Abstract
The scale, severity, and synchronicity of recent outbreaks of forest pests such as bark beetles (Coleoptera, Scolytinae) and defoliators (Lepidoptera, Choristoneura) within coniferous forest ecosystems of North America, Europe, and Asia are widely regarded as ‘unprecedented’. Despite such devastating outbreak occurrence in recent times, very little is known about historic outbreak occurrence. Traditional methods of reconstructing historic outbreak dynamics, including dendroecology, pollen analysis, and the identification of fossilised pest remains, all have critical weaknesses in their ability to reconstruct such outbreaks accurately, notably non-standardised methodologies, varying parameters for identifying outbreak periods within proxy records, and a bias towards the detection of large-scale, highly destructive outbreaks only. The development of a more accurate detection tool to reconstruct historic outbreak dynamics within the palaeoecological record has been prioritised as one of the top 50 areas of research within Quaternary science. This paper assesses the current methodologies, before presenting the potential role of DNA-based methodologies can play in overcoming some of these limitations and providing more comprehensive reconstructions, and critically, direct detection of historic forest pathogen outbreaks.
Introduction
The detection of insect and disease outbreaks within the palaeoecological record has been identified as one of the top 50 priority research areas within the field of Quaternary science (Seddon et al., 2014), emphasising the demand for widely applicable, standardised methodologies capable of the direct detection of historic outbreaks (Brunelle et al., 2008; Morris et al., 2010; Morris and Brunelle, 2012; Raffa et al., 2008). The need for a deeper understanding of historic forest pathogen dynamics has been heightened by the increased scale, severity, and synchronicity of recent outbreaks of agents such as bark beetles (Coleoptera, Scolytinae) and defoliators (Lepidoptera, Choristoneura) within coniferous forest ecosystems of North America, Europe, and Asia. These outbreaks are regarded by forest managers and conservationists, as well as within the wider ecological literature, as ‘unprecedented’ (Bentz et al., 2010; Brunelle et al., 2008; Mitton and Ferrenberg, 2012; Morris et al., 2015; Raffa et al., 2008), yet this determination lacks critical evaluation based on palaeoecological evidence. As relatively little is known of historic outbreak dynamics (Morris et al., 2015), understanding of the spatial and temporal dynamics of past forest disturbances and of the naturally occurring resistance and resilience of these forest ecosystems remains limited (Hessburg et al., 2019; Morris et al., 2017).
Past dynamics of some key disturbance agents, such as fire, are relatively well understood (Bergeron et al., 2010; Brunelle et al., 2008; Flannigan et al., 2001; Long et al., 1998, 2007) and methodically assessed within the palaeoecological record, for example, the presence and morphology of charcoal particles, which can indicate occurrence and severity of historic fire events. However, unlike wildfires, unequivocal evidence of insect pest outbreaks is rarely preserved (Hebertson and Jenkins, 2008; Jenkins et al., 2008; Morris et al., 2013) and only the inferred indirect effects these agents inflict on forest communities can typically be measured. In addition, the reconstruction of historic insect outbreaks has been largely neglected owing to the absence of an effective detection tool (Montoro Girona et al., 2018). Consequently, most of the information is derived from proxy evidence of recent known, large-scale, highly destructive events, such as the 1940s spruce beetle (Dendroctonus rufipennis) outbreak in Colorado, USA, which killed over 90% of Engelmann spruce (Picea engelmanii) in White River National Forest (Eisenhart and Veblen, 2000). This bias towards large-scale events results in a skewed understanding of forest pathogen occurrence and impact (Blomquist et al., 2016).
The knowledge gaps associated with historic forest pathogen occurrence are compounded by a lack of standardised detection methods, which lack commonly accepted quantitative thresholds to identify periods of outbreak, and a bias towards calibration on large-scale outbreaks only. The ability of these methods to detect smaller, more localised outbreaks that still affect the overall forest ecology or create ideal conditions for subsequent outbreaks is unknown. The development of more sensitive, direct detection of disturbance agents over longer timescales is required to understand historic outbreak dynamics. Here, we assess the current methods of detecting and reconstructing historic forest insect outbreaks, focussing on two key aspects of outbreak dynamics – to what degree current methods are capable of detecting varying levels of disturbance and whether they are able to distinguish the responsible agent(s). This is followed by an exploration of the potential role of DNA-based methodologies within the field of palaeoecology.
Current methods of reconstructing historic forest insect outbreaks
Baseline knowledge of historic bark beetle dynamics comes from traditional palaeoecological-based methods such as dendroecology (Alfaro et al., 2010; Berg and Anderson, 2006; Smith et al., 2012), palynology (Morris et al., 2010; Morris and Brunelle, 2012), and the identification of subfossil insect remains (Brunelle et al., 2008; Clark and Edwards, 2004; Girling, 1988; Girling and Greig, 1985; Morris et al., 2015; Schafstall et al., 2020). These methods provide information on historic occurrence events and inferred impacts on vegetation communities over longer timescales (Figure 1) (Meddens and Hicke, 2014). Each of these methodologies has associated limitations (Figure 2), and rarely pre-date the 20th Century, with the majority of high-quality reconstructions focussing on the 20th Century itself (Brunelle et al., 2008; Morris et al., 2015). Fossil pollen can provide long-term records of ecological disturbance; however, fluctuations in pollen abundance provide only evidence of inferred impacts of forest insect pests, while dendroecological assessments require trees to survive and subsequently recover, which is a rare occurrence during particularly severe outbreaks. These studies highlight an important principle, that when insect outbreaks reach epidemic levels, they can be ‘directly’ (through insect remains) or ‘indirectly’ (through their impacts) detectable within environmental samples (Brunelle et al., 2008). Yet, owing to the absence of effective direct detection tools (Montoro Girona et al., 2018), the reconstruction of insect outbreaks, including bark beetles and defoliators, has been greatly underrepresented, with only a small number of notable examples (Brunelle et al., 2008; Clark and Edwards, 2004; Girling and Greig, 1985; Schafstall et al., 2022; see also Schafstall et al., 2022 and references therein). It should be highlighted that GIS/ remote sensing methodologies can also provide information on forest pathogen occurrence and dynamics; however, owing to their inability to reconstruct such outbreaks before the mid- to late-20th century, they are not discussed here. This paper aims to provide a review of the most widely applied methodologies of reconstructing forest insect pest outbreaks and assess to what extent the lack of standardised methods and an absence of quantitative thresholds indicative of periods of infestation have impeded understanding of insect outbreaks beyond recorded history.
