Abstract
Objectives:
Domain shift has been shown to have a major detrimental effect on AI model performance however prior studies on domain shift for MRI prostate cancer segmentation have been limited to small, or heterogenous cohorts. Our objective was to assess whether prostate cancer segmentation models trained on local MRI data continue to outperform those trained on external data with cohorts exceeding 1000.
Methods:
We simulated a multi-institutional consortium using the public PICAI dataset (PICAI-TRAIN:
Results:
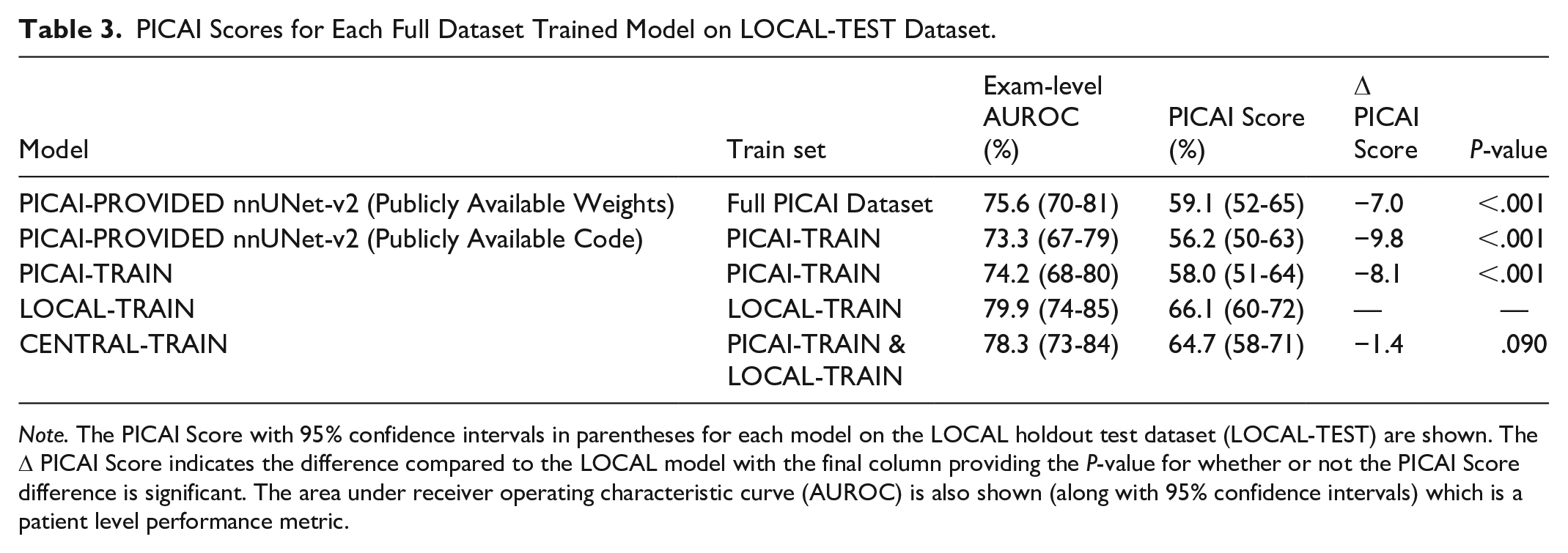
Just 22% (309/1400) of LOCAL-TRAIN exams would be sufficient to match the performance of a model trained on PICAI-TRAIN. The CENTRAL-TRAIN performance was similar to LOCAL-TRAIN performance, with PICAI Scores [95% CI] of 65 [58-71] and 66 [60-72], respectively. Both of these models exceeded the model trained on PICAI-TRAIN alone which had a score of 58 [51-64] (
Conclusion:
Domain shift limits MRI prostate cancer segmentation performance even when training with over 1000 exams from 3 external institutions. Use of local data is paramount at these scales.
Introduction
Magnetic Resonance Imaging (MRI) is the recommended imaging test for clinically significant prostate cancer (csPCa) detection.1,2 However, MRI requires interpretation by highly trained radiologists, with significant variation in performance. 3 Artificial intelligence (AI) may help in improving radiologist performance and consistency. 4 The ideal would be a single AI model that can identify and localize suspicious lesions with expert-level performance despite inherent variations such as patient populations, scanner types, and MRI acquisition protocols.
