Abstract
Background
Infective pneumonia is a major cause of respiratory failure and ICU admission, but non-infectious mimics—such as autoimmune disease, malignancy, and organizing pneumonia—are also common. Multiplex PCR pneumonia panels may help differentiate these entities.
Methods
We conducted a prospective before-and-after study (1 Jan 2024-30 Jun 2025) evaluating the FilmArray Pneumonia plus panel (FAPN) in adults with respiratory failure admitted to two district general hospital ICUs. Consecutive patients undergoing FAPN testing (n = 116) were included. In the post-intervention cohort (n = 56), a consensus panel determined the cause of respiratory failure in patients with suspected pneumonia (PF ratio <300 mm Hg and ≥1 radiographic quadrant involvement). Agreement between FAPN and sputum culture—including the diagnostic value of a negative FAPN—was assessed.
Results
Among post-intervention patients with suspected pneumonia (n = 45), the combined finding of “negative FAPN and negative sputum culture” identified non-infectious mimics with a sensitivity of 75% (95% CI 50-100%) and specificity of 91% (95% CI 81-100%). In multivariable analysis, negative FAPN results were associated with reduced antibiotic escalation (OR 0.70; p < .01). The positive predictive value of individual FAPN bacterial targets ranged from 0%–100% (median 35%). Culture confirmation increased with higher FAPN quantitative bins: 8% at 104 CFU/mL, 7% at 105 CFU/mL, 48% at 106 CFU/mL, and 57% at 107 CFU/mL.
Conclusions
In ICU patients with suspected pneumonia, a combined negative FAPN and sputum culture result should prompt evaluation for non-infectious mimics. Negative FAPN findings may help reduce unnecessary antibiotic escalation, although their positive predictive value for culture-confirmed bacterial infection is limited.
Background
Globally, respiratory failure is a common cause of admission to the intensive care unit (ICU). In addition to organ support—ranging from oxygen therapy and invasive mechanical ventilation to extracorporeal membrane oxygenation—targeted treatment of the underlying cause is essential.
While pneumonia, particularly of bacterial origin, accounts for many cases of respiratory failure and is a leading cause of acute respiratory distress syndrome (ARDS), a substantial proportion of patients instead have non-infectious mimics of pneumonia. Autoimmune diseases and malignancies are well-recognised mimics, 1 along with other conditions such as cryptogenic organizing pneumonia and various interstitial lung diseases. 2 Distinguishing between infectious and non-infectious causes is challenging, as clinical symptoms, laboratory findings, and radiological features frequently overlap. These mimics commonly present in the ICU as non-resolving pneumonia, with reported non-response rates of 15%–30%. Suspicion for non-infectious causes typically arises only days to weeks into antimicrobial therapy.3,4 Continued antimicrobial use in such cases is not only ineffective but potentially harmful, contributing to antimicrobial resistance and delaying identification of the true underlying pathology.
Multiplex polymerase chain reaction (PCR) panels performed on respiratory samples have been proposed as a diagnostic aid in this setting, offering rapid pathogen detection and potential benefits for antimicrobial stewardship. Their use in pneumonia has been evaluated in several randomized controlled trials (RCTs), with variable effects on antimicrobial escalation, de-escalation, and clinical outcomes.5–7 However, these panels often detect organisms such as Staphylococcus aureus, Enterobacterales, and non-fermenters, which may represent colonization rather than true infection. 8 To our knowledge, no studies have specifically evaluated the utility of these tests in patients with non-infectious mimics of pneumonia.
In this study, we evaluate the clinical utility and impact of a multiplex PCR pneumonia panel in ICU patients with respiratory failure, particularly its role in distinguishing infectious pneumonia from non-infectious mimics.
Patient Population
This study was conducted in the ICUs of two district general hospitals in Hong Kong. The BioFire FilmArray Pneumonia Plus (FAPN) panel (BioMérieux, Marcy-l’Étoile, France) was implemented in both units on 1 October 2024. Prospectively, all consecutive adult patients admitted with respiratory failure between 1 October 2024 and 31 May 2025 who underwent FAPN testing were included for microbiological analysis. Clinical outcome analysis was limited to patients with a PaO2:FiO₂ (PF) ratio <300 mm Hg and radiographic involvement of ≥1 quadrant on chest X-ray, criteria consistent with suspected ARDS.
Respiratory Specimen Culture and Antibiotic Susceptibility Testing
All microbiological processing was performed at the Department of Microbiology, Prince of Wales Hospital, and the Department of Pathology, Alice Ho Miu Ling Nethersole Hospital. Specimen types included sputum, tracheal aspirate, bronchial aspirate, and bronchoalveolar lavage (BAL).