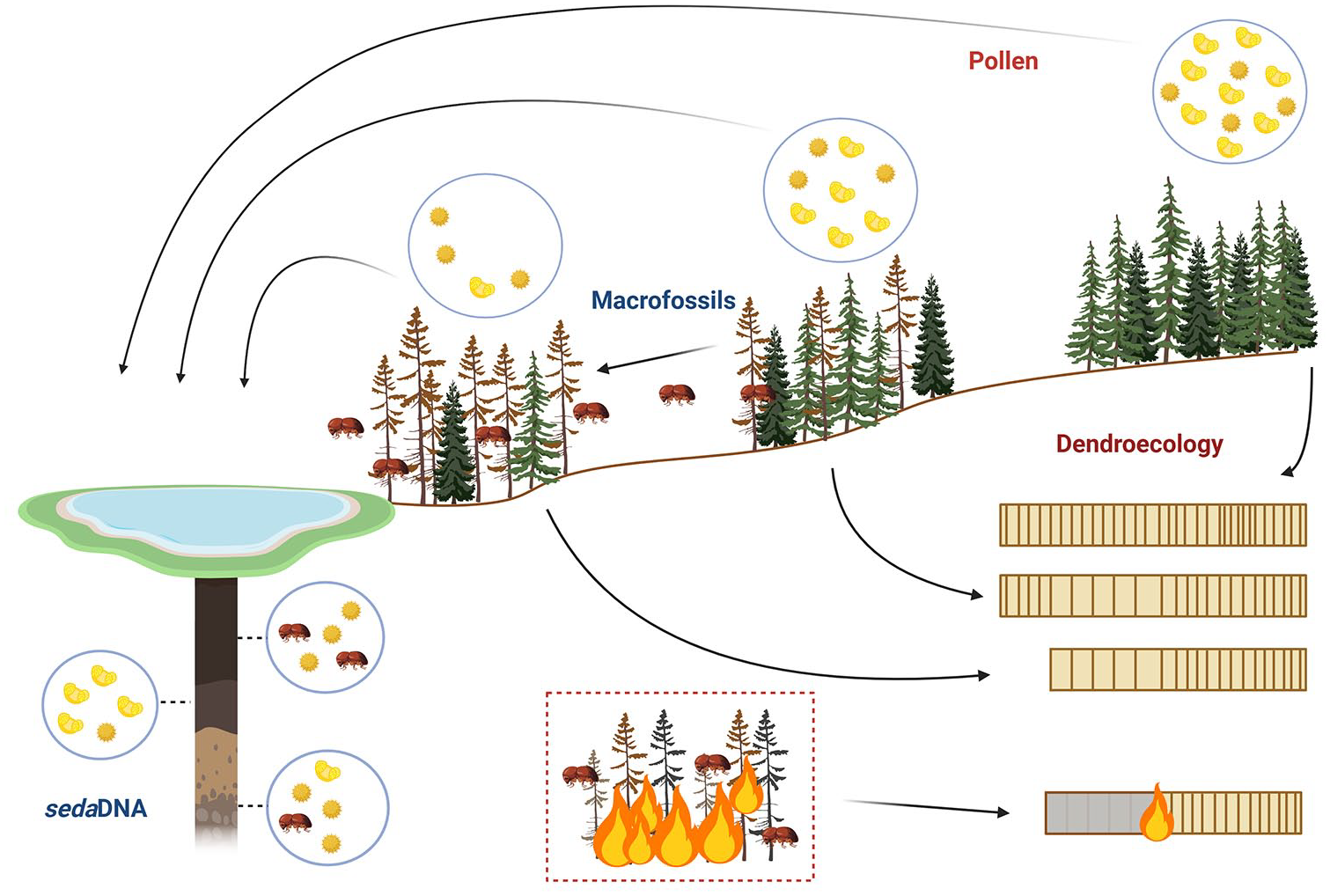
Current methods of detecting and reconstructing historic insect outbreaks, highlighting how signals are obtained for each method.
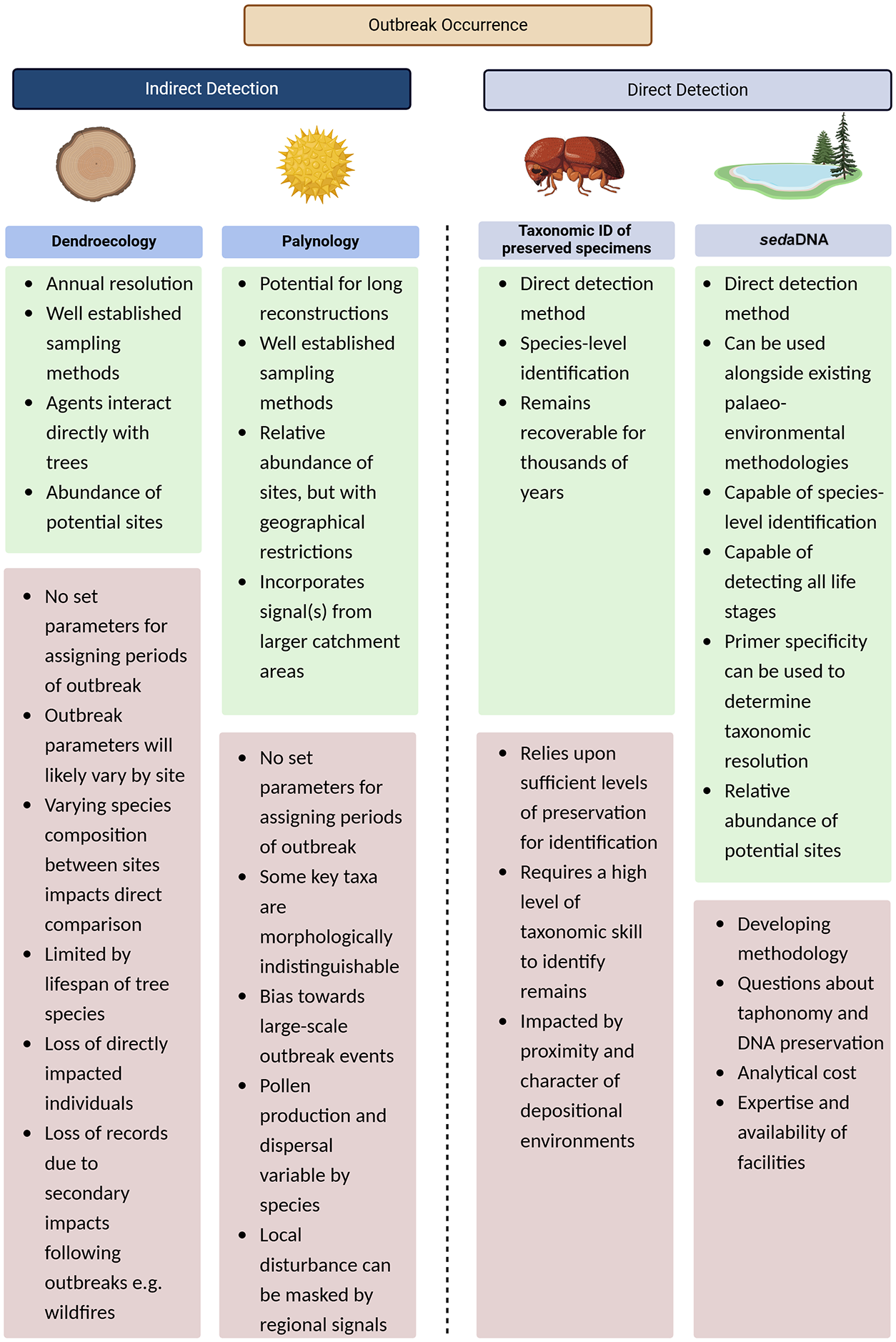
A summary of advantages and disadvantages of each common method used to detect and reconstruct historic insect outbreaks.