A long-standing belief is that training a single model on as much multisite data as possible will, given enough samples, eventually achieve expert-level performance. 5 While this assumption generally holds true, how much data is sufficient in the prostate MRI domain remains uncertain. 5 Moreover, emerging research in bi-parametric MRI (bpMRI) csPCa segmentation suggests that performance gains plateau when extending data beyond a single institution using data sharing, and that such models struggle to generalize to institutions that did not contribute data.6 -9 This decline is largely attributed to domain shift—where subtle differences in imaging protocols, scanner types, and patient demographics degrade model performance on unseen data.6 -10 AI model development assumes that the training set is representative of all technical and population variations for sufficient generalizability. Covering all variations is a major challenge and tuning with local data may be more efficient than trying to create such a diverse data set.
Given the availability of models based on multi-institutional data, this raises the question of whether training with local data is still necessary. Previous studies exploring this question for csPCa segmentation have been limited by small datasets, custom models, inconsistent metrics, and mixtures of endorectal and non-endorectal coil MRI protocols the latter being an obvious cause of domain shift.6 -9 Given the availability of an established open-source model (nnUNet-v2) 11 a sizable open-source multi-institutional dataset and metric library shared by the PICAI group, 12 we can study the effect of local data on performance with contemporary prostate MRI at a larger scale. The purpose of this study was to assess, using standardized metrics and methodologies, whether csPCa segmentation models trained on local MRI data continue to outperform those trained on external data at dataset sizes exceeding 1000 exams.
Materials and Methods
Study Design
This retrospective study employed 2 bpMRI datasets to compare local versus external data for csPCa segmentation: PICAI and LOCAL. Exclusion criteria are detailed in Figure 2. The study was approved by the institutional review board and informed consent was waived.