Sputum and tracheal aspirates were homogenized using an equal volume of Sputasol (Remel, Thermo Fisher, Massachusetts, USA). BAL and bronchial aspirate samples did not require homogenization. Specimens were inoculated onto horse blood agar (HBA), chocolate blood agar (CBA), MacConkey agar (MAC), and Sabouraud agar (SAB). HBA and CBA were incubated at 35–37 °C in atmospheric conditions for 48 h, MAC for 24 h, and SAB for 5 days, with daily examination.
Organisms were identified to species or species-complex level using MALDI-TOF mass spectrometry (Bruker, Massachusetts, USA). Streptococcus pneumoniae was confirmed using MALDI-TOF and optochin susceptibility. Antibiotic susceptibility testing followed CLSI M100, 34th Edition (2024).
Multiplex PCR Pneumonia Panel
FAPN was performed on respiratory specimens—including sputum, tracheal aspirate, bronchial aspirate, and bronchoalveolar lavage—according to the manufacturer's instructions. In short, sample was added to the sample buffer by using the sample swab provided, then the sample-buffer mix and hydration solution were added to the corresponding ports of the sample pouch. The pouch was then loaded into the BioFire FilmArray TORCH PCR system (BioMérieux, Marcy-l’Étoile, France). Upon completion of the PCR process, results were automatically generated by the system and reported to clinicians in accordance with the manufacturer's guidelines.
Clinical Information
Patient records were collected using a standardized form, supplemented by electronic medical records, to obtain baseline epidemiological data, Acute Physiology and Chronic Health Evaluation (APACHE) II scores, and clinical information. This included temperature, hemodynamic parameters, use of mechanical ventilation, PaO2:FiO₂ (PF) ratio, positive end-expiratory pressure (PEEP), left ventricular ejection fraction (LVEF) on echocardiography, microbiological culture results, medical comorbidities, antibiotic use, immunosuppressant use, radiological imaging findings, and biochemical markers such as procalcitonin, C-reactive protein, urea, creatinine, and white cell count. All data were reviewed and anonymized for clinical evaluation of the etiology of respiratory failure.
Clinical Evaluation of the Etiology of Respiratory Failure
The acute cause of respiratory failure in each patient in the post-intervention group was determined by a consensus panel of three specialist physicians: a respiratory medicine specialist (CWY), an intensive care medicine specialist (LTH), and a clinical microbiology and infection specialist (ALHL). The consensus panel was blinded to the diagnosis given by the attending clinicians, and determined the causes of respiratory failure as per the categories below based on the clinical information gathered in the above section. Only the clinical information available within 48 h of the respiratory specimen taken were available to the panel. The consensus panel categorized independently on initial assessment, and were required to categorize until unanimous agreement was reached. The causes of acute respiratory failure were categorized as follows:
Infectious pneumonia Non-infectious mimics of pneumonia No primary lung disease
Microbiological results of each patient were classified into two categories:
Positive for any FAPN targets Negative for both FAPN
After excluding patients in category 3 (“no primary lung disease”), diagnostic performance of category B for identifying non-infectious mimics (category 2) was assessed using a 2 × 2 contingency table. The sensitivity and specificity of
Evaluation of Antibiotics use
Antibiotic use before and after FAPN was assessed using a quasi-experimental before-and-after design. The intervention group included patients tested with FAPN from 1 October 2024 onward. A historical cohort comprised patients with PF ratio <300 mm Hg and ≥1 CXR quadrant involvement admitted between 1 January and 30 September 2024.
For each patient, antibiotic regimens on the day before and the day after test result availability were reviewed. Changes were categorized as escalation, de-escalation, or no change based on the antibiotic spectrum score. 9
Univariable logistic regression identified factors associated with escalation; significant variables and clinically relevant covariates were entered into a multivariable logistic regression model.
Microbiological Assessment of multiplex PCR Accuracy
Conventional respiratory culture served as the reference standard. For each bacterial and resistance marker, FAPN performance was evaluated using positive percent agreement, negative percent agreement, sensitivity, and specificity. Overall concordance and detection of out-of-panel organisms were also recorded.
Statistical Analysis
Statistical analysis was performed using R software version 4.4.1 (R Foundation, Vienna, Austria). Descriptive statistics were used for exploratory data analysis. Categorical variables were compared using the chi-square test, and continuous variables using the Wilcoxon rank-sum test. A p-value <.05 was considered statistically significant.
Results
Clinical Characteristics
A total of 112 consecutive patients underwent FAPN testing. After excluding patients with a PF ratio >300 mm Hg or a clear chest X-ray, 56 patients with severe respiratory failure were included for analysis of the causative factor. The patient inclusion flow is shown in Figure 1, and baseline characteristics are presented in Table 1. The mean age was 65 years, and 38 (65%) were male. The mean APACHE II score was 25, and the average ICU length of stay was 14 days. Regarding respiratory severity, an average of 2.1 quadrants were involved on CXR, the mean PF ratio was 201 mm Hg, the mean PEEP was 9.4 cmH2O, and 50 patients (89%) required invasive mechanical ventilation.
