Abstract
The identification of biomarkers for toxicity prediction is crucial for drug development and safety evaluation. The selective and specific biomarkers for antihistamines-induced cardiotoxicity is not well identified yet. In order to evaluate the mechanism of the life-threatening effects caused by antihistamines, we used DNA microarrays to analyze genomic profiles in H9C2 rat cardiomyocytes that were treated with antihistamines. The gene expression profiles from drug-treated cells revealed changes in the integrin signaling pathway, suggesting that cardiac arrhythmias induced by antihistamine treatment may be mediated by changes in integrin-mediated signaling. It has been reported that integrin plays a role in QT prolongation that may induce cardiac arrhythmia. These results indicate that the integrin-mediated signaling pathway induced by antihistamines is involved in various biological mechanisms that lead to cardiac QT prolongation. Therefore, we suggest that genomic profiling of antihistamine-treated cardiomyocytes has the potential to reveal the mechanism of adverse drug reactions, and this signal pathway is applicable to prediction of in vitro cardiotoxicity induced by antihistamines as a biomarker candidate.
Introduction
Certain non-antiarrhythmic drugs produce undesirable effects by delaying cardiac repolarization, which can be defined as a prolongation of the QT interval on a surface electrocardiogram (ECG). 1 The QT interval represents the duration of ventricular depolarization and subsequent repolarization and is measured from the beginning of the QRS complex to the end of the T wave. QT prolongation and proarrhythmias have been the most common reasons for the withdrawal of a wide variety of therapeutic drugs from the market. Identification of toxicity biomarkers may potentially provide a solution to these challenges; however, the number of effective and reliable cardiovascular biomarkers currently available is low. Therefore, there is a compelling need to find suitable biomarkers for drug safety prediction and evaluation.
First-generation sedative antihistamines, such as diphenhydramine, are still widely used in several countries where they are the major cause of drug toxicity, regarding their marked central nervous system and anticholinergic adverse effects, 2 where they are a major cause of drug toxicity. Second-generation, nonsedating histamine H1-receptor antagonist antihistamines were introduced in the 1980s. 3 However, after a decade of worldwide usage, the second-generation drugs terfenadine and astemizole were found to cause prolonged QT interval measured on ECGs, leading to their withdrawal from the market. 4,5 The side effects of the drugs were primarily attributed to blockade of cardiac potassium channel currents, resulting in impaired repolarization of the heart and increased risk of ventricular tachyarrhythmia (torsades de pointes (TdP)). Another second-generation antihistamine, loratadine, has been studied extensively in vitro and in the clinic to assess whether it has any risk of QT prolongation. 6,7
The molecular mechanism underlying the adverse reactions of the QT-prolonging drugs is thought to be via the blockade of the delayed rectifier potassium channels (IKr), encoded by the human ether-a-go-go-related gene (
In this study, we analyzed genomic profiles of rat cardiomyocytes for the screening of putative biomarker candidates, which are specific to adverse effects caused by astemizole, loratadine, and terfenadine.
Materials and methods
Chemicals and reagents
Astemizole, loratadine, and terfenadine were purchased from Sigma Chemical Co. (St Louis, Missouri, USA). Microarray chips and reagents for rat genome were purchased from Applied Biosystems (Foster city, California, USA). Isopropanol, ethanol, and other routine chemicals were purchased from Sigma Chemical Co. unless mentioned otherwise.