Dendroecology
Bark beetles spend most of their life history within host trees (Raffa et al., 2008). As such, measuring the effects beetles induce on attacked trees and the surrounding vegetation forms a significant proportion of research concerning modern and historic bark beetle dynamics. The methods associated with dendroecological sample collection are relatively standardised (Alfaro et al., 2010; Davies and Loader, 2020; McCarroll and Loader, 2004; Young et al., 2010). It is, therefore, the interpretation of the tree-ring measurements where some level of disparity within these methodologies arises.
To identify historic disturbance events, tree-ring data are typically deconstructed into a background component and a peak component. The background component usually consists of a statistical analysis to identify the expected levels of tree growth, while the peak component is calculated using an index to assess the rate of growth in relation to expected levels (Alfaro et al., 2010; Axelson et al., 2010). These indices identify periods whereby tree growth diverges from expected, that is, values > 1 indicate a period when tree growth is more than expected, while values < 1 indicated periods where it is less than expected. These are termed accelerated or restricted growth, respectively. This type of analysis constitutes the most widely implemented measurement of forest pathogen outbreak events, as measurements serve as a proxy for canopy disturbance and associated growth response of surviving trees (Alfaro et al., 2003; Reid and Robb, 1999). Periods of accelerated growth, termed release, occur as disturbance creates openings within which surviving host trees, non-attacked individuals of the host species, non-host canopy species, or understory species can thrive (Morris et al., 2010). As surviving hosts are rare, particularly during severe outbreaks, most assessments focus on heightened growth rates in non-host species or non-attacked individuals (Alfaro et al., 2003; Campbell et al., 2007; Zhang and Alfaro, 2002).
In most dendroecological reconstructions of bark beetle outbreaks (Table 1), each set of growth rate parameters did accurately identify known outbreaks in the 20th century or several previously unknown outbreaks, but the parameters chosen exhibit high levels of variation that may have led to key ecological data being lost. For example, Alfaro et al. (2004), a study examining mountain pine beetle (Dendroctonus ponderosae) outbreaks in lodgepole pine (Pinus contorta) forests of Chilcotin Plateau, BC, Canada, dismissed release periods less than 5 years in duration, suggesting severe outbreaks would likely cause opening of the canopy for at least 5 years. Using this approach, it is possible that low-level, short duration, or more isolated/scattered outbreaks or outbreaks within mixed species stands would remain undetected, reemphasising the bias towards large-scale outbreaks.
Dendroecological reconstructions of bark beetle outbreaks, emphasising the disparity in agreed definitions for growth release.
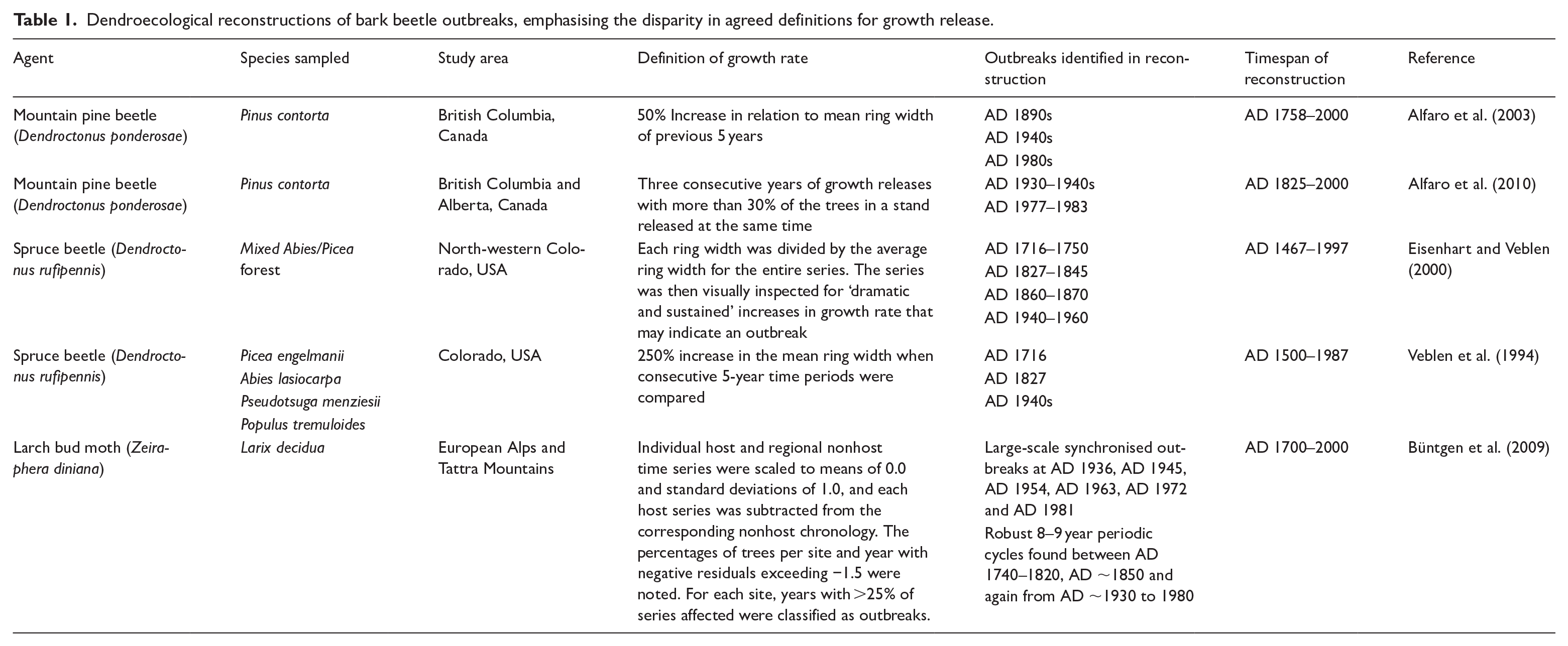
During a reconstruction of mountain pine beetle (D. ponderosae) outbreaks in lodgepole pine (P. contorta) across British Columbia and Alberta, Canada, a series of 121 host and non-host stand chronologies was constructed (Alfaro et al., 2010). Growth release rate was defined as a period in which 30% of trees within a stand exhibited a 25% increase in growth in mean ring width for three consecutive years. Outbreaks in this region were known to occur between AD 1977 and AD 1983 (from aerial detection survey data) and in the late 1930s (from historical records) and were reflected in relative growth rates (Table 1). No other periods of outbreaks were identified within the time series.
A reconstruction of spruce beetle (Dendroctonus rufipennis) outbreaks in Engelmann spruce (Picea engelmannii), in Colorado, USA used a minimum 200% threshold increase in mean ring-width between consecutive 5-year increments (Eisenhart and Veblen, 2000) to determine growth release events attributed to spruce beetle outbreaks. Using these parameters, several outbreaks were identified (in AD 1716–1750, AD 1827–1845, AD 1860–1870, and AD 1940–1960). In a reconstruction of spruce beetle outbreaks in Engelmann spruce, White River Plateau, Colorado, growth release was defined as a 250% increase in the mean ring width when consecutive groups of 5 years or more are compared against the mean ring width for the entire series (Veblen et al., 1994), which varies from the decomposition of the series outlined in the previous study. Standing dead trees were also sampled and successfully cross-dated, extending the length of the chronology to 500 years BP. Evidence of outbreaks were found in AD 1716, AD 1827, and AD 1949. Other studies use tree species-specific growth ring width parameters to identify periods of release, for example Kuosmanen et al. (2020) used a threshold value of 0.55 mm absolute growth increase between subsequent 10-year means for Norway spruce.