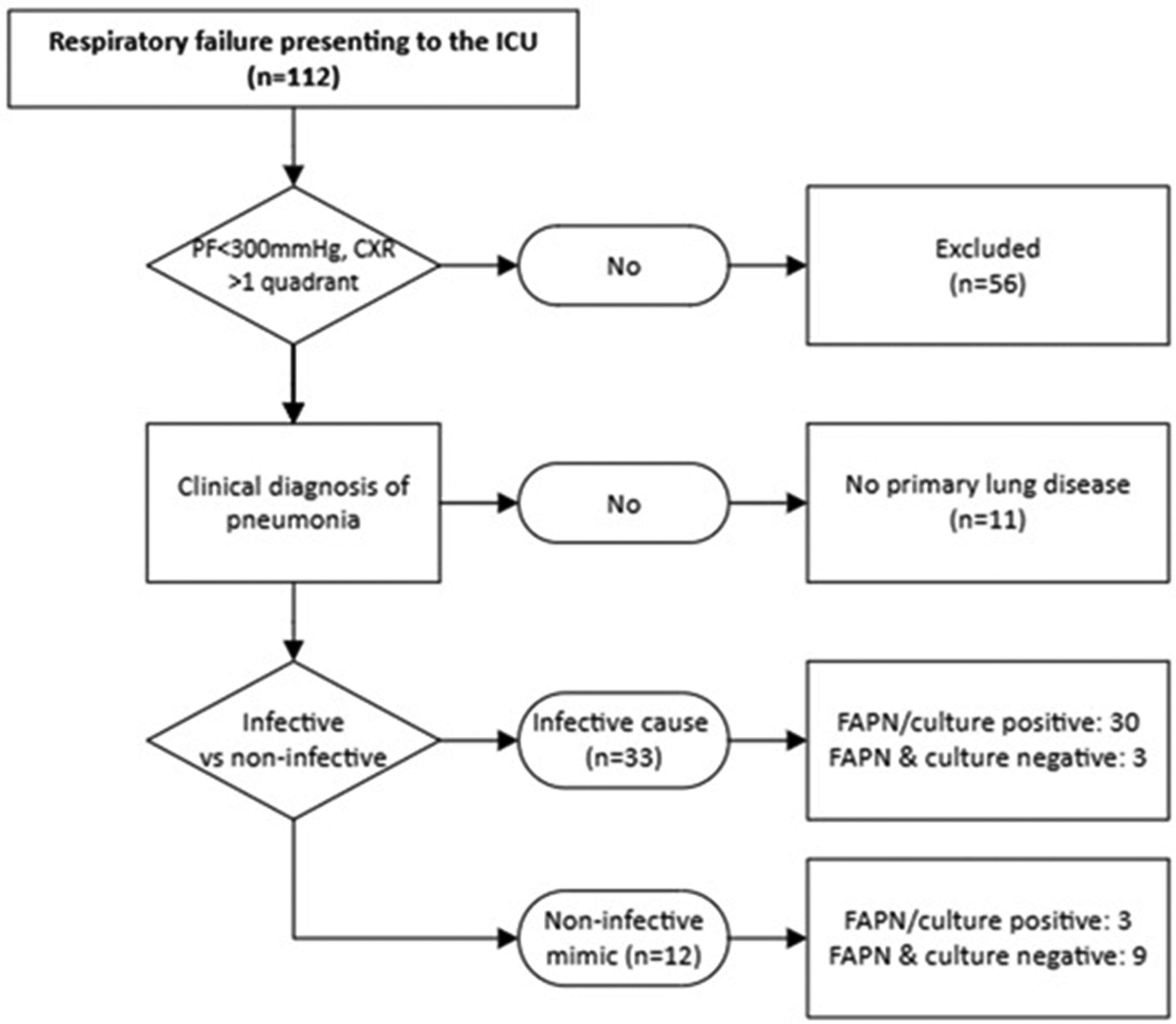
Flowchart describes a clinical algorithm incorporating FAPN and standard respiratory specimen cultures for the diagnosis of pneumonia compared to true pneumonia as determined by a consensus panel.
Baseline Characteristics of Patients Included in the Post-intervention Multiplex PCR Analysis.
