Abstract
Carbon tetrachloride (CCl4) is a common hepatotoxin used in experimental models to elicit liver injury. To identify the proteins involved in CCl4-induced hepatotoxicity, two-dimensional gel electrophoresis was employed followed by mass spectrometry - mass spectrometry (MS/MS) to study the differentially expressed proteins during CCl4 exposure in the Fischer 344 rat liver proteome for 5 weeks. Ten spots with notable changes between the Control and CCl4 groups were successfully identified. Among them, four proteins with significant up-regulation, namely calcium-binding protein 1, protein disulfide isomerase, mitochondrial aldehyde dehydrogenase precursor, and, glutathione-S-transferase mu1 and six proteins with significant down-regulation, namely catechol-
Introduction
Carbon tetrachloride (CCl4)-induced hepatotoxicity is the standard use of CCl4 as a liver toxin to induce liver injury. It is a well-investigated experimental model used to study the principles of hepatotoxic-induced lipid peroxidation. 1 CCl4 is a commercial solvent used in machine cleaning and as a household agent for the removal of stain in the past and was found to cause intoxification. 2 The hepatotoxicity was found to be the result of inflammation initiated by oxidative stress, free radicals and lipid peroxidation. 3 CCl4-induced hepatotoxicity mediates oxidative stress by cytochrome P450 isoenzymes, resulting in trichloroperoyl free radical metabolites formation from a chain of events involving plasma membrane in lipid peroxidation. 3 The free radical metabolites form from the polyunsaturated fatty acids of the cell membranes will adversely affect the structural component of the membrane, resulting in alterations in fluidity or permeability of membranes, cell energy processes destruction and protein synthesis. 4-7 Such mechanisms are believed to be the cause of liver injury by CCl4.
Proteomics has gained increasing popularity in the investigation of quantitative change in protein expression in biological systems. 8,9 Such techniques provide bioinformatics for obtaining a better understanding of the disease pathogenesis. Differences in protein expression have been recorded in liver cancer with the use of proteomic analysis. 10 In the current study, a two-dimensional gel-based proteomic approach was adopted to examine the liver proteome changes in response to chronic treatment with CCl4.
Materials and methods
Animals
Adult male Fischer 344 rats (initial body weights = 250−280 g;
Generation of liver injury model
All rats were allowed to acclimatize for a week before the experiment. The rats were randomly assigned into Control and CCl4 groups. The Control group received subcutaneous injection with pure olive oil twice a week for 5 weeks. The CCl4 group received subcutaneous injection with CCl4 (Merck KGaA, Darmstadt, Germany) diluted 1:1 (v/v) in pure olive oil at a dose of 0.2 mL/100 g of body weight twice a week for 5 weeks. This hepatotoxicity animal model was described previously, with modification. 12
Tissue collection
At the end of experiment, animals were anesthetized with ether and sacrificed. Blood samples were taken by cardiac puncture and the collected serum was used for biochemical analysis. The liver tissue specimens were removed immediately and approximately 1 cm sections of the right lobe were harvested, snap-frozen in liquid nitrogen and stored at –80°C for liver enzyme assays and proteomic analysis.
Biochemical assays
Alanine aminotransferase
Sample preparation for proteomic analysis
Protein extraction method has been described previously.
8
The liver tissue samples were disrupted with a tissue teaser (Biospec Products, Oklahoma, USA) in a cocktail buffer (1 % Triton X-100, 25 mmol/L HEPES, 150 mmol/L NaCl, 1 mmol/L EDTA disodium salt, 1 mmol/L dithiothreitol [DTT]) with added Protease Inhibitor Cocktail Set III (Bio-Rad, California, USA). The superfluous salt in the supernatant was removed by incubating with trichloroacetic acid (TCA)-acetone solution (20 % TCA, 20 mmol/L DTT in acetone) for 4 h at –40°C. The protein pellet was obtained by centrifugation at 15,800 x
Two-dimensional gel electrophoresis and MS/MS analysis
The two-dimensional gel electrophoresis (2-DE) procedures used here have been previously described by us with modifications.
