Abstract
Background
Transmission of HIV and hepatitis C virus (HCV) are heavily influenced by complex interactions within sexual or injecting networks where risk behaviors occur. In Ukraine, women engaged in sex work (WSW) and men who purchase sex (MWPS) are disproportionately affected by both viruses. The aim of our study was to the investigate the influence of underlying networks on transmission of HIV and HCV.
Methods
A cross-sectional integrated bio-behavioural survey was implemented among 560 WSW and 370 MWPS representative of sex work hotspots in Dnipro, Ukraine (December 2017 to March 2018). A portion of the HIV reverse transcriptase gene (n = 13; 62% WSW, 38% MWPS) and HCV NS5B gene (n = 46; 70% WSW, 30% MWPS) were sequenced from dried blood spot specimens. Tip-to-tip distances on phylogenetic trees were used to infer phylogenetic clusters for identifying potential transmission clusters.
Results
Phylogenetic analyses identified two HIV clusters containing four sequences (50% WSW; 50% MWPS) and 11 HCV clusters containing 31 sequences – the majority comprising infections in WSW (83.9%). Nearly half (45.4%) of HCV clusters contained at least one WSW with a history of injecting drugs.
Conclusions
Joint analyses of HIV and HCV signal overlap in sex work and injecting networks in Ukraine, suggesting implications for the comprehensive coverage of prevention programs for WSW including harm reduction services. Conducting phylogenetic analyses with HCV may provide a more complete appraisal of underlying transmission networks than HIV alone, particularly in the context of high HIV treatment coverage yielding viral suppression.
Introduction
Women engaged in sex work (WSW) have increased vulnerability to HIV and other sexually transmitted and bloodborne infections (STBBIs) due to structural factors, including violence, unsafe working conditions, stigma, and criminalization, which constrain accessibility and uptake of testing and prevention services.1,2 Men who purchase sex (MWPS) experience vulnerability to HIV through multiple sexual partners, coinfection with other STBBIs, and a history of injection drug use (IDU) 3 in some instances. Barriers to HIV testing, such as fear of stigma, gender norms of masculinity, strength, and control, a lack of HIV knowledge, and an overall shortage of interventions designed to reach this population can exacerbate this risk. 4 Transmission of HIV and hepatitis C virus (HCV) are influenced by complex interactions within sexual or injecting networks5,6 in which HIV risk behaviors occur among highly connected members 6 and become normalized, including inconsistent condom use, 7 multiple, concurrent sexual partners, 8 and needle and syringe sharing. 9
In Ukraine, individuals involved in sex work are disproportionately affected by HIV and HCV, with prevalence estimated to be 3% and 8% respectively among WSW in 2022 10 , and 5% and 13% respectively among MWPS in 2015. 11 Research in Ukraine associates most new HIV and HCV infections with IDU. 12 However, an increase in heterosexual transmission of HIV has been observed since 2008 13 , particularly among sexual partners of people who inject drugs (PWID). 14 Treatment and prevention of HIV and HCV remains challenging due to low awareness of non-governmental organizations offering services, 15 discontinuity of programs and harm reduction initiatives due to unsustainable funding and healthcare reform, 16 and previous conflicts and wars – specifically the protracted conflict in the eastern Donbas region of Ukraine since 2014 17 and Russia’s full-scale war in Ukraine beginning in 2022. 18
Here, we characterize HIV and HCV transmission networks using viral phylogenies among WSW and MWPS in Dnipro, Ukraine. Most Ukrainian phylogenetic studies focus on HIV or HCV networks exclusively among PWID,19–23 and to our knowledge, we are the first to infer HIV and HCV transmission networks among WSW and their clients. We conducted HIV and HCV next generation sequencing with the intent of constructing phylogenies to infer potential transmission clusters. Given shared risk behaviors and overlap in social and sexual networks for both viruses, conducting our phylogenetic study with both viruses allows us to investigate the influence of underlying networks on viral transmission.