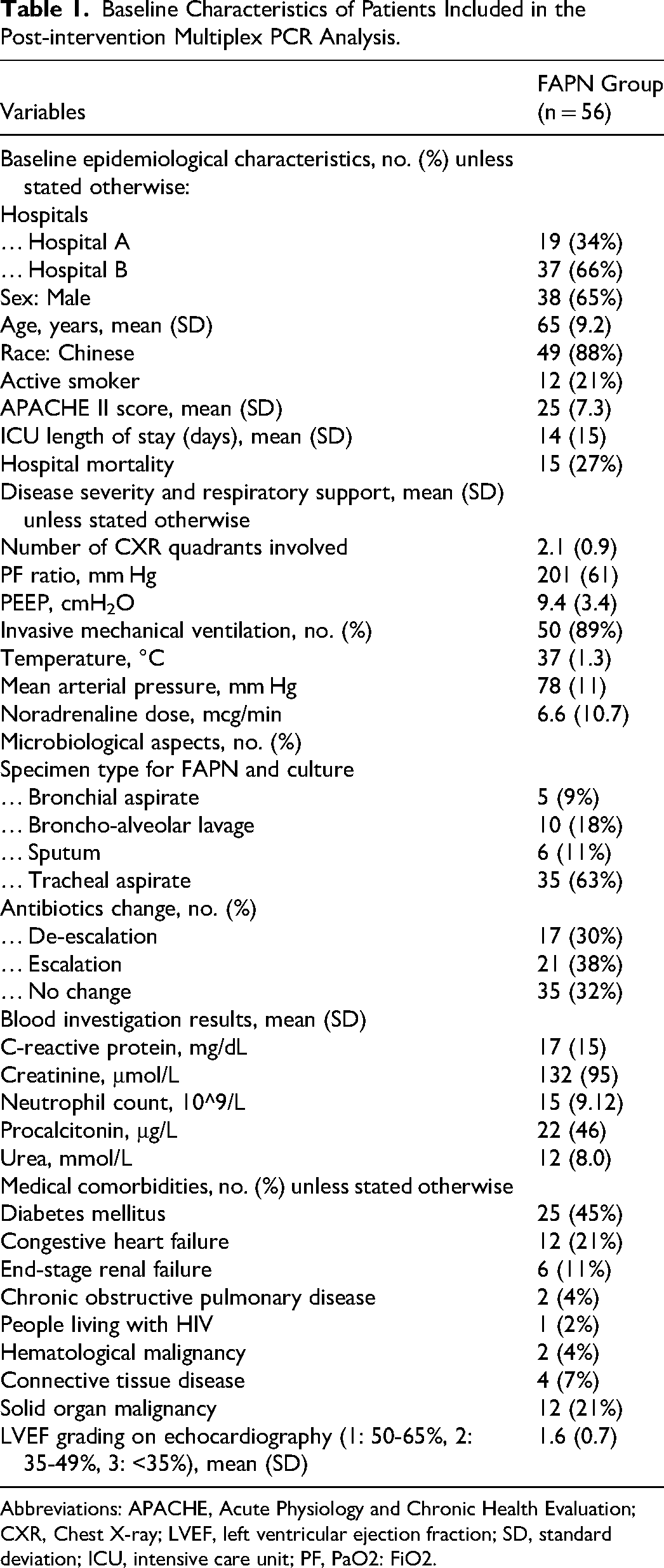

Abbreviations: APACHE, Acute Physiology and Chronic Health Evaluation; CXR, Chest X-ray; LVEF, left ventricular ejection fraction; SD, standard deviation; ICU, intensive care unit; PF, PaO2: FiO2.
Standard Cultures Combined with FAPN Identifies non-Infectious Mimics of Pneumonia
Among the 56 patients in the post-intervention cohort, 45 (80%) were clinically diagnosed with pneumonia, including infectious pneumonia (n = 33, 73%) and non-infectious mimics of pneumonia (n = 12, 27%). The correlation between clinical diagnosis and combined microbiological findings is shown in Table 2. Using “negative FAPN and sputum culture” to detect non-infectious mimics yielded a sensitivity of 75% (95% CI: 50-100%) and a specificity of 91% (95% CI: 81-100%).
