, persistent asthma, and BHR. Atopic children had a significantly higher mean soluble CD14 protein, however, compared with non-atopic children (9.5 ng/mL vs. 6.8 ng/mL, p = 0.04; Table 4).
Demographic, phenotypic and genotypic characteristics of study population (n = 71).

Summary of ambient pollutant levels (24-h averages) in Durban during 2004–2005.

SO2: sulfur dioxide; NO2: nitrogen dioxide; NO: nitric oxide.
Association of CD14 (-159) gene polymorphism with respiratory outcome.

OR: Odds ratio; CI: confidence interval; BHR: bronchial airway hyperreactivity.
aLogistic regression models adjusted for age and gender.
Soluble CD14 protein from blood plasma stratified by genotype and respiratory phenotype (n = 71).
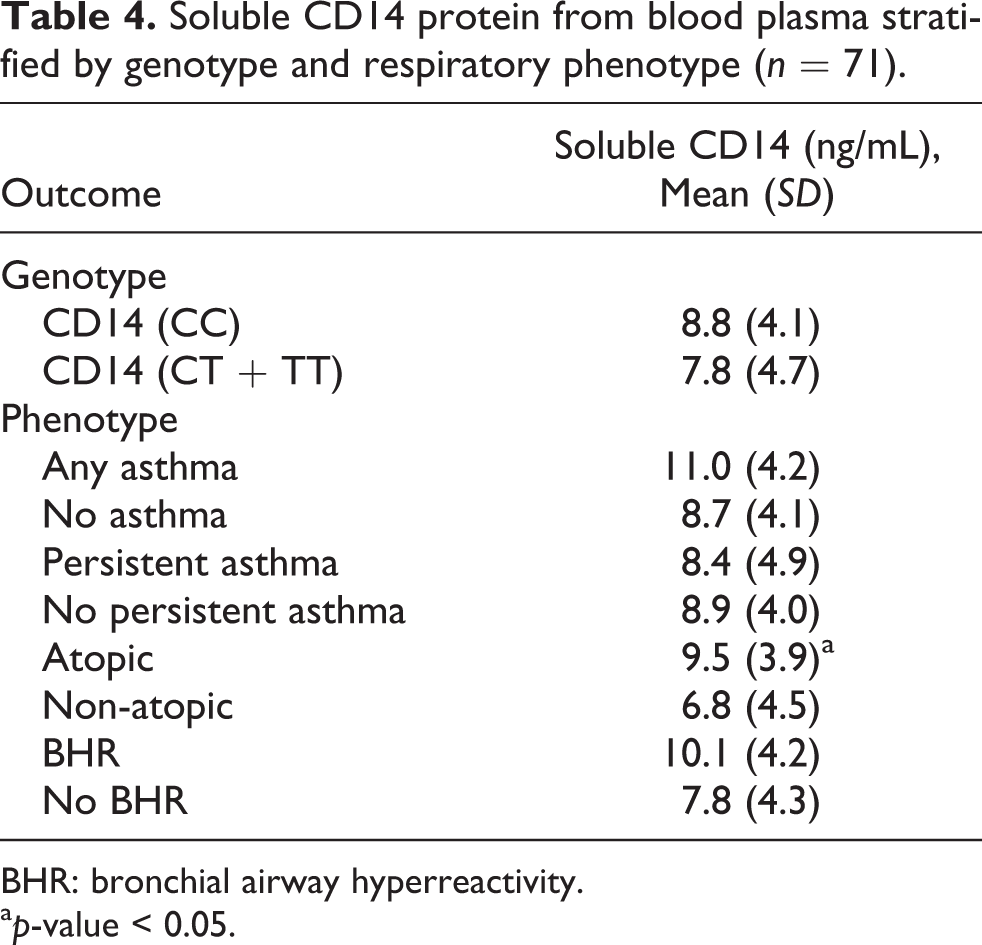
BHR: bronchial airway hyperreactivity.
ap-value < 0.05.
Effect estimates (ignoring the CD14 (-159) genotype) for within-day variability in FEV1 associated with PM10, SO2, NO, and NO2 (tested independently) were inconsistent across different lagged times (Table 5). None of the pollutant-lag combinations were statistically significant. Table 6 presents models for potential effect modification by the CD14 (-159) genotype on the association between air pollutant exposures and FEV1 intraday variability. Among the CD14 (-159) CC children, a statistically significant association of exposure with FEV1 intraday variability was seen for SO2 (β = −1.50, CI: −0.36, 3.37; p = 0.01, p(int) = 0.01) and PM10 (β = 1.26, CI: 0.02, 2.50; p = 0.05) at lags 1 and 2, respectively. Conversely, there was a significant association between intraday variability and NO2 and NO among participants carrying the CD14 CT/TT genotype for lags 1, 2, and the 5-day average. The effect estimates for children who carried the homozygote wild type were higher than the polymorphic genotype, reaching a significant gene–pollutant interaction with intraday variability in FEV1 at lags 1, 2, and 5-day average post exposure for NO2. Similarly, statistically significant gene–pollutant interactions (p < 0.05) were found with NO and CD14 CT/TT at lag 2 and for the 5-day average. The CT/TT genotype appeared to be protective to increased exposure to NO2 and NO.