Cell culture
H9C2 rat cardiomyocytes (CRL-1446, ATCC; American Type Culture Collection, Manassas, Virginia, USA) were cultured with Dulbecco’s modified Eagle’s medium (DMEM; Gibco BRL, Grand Island, New York, USA) supplemented with 10% fetal bovine serum, 2 mM
RNA isolation and DNA microarray
After 24 h treatment of vehicle (DMEM with dimethyl sulfoxide 0.1%), astemizole, loratadine, or terfenadine to rat cardiomyocytes at concentrations of 0, 0.5, 1, and 5 µM, RNA isolation was performed. Astemizole (3.1 µM) was reported to cause between 20% and 30% cytotoxicity after 24 h treatment in H9C2 cardiomyocytes. 11 Therefore, we choose concentration range as above to study and compare the effects of antihistamines on gene expression. Total RNA was extracted using a Trizol (Invitrogen, Carlsbad, California, USA) in accordance with the manufacturer’s procedures. Quantity and purity (260/280 ratio) of RNA were monitored using the ultraviolet–visible spectrophotometer (ND-1000, NanoDrop Technologies, Wilmington, Delaware, USA). Differential gene expressions were profiled using rat genome survey array chips (Applied Biosystems, Foster City, California, USA) containing 60-mer oligonucleotide probes representing a set of 27,088 individual rat genes and more than 1000 control probes. Microarray experiments were performed according to manufacturer’s instructions. Digoxigenin (DIG)–uridine triphosphate-labeled complementary RNA (cRNA) was generated from 5 µg of total RNA and amplified using a chemiluminescent reverse transcription in vitro transcription labeling kit (Applied Biosystems). Briefly, each microarray was prehybridized in hybridization buffer with blocking reagent at 55°C for 1 h. DIG-labeled cRNA targets (10 µg) were fragmented to 100–400 bp and hybridized with each prehybridized microarray at 55°C for 16 h. Arrays were washed with hybridization wash buffer, followed by chemiluminescence rinse buffer. Chemiluminescent signals were generated by incubating arrays with anti-DIG–alkaline phosphatase and chemiluminescence substrate. Images were collected for each microarray using a Model 1700 Chemiluminescent Microarray Analyzer (Applied Biosystems).