Methods
Study setting
Through the Dynamics Study, we implemented a cross-sectional bio-behavioral survey among WSW and MWPS in Dnipro, Ukraine from December 2017 to March 2018. 24 Following initial geographic mapping, a two-stage sampling design was used to recruit participants at sex work hotspots, as previously described elsewhere. 24 WSW aged 14 years or older who self-identified as currently engaged in sex work and MWPS aged 18 years or older who had ever purchased sex were eligible to participate.
Following informed consent, participants completed a questionnaire pertaining to sociodemographics, sex work practices and experiences, HIV and HCV risk behaviours, and utilization of prevention and care services. 24 Participants then underwent rapid testing for HIV (SD Bioline HIV 1/2 3.0, Wantai Rapid Test for Antibody to HIV 1 + 2, and Profitest Rapid Anti-HIV 1/2 Test) and HCV (Wantai Rapid Test for Antibody to HCV), before providing a dried blood spot (DBS) specimen.
Serological testing
All DBS were shipped to the National Sexually Transmitted and Blood Borne Infection Laboratory (Public Health Agency of Canada, Winnipeg, Canada) for subsequent analysis, where serological testing using the AVIOQ HIV Microelisa System (Avioq Inc., Durham, NC) and ORTHO® HCV Version 3.0 ELISA Test System (Ortho-Clinical Diagnostics, Inc., Raritan, NJ, USA) was performed according to the manufacturer’s instructions for the purpose of quality control and validating rapid testing results. We attempted viral genomic sequencing on all HIV- and HCV-reactive DBS.
Next generation sequencing and bioinformatic analysis
A portion of the HIV pol gene (position 2,074-3,334 on HXB2 accession no. K03455) was amplified in overlapping protease (PR) and reverse transcriptase (RT) regions using an in-house HIV drug resistance genotyping assay described in supplementary materials. A portion of the HCV NS5B gene (position 8,182-8,610 on strain H77 accession no. AF009606) was amplified using an assay described elsewhere. 25
Sequencing libraries were prepared using the Nextera XT DNA library preparation kit (Illumina, San Diego, USA) and conducted using an epMotion 5075t liquid handling station (Eppendorf, Hamburg, Germany). Sequencing was performed on a MiSeq platform (Illumina) with the MiSeq Reagent Kit v2 (300 cycles; Illumina) according to the manufacturer’s instructions. Bioinformatics analysis was performed in Geneious 2023.0.4 (https://www.geneious.com) as described in supplementary materials.
Subtyping and phylogenetic cluster analysis
Subtyping of HIV and HCV sequences was performed using IQ-TREE v2.2.0.3 26 with references and parameters outlined in supplementary materials.
For each subtype, MAFFT v7.490 27 was used to prepare alignments before visualizing in MEGAX v10.0.05. 28 Maximum-likelihood tree reconstruction was performed in IQ-TREE v2.2.0.3 26 using the automatic best-fitting substitution model and UFBoot 29 re-sampling with 100,000 replicates. Accession numbers for background sequences collected from the Los Alamos HIV (https://www.hiv.lanl.gov/) and HCV (https://www.hcv.lanl.gov/) databases, GenBank, 30 and BLASTn 31 are provided as supplementary material (Table S2). Phylogenetic clusters were inferred by tip-to-tip (patristic) distance between sequences on phylogenetic trees. A distance threshold of <0.02 substitutions per site and >50% bootstrap support was used to infer transmission clusters. 32 Clusters were visualized and annotated using Cytoscape. 33
Ethics
Ethical approvals were obtained from the Health Research Ethics Board at the University of Manitoba (HS20653 [H2017:097]), Canada; the Ethical Review Committee of the Sociological Association of Ukraine; and the Committee of Medical Ethics of the L. Gromashevsky Institute of Epidemiology and Infectious Diseases at the National Academy of Medical Sciences of Ukraine.