The methods used in these examples emphasise a lack of standardised thresholds for several parameters associated with growth release. The amount of growth, the proportion of trees affected, and the period over which these effects must be observed to be deemed a period of disturbance all vary between these studies. Variations in localised conditions, community structure and the subject of the study (host or non-host species), make standardised methods difficult, and subsequent comparisons problematic.
Büntgen et al. (2009) combined tree-ring width data with climate data to produce a 300-year record of larch bud moth (Zeiraphera diniana; LBM) outbreaks across the European Alps and Tatra Mountains. Assessments of residuals of host and non-host chronologies were analysed to identify periods of disturbance. Insect outbreaks were attributed to periods whereby >25% of the series were affected. Cyclic outbreaks of LBM were identified every 8–9 years over the period studied. In addition to the typical assessment of host to non-host growth rates, the introduction of seasonal temperature means allowed the separation of growth response to climate from that caused by insect outbreaks. In addition to the moderation of tree response attributed to outbreak–induced growth release, as described by Büntgen et al. (2009), inclusion of climate data could also potentially be used in historic reconstructions to identify periods of high climatic suitability for insect population increase – for example, warmer winters leading to greater overwintering numbers or an earlier flight season leading to greater fecundity.
Tree-ring records are suitable for reconstructing disturbance events, as they offer annual resolution and therefore the possibility of identifying single events over multiple centuries, beyond recorded history (Morris et al., 2017). Trees of the family Pinaceae, which form the primary host of bark beetles, are relatively long-lived, although there is a high level of variation in maximum age among several key bark beetle host taxa. Douglas-fir (Pseudotsuga menziesii), for example, the primary host of the Douglas-fir beetle (Dendroctonus pseudotsugae), is not only a long-lived species, typically averaging ~1200 years, but is also resistant to low- to moderate-level fires owing to its thick bark. Long-term survival of the tree is critical for the longevity of the dendroecological record. In contrast, the mountain pine beetle (D. ponderosae) favour lodgepole pine (P. contorta), a much shorter-lived, typically around 200 years, fire-dependent species (Lotan et al., 1985). Communities comprising fire-dependent species that exhibit stand-replacement fire behaviour, such as lodgepole pine, affect the ability to reconstruct disturbance events as there are frequently no surviving trees available to produce records.
One method of increasing the temporal resolution of dendroecological methods, involves matching diagnostic features, such as a unique ring-width pattern or a fire scar, shared between dead standing trees and live trees (Fritts, 1976), extending the chronologies far beyond recorded history. One notable example of successful cross-dating (Perkins and Swetnam, 1996) facilitated the reconstruction of mountain pine beetle (D. ponderosae) in white bark pine (Pinus albicaulis) in central Idaho USA over the last 1000 years. A series of narrow ring structures (the diagnostic feature within the series which allowed matching of tree rings) in the late 19th Century enabled successful cross-dating of records of multiple age. To identify outbreak periods, each ring width was divided by the mean width for the entire series (the expected growth) to identify peak components indicative of outbreak events. Three outbreak periods were identified, one well-documented period in the AD 1920s and two previously unknown occurrences in AD 1730 and AD 1887. This example is, to our knowledge, the longest dendroecological reconstruction of bark beetle outbreaks.
Mountainous areas where bark beetle outbreaks occur are typically also affected by a variety of other disturbance agents (fire, snow, avalanche, windstorms, flooding) and climatic influences (Kuosmanen et al., 2020). While indirect methods, such as dendroecological reconstruction, imply periods of outbreak, the effects of these other agents cannot always be discounted, nor can the potential for combination of several agents to act in unison in affecting forest structure and, therefore, growth dynamics. These examples show how variation in the ecological dynamics among tree species, along with variation in event duration, host, stand composition, agent, and fire regime complicate uniform analysis, highlighting the difficulty in assigning standardised threshold for determination of growth release events for all stands.
Palynology
Unlike fire, which is a largely non-selective agent, forest pathogens often adapt to exploit a single host species (Maroja et al., 2007). Measurements of single host species decline, inferred from fossil pollen abundance, allow the identification of shifts in vegetation, and therefore, possible periods of pathogenic outbreaks through time. One major advantage of palynology is that under favourable conditions, pollen seemingly preserves indefinitely (Bennett and Willis, 2002), with pollen in sedimentary archives often providing the longest continuous records of environmental change, as well as enabling the inferred identification of significant historic disturbances. Notable examples include the mid-Holocene elm decline, during which widespread decline was observed in the levels of elm (Ulmus spp.) pollen across much of northwest Europe (Andersen and Rasmussen, 1993; Garbett, 1981; Hirons and Edwards, 1986; Huntley and Birks, 1983; Parker et al., 2002; Peglar, 1993; Ten Hove, 1968) and the mid-Holocene hemlock decline observed as a synchronous decline of Tsuga canadensis across much of eastern North America around 5000 BP (Bennett and Fuller, 2002; Davis, 1965; Davis et al., 1980; Davis, 1981; Fuller, 1998; Parker et al., 2002). These have both since been attributed to pathogenic outbreaks likely carried by bark beetle vectors (Parker et al., 2002). More recently, the introduction of balsam woolly adelgid (Adelges picea) to eastern North America in the early 20th Century, has led to a rapid and widespread decline in Tsuga canadensis which is reflected in pollen records across this region (Allison et al., 1986; Calcote, 2003).
While not all events result in such dramatic fluctuations as those outlined above, quantification of these vegetational shifts, and the associated pollen fluctuations, during periods of known outbreaks, allow the identification of these characteristic signatures through time, extending the knowledge of historic occurrences (Parducci et al., 2017). These fluctuations can be assessed through qualitative (relative abundances of pollen) or quantitative (set thresholds for periods of infestation) measurements, both of which are employed in the current literature.
A calibration of a known AD 1986 spruce beetle outbreak in Wasatch Plateau, Utah, (Morris et al., 2010), against pollen signatures retrieved from lake sediments, aimed to identify periods of outbreak qualitatively using a sharp decline in pollen of host Picea engelmanii, coinciding with an increasing relative abundance non-host Abies and understory species, as a parameter for identifying periods of infestation. While the AD 1986 outbreak was identified within the pollen sequence, no other dramatic fluctuation was observed within the 750-year sedimentary sequence (Morris et al., 2010). It is possible that purely qualitative parameters, for example, sharp declines in host species, may fail to identify lower-level outbreaks, as not all outbreaks result in such observable dramatic shifts in pollen abundances.