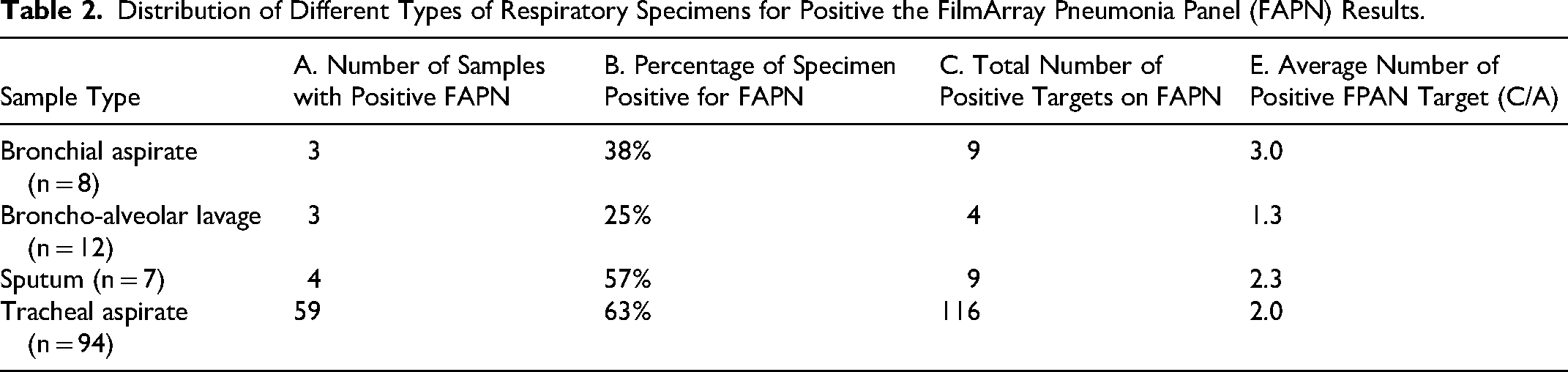
Distribution of Different Types of Respiratory Specimens for Positive the FilmArray Pneumonia Panel (FAPN) Results.
Use of FAPN in Antimicrobial Stewardship: Reduced Escalation Observed
To evaluate the impact of FAPN on antibiotic prescribing, we compared the intervention cohort with a historical cohort that did not undergo FAPN testing. In univariable logistic regression, the use of FAPN, age, and the presence of connective tissue disease were significantly associated with antibiotic escalation following release of test results (Appendix Table 1).
In multivariable analysis (Figure 2), FAPN use remained independently associated with a lower likelihood of escalation (OR 0.70, 95% CI 0.60-0.81; p < .01). Conversely, connective tissue disease was associated with higher odds of escalation (OR 1.29, 95% CI 1.02-1.64; p = .03). Other variables—including age, APACHE II score, CURB-65 score, PF ratio, and number of CXR quadrants involved—were not significantly associated (Appendix Table 2).
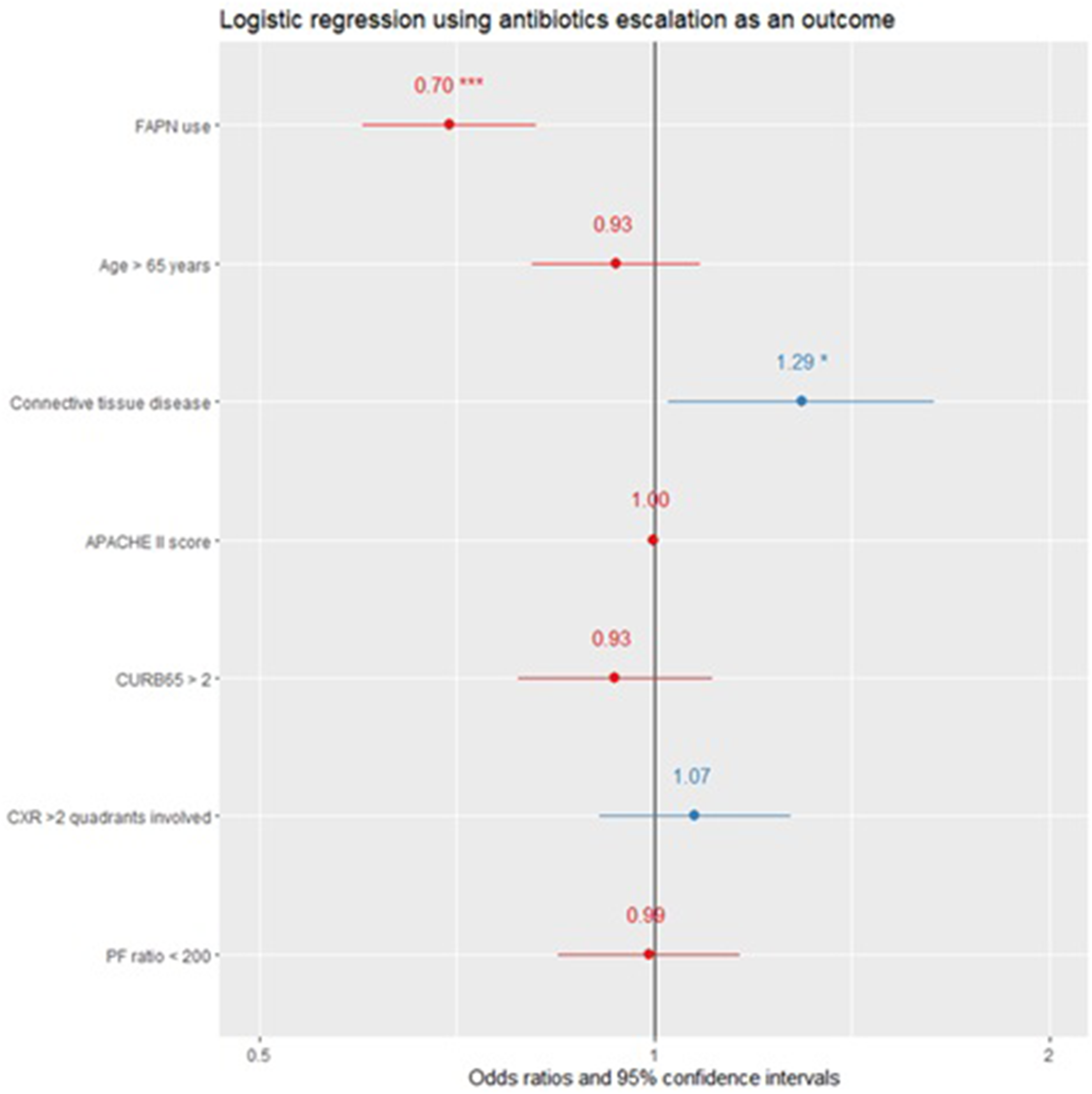
Forest plot of multivariable logistic regression using antibiotics escalation as an outcome.