Results
Study population and sequence dataset
A total of 560 WSW (86.0%) and 370 MWPS (88.9%) consented to complete the questionnaire, with all participants except one (99.9%) consenting to provide a DBS sample (Figure 1). Participant characteristics are presented as supplementary material (Table S1) and elsewhere.4,15,24,34 Dynamics Study HIV and HCV phylogenetic analysis workflow.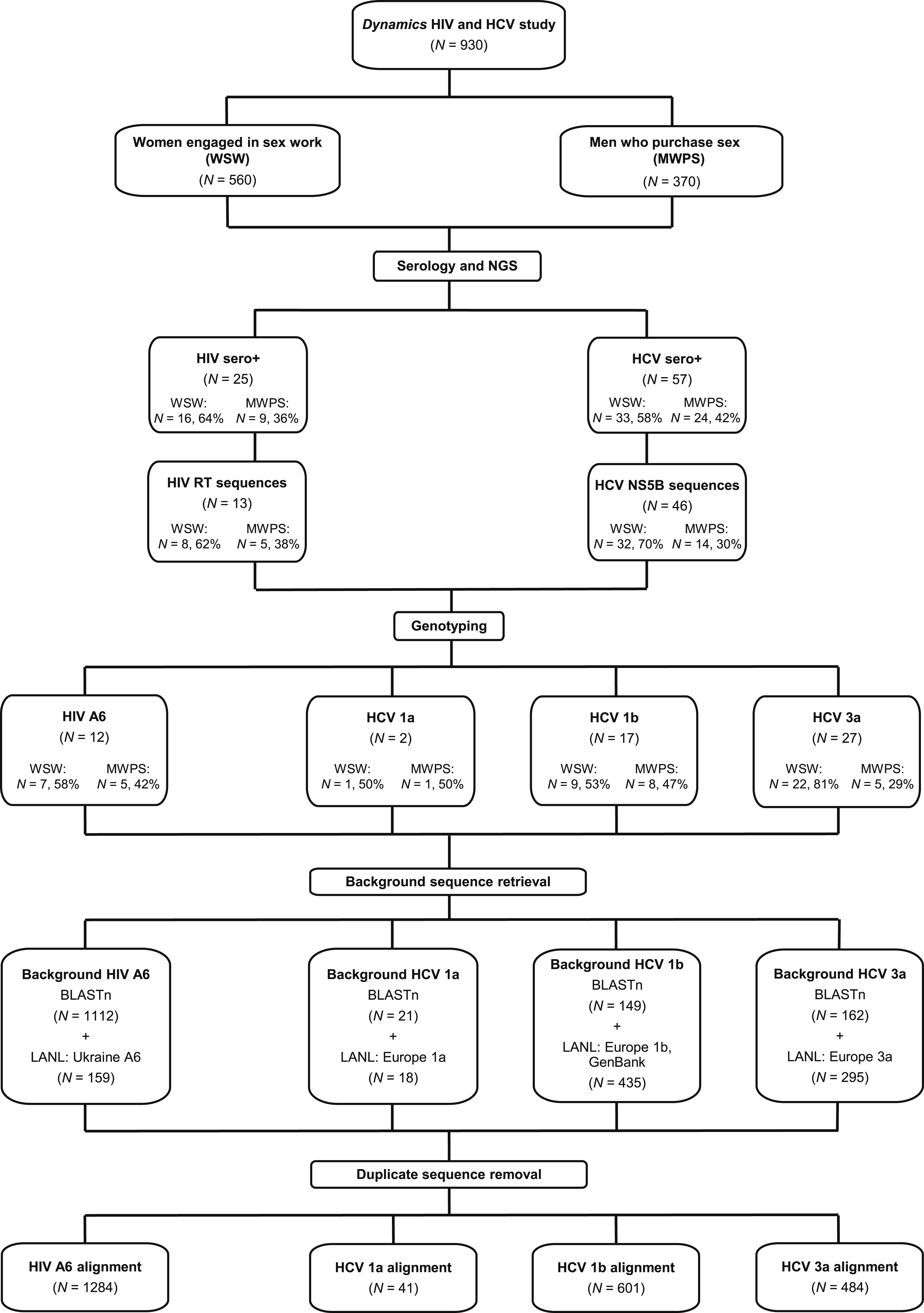
Serology identified 25 (2.7%) participants living with HIV, consisting of 16 (64.0%) WSW and 9 (36.0%) MWPS, which was consistent with rapid test results. The HIV PR and RT genes were successfully sequenced from 10 (40.0%) and 13 (52.0%) HIV-reactive DBS specimens, respectively. To maintain largest sample size, RT sequences were used for subsequent analyses. All sequences except one (n = 12, 92.3%) classified as sub-subtype A6, being a singular A1/A2 recombinant omitted from cluster analysis.