9
A total of 10 gels (5 for Control and 5 for CCl4) were conducted, with one gel per animal from each of the time points. For the first dimensional electrophoresis of proteins, a fixed amount of 125 µg protein samples were loaded onto an 18 cm ReadyStrip immobilized pH gradient (IPG) Strips (p
Image acquisition and analysis
Image analysis and protein identification
The Molecular Imager PharosFX Plus System (Bio-Rad) was used to scan stained 2-DE gels. PDQuest 8.0 for Windows (Bio-Rad) software was used for matching and analyzing protein spots on the 2-DE gels, with the function on background subtraction, spots detection and volume normalization. Detected spots was normalized and assigned with a number referenced by a ‘virtual gel’ or a reference gel (the best gel of the 10 gels) automatically. Any under-detected spots were manually assigned with the number according to the reference gel by the researcher. The expression level, expressed as percentage volume (% vol), was exported for statistical analysis. Protein spots with differential expression between Control group and CCl4 group (
Protein identification by MS/MS
Spots with differential expression (
Western blotting analysis
Liver protein extract was mixed with sample buffer (62.5 mmol/L Tris, pH 6.8, 25% glycerol, 2% SDS, 350 mmol/L DTT, 0.01% bromophenol blue). Thirty µg protein of each sample was applied for electrophoresis on 12.5% SDS-PAGE gels with constant voltage (125 V) and transferred to polyvinylidene diflouride membrane. The membranes were incubated with primary antibodies and subsequently with secondary antibodies as listed below: rabbit polyclonal against Catalase (CAT) or protein disulphide isomerase (PDI; secondary antibody: goat anti-rabbit conjugated with horseradish peroxidase); goat polyclonal against catechol-
Statistical analysis
All data were analyzed by SPSS 11.5 software and reported as means ± SEM using one-way ANOVA. Student’s t-test was used for between-groups comparison.
Results
Body weight and liver weight
At the end of experiment, the mean body weight of the rats in the CCl4 group was 287.4 ± 19.96 g and in the Control group was 312.4 ± 9.6 g, with no significant difference (Figure 1A). In contrast, the mean liver weight in the CCl4 group (22.9 ± 0.72 g) was significantly heavier than those in the Control group (16.55 ± 0.32 g;
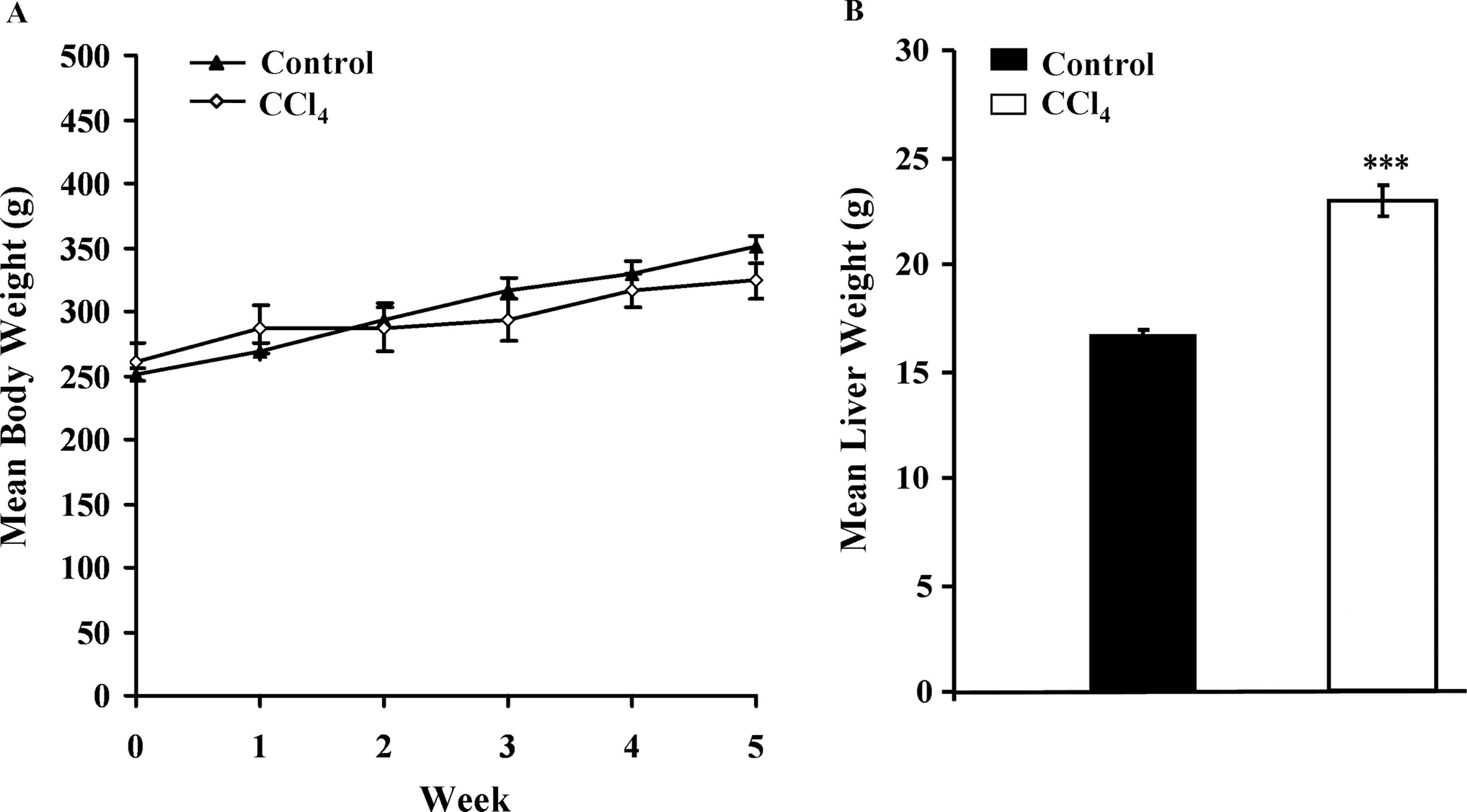