Analysis of microarray expression data
Signal intensities were imported into Avadis software (Strand and Stratagene, Bengaluru, Karnataka, India). Interarray quantile normalization was performed in order to minimize the effect of external variables. Average values of gene expression ratios obtained from three replicates were calculated. Differentially expressed genes (DEGs) were selected based on fold changes of ≥2 and
Statistical analysis
Statistical analyses were performed using the Avadis softwar (Strand and Stratagene). Welch’s
Results
Differentially expressed genes in astemizole-, loratadine-, and terfenadine-treated H9C2 rat cardiomyocytes
The effects of astemizole, loratadine, and terfenadine on gene expression in rat cardiomyocytes were examined. Total RNA, derived from H9C2 rat cardiomyocytes treated at various doses and the untreated control as well, was subjected to cRNA synthesis. Table 1 shows the number of differentially expressed genes (fold change ≥2,
The number of differentially expressed genes by treatment of astemizole, loratadine, and terfenadine.a

aThe numbers indicate differentially expressed genes (fold change ≥2,
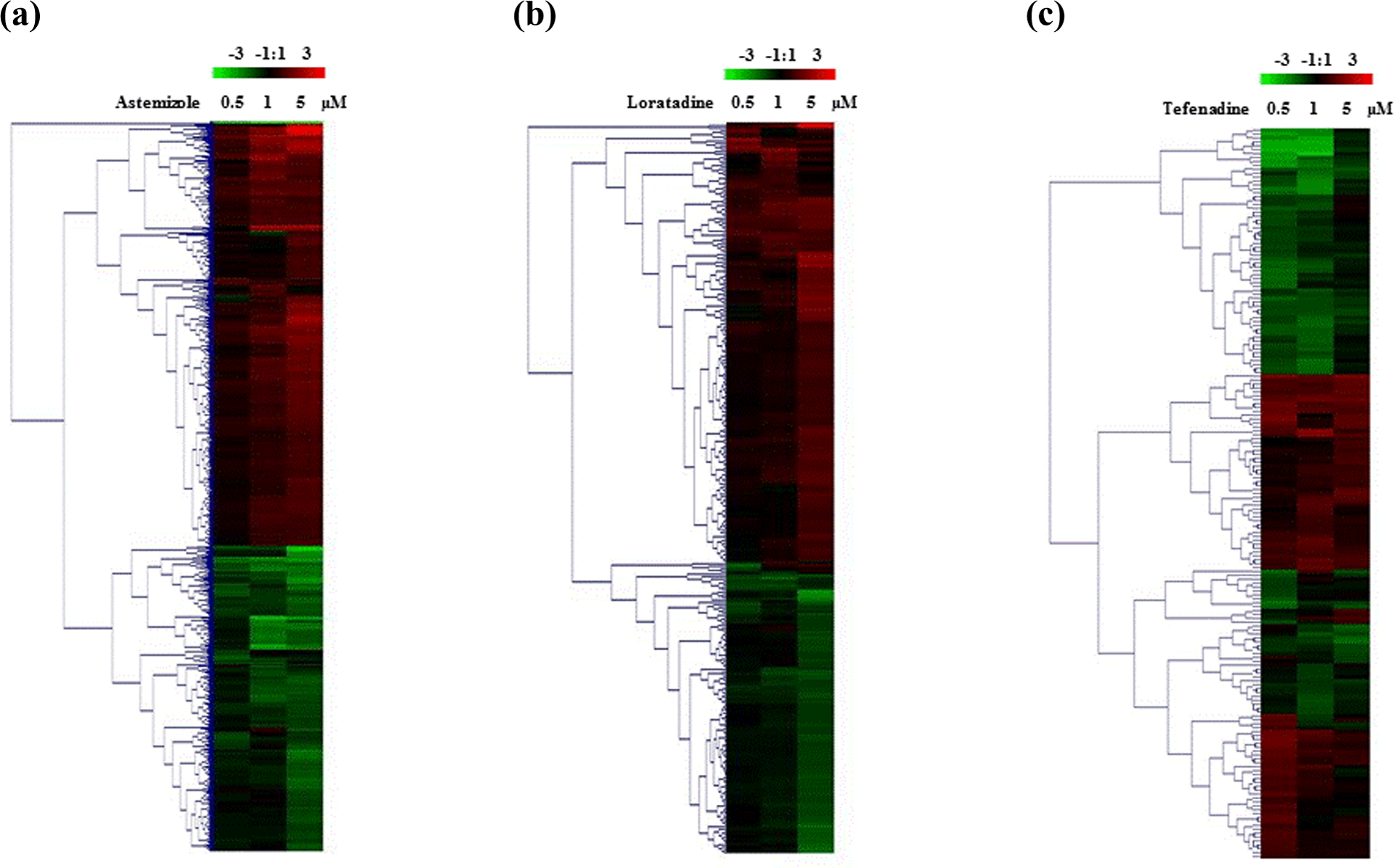
Hierarchical clusterings of astemizole (a), loratadine (b), and terfenadine (c)-treated gene expression in H9C2 rat cardiomyocytes. H9C2 rat cardiomyocytes were seeded at a density of 3 × 106 cells in 75-cm2 tissue culture flasks. Cells were treated with 0, 0.1, 0.5, 1, and 5 µM of astemizole, loratadine, and terfenadine, respectively. The cells were harvested 24 h after treatment. Data were measured using a cRNA microarray with elements representing 27,088 distinct rat genes. All measurements are relative to nontreated control. Expression values are displayed within the rage [−3, 3] with levels above, equal to, or below the mean displayed in red, black, and green, respectively. Each gene is represented by a single row of colored boxes; each concentration is represented by a single column. cRNA: complementary RNA.