Low Concordance Between Bacterial Targets of multiplex PCR and Bacterial Culture
All 112 patients who underwent FAPN testing were included in the concordance analysis. The distribution of sample types is shown in Table 2. Most specimens were tracheal aspirates (n = 94, 77.7%), followed by bronchoalveolar lavage (n = 12, 9.9%), bronchial aspirates (n = 8, 6.6%), and sputum (n = 7, 5.8%). Tracheal aspirates had the highest proportion of FAPN-positive results (59/94, 63%), with an average of 2.0 bacterial targets per positive sample. Bronchial aspirates and BAL specimens were positive in 38% and 25% of cases, with average target counts of 3.0 and 1.3, respectively.
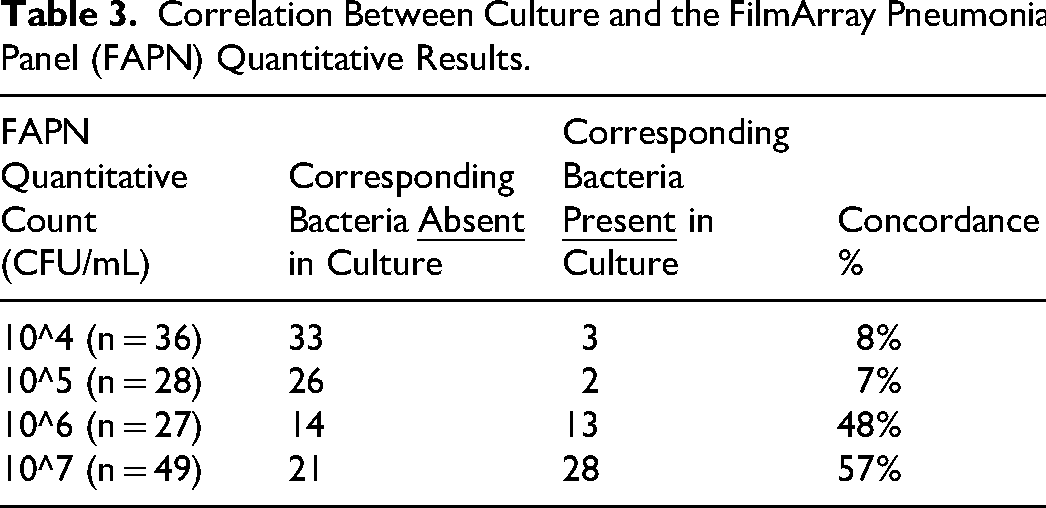
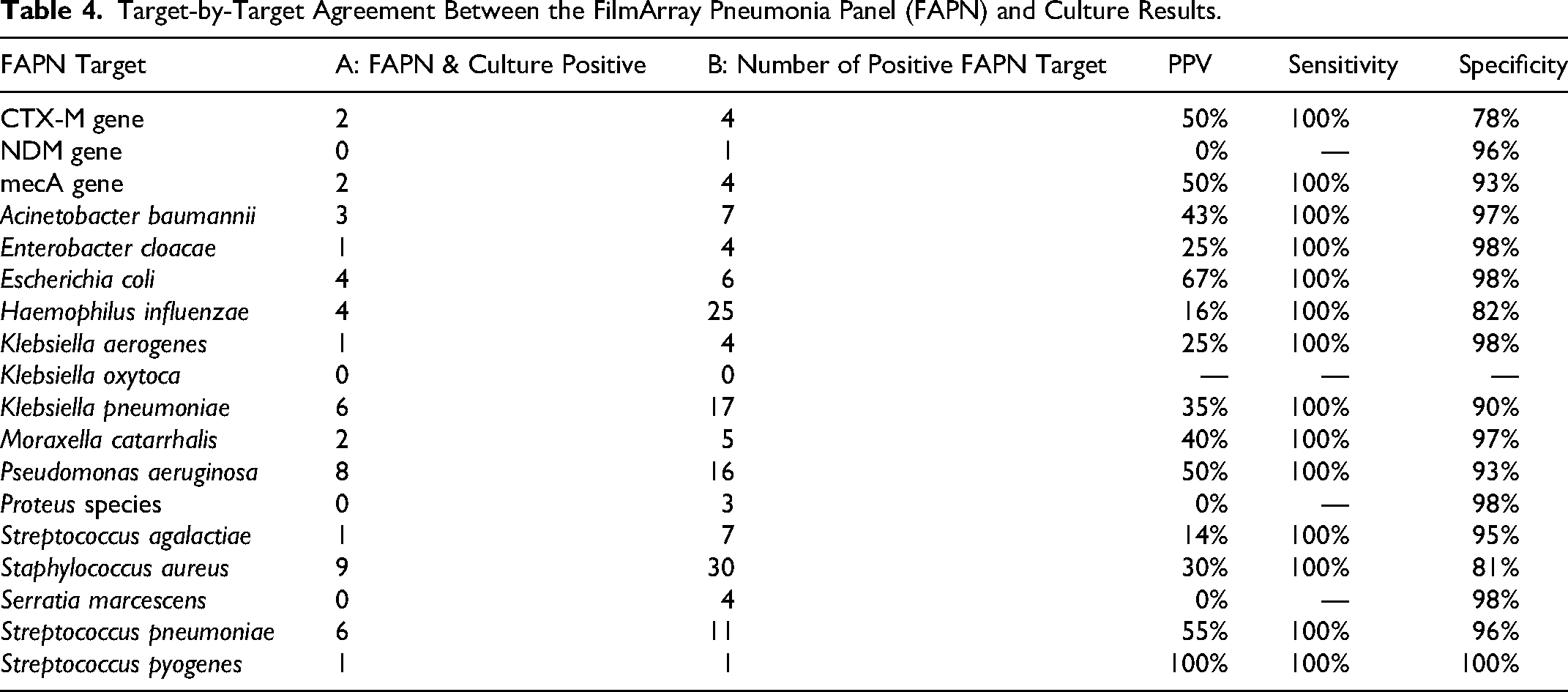
Quantitative concordance between FAPN and culture differed significantly across bacterial load categories (p < .001; Table 3). For detections at 104 and 105 CFU/mL, only 8% and 7% were culture-confirmed. At higher loads of 106 and 107 CFU/mL, concordance increased to 48% and 57%. Target-specific performance data are shown in Table 4. PPVs ranged from 0% to 100%, with a median of 35%. Sensitivity was 100% for all targets; specificity ranged from 78% to 100%; and NPV was 100% for all targets. Half of CTX-M and mecA gene detections were culture-confirmed, whereas the single NDM detection had no corresponding isolate.
Correlation Between Culture and the FilmArray Pneumonia Panel (FAPN) Quantitative Results.

Target-by-Target Agreement Between the FilmArray Pneumonia Panel (FAPN) and Culture Results.
Fourteen bacterial isolates (11.6%) identified by culture were not included in the FAPN panel. These included Stenotrophomonas maltophilia (n = 4), Citrobacter koseri (n = 2), Corynebacterium striatum (n = 2), Pseudomonas spp. other than P. aeruginosa (n = 2), Bordetella spp. (n = 2), and Burkholderia cepacia complex (n = 2). Additionally, 18 Candida isolates (16.1%) were recovered by culture but not detected by FAPN.