Effects of carbon tetrachloride (CCl4) on mean body weight (g) (A) and mean liver weight (g) (B). Male F344 rats were subcutaneously injected with or without CCl4 (0.2 mL/100g/twice a week) for 5 weeks. Values are mean ± SEM for five rats per group. ***
Biochemical assay of liver function CCl4-induced hepatotoxicity
The data revealed that the administration of CCl4 compared to Control significantly increased the serum activity of ALT by 40-fold (2798.2 ± 193.9 U/L in CCl4 group and 65.02 ± 3.2 U/L in Control group;
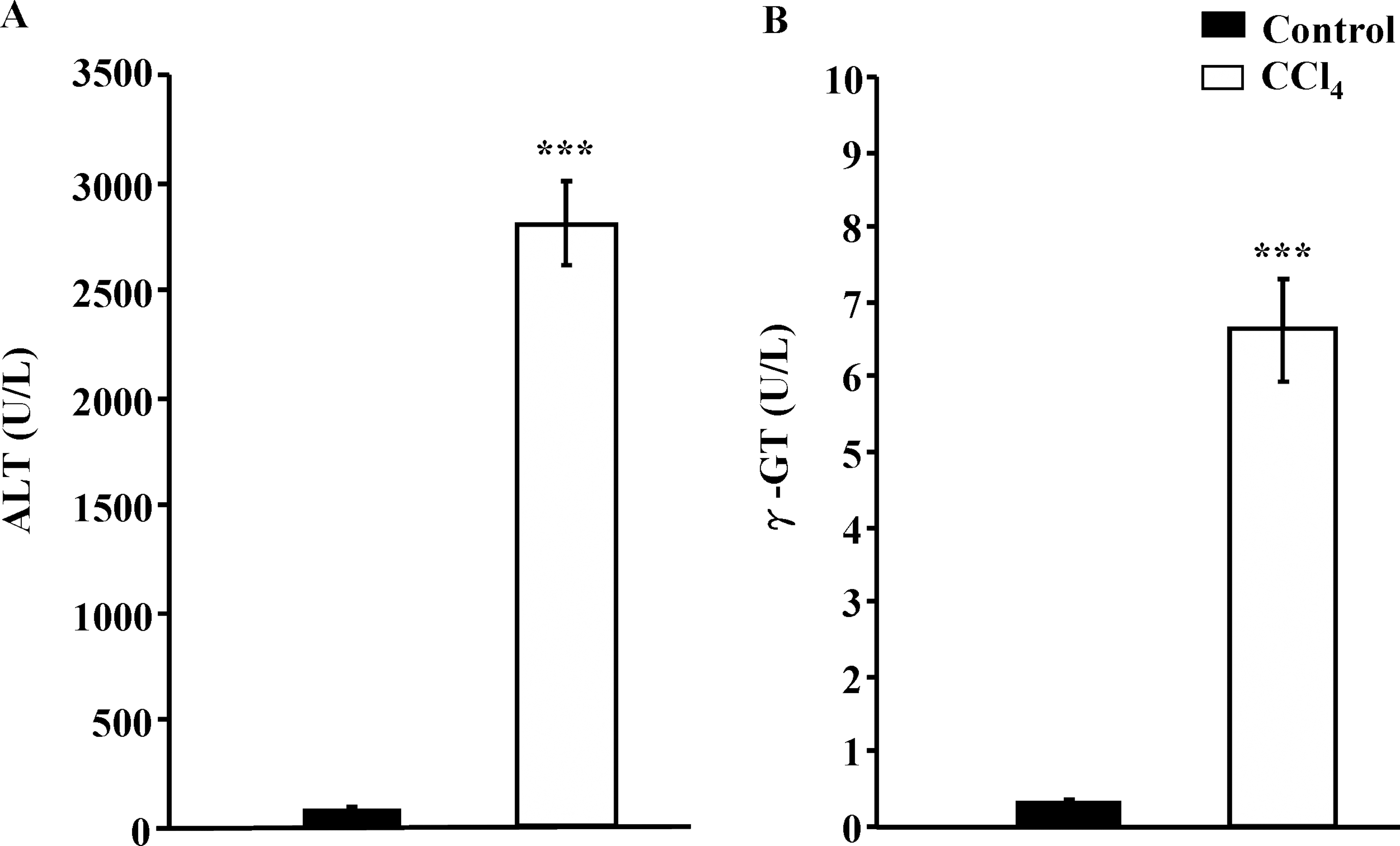
Serum biochemical assays of alanine aminotransferase (ALT) (A) and γ-glutamyl transferase (γ-GT) (B) liver function in response to carbon tetrachloride (CCl4)-induced hepatotoxicity. Rats were treated with or without CCl4 for 5 weeks. Serum samples were assayed for the activities of ALT and γ-GT. Values are mean ± SEM for five rats per group. ***
GSH and GST assays on glutathione metabolism in CCl4-induced hepatotoxicity
Subcutaneous administration of CCl4 significantly depleted hepatic GSH concentration by 74% from normal rats 6.68 ± 0.05 µmol/g to 1.75 ± 0.33 µmol/g (