Pollen-based methods often calculate ratios between key taxa to identify vegetational shifts indicative of disturbance. A high-resolution pollen analysis of the AD 1939-1951 spruce beetle outbreak at Antler Pond, CO, which killed >90% of Picea engelmanii by volume in the area, was used to calibrate a Picea/Abies ratio during severe outbreaks (Anderson et al., 2010). A pre-outbreak ratio of 1.36 was reduced to 0.45 for the 40 years (1955–1996) following the known outbreak. This ratio of 0.45 was used as a threshold in the determination of outbreak periods preceeding AD 1939, but no other periods meeting this criterion for infestation were identified in the record. Morris and Brunelle (2012) scaled up the basin-level approaches of Morris et al. (2010) and Anderson et al. (2010) to determine whether palynological responses are robust at landscape-level assessments. In addition to the Wasatch Plateau study (Morris et al., 2010), Morris and Brunelle (2012) assessed a further six sites across Utah. Combinations of pollen percentage data, influx data and pollen ratios were assessed to provide the closest association with changes in vegetation during known period of outbreak during the 20th century (a decline in spruce coinciding with an increase in fir and understory species). Within this study, the ratio of spruce and fir to total non-arboreal pollen (NAP) most closely reflected these changes, yet no quantitative threshold (i.e. no specific ratio) was determined to identify historic outbreaks.
There are several factors that complicate the direct relationship between pollen abundances/percentages and vegetation biomass (Birks and Berglund, 2018), the first being the inability to distinguish pollen grains to species level across all taxa. As such, there are several groups of taxa, particularly within coniferous species, that have indistinguishable pollen grains (Birks and Birks, 2000), some of which are only differentiated through the presence/absence of very subtle features that may or may not be preserved (Moore et al., 1991). This is the case for several species of Pinus, a principal host of bark beetles. As such, they are often grouped into two pollen types: diploxylon-type and haploxylon-type. For example, in western North America, where there are numerous Pinus species, the diploxylon-type groups together lodgepole pine (P. contorta), Ponderosa pine (P. ponderosae) and Jeffrey pine (P. jeffreyi), while the haploxylon-type groups together white bark pine (P. albicaulis), western white pine (P. monticola), limber pine (Pinus flexilis) and sugar pine (P. lambertiana) (Brunelle et al., 2008). While these species mostly occupy distinct ecological zones, there are communities where multiple species are found, and consequently, reconstructing historic outbreaks in these areas based on palynological assessments alone are problematic. For instance, if an outbreak occurs in a single host, the pollen abundance from this single species declines, yet the surrounding vegetation, including a palynologically indistinguishable non-host species, increases as a result of the newly available canopy space. The overall net pollen production for that pollen type may therefore remain consistent through the period of infestation and the outbreak remains undetected. A mountain pine beetle outbreak reconstruction in the northern Rocky Mountains (Brunelle et al., 2008), measured the ratio of host lodgepole pine pollen (diploxylon-type) to non-host white bark pine pollen (haploxylon-type) to identify periods of infestation. During the analysis, the authors concluded that the possibility of Ponderosa pine contributing to diploxylon-type, and western white pine and limber pine (Pinus flexilis) contributing to haploxylon-type pollen, cannot be completely dismissed as they all occur in the northern Rocky Mountain region.
The amount of pollen produced also varies considerably between species (Andersen, 1970; Birks and Birks, 2000), with anemophilous (wind-dispersed) species generally producing significantly more pollen than entomophilous (insect-dispersed) species. As such, pollen deposited within an environment can be over- or under-representative of the contemporary vegetation. It is therefore vital to consider the source of pollen (local signal vs regional signal). Pollen deposition can vary by catchment characteristics and vegetation composition that is, dense forests comprise more vegetation typically representing local taxa, while open tundra, with very little vegetation, depicts more regional, wind-blown signals. Bark beetle outbreaks occur in dense coniferous ecosystems, so it is possible that the signals of low-level outbreaks, a decline in a local population, may be masked by the regional signal. This was demonstrated by Kuosmanen et al. (2020), whereby a multiproxy approach examining fossil bark beetle remains, charcoal, and geochemical erosion markers all clearly indicated periods of disturbance, yet the pollen signal failed to show a clear response. The authors conclude that opening of the forest canopy may have been masked by regional pollen signals, leading to failure in the detection of local disturbance. In addition, many bark beetle hosts, including Tsuga, Abies, and Picea are fast-growing species. As such, pollen production (associated with the death of a local population or supressed pollen production during periods of attack) may recover to full production in less than 20 years (Cruz and Alexander, 2010). If sampling resolutions are too coarse, they can essentially ‘miss’ periods of supressed pollen production associated with the disturbance event.
An additional problem with the use of pollen and other proxies derived from sedimentary archives to ‘calibrate’ against known, and therefore recent, outbreaks is that the dating chronology is not sufficiently resolved. This refers not only to the precision associated with dating techniques such as 14C, but also to a systematic error in the 14C framework. Dates from events that post-date AD ~1650 are influenced by the industrial revolution (the Suess Effect) and may be ambiguous (Keeling, 1979). Carbon assimilation into some archives is not instantaneous and the ‘target date’ has associated imprecision. Reducing ambiguity, to the extent that it is possible, requires additional radiocarbon dating beyond that typically employed in standard palaeoecological investigations. Additional dating requirements are dependent on where the sequence sits in the Suess period, but typically each event may require three to five dates to improve resolution. Chronologies can further be improved with incorporation of a priori information, normally stratigraphy or other forms of corroboration, but even then, are likely only to achieve decadal-scale resolution at best.
Taxonomic identification of preserved macrofossil remains
Assessments of tree-ring growth rates and pollen assemblages are indirect measurements of bark beetle outbreak events, as they detect the effects of the agent on the surrounding vegetation. Taxonomic identification of sufficiently preserved bark beetle remains within depositional environments is, to date, the only reconstructive method of analysis aimed at the direct detection of this disturbance agent. Both the deposition and preservation of remains is influenced by a range of environmental, taphonomic, and species-specific factors. Dispersal of bark beetles across North America, for example, typically occurs in June, and is particularly brief, with adult beetles leaving the inner bark for only several days within their 1- to 2-year life cycle (Alfaro et al., 2010; Raffa et al., 2008) and typically travelling less than 300 m from their original host (Seidl and Rammer, 2017). Unfavourable environmental conditions during this short window, such as heavy rainfall, can lead to interrupted flight patterns, and the subsequent incorporation of beetle remains within depositional environments (Brunelle et al., 2008). For example, the AD 1949 spruce beetle epidemic on the White River Plateau, Colorado, coincided with the wettest month of June on record (NCDC, 2006), resulting in a significant accumulation of adult bark beetle carcasses reported to be 15 cm deep, 2 m wide, and spanning more than 1.6 km along the eastern shore of Trappers Lake (Frye et al., 1977; Morris et al., 2010).