The LOCAL dataset represented a single local institution and the PICAI dataset represented an external multi-institutional cohort. Each dataset was split into training and testing cohorts resulting in 4 subset datasets (LOCAL-TRAIN, LOCAL-TEST, PICAI-TRAIN, PICAI-TEST). We defined a 5th dataset (CENTRAL-TRAIN) as the union of LOCAL-TRAIN and PICAI-TRAIN (Figure 1). Using identical model architectures and training procedures, we trained 3 models: one on local data (LOCAL-TRAIN), one on the external PICAI data (PICAI-TRAIN), and one on the combined local and external data (CENTRAL-TRAIN). This process was repeated with reduced dataset sizes (40%, 20%, and 10% of the original PICAI and LOCAL training sets), resulting in 12 total models (3 per subset size). To further contextualize results, we compared performance to the nnUNet-v2 model provided from the PICAI competition (PICAI-PROVIDED)—a semi-supervised model that ranked 11th of 26 in the open phase and 8th of 12 in the closed phase. We reported results using both the official pretrained weights 13 on the full publicly available PICAI dataset (1500 exams) and our own version trained on the PICAI-TRAIN cohort (subset of 1241 exams) using the authors’ publicly released code. 14 All models were evaluated on the LOCAL-TEST dataset. Secondary testing on the PICAI-TEST set is provided in Supplemental Material.
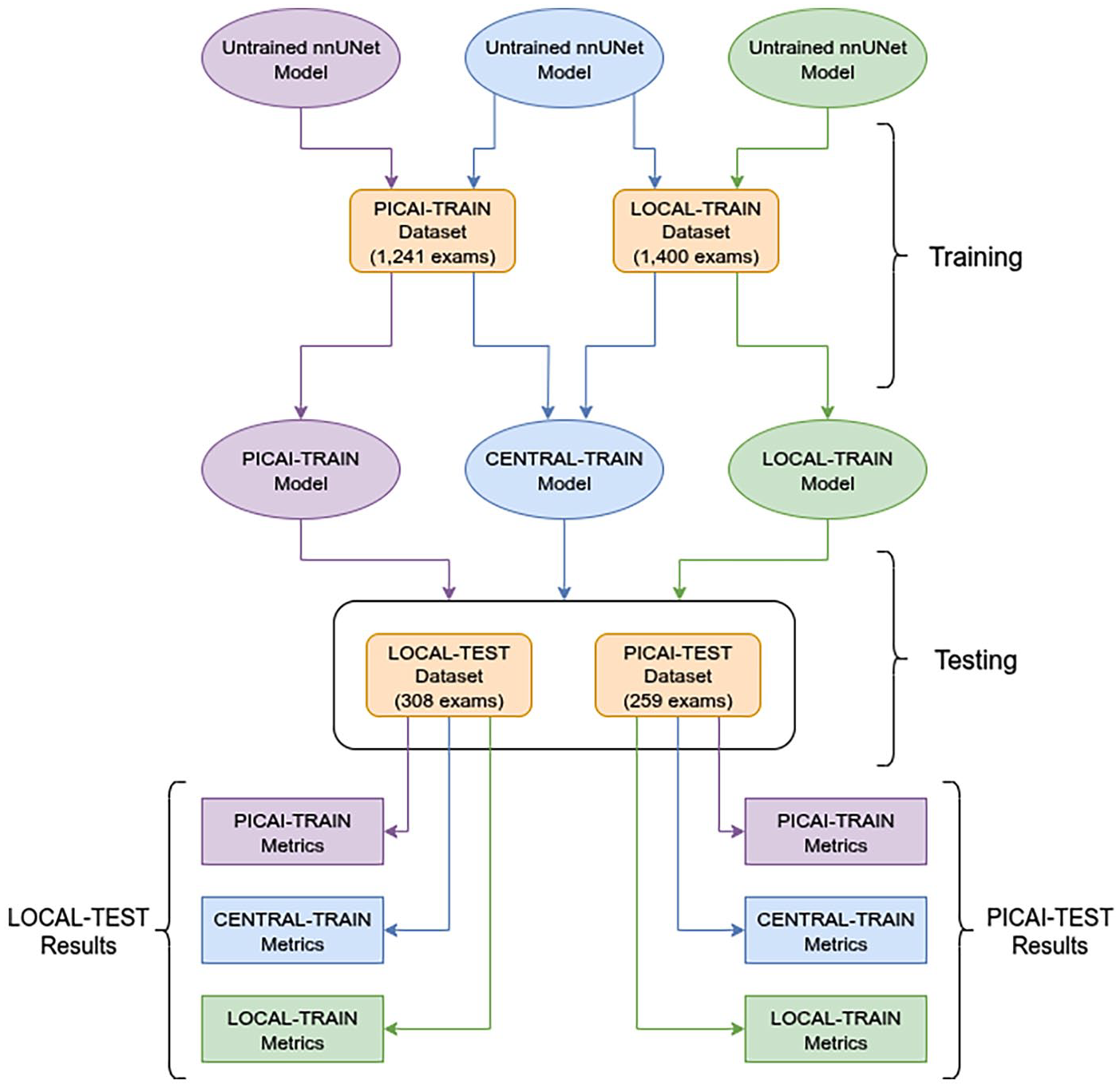
Dataset combinations used for training and naming of the corresponding models. Datasets are indicated in rounded corner rectangles, models in ellipses, and metrics in rectangles. The primary test dataset for all models was the LOCAL-TEST dataset. Additional results on the PICAI-TEST dataset are in Supplemental Material. The training cohorts consisted of the LOCAL training cohort, the PICAI training cohort of the combination of both termed the CENTRAL training cohort. This same experimental design was repeated using 40%, 20%, and 10% of the PICAI-TRAIN and LOCAL-TRAIN datasets. 100% of the LOCAL and PICAI TEST datasets were always used during evaluation.
Participants
The PICAI dataset comprised 1500 publicly available prostate MRI exams from 1476 men suspected of having csPCa based on elevated prostate-specific antigen (PSA) levels or abnormal digital rectal examination. These exams were acquired between 2012 and 2021 across 3 centers in the Netherlands. 12 For context PICAI only publicly released a portion of their full dataset which included an additional 8707 exams from 4 institutions. The LOCAL dataset included 2771 consecutive exams acquired between 2012 and 2020 from treatment-naive patients referred for prostate MRI at our institution due to elevated risk. After applying the exclusion criteria (Figure 2) to the LOCAL institution data, 1708 MRI examinations from 1514 patients were included.