Serology indicated 57 (6.1%) participants ever-having acquired HCV, of whom 33 (58.0%) were WSW and 24 (42.0%) were MWPS, consistent with rapid test results. The HCV NS5B gene was successfully sequenced from 46 (80.7%) HCV-reactive DBS specimens, comprising 32 (69.6%) WSW and 14 (30.4%) MWPS. Sequences classified as subtypes 3a (n = 27, 58.7%), 1b (n = 17, 37.0%), and 1a (n = 2, 4.3%). No recombinants were detected.
Sociodemographic characteristics and phylogenetic clusters
Of all Dynamics study participants, more WSW (n = 44, 7.9%) than MWPS (n = 14, 3.8%) reported a history of injecting drugs, of which clustering WSW (n = 6, 21.4%) and MWPS (n = 1, 14.3%) exhibit a higher proportion (Table S1).
Of the 13 HIV sequences, four sequences formed two clusters (Figure 2(a)), one comprising only sequences from WSW, and the other comprising only sequences from MWPS. All four (100%) participants reported no history of IDU (Table S1). Of the 46 HCV sequences, 31 formed 11 statistically significant clusters ranging in size from two to eight sequences (Figure 2(b)). Most (9/11, 81.8%) HCV phylogenetic clusters contained exclusively WSW or MWPS sequences, except for two dyad/pairs of HCV subtypes 1b and 1a, consisting of a WSW and MWPS sequence. Seven clusters consisting of 26 WSW sequences comprised the entirety of HCV subtype 3a clusters detected, including the largest eight-sequence cluster detected in our study. Of the 11 HCV clusters, five (45.5%) contained at least one participant who reported IDU in their lifetime. HIV subtype A6 and HCV subtypes 3a, 1b, and 1a phylogenetic transmission clusters derived from RT and NS5B genes. Phylogenetic clusters from principal maximum likelihood phylogenetic trees of (a) HIV subtype A6 and (b) HCV subtype 3a, 1b, and 1a clusters according to history of IDU. Nodes shapes indicate sequences derived from women engaged in sex work (triangles) or men who purchase sex (circles). Node color designates a history of IDU in lifetime (red) or no history of IDU (white).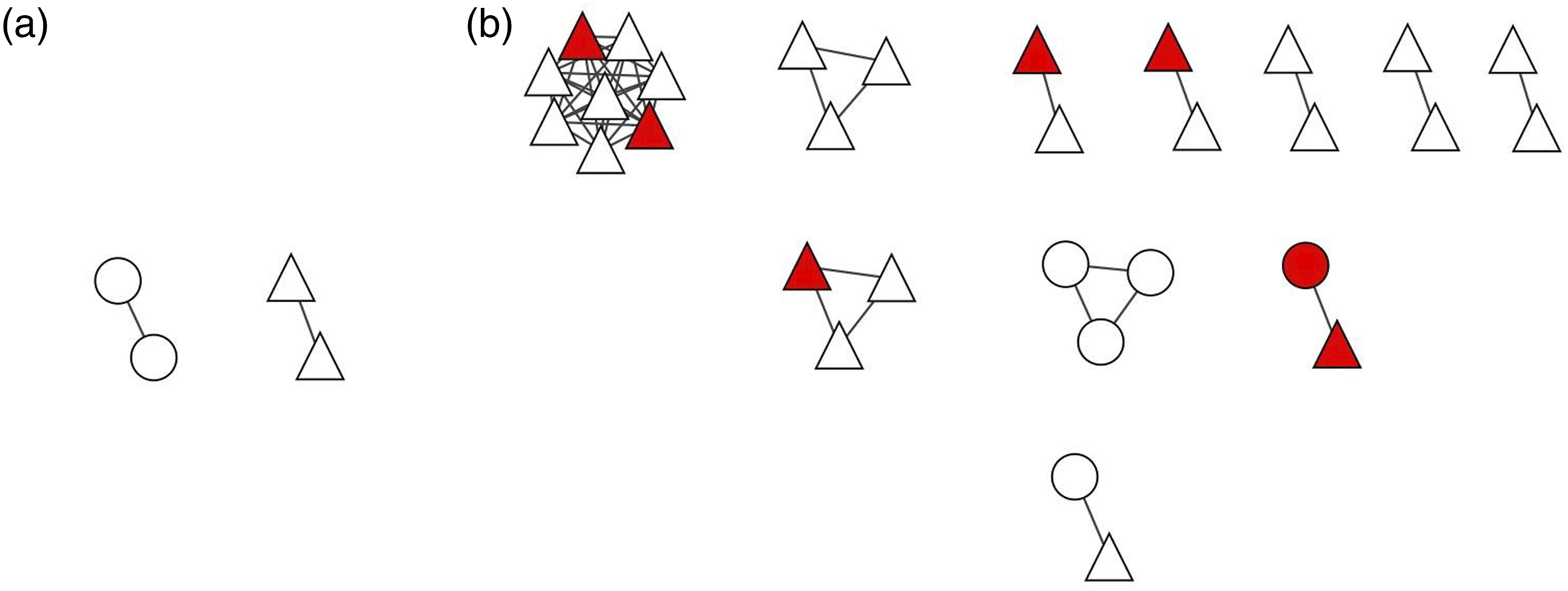
Discussion
Using cross-sectional data, we have characterized phylogenetic clusters and identified HIV and HCV phylogenies among WSW and MWPS. We found Dynamics participants did not cluster based on HIV sequences, but formed phylogenetic linkages based on HCV sequences. Although low HIV prevalence and number of sequences might explain the lack of phylogenetic clusters, the recruitment of WSW and MWPS from similar hotspots would suggest any observed linkages indicate closely related transmission events versus random sampling.