It seems, however, that the conditions required for large masses of beetle remains to be incorporated, preserved, and recovered to a level that is taxonomically identifiable are rare. In a comprehensive review of published geological and archaeological records throughout Europe and North America, Schafstall et al. (2020) report only 53 sites containing fossil bark beetle remains, as well as the notable absence of numerous key forest pest species. Although sparse, macrofossil records do significantly pre-date other types of reconstructive archives and provide the earliest examples of physical bark beetle presence in both North America and Europe, with a number of late Glacial and early Holocene occurrences (see Ashworth et al., 1981; Brunelle et al., 2008; Ponel et al., 2003; Schafstall et al., 2020). Scattered occurrence of elm bark beetle (Scolytus scolytus) remains have provided evidence supporting the species’ potential role as a pathogen vector in the mid-Holocene elm decline across northwest Europe (Clark and Edwards, 2004; Girling, 1988; Girling and Greig, 1985; Robinson, 2000); however, the abundance of macrofossils recovered is low. Girling and Greig (1985), for example, report elytra from two individuals discovered within elm decline deposits. Clark and Edwards (2004) analysed large volume sediment samples (reported as 6 L) from peat monoliths at two sites in northeast Scotland: one site yielded a single Scolytus sp. elytron, too fragmented to identify to species level; the other site revealed Scolytus scolytus macrofossil presence in six samples occurring throughout the 3000-year period preceding the elm decline, but the abundance of these beetle remains was not documented.
Macrofossil records of bark beetle remains throughout the historic period are surprisingly rare. Schafstall et al. (2020) note that only 29% of samples from 21 European sites containing bark beetle remains were younger than 1000 years and recovery of more recent remains in North America is even more infrequent. There are some notable examples of successful recovery of taxonomically identifiable remains (head and elytra) of adult bark beetles from lake sediments that provide information on historic occurrence. Kuosmanen et al. (2020) found small numbers of primary bark beetle remains from five different species to be present throughout the period AD 1250–1950 in a lake sequence from the Czech Republic, although typically only one individual beetle was reported to be recovered per sampling level. More temporally consistent occurrence, but still only in low abundance, were found between AD 1620 and AD 1820, interpreted possibly to be indicative of increasing frequency of disturbance from bark beetles. The authors speculate that low sediment sample volumes were a factor in recovery of remains (Kuosmanen et al., 2020). Schafstall et al. (2022) report three peaks in bark beetle remains (post AD 2004, AD 1140–1440, AD 930–1040 recovered from forest hollow sequences at a site in the Tatra Mountains, Slovakia. Pooled samples derived from combining 13 proximal cores were analysed in order to obtain higher sediment volumes (~2 L per 10 cm profile depth) for macrofossil counts. This study documents recovery of some of the largest numbers of bark beetle remains reported in the literature to date; however, when standardised to 100 ml sample sediment volumes, counts were found to range from 0.7 to 4.4 individuals per sample across nine different species of primary bark beetles (Schafstall et al., 2022). The authors note differences in preservation between bark beetle species, with only low numbers recovered for Ips typographus, the key disturbance agent in the region. Single occurrence of a pest species in fossil sequences is common (Schafstall et al., 2020). Increasing sediment volumes for recovery of bark beetle remains within palaeoecological sequences (Schafstall et al., 2022), in conjunction with increased spatial sampling, are key methodological improvements required to examine historic beetle outbreaks more critically, based on analysis of macrofossils.
Brunelle et al. (2008) recovered mountain pine beetle (D. ponderosae) remains in a sedimentary sequence from Baker Lake, Montana USA, which corresponded to the highly cited 1920/30s outbreak in the region, in addition to revealing three early-Holocene outbreaks at the site (8331 (8331–8726), 8410 (8392–8801), and 8529 (8483–8914) cal yr BP) and other early Holocene occurrence (7954 (7782–7954) and 8163 (7850–8213) cal yr BP) at Hoodoo Lake Idaho, another site in the Bitterroot Range of the northern Rocky Mountains..
However, a comprehensive assessment of 30 sedimentary sequences from sites throughout the Intermountain West region of North America (Montana, Idaho, Wyoming, Utah, Colorado, and British Columbia) for bark beetle macrofossils, recovered no diagnostic remains for primary bark beetles, which seems to confirm the rare nature of preservation (Morris et al., 2013). A major limitation of this method is that, even when remains are found, they can often be too poorly preserved for taxonomic identification. Taxonomic analysis of the remains, consisting of beetle elytra and head capsules, were deemed ‘most likely’ to be associated with adult mountain pine beetle. This raises two points that are worth elaborating. First, bark beetles within the genus Dendroctonus are morphologically very similar to each other and, therefore, require a high level of taxonomic skill to distinguish, and second, diagnostic features are not always preserved. Morris et al. (2010) recovered chitinous remains that temporally correspond to very recent outbreaks, but preservation levels were too poor to allow taxonomic identification.
While the examples of taxonomically identifiable remains are rare, they highlight a fundamental principle, that bark beetle remains can become incorporated into depositional environments. This demonstrated potential for incorporation of bark beetles within sedimentary material, but lack of preservation of the features required for definitive identification or the availability of required sediment volumes through traditional palaeoecological methods, highlights the potential that DNA-based methods could be used to identify historic forest pathogen occurrence.
The potential role of DNA-based methodologies to detect historic pathogen occurrence
Since the revolutionary work on the retrieval, extraction, and analysis of ancient DNA (aDNA) in the 1980s (Pääbo, 1985, 1989), the field of DNA-based methodologies to address key environmental and ecological questions has expanded exponentially (Bennett and Parducci, 2006; Crump, 2021; Edwards, 2020; Ficetola et al., 2008; Giguet-Covex et al., 2014; Parducci, 2019; Parducci et al., 2017; Pedersen et al., 2015; Taberlet et al., 2018; Thomsen and Willerslev, 2015). aDNA is characteristically found in low abundance, is subject to degradation through time, as a result of environmental conditions, and requires specific and rigorous manipulation procedures to avoid contamination (Orlando et al., 2021). Each of these traits presents analytical challenges. The breakthrough technique that facilitated the analysis of aDNA was the polymerase chain reaction (PCR), an enzymatic reaction that exponentially increases the number of copies of DNA molecules (Kelly et al., 2019; Mullis and Faloona, 1987; Nichols et al., 2019).
A branch of DNA-based methodologies particularly relevant for palaeoecology involves the extraction of aDNA from sediments, referred to as sedaDNA. Despite presenting challenges akin to aDNA research, sedaDNA has become a rapidly developing field of study in recent years (Coolen and Overmann, 1998), with most of the major methodological developments occurring only in the last 15 years (Crump, 2021; Edwards, 2020; Pedersen et al., 2015). sedaDNA is composed of molecules released from a wide range of sources and persists in the sediment for periods of several hours to several millions of years, depending on depositional conditions (Bohmann et al., 2014; Taberlet et al., 2012, 2018; Thomsen and Willerslev, 2015; Willerslev et al., 2007). The analysis of DNA from sediments is still a relatively novel and expanding field within Quaternary science (Edwards, 2020; Nichols et al., 2019), with the majority of research focussed on the identification of plant taxa and the reconstruction of plant communities (Alsos et al., 2016; Ariza et al., 2023; Bennett and Parducci, 2006; Boessenkool et al., 2014; Hartvig et al., 2021; Jørgensen et al., 2012; Kjær et al., 2022; Sjögren et al., 2016; Willerslev et al., 2014), and is still greatly underutilised in the detection of forest pest dynamics. Macrofossil remains of forest pests, including bark beetles, have been successfully recovered from lake sediments (Brunelle et al., 2008; Kuosmanen et al., 2020; Morris et al., 2015; Schafstall et al., 2022) and are currently the oldest direct record for historic presence of these species. Therefore, their preservation suggests a strong potential for the successful recovery of aDNA from decomposed and/or indistinguishable remains from sedimentary archives, aiding in the understanding of historic outbreak occurrence. Lake sediments are already a well-established source of long-term biological and environmental information and have been widely used to provide proxy evidence of past biological communities and environmental change (Birks and Birks, 2000; Parducci et al., 2017), as evidence of this change is preserved for millions of years. Lake sediments typically contain varying levels of organic matter (Meyers and Teranes, 2002; von Wachenfeldt and Tranvik, 2008), sourced from autochthonous biological processes, as well as allochthonous material from the surrounding catchment and the atmosphere (Giguet-Covex et al., 2019). In addition, lakes are found in almost all environments, making these methodologies widely applicable.