FEV1: forced expiratory volume in 1 second; SO2: sulfur dioxide; NO2: nitrogen dioxde; NO: nitrous oxide; SO2: sulfur dioxide; GEE: generalized estimating equation.
1The percent change value shown is for an increase of one inter-quartile range in each respective pollutant: NO2: 8.19 ppb, NO: 29.7 ppb, PM10: 29.4 µg m−3, SO2: 9.8 ppb.
2Intraday variability for FEV1 is defined as: 100 (maximum best FEV1 − minimum best FEV1)/maximum best FEV1; where the ‘best FEV1’ is the highest valid, error-free value for the specific time of day (08h00, 09h45, 11h30, 13h20). An increase in the estimate for intraday FEV1 is indicative of a negative impact on lung function.
3Pollution levels used in regression models combine measured and imputed values. Covariates in each model: race, school, caregiver smoking, caregiver education, household income, season.
FEV1: forced expiratory volume in 1 second; SO2: sulfur dioxide; NO2: nitrogen dioxde; NO: nitrous oxide; SO2: sulfur dioxide; GEE: generalized estimating equation; CI: confidence interval.
1The percent change value shown is for an increase of one inter-quartile range in each respective pollutant: NO2: 8.19 ppb; NO: 29.7 ppb, PM10: 29.4 μg m−3; SO2: 9.8 ppb.
2Intraday variability for FEV1 is defined as: 100 (maximum best FEV1-minimum best FEV1)/maximum best FEV1; where the ‘best FEV1’ is the highest valid, error-free value for the specific time of day (08h00, 09h45, 11h30, 13h20). An increase in the estimate for intraday FEV1 is indicative of a negative impact on lung function.
3Pollution levels used in regression models combine measured and imputed values. Covariates in each model: race, school, caregiver smoking, caregiver education, household income, season, interaction between genotype and pollutants.
4p value for genotype-pollutant interaction term ≤0.05.
Discussion
A number of studies have provided strong evidence that innate immune responses modulate the acute response to pollutants. While numerous studies have been published on the association of endotoxin exposure with the polymorphic CD14 (-159) T-allele, data exploring the gene–environment interaction with air pollutant exposures is limited. This study evaluated the risk conferred by the CD14 (-159) polymorphism on respiratory outcomes and any effect modification on lung function and exposure to pollutant in a cohort of South African children. The CD14 (-159) T-allele was relatively common in our African population (37%) with an allelic frequency of 0.23. Almost 39% of the population reported having any category of asthma. In this study, atopic participants had significantly higher sCD14 levels than non-atopic subjects. The effect of PM10, SO2, NO, and NO exposures on lung function measures was significantly modified by the CD14 (-159) polymorphism. Adverse effects of exposure to nitrogen oxides were limited to individuals carrying the CD14 (-159) CC genotype and the CT/TT genotype appeared to be protective to increased exposure to NO2 and NO.