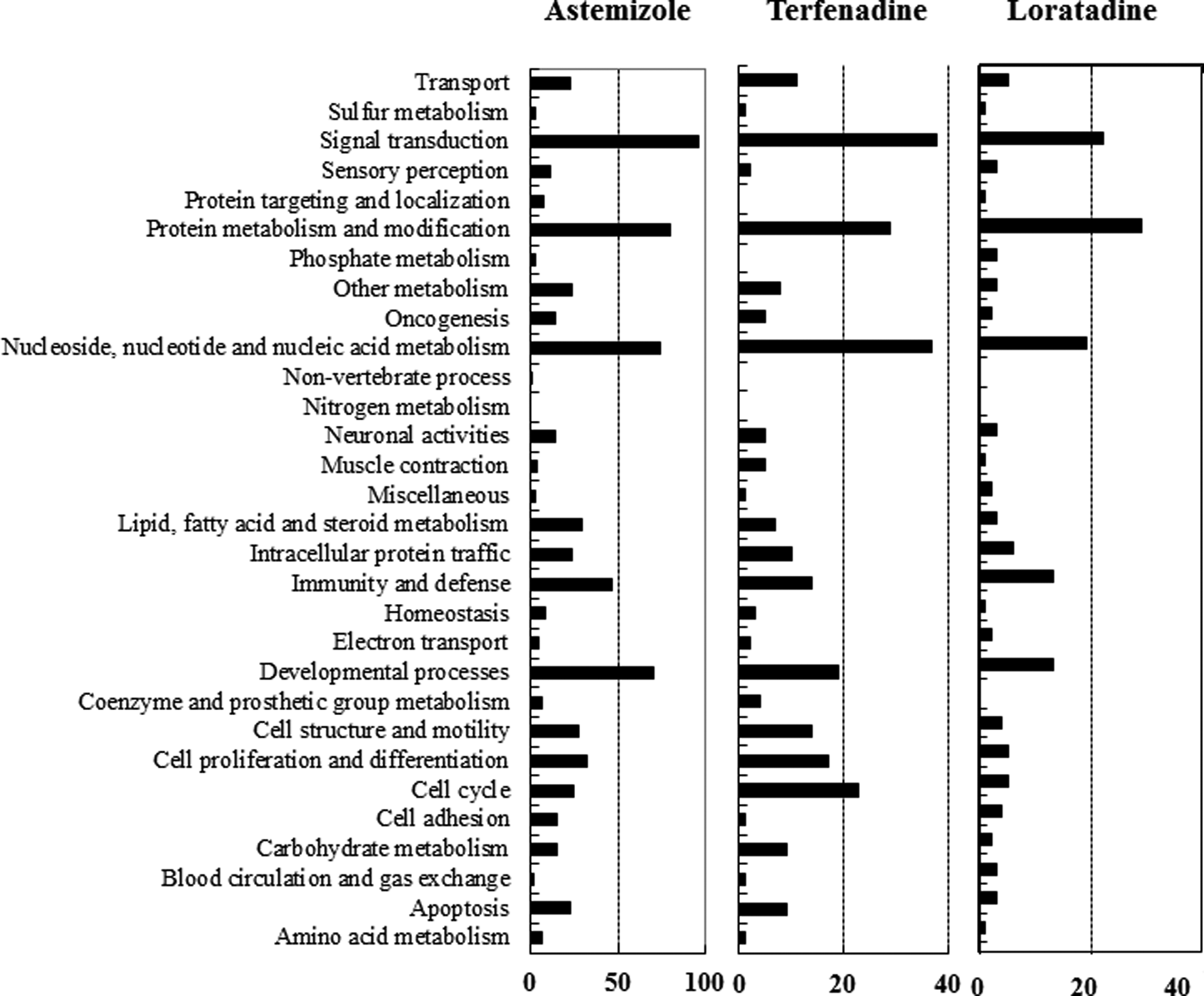
PANTHER biological process classification of genes differentially expressed by astemizole, loratadine, and terfenadine in H9C2 rat cardiomyocytes. This figure represents various gene families that are overrepresented or underrepresented by antihistamine treatments. The number of genes for each gene family that were differentially represented are shown in the figure. Signal transduction is the most affected gene family in antihistamine treatment cells.