Observing few HIV clusters overall, none of which contained both a WSW and MWPS, might be attributable to phylogeny reconstruction issues. HIV’s prolonged infectivity period and low transmission risk per contact can limit sexual contact-based clusters detection, compared to better detection of injection-based clusters due to the rapid growth of epidemics among PWID. 35 This might also be explained an overall decline in new HIV infections reported among young PWID and women in Ukraine, 14 including among WSW. 36
We observed most (n = 8/11, 72.7%) HCV clusters to contain exclusively WSW sequences, suggesting non-sexual transmission events. Underlying injection networks might be present based on the five HCV phylogenetic clusters containing at least one WSW reporting IDU in their lifetime, in agreeance with reports of overlapping WSW and PWID populations.37–39 Research has shown high prevalence of HCV (31–63%) among Ukrainian WSW reporting IDU within the last 12 months.14,40 We expect some under-reporting of IDU among participants due to social desirability bias, which was shown in previous Ukrainian integrated bio-behavioral surveillance studies, 14 as HCV phylogenetic networks constructed from PWID have been shown to correlate well with their social injecting networks. 41 Dynamics participants may have underreported IDU due to criminalization, stigmatization, and negative reactions associated with breaching gender roles as women who inject drugs. 42
Through examining both HIV and HCV, we identify phylogenies indicative of overlapping sex work and injecting networks among WSW in Dnipro, Ukraine. These findings stress the importance of considering additional pathogens to assess the social networks in which STBBIs travel through, particularly while concurrent and synergistic HIV and HCV epidemics are an obstacle in Ukraine.43–45 Further, dual STBBI analyses such as ours present alternative ways to conduct molecular studies across contexts in the era of highly effective antiretroviral therapy (ART), which effectively mitigates HIV transmission46,47 and limits collection of HIV sequences for phylogenetic analyses. Previous reports suggest a well-established HIV treatment cascade among Ukrainian WSW, with 83% being aware of their HIV-positive status, 94% receiving ART, and 80% achieving an undetectable HIV viral load, 40 however, this may have shifted in response to conflict. Unfortunately, similar data is not available for MWPS. Numerous ongoing challenges to HCV elimination in Ukraine have been described, including an overall lack of awareness, 48 inadequate surveillance and infection estimates,45,49 and barriers to accessing diagnostics and treatment. 43 Given transmission routes shared by HIV and HCV, clusters inferred from HCV sequences also reflect possible HIV transmission networks, providing valuable insight for STBBI prevention. If HCV sequences were omitted from our study, the vast majority of phylogenetic clusters would not have been observed and important linkages missed, such as HCV clusters among WSW.