The frequency of the CD14 (-159) CT+TT genotype among children reporting any asthma in this study was 37%, which was markedly lower than the frequency found in a Tunisian population of asthmatics (78%). 19 Although this was the only comparable African study, it should be noted that the participants in the Tunisian study were Caucasian as compared with our Black African sample group. The polymorphic T-allele may be rarer in African populations as compared with other populations (East Asian 57.5%, Indian 62.1%, and European 45.7%). An allele frequency of 29.3% for the T-allele was found among the Yoruba in Ibadan, Nigeria 18 that is closely related to our results. Studies are required to contribute to data on the frequencies of this polymorphism among African populations. We found that CD14 (-159) C/T polymorphism was not significantly associated with asthma, and its related respiratory phenotypes such as atopy and BHR. Although a recent case control study among Chinese children has reported an association between the TT genotype and an increased risk of atopic asthma, 22 our findings are similar to those by Zhao and Bracken who published a meta-analysis of 23 studies, which yielded a non-significant overall association between the CD14 (-159) C/T gene polymorphism and asthma. 18 There have been variable results published from different studies. After restricting analysis to selected studies, these authors found that carriers of the TT and CT genotypes were about 33% less likely (Odds ratio (OR) = 0.67, 95% confidence interval (CI): 0.54–0.84) and about 20% less likely (OR = 0.80, 95% CI: 0.66–0.95), respectively, to have atopic asthma compared with carriers of the CC genotype. They then concluded that there was a protective relationship between the CD14 (-159) T-allele and atopic asthma susceptibility. 18
Participants who were atopic had significantly higher sCD14 levels compared with non-atopic subjects (p = 0.04). In contrast, other studies have found lowered sCD14 levels among atopic subjects compared with non-atopic subjects before endotoxin inhalation. 23 A study to determine whether infant and maternal levels of sCD14 were associated with the atopic outcomes of infants found that there was no difference in plasma sCD14 levels at birth in children with a high risk of development of atopy, when compared with those having a low risk. 24 Generally, T-allele carriers have been found to have higher serum CD14 levels than carriers of the C-allele 25 and this single nucleotide polymorphism (SNP) may therefore act as modifier of asthma risk in individuals with differing environmental exposure. Functional genomic studies have supported this finding by showing increased transcriptional activity of the CD14 (-159) T-allele. 14 Homozygotes of the T-allele appear to be protective for asthma at low levels of endotoxin exposure, but may increase asthma risk at high levels of endotoxin exposure. 25
In the Durban South industrial basin, the oil refineries, paper producer, and sugar refinery are responsible for 80% of the SO2 pollution load. 26 In addition, there is a high traffic volume in this area, which has been associated with asthma, cough, and wheeze. 27 Primary sources of nitrogen oxide emissions include motor vehicles on the various motorways which converge in the South Durban basin. 28 Nitrogen oxides react with other pollutants to form ground-level ozone. This irritates the nose and throat, especially in people with asthma, and appears to increase susceptibility to respiratory infections. Generally, air concentrations of SO2 are highest near large industrial complexes. The average concentrations of pollutants per region have been described elsewhere. 20
Despite not detecting an association between the CD14 gene polymorphism with respiratory outcome or protein expression, we found significant gene–environment interactions between ambient pollutants (NO2, NO), CD14 (-159) C/T gene polymorphism and lung function measures. Children carrying the CT/TT genotype presented with a statistically significant decreased intraday variability of FEV1 for NO and NO2 at lags 1, 2, and 5 days. Published data about exposure to house-dust endotoxin or to domestic sources of microbial exposure, which was assessed concomitantly with CD14 (-159) C/T, have shown a consistent, replicable gene–environment interaction. Specifically, results suggest that the C-allele is a risk factor for allergic phenotypes at low levels of exposure, whereas the T-allele is a risk factor at high levels of exposure. 25,29 At first glance these results suggest a disappointing inconsistency; however, a far more complex picture emerged from further studies. 24 In a founder population in the United States, findings suggested that there could be an antagonistic interaction between environmental conditions and CD14 (-159) C/T as determinants of allergic sensitization: the T-allele could be either protective or a risk factor, depending on the degree of exposure to environmental microbial products. 30 It would appear that multiple mechanisms are involved. To date, the best supportive evidence implicates both heightened innate and adaptive immune responses. Since acute endotoxin-rich bioaerosol exposure is a known cause of acute lung disease and asthma exacerbation, myeloid differentiation primary response gene 88 (MyD88)-dependent TLR signalling is a possible candidate mechanism for the inflammatory actions of pollutants. Innate immune mechanisms can be activated by damage-associated molecular patterns (DAMPs) and pollutant exposure results in increased expression of CD14 and TLR4 in airway macrophages, thus increasing both the DAMP ligand levels and appropriate effector cell numbers in the airway. 2
In summary, the present study suggests that the CD14 polymorphism may modify the effects of air pollutants. While there was no association with any respiratory phenotype (as determined by symptoms), the CD14 CT/TT genotype appeared to be protective to increased exposure to NO2 and NO. A limitation of this study is that each of the effects of the four pollutants was studied independently, using the approach of one gene with one exposure. This model does not account for interaction of pollutants and modifying the effect on lung function. Additionally, the sample size was small although power was enhanced by characterization of individual exposure and repeated measures over time. In addition, the study population was confined to defined areas with its own monitoring site, allowing a more precise estimation of exposure. Current research has focused on epigenetic regulation and gene–environment–epigene interactions. In future, this may lead to a more accurate estimation of respiratory risk among susceptible populations.
Footnotes
Acknowledgements
We wish to thank the children and parents of the South Durban Health Study cohort for their dedicated cooperation and commitment. Mrs K Asharam, Dr Moodley, and Dr Alisa Phulukdaree are gratefully acknowledged for their assistance. We also thank the following sponsors: Grant sponsor: National Research Foundation (NRF) supplemented by Durban University of Technology (DUT); Grant no: 2068195 Thuthuka; Grant sponsor: Ethekwini Municipality, KwaZulu-Natal, Durban, South Africa, Grant no: 1a-103 – Health study.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.