Liver enzyme assays of reduced glutathione (GSH) (A), glutathione-S-transferase (GST) (B), catalase (CAT) (C) and glutathione peroxidase (GPx) (D) in response to carbon tetrachloride (CCl4)-induced hepatotoxicity. Rats were treated with or without CCl4 for 5 weeks. Liver homogenate samples were assayed for the activities of GSH, GST, CAT and Gpx. Values are mean ± SEM for five rats per group. **, ***
CAT and Gpx activities in CCl4-induced hepatotoxicity
CCl4 significantly reduced hepatic CAT activity by 74% from normal rats of 5.14 ± 0.21 mmole H2O2/min/mg to 1.34 ± 0.18 mmole H2O2/min/mg (
Liver proteome profile and protein identification
There were approximately 500 well-resolved protein spots detected on each 2-DE gels. Ten spots with notable changes between the Control and CCl4 groups were successfully identified (Figure 4). The subcellular locations were examined and the basic information on the identified proteins with a lower or higher expression level (

Representative 2-DE gels of a rat liver proteome map, with or without the administration of carbon tetrachloride (CCl4) for 5 weeks (
Rat liver proteins exhibiting differential expression with or without the administration of carbon tetrachloride (CCl4)
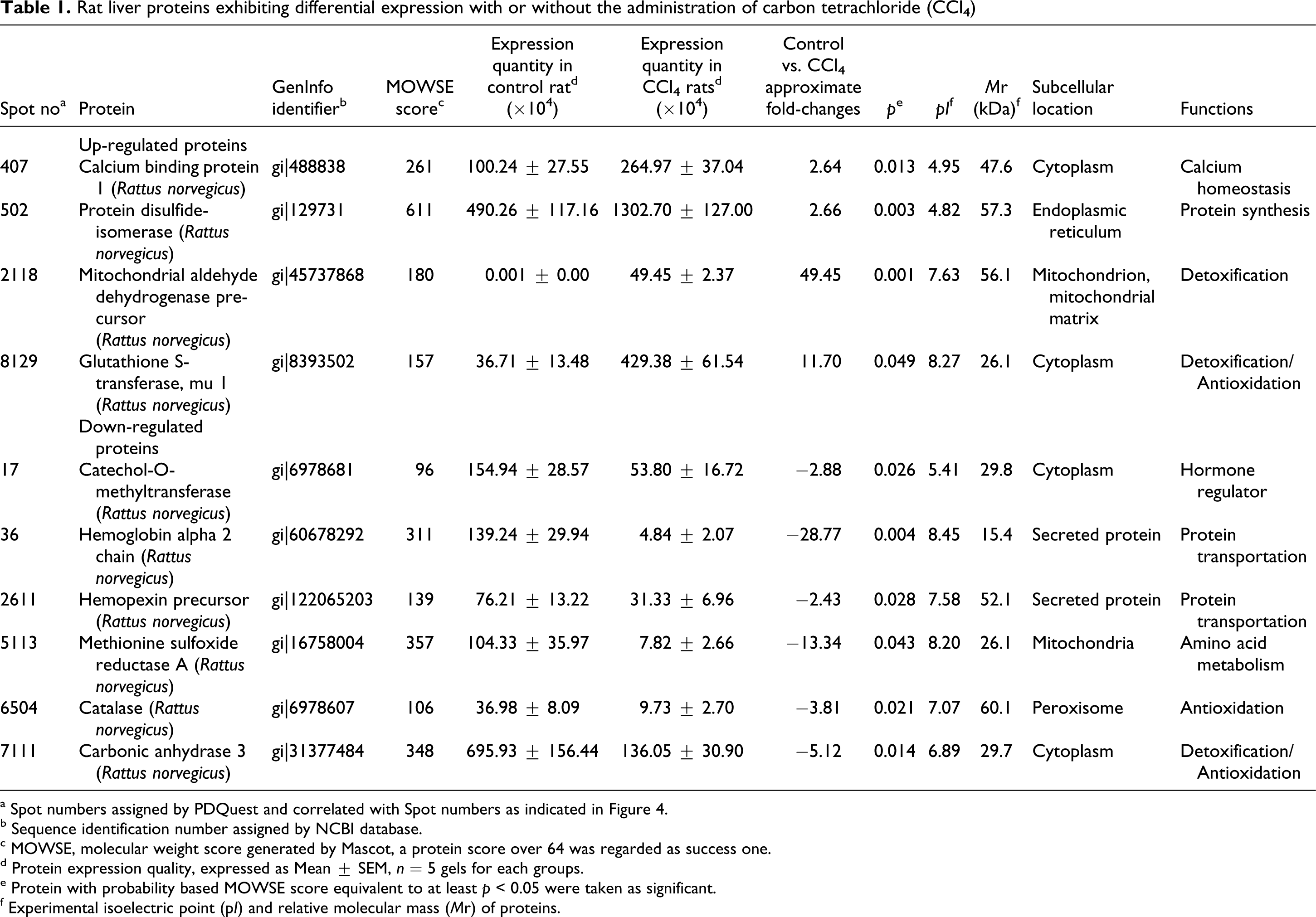
a Spot numbers assigned by PDQuest and correlated with Spot numbers as indicated in Figure 4.