Discussion
In this study, we evaluated the clinical utility of the FilmArray Pneumonia Panel (FAPN) in critically ill patients with severe respiratory failure, focusing on diagnostic performance, concordance with conventional microbiology, and impact on antimicrobial stewardship. Several important observations emerged.
First, among ARDS patients with suspected pneumonia, combining FAPN with standard sputum cultures proved useful in identifying non-infectious mimics of pneumonia. A negative combined FAPN–culture result showed good sensitivity (75%) and excellent specificity (91%) for detecting non-infectious causes, suggesting that negative multiplex PCR results can meaningfully exclude a bacterial etiology when interpreted alongside conventional microbiology. Without these negative results, approximately one-quarter of clinically confirmed non-infectious mimics might have been misclassified, highlighting the value of rapid molecular diagnostics not only for pathogen identification but also for ruling out bacterial infection in complex ICU presentations.
Second, FAPN use was associated with a significant reduction in antibiotic escalation. After adjustment for confounders, FAPN remained independently associated with lower odds of escalation (OR 0.70, p < .01), indicating that rapid diagnostic data influenced prescribing behavior beyond traditional markers of illness severity. In contrast, connective tissue disease increased the likelihood of escalation, likely reflecting clinician concern about immunosuppression or atypical presentations. Together, these findings reinforce an important stewardship benefit of FAPN: timely, actionable information that helps clinicians avoid unnecessary antimicrobial broadening.