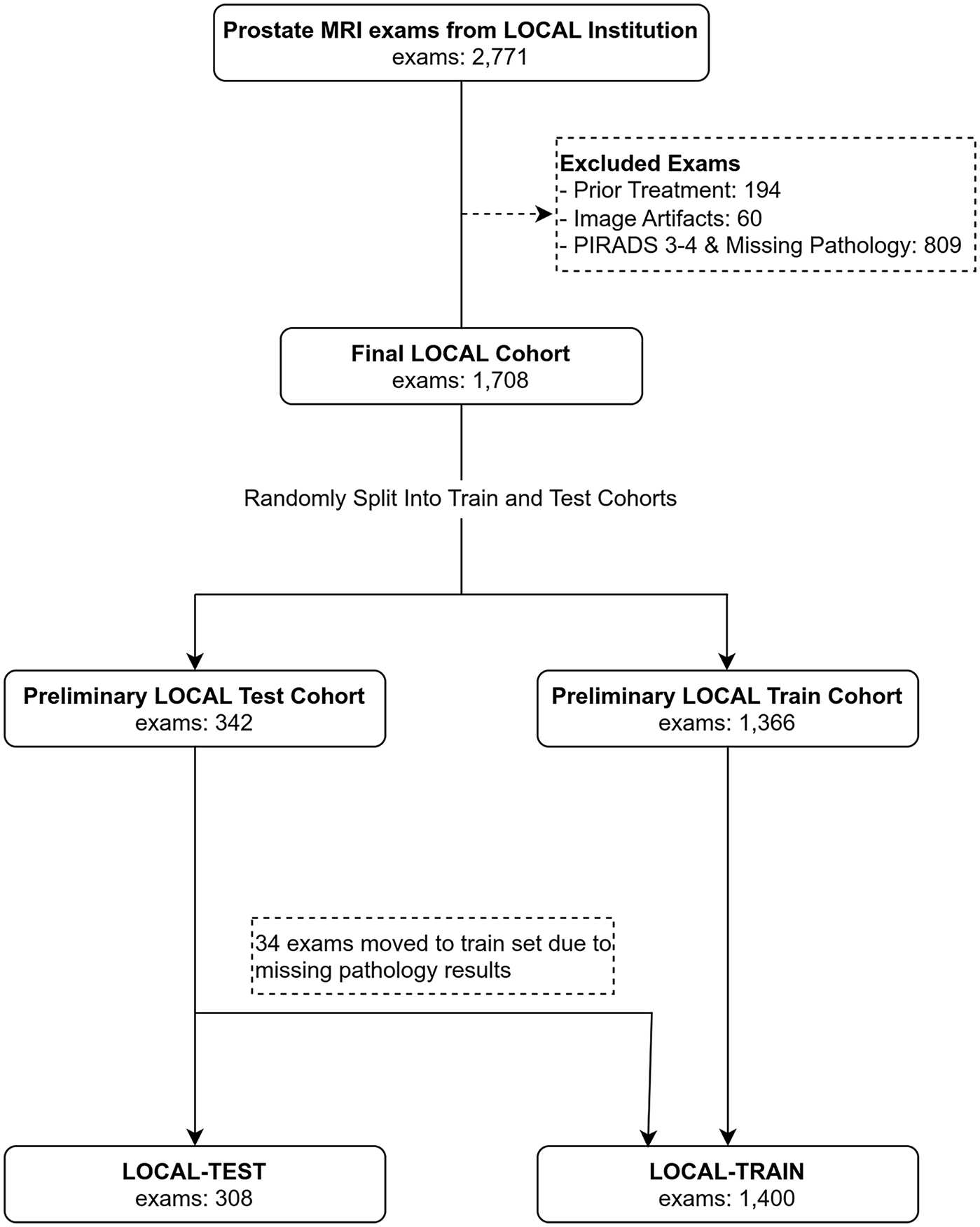
Flow diagrams showing exam selection from LOCAL Institution for LOCAL Dataset.