b Sequence identification number assigned by NCBI database.
c MOWSE, molecular weight score generated by Mascot, a protein score over 64 was regarded as success one.
d Protein expression quality, expressed as Mean ± SEM,
e Protein with probability based MOWSE score equivalent to at least
f Experimental isoelectric point (p
Western blotting confirmation
The results are similar to those of the 2-DE proteomic analysis (Table 1), showing both COMT and CAT were reduced (

Western blot analysis of liver proteins, catechol-
Discussion
In this study, we demonstrated that the injection of CCl4 for 5 weeks in rats led to a marked elevation in the levels of serum ALT and γ-GT. Both the cytoplasmic ALT and γ-GT released into the circulation after cellular damage have been considered as hepatic damage biomarkers in the diagnosis of liver injury. 18
Our data (Figure 3) indicates the presence of a suppression of the antioxidant system in the liver of rats treated with CCl4 since GSH is an antioxidant that helps protect cells from reactive oxygen species (ROS), such as free radicals and peroxides. In the liver, CCl4 is metabolized by cytochrome P450 in the endoplasmic reticulum to produce the highly reactive trichloromethyl radical, which is further converted to the peroxytrichloromethyl radical. This free radical can extract hydrogen from different molecules, thus initiating the oxidation of lipids, proteins and DNA in liver injuries. 19
In order to construct a database to improve our understanding of hepatotoxicity and of potential diagnostic or therapeutic protein target discovery, we applied a comprehensive analysis of proteins associated with the hepatotoxicity induced by chronic exposure of rats to CCl4 by using a combination of two-dimensional electrophoresis (2-DE) and mass spectrometry. The 2-DE proteomic technique is useful for systematic analyses of global protein abundance profiles of normal and diseased tissues, often providing novel insights into the pathogenic mechanisms of human disease as well as disease-associated targets. In the present study, we have successfully identified 10 proteins from the NCBI database, and their importance in the pathophysiology of CCl4-induced hepatotoxicity is discussed below under the following clusters: detoxification and antioxidation, hormone regulation, amino acid metabolism, protein transportation and protein synthesis and calcium homeostasis (Table 1).
Detoxification and antioxidation
Carbonic anhydrase 3 (CAR3) is a member of the carbonic anhydrase family of zinc metalloenzymes. Apart from aiding the transport of carbon dioxide out of tissues, it may also serve as an oxygen radical scavenger to protect hepatocytes from oxidation damage. The enzyme contains two reactive sulfhydryl groups (Cys 181 and Cys 186) that readily form disulfide linkages with GSH, a process termed

Proposed inter-relationships of the differentially expressed liver proteins in carbon tetrachloride (CCl4)-induced hepatotoxicity
Glutathione-
Mitochondrial ALDH2 was significantly increased in CCl4-treated rats. This protein is responsible for eliminating xenobiotic aldehydes and toxic biogenic agents 34-36 and also has a major role in acetaldehyde detoxification. 37 It is believed that it may play a protective role in the mechanism by which hepatocytes combat sudden increases in aldehyde products. The 49-fold increase of ALDH2 is a strong indication of the presence of increased lipid peroxides in the CCl4-treated liver. In this pathway, the intermediate structures can be toxic and health problems may arise when those intermediates cannot be cleared 38 (Figure 6).
Catalase (CAT) is a ROS scavenger responsible for the breakdown of H2O2. The reduced expression of catalase and glutathione peroxidase (GPx) in hepatocytes is generally correlated to the decrease in the GSH/GSSG ratio and increase in lipid peroxidation. 39 In the present study, the hepatic CAT activities was significantly decreased by 74% (Figure 3C) and there was a reduced expression of hepatic GPx activity by 31% when CCl4 was administered (Figure 3D). We conclude that the fourfold decrease in CAT in liver proteome could lead to an increase in hepatocyte lipid peroxidation in the animals (Figure 6).