Gene expression profile analysis in astemizole-, loratadine-, and terfenadine-treated H9C2 rat cardiomyocytes
To identify genes commonly and specifically expressed in response to treatment with the three antihistamines, Venn diagram analysis was used (Figure 3). Results indicated that the numbers of commonly modulated genes within each drug pair were 53 (astemizole and loratadine groups, Table 2), 9 (astemizole and terfenadine groups, Table 3), and 7 (loratadine and terfenadine groups, Table 4). However, there was only one gene commonly modulated by all these three drugs (Table 5). All genes modulated by astemizole, loratadine, and terfenadine were classified into functional pathways using the PANTHER classification system. As shown in Figure 4, the genes modulated by astemizole, loratadine, and terfenadine were mostly involved in the integrin signaling pathway, interleukin signaling pathway, angiogenesis, inflammation-mediated signaling pathway, and Wnt signaling pathway. The genes that were functionally categorized in the integrin signaling pathway were further analyzed by KEGG pathway mapping. The results from the pathway mapping are shown in Figure 5.

Venn diagram of commonly modulated genes by treatment with astemizole, loratadine, and terfenadine in H9C2 rat cardiomyocytes at concentrations of 0.5, 1, and 5 µM. Number indicates number of genes.

PANTHER pathway analysis of commonly modulated genes by treatment with astemizole, loratadine, and terfenadine in H9C2 rat cardiomyocytes. The number of genes for each pathway that were differentially represented are shown in the figure. Intergrin signalling pathway is the most affected signal transduction in antihistamine treatment cells.

The KEGG pathway map of gene expression data from astemizole, loratadine, and terfenadine in H9C2 rat cardiomyocytes. A rectangle and a circle represent, respectively, an enzyme and a compound. The enzymes whose genes are identified in the genome are shown by colored (shaded) rectangles. KEGG: Kyoto Encyclopedia of Genes and Genomes.
Commonly modulated genes in astemizole and loratadine groups.a

aResults indicate the numbers of commonly modulated genes within each drug pair. Genes without any known function are not shown.
bPredicted gene symbols.
cPredicted gene names.
d
Commonly modulated genes in astemizole and terfenadine groups.a

aResults indicate the numbers of commonly modulated genes within each drug pair. Genes without any known function are not shown.
bPredicted gene symbols.
cPredicted gene names.
d
Commonly modulated genes in loratadine and terfenadine groups.a
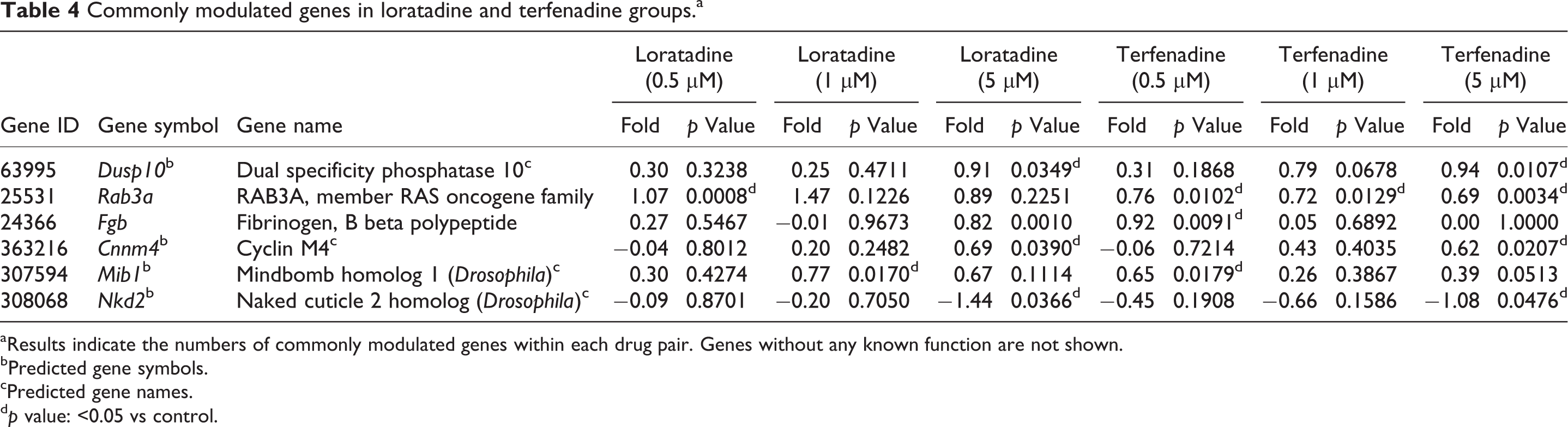
aResults indicate the numbers of commonly modulated genes within each drug pair. Genes without any known function are not shown.