sedaDNA analytical techniques
Most published palaeoecological sedaDNA analyses have focussed on the reconstruction of whole communities using high-throughput sequencing (HTS) techniques, such as DNA metabarcoding, that identify multiple species using ‘universal’ primers, which use conserved regions of a gene or genes to amplify groups of species (Deiner et al., 2017). Often used in biodiversity assessments, population genetics, analyses of diet/foodwebs, this type of data can also be used to influence species-specificprimer assays. In the case of sedaDNA, these studies have typically focussed on plant and vertebrate communities (Compson et al., 2020; Deiner et al., 2017; Harper et al., 2018; Hatzenbuhler et al., 2017), but the inclusion of invertebrate communities would aid in future studies identifying historic disturbances by biological agents and their impacts on plant communities. DNA metabarcoding has the advantage of providing integrated, community-level information; however, it can be expensive for analysing the number of samples required for high temporal-resolution reconstructions. Additionally, its accuracy depends on the taxonomic coverage and resolution as well as on the completeness of the reference database, which can be very variable for invertebrates (Liu et al., 2022; Lunghi et al., 2022).
An alternative DNA-based methodology is quantitative or real-time PCR (qPCR), commonly used for species-specific analyses of contemporary eDNA (Tsuji et al., 2019). qPCR is highly sensitive and specific and has little to no post-amplification processing requirements (Cao and Shockey, 2012; Tsuji et al., 2019), representing a potentially more cost-effective method than metabarcoding of assessing the presence of single species or small groups of known target organisms within palaeoecological records, and a promising technique for producing high temporal-resolution records of specific forest pathogens over time (see Watkins, 2022). This technique has been shown to have higher sensitivity in detecting rare taxa within eDNA than traditional PCR (Harper et al., 2018; Robinson et al., 2019) and can be used for quantification of abundance (Benoit et al., 2023), making it potentially useful for inference of past relative abundance of target species within palaeoecological records. The quantitative aspect of qPCR is achieved by measuring the level of amplification-dependent fluorescence emitted after each cycle of the qPCR reaction, through the use of fluorescent dyes (such as SYBR Green) or probes (such as Taqman) that bind to DNA product.
High-resolution melt curve analysis (HRM), qPCR can be used to identify different species within the same sample (e.g. it can be used to differentiate species within a target genus such as Dendroctonus) based on variation in the melting temperatures (Tm) of the species-specific amplicons (Minett et al., 2021; Robinson et al., 2018). Thus, qPCR protocols can be designed to target single or multiple species, the main limitation being the identification of variability in the short fragments (mini-barcodes) required for the successful amplification of any type of degraded DNA (Roy et al., 2018), as in the case of sedaDNA. At study sites where species presence and abundance are largely unknown, it may be useful to opt initially for more general DNA-based methodologies, such as metabarcoding analysis. As the field develops, increasing number of published primer sets will enable more widespread implementation of sedaDNA investigations of forest insect pathogen dynamics.
While the combination of the wide range of depositional environments on a range of timescales makes DNA-based methodologies suitable for addressing palaeoenvironmental questions, these techniques have remained underutilised for historic pathogen detection or reconstructive capacities. To date, there have been no published investigations using DNA-based methodologies to detect forest pathogen outbreaks, current or historic. Of the few DNA-based studies of key forest pathogens, such as bark beetles, that exist, most concern phylogenetic relationships within and between species and populations (Godefroid et al., 2019; Maroja et al., 2007; Peters et al., 2014; Schrey et al., 2005; Stauffer et al., 2001). Analysis of mtDNA COI sequences and allele frequency for nine microsatellite loci across the contemporary populations of spruce beetle has been used to assess timing of divergence and possible introduction, the results of which suggest these pathogens have been affecting North American forest ecosystems far longer than previously thought (Maroja et al., 2007). Lineage differentiation was identified between Newfoundland and Alaska, where white spruce is a primary host, with a third lineage located in the Rocky Mountains where Engelmann spruce is favoured. The two northern lineages were 3%–4% divergent from each other and from the Rocky Mountain population. Together with pollen records used to confirm the historic availability of suitable hosts, these results indicate that it is possible that spruce beetles have been present in North America for the last 1.7 million years (Maroja et al., 2007). Studies such as these highlight the great potential for DNA methodologies to accompany traditional palaeoecological techniques to provide highly informative reconstructions of past dynamics.
Advantages of DNA studies over traditional detection methods of historical insect outbreaks
Most existing methods of detecting historic insect pathogen outbreaks rely on indirect detection, observing the effects they inflict on plant communities, rather than direct detection of the agent themselves. As such, these signals need to be of enough magnitude to alter the forest communities significantly and produce an ‘outbreak’ signature within the chosen proxy. This has led to an overall bias within the literature towards the detection of highly severe, large-scale outbreaks. The greatest advantage of sedaDNA-methods is that they involve the direct detection of the agent or agents, that is, if DNA from the target species is recovered, the presence of that species can be confirmed in that location, at that point in time. As the agent is being detected directly, they can be found at lower abundances than those observed indirectly, as there is no reliance on the production of outbreak signatures in other proxy datasets.
With regards to the detection or reconstruction of insect populations from sedaDNA samples, there are currently no standardised methods to screen species diversity or assess abundance (Elbrecht and Leese, 2015). qPCR has, however, been applied to the determination of detection probabilities in modern terrestrial eDNA samples for some insect pests (Allen et al., 2021). This type of information would differentiate periods of ‘expected’ population levels and ‘outbreak’ periods. Other fields have presented successful attempts to calibrate eDNA quantity and abundance and derive biomass estimates for target fish and amphibian freshwater species (Lacoursière-Roussel et al., 2016; Lodge et al., 2012; Mahon et al., 2013; Pilliod et al., 2013; Takahara et al., 2012; Thomsen and Willerslev, 2015). These studies showed that eDNA concentration was most significantly correlated with relative species abundance (Lacoursière-Roussel et al., 2016) than environmental conditions, as previously expected. The mechanisms associated with how much DNA each species typically deposits, via cellular and extracellular sources within the environment, known as shedding, forms the basis for interpreting biomass recovered in eDNA samples (Altermatt et al., 2023; Sassoubre et al., 2016). Species morphology and behaviour are crucial considerations in relation to DNA shedding (Andruszkiewicz Allan et al., 2021). In reconstructing forest pest outbreaks via sedaDNA methodologies, it may be best practice to compare one species at a time as levels of DNA shedding will likely vary between agents for example, soft bodied organisms such as Choristoneura sp. versus hard bodied agents such as Dendroctonus spp. bark beetles.