Amino acid metabolism
The present study shows that CCl4 damage to the rat liver led to a decrease in the abundance of methionine sulfoxide reductase A (MSRA), which is responsible for catalyzing the thioredoxin-dependent reduction of free and protein-bound methionine sulfoxide [Met(O)] residues to methionine. According to previous studies, it was indicated that mouse lacking the MSRA gene exhibited enhanced sensitivity to oxidative stress when compared with the wild type. It is believed that MSRA may possibly play an important role as an antioxidant in mammals. 40 The 13-fold depletion of MSRA found in this study may have resulted in a decrease in the reduction of Met(O) residues to methionine, a ROS scavenger, which could increase the susceptibility of the rat liver to oxidative stress imposed by ROS (Figure 6).
Hormone regulator
A threefold depletion of catechol-
Protein transportation and protein synthesis
We observed that CCl4 treatment caused twofold decrease in Hemopexin precursor (HPX) and 29-fold decrease in Hemoglobin alpha 2 chain (HBα2). Since both of them are transporters of heme to liver parenchymal cells and active in heme scavenging,
43
the reduction of HPX and HBα2 could result increased level of free heme, which is highly toxic as it promotes the generation of ROS and thus promoting oxidative damage.
44,45
Moreover, heme is a positive regulator of cytochrome P450 gene transcription.
46
It is believed that free radicals (CCl3
- and CCl3OO-) generated by the cytochrome P450-dependent detoxification step could induce liver injury by lipid peroxidation (Figure 6). The increase in cytochrome P450, however, cannot be shown by proteomic analysis in the present study, due to the fact that cytochrome P450 proteins are endoplasmic reticulum membrane proteins with a very high isoelectric point (
The PDI, which exhibited a threefold increase in the present study, is a multifunctional protein. PDI plays important roles in corrective protein folding and as a cellular chaperone. 49 It is also involved in the maturation of collagen helix, which is a landmark of liver fibrosis which is increased in liver injury. The intrinsic function of this protein, however, is believed to be the rearrangement of disulfide bridges and the catalysis of oxidation, reduction and corrective disulfide isomerization so as to prevent unnecessary aggregation. 49 Under the oxidative stress induced by CCl4 consumption, proteins are highly susceptible to cysteine oxidation and the incorrect formation of disulfide bridges, leading to detrimental change in the 3-dimensional structure of the protein and aggregation. 49 Moreover, it is hypothesized that under significant oxidative stress in the injured hepatocytes, the increased abundance of PDI production is to compensate for the demand of corrective disulfide isomerization, which is needed to stabilize the proteins susceptible to aggregation (Figure 6).
Calcium homeostasis
CABP1 is a high-affinity calcium-binding glycoprotein of the rat liver endoplasmic reticulum. 50 Studies have indicated the cytosolic calcium level is elevated after exposure to hepatotoxins such as CCl4. This elevated cytosolic Ca2+ is believed to be the result of an activation of calcium-releasing channels. 51-53 The threefold increase in CABP1 found in the present study supports the fact that CCl4 metabolism may play a role in the activation of calcium release into the cytosol (Figure 6). The high levels of calcium in the cytosol activate calcium-dependant proteases that exacerbate membrane damage. The increase in intracellular calcium can activate endonucleases that inflict chromosomal damage that in turn contributes to cell death and cancer. 54
In summary, CCl4 is a hepatotoxicant that generates ROS and initiates lipid peroxidation and induces significant liver injury in the rats. We propose that the hepatotoxicity mechanism by which CCl4 induced liver injury in the rats is mostly due to the suppression of the antioxidant and detoxification system. Also, in this study, we showed that proteomic technology can serve as a powerful tool for the rapid identification and confirmation of proteins with significant roles in the pathogenesis of liver injury during toxic chemical exposure. The liver proteomic analysis approach hastened the discovery of significant proteins that are involved in the pathogenesis of the liver injury induced by the hepatotoxicant. MS/MS-based sequencing of peptides derived from these differentially expressed proteins has helped identify both protein markers that had previously been cited as possible indicators as well as potentially novel associations with hepatic effects for proteins already established by other investigations. The present data facilitate not only the molecular mechanistic investigation of CCl4-induced liver injury but also provide new direction in the study of disease pathogenesis.
Footnotes
This work was supported by the grants of the Liver Strategic Research Grant by The University of Hong Kong.