bPredicted gene symbols.
cPredicted gene names.
d
Commonly modulated genes in astemizole and loratadine groups.a

aResults indicate the numbers of commonly modulated genes within each drug pair.
bPredicted gene symbols.
cPredicted gene names.
d
Discussion
Antihistamines, such as astemizole and terfenadine, are associated with TdP-type arrhythmias, arrhythmias syncope and even sudden death. The main mechanism of antihistamines-induced cardiovascular toxicity is the iKr current block, which play a role in QT interval prolongation, a biomarker of multiple arrhythmias. However, it is well recognized that a QT alone is not adequate for assessment of drug-induced cardiac toxicity, because QT prolongation has a poor correlation with occurrence of TdP.
13
In addition, it has also been reported that hERG block is not necessarily related to increased arrhythmic risk. Thus, a large body of research has been directed at identifying new biomarkers of drug cardiotoxicity.
14
In this study, DNA microarrays were used for identifying and analyzing putative biomarkers that are regulated by the treatment of cardiomyocytes with astemizole, loratadine, and terfenadine. PANTHER analysis revealed that genes of the integrin signaling pathway were most commonly modulated by astemizole, loratadine, and terfenadine treatment. Integrins play an important role in directing membrane localization and functional association of ion channels,
15,16
especially HERG potassium channels and L-type calcium (Ca2+) channels,
17
which are known to be involved in QT prolongation during cardiac arrhythmia. Furthermore, a study of gene expression profile of the myocardial response to QT prolonging drug reveals integrin signaling as novel pathways of action.
18
From our DNA microarray studies, we identified 53 genes that were commonly modulated by astemizole and loratadine in H9C2 rat cardiomyocytes, of which 3 were involved in integrin signaling. Spermidine/spermine N1-acetyltransferase 1 is a rate-limiting enzyme in polyamine metabolism
19
that binds specifically to the integrin cytoplasmic domains and enhances integrin-mediated cell migration in leukocytes.
20
Ras-associated binding guanosine triphosphatases (Rab-GTPases) associate with integrin subunits and play an important role in integrin trafficking, which regulates membrane protrusion activity.
21
Rab33a, a member of the Ras oncogene superfamily of small GTPases, localizes to the Golgi apparatus and mediates vesicular transport in rat embryonic hippocampal neurons.
22
The Golgi apparatus is important for normal protein processing, which includes processing of the hERG protein that contributes to the pathogenesis of long QT syndrome.
23–25
G protein-coupled receptor 30 (GPR30), a member of the rhodopsin-like family of G protein-coupled receptors, is a multi-pass membrane protein that localizes to the endoplasmic reticulum.
26
GPR30 plays an important role in cardiac arrhythmic pathogenesis by binding with estrogens, which are a group of steroid hormones that regulate the activity of cardiac ion channels and modulate cardiac repolarization, causing QT prolongation.
27
GPR30 also coordinates estrogen-mediated fibronectin matrix assembly and promotes the formation of Src-dependent, Shc-integrin α5β1 complexes,
28
which modulate L-type calcium channels.
29
One of the genes that were commonly modulated with astemizole and terfenadine encoded a chemokine ligand, chemokine (C–X–C motif) ligand 1 (CXCL1), which is also involved in integrin signaling. CXCL1 is a member of the CXC subfamily of chemokines, which binds the G protein-coupled receptor chemokine CXC receptor 2 (CXCR2)
30
and facilitates the activation of β2 integrins.
31
Loratadine and terfenadine treatment commonly modulated the
Footnotes
Conflict of interest
The authors declared no conflicts of interest.
Funding
This work was supported by Ministry of Food and Drug Safety grant.