Several other potential methods of semi-quantifying abundance that could be used as a proxy for ‘outbreak’ versus ‘normal’ population sizes in long-term records include comparing amplification from modern sedimentary material within areas of varying forest pathogen outbreak intensity. This quantification can be in the form of the standardised number of qPCR replicates with positive amplifications or the number of reads within metabarcoding samples. Thus far, the majority of research associated with attempt to ‘calibrate’ eDNA signals have involved plant biodiversity assessments (Ariza et al., 2023; Kestel et al., 2022; Schultz and Lance, 2015), with only a few examples concerning insect communities (Camila et al., 2021; Iwaszkiewicz-Eggebrecht et al., 2024).
Changes in insect community composition over time (sensu Talas et al., 2021) obtained from metabarcoding analyses, could also be used to identify changes in pest prevalence and so to infer outbreaks, for example progression from primary bark beetles, to secondary bark beetles, to other taxa associated with deadwood. Where temporal control is adequate, a range of DNA methods could be combined to maximise cost efficiency and information content, for example, high-temporal resolution targeted assay for insect pests, potentially followed by metabarcoding for the examination of community compositional change during identified periods of predicted outbreaks. Additional methodological work is needed to understand depositional processes and incorporation of insect DNA within both lacustrine and terrestrial (peat/forest hollow) sedimentary records, including outbreak severity, catchment topography, eDNA abundance in both modern freshwater environments and sediment surface layers, spatial variability in incorporation into the sedimentary layer, and number of replicate extracts required from each sampling level.
The methods associated with DNA recovery (i.e. coring, subsampling, sampling strategies) are reasonably similar to existing palaeoecological methods that analyse charcoal, pollen and macrofossils. Samples obtained for DNA extraction can be collected in generally the same way as, for example, bulk sediment samples taken for radiocarbon dating (Bennett and Parducci, 2006), but key methodological differences include enhanced emphasis on sterile conditions when coring and subsampling to limit contamination (Feek et al., 2011) and recommended freezing of sedimentary material and subsampling to avoid multiple freeze/thaw events. When correct sedaDNA sampling methods are followed, samples intended for DNA-based methodologies are of no greater risk of contamination than any other type of analyses (Bowers et al., 2021; Edwards, 2020).
As with the other proxy records derived from sedimentary archives (e.g. pollen and macrofossils), the potentially short duration of insect outbreaks means that high-resolution temporal sampling and increased dating resolution is required. The high-resolution sub-sampling required to examine forest pathogen dynamics critically (preferably at least decadal) means that only very low sediment volumes will be available for analysis from traditional cores. Schafstall et al. (2020) have highlighted that in order to produce meaningful reconstructions of forest pests via the retrieval of subfossil remains, large amounts of sediment are required. DNA-based analyses, requiring significantly less sample volume (0.25 g with a QIAGEN PowerSoil Kit, for example), have a clear methodological advantage in this regard. Nevertheless, the tradeoff between temporal resolution and sample requirements favours fast accumulating circumstances.
DNA-based analyses of insect occurrence in palaeoecological records also have the potential to detect different life stages of the target organism and allow the analysis of morphologically poorly preserved samples. Some forest insect species are typically only taxonomically distinguishable as adults, and even then, species-specific distinction is not always possible as adults, even in modern specimens (Pochon et al., 2013). Bark beetles, for example, develop across four main life stages – egg, larvae, pupae, and adult – within which there are four to five stages of instar development during maturation. Species are, therefore, morphologically similar until they reach the adult stage and diagnostic features develop. Methods that can detect and identify specimens at larval or instar stages as well as poorly preserved adults provide results with higher resolution than those identified taxonomically (Andruszkiewicz Allan et al., 2021; Deiner et al., 2017; Parducci et al., 2017). DNA-studies do not rely on the levels of morphological preservation required for traditional taxonomic identification and do not require taxonomic identification expertise (Deiner et al., 2017).
In addition, all aspects of DNA-based methodologies, including extraction protocols, primer sets, PCR conditions, number of replicates, detection thresholds, post-PCR bioinformatics have the potential to be standardised (Loeza-Quintana et al., 2020), whether through metabarcoding, single, or multiplex qPCR assays and the creation of named/standardised protocols for species or groups of species. The implementation of a DNA ‘workflow’ that is, a set parameters for sample collection, preservation conditions, methods of extraction and amplification, sequencing protocols, and bioinformatics (Trujillo-González et al., 2021) or internationally recognised accreditation (ISO/IEC) has been discussed as a possible approach to increase consistency and reliability. This would also address the major disadvantage of palaeoecological science of lacking standardised protocols and thresholds for disturbance, which fail to facilitate the comparison of results across studies and/or sites. However, while there have been calls for standardisation of DNA methodologies (Harper et al., 2018; Loeza-Quintana et al., 2020), it seems that the abundance of variables associated with each research study (target species or group of species, sample type, budget) could limit the potential for standardisation. Other paleoenvironmental methods, such as pollen analysis, have become standardised in their preparation methods (see Erdtman, 1954; Moore et al., 1991) and it is hoped that the level of standardisation in sedaDNA analysis that is seen in other areas of the field will be realised. Alongside the move towards standardisation of protocols, the field of palaeoecology would also greatly benefit from the increased transparency of data associated with historic environmental change. While there exists Tephrabase for tephra geochemical data and GenBank for genetic data, there is less availability of raw palynological or dendroecological data. This inaccessability of data greatly reduces the capacity for standardisation and reproducible analyses.
Conclusion
DNA-based methodologies are revolutionising the field of Quaternary science through the retrieval of target species DNA from a wide range of depositional environments, overcoming the need for sufficient levels of preservation and taxonomic skills required for species identification from macrofossils. However, the application of environmental DNA, ancient DNA, and sedimentary DNA research to the reconstruction of historic forest pest dynamics is still in its infancy (Dickie et al., 2018; Klymus et al., 2020). This review highlighted how traditional methods of detection greatly rely on the production of ‘outbreak’ signatures within proxy records, which must then be calibrated uniquely in order to identify periods of outbreak, a critical weakness in the current understanding of historic forest pathogen dynamics. DNA-based methodologies, however, hold the potential to detect the presence of the responsible agent directly at all life stages. This cannot be underestimated as a critical advantage of this technique, as positive amplifications of target species within a sampled depth directly confirm the presence of that species at that point in time, without the need to assign ‘outbreak’ parameters. DNA-based methodologies are becoming increasingly incorporated into paleoenvironmental reconstructions as they have been shown to be highly complementary to existing proxy records and produce many robust inferences of environmental change across key areas of Quaternary science (Edwards, 2020).
Footnotes
Author contribution(s)
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported by the Quaternary Research Association (QRA) through a QRA New Research Worker’s Award (NRWA) obtained by the first author.