Third, we observed substantial discordance between FAPN and standard bacterial cultures across specimen types. Although tracheal aspirates produced the highest proportion of FAPN-positive results, culture concordance was poor at lower PCR bins (10^4-10^5 CFU/mL), where fewer than 10% of detections were confirmed. Concordance improved at higher loads, consistent with existing evidence that low qPCR signals often indicate colonization rather than true infection. The median PPV of 35% for individual targets further underscores the risk of over-interpreting positive PCR results. Importantly, sensitivity and NPV remained 100%, reinforcing FAPN's strength as a rule-out tool rather than a stand-alone rule-in test.
The discrepancy between FAPN and culture is likely multifactorial. Differences in detection thresholds, the presence of non-viable organisms—particularly in antibiotic-exposed patients—and airway colonization, especially in the mechanically ventilated, contribute to discordance.10,11 Conversely, 14 isolates not covered by the FAPN panel (eg, Stenotrophomonas maltophilia, Citrobacter koseri, Burkholderia cepacia complex) were identified only by culture, highlighting the incompleteness of even broad multiplex panels. This limitation is especially relevant in units where non-panel organisms are prevalent or clinically significant.
These findings have important clinical implications. The strong negative predictive performance of FAPN supports its use for early de-escalation and avoidance of unnecessary antibiotic escalation. However, positive detections—particularly multiple organisms or low bacterial loads—must be interpreted holistically with clinical context, radiology, biomarkers, and culture results. Over-reliance on multiplex PCR risks misclassifying colonization as infection and driving overtreatment. Our findings suggest that FAPN is optimally used as an adjunct to rather than a replacement for standard microbiological methods.
This study has strengths, including real-world implementation of FAPN, detailed culture correlation, and a comparative historical cohort evaluating stewardship outcomes. Limitations include modest sample size, potential confounding inherent in a quasi-experimental design, and reliance on culture as an imperfect gold standard. Microbial epidemiology and stewardship practices may also limit generalizability.
In conclusion, FAPN offers valuable diagnostic information for critically ill patients with suspected pneumonia, particularly through its high negative predictive value and its association with reduced antibiotic escalation. However, limited concordance with culture at lower bacterial loads and incomplete coverage of clinically relevant organisms emphasize the need for thoughtful, context-dependent interpretation. When integrated appropriately, FAPN can enhance antimicrobial stewardship and improve diagnostic clarity in severe respiratory failure.
Supplemental Material
sj-docx-1-jic-10.1177_08850666261442190 - Supplemental material for Prospective Multicenter Evaluation of a multiplex PCR Pneumonia Panel to Distinguish Infectious Pneumonia from Non-Infectious Mimics in ICU Respiratory Failure
Supplemental material, sj-docx-1-jic-10.1177_08850666261442190 for Prospective Multicenter Evaluation of a multiplex PCR Pneumonia Panel to Distinguish Infectious Pneumonia from Non-Infectious Mimics in ICU Respiratory Failure by Lap Tin Ho, Alfred Lok Hang Lee, Chin Wing Yu, Eddie Chi Man Leung, Man Ting Yuk, Thomas Yuk Bun Lau, Ingrid Yu Ying Cheung, Viola Chi Ying Chow, Duncan Ka Hing Lam, Philip Koon Ngai Lam and Kwok Ming Ho in Journal of Intensive Care Medicine
Footnotes
Acknowledgements
The authors gratefully acknowledge the dedication and collaboration of our colleagues from the four participating teams: the Intensive Care Unit of North District Hospital, the Intensive Care Unit of Alice Ho Miu Ling Nethersole Hospital, the Microbiology Laboratory of Prince of Wales Hospital, and the Microbiology Laboratory of Alice Ho Miu Ling Nethersole Hospital. Their commitment and hard work made this study possible.
ORCID iDs
Ethics Approval and Consent to Participate
Ethics approval was obtained from the Joint Chinese University of Hong Kong–New Territories East Cluster Clinical Research Ethics Committee (reference numbers: 2025.172 and 2025.173). The committee waived the requirement for patient consent. All study procedures were conducted in accordance with the Declaration of Helsinki.
Consent for Publication
Not applicable
Authors’ Contributions
LTH and ALHL were responsible for conception, study design, clinical case review, data acquisition and analysis, and manuscript preparation. CWY was responsible for clinical case review and manuscript review. ECML was responsible for laboratory testing and laboratory data acquisition. TLYB, DKHL, MTY, IYYC, VCYC, PKNL, and KMH were responsible for manuscript review, clinical oversight, and institutional support.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