Our data suggests implications for comprehensive coverage of intervention efforts, as STBBI prevention efforts in Ukraine for WSW traditionally aim to circumvent heterosexual transmission, such as male condom and lubricant provisions, HIV/STBBI testing, and counselling and support services,17,50,51 leaving out broad-based harm reduction prioritized among PWID – namely needle and syringe programs. 51 Moreover, our results agree with suggestions to enhance programming for Ukrainian MWPS. Recent studies highlight the elevated HIV and STBBI risk facing MWPS,3,52 compounded by the rarity of STBBI prevention programs designed for men across global contexts.4,52–54 Innovative programming is needed to reach men who do not identify into key population groups, such as PWID, which requires addressing masculinity and other barriers to accessing prevention and testing services. 4 Monitoring of Ukrainian epidemics remains important, albeit difficult, while the ongoing war destabilizes public health infrastructure and displaces millions of people,24,44 with severely affected conflict areas experiencing disruptions to healthcare services, 43 ART delivery,55,56 harm reduction and HIV testing services. 57 Post-conflict, it will be critical to prioritize STBBI programming for Ukrainians due to the restructuring of social, sexual, and injecting networks in response to conflict, which may heighten HIV and HCV vulnerability.4,44
In this paper we report, using viral sequences, that HCV phylogenies highlight overlap between sex work and injecting networks in Ukraine. Ukrainian WSW might benefit from more comprehensive STBBI prevention programming, including harm reduction and broader sexual and reproduction health services that expand on current programs. Efforts must continue to reach Ukrainian MWPS not in key population groups with innovative prevention and testing programming which address socio-structural barriers to engagement. In the era of highly effective ART which achieves HIV viral suppression, conducting our phylogenetic analysis with HCV provides a better appraisal of transmission networks which may have been overlooked by relying on HIV sequences alone.
Supplemental Material
Supplemental Material - Joint HIV and HCV phylogenetic analyses signal network overlap among women engaged in sex work and men who purchase sex
Supplemental Material for Joint HIV and HCV phylogenetic analyses signal network overlap among women engaged in sex work and men who purchase sex by Stephanie Melnychuk, Olga Balakireva, Daria Pavlova, Anna Lopatenko, Leigh M McClarty, Lisa Lazarus, Nicole Herpai, Michael Pickles, Sharmistha Mishra, Marissa l Becker, Paul Sandstrom, François Cholette and on behalf of the Dynamics study team in International Journal of STD & AIDS
Footnotes
Acknowledgements
We acknowledge the Dynamics Study Team including the authors of this manuscript, as well as Sevgi Aral, James Blanchard, Tetiana Bondar, Eve Cheuk, Christina Daniuk, Evelyn Forget, Shajy Isac, Emma Lee, Robert Lorway, Huiting Ma, Lyle McKinnon, Stephen Moses, Maureen Murney, Nam-Mykhailo Nguien, Ani Shakarishvili, and Tatiana Tarasova for their work on the study. We would also like to thank the field research team and Road to Life and DEF Group (Dnipro) for their support. The work described here was conducted in partnership with the NGO Ukrainian Institute for Social Research after Oleksandr Yaremenko, the Alliance for Public Health in Ukraine, the Dnipro Oblast AIDS Centre and the Center for Public Health, the Ministry of Health in Ukraine.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Canadian Institutes for Health Research (PJT-148876). Sharmistha Mishra is supported by a Tier 2 Canada Research Chair in Mathematical Modeling and Program Science (grant number 950-232643). The funders had no role in study design; in the collection, analysis, and interpretation of data; in the writing of the report; and in the decision to submit the paper for publication.
Ethical statement
Data availability statement
Due to the sensitive nature of molecular surveillance among marginalized communities, sequences from our study have not been deposited in public repositories (ex: GenBank). However, requests to access the data can be made by contacting the principal investigators (Sharmistha Mishra and Marissa L. Becker).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