Image Acquisition
All exams across both datasets consisted of an axial T2-weighted (T2W) image, a high b-value image (BHIGH) and an apparent diffusion coefficient (ADC) map derived from the diffusion weighted images. All images in the LOCAL cohort were acquired on Siemens 3T systems (Avanto, Vario, Skyra, Siemens Healthineers, Erlangen, Germany). Protocols varied over time however the following common protocol parameters were used: a surface phased array coil without an endorectal coil, 3 mm slice thickness, field of view from 16 to 20 cm, and b values ranging from 0-100 s/mm2 for the lowest b-value to 900-1600 s/mm2 for the highest b-value. For BHIGH, extrapolated b-value maps were calculated and used at
Labels
For both the LOCAL and PICAI datasets, available labels were the segmentation masks, PIRADS v2.1 scores assigned by radiologists 15 and biopsy-confirmed pathology results. Patients with multiple exams were treated independently. In the LOCAL dataset, segmentation masks were drawn using ITK-SNAP v4.0.2 16 by an abdominal radiologist under the supervision of a uroradiologist with more than 25 years of experience reporting more than 1000 exams with pathology correlation. For the PICAI dataset training we used the official annotations provided by Saha et al. 12 At the voxel-level these included 220 annotations completed by a trained investigator or expert radiologist, while the annotations for the remaining 205 csPCa-positive exams were AI-generated using a model by the PICAI group. 17
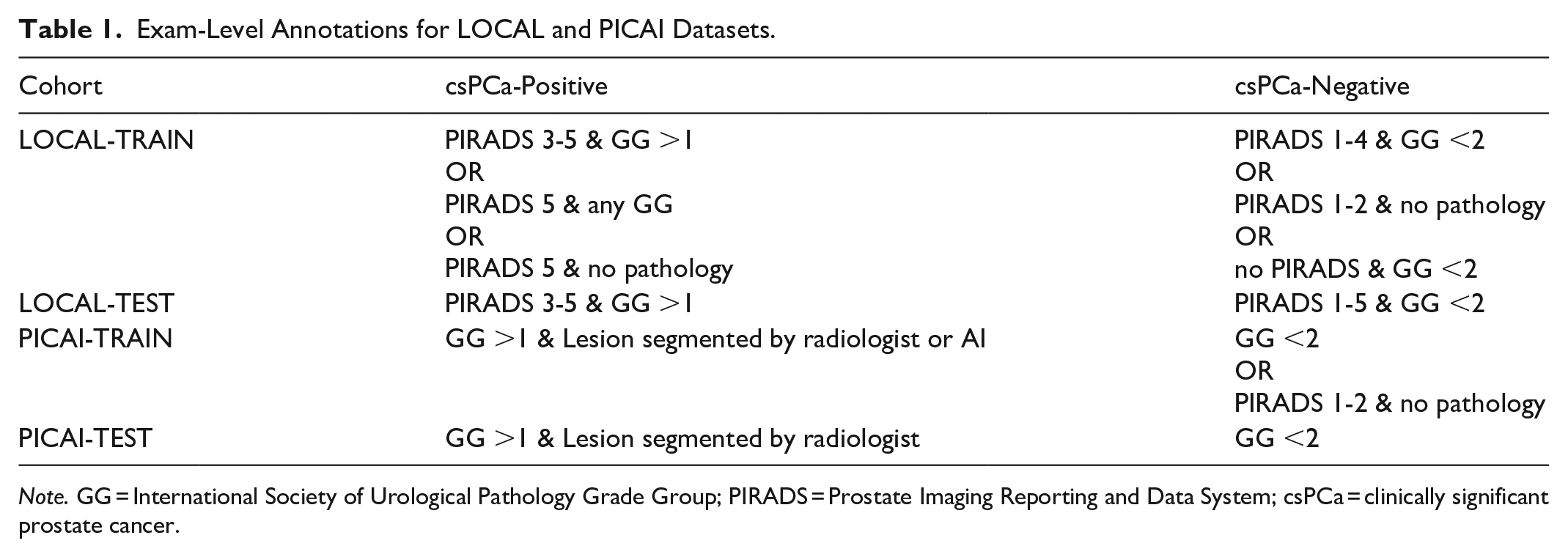
Both the PICAI and LOCAL datasets were randomly split into training (≈80%) and testing (≈20%) cohorts resulting in 4 dataset subsets (PICAI-TRAIN, PICAI-TEST, LOCAL-TRAIN, LOCAL-TEST). There were 171 and 23 patients with multiple exams in LOCAL and PICAI respectively. To ensure independence between subsets, patients with multiple MRI exams were assigned entirely to either training or testing. This was consistent with the approach taken by the PICAI group. 12 It is worth noting that individual lesion PIRADS scores were not publicly released for the PICAI dataset. As a result slightly different label criteria were used in training and testing. A summary of the annotation criteria used for training and testing is provided in Table 1. Pathology labels were available for all exams in the PICAI dataset, some LOCAL training data and all LOCAL test data. A detailed overview of the datasets and labeling criteria are in Table 2. Exams with PIRADS scores 1, 2, or 5 that lacked pathology were included only in training with 1 and 2 considered negative and 5 considered positive. PIRADS 3 and 4 exams without pathology labels were not used. For LOCAL-TEST, only pathologically proven cases were included with Grade Group (GG) greater than 1 being considered positive for csPCa. For model training, standard 5-fold cross-validation was used.
Exam-Level Annotations for LOCAL and PICAI Datasets.
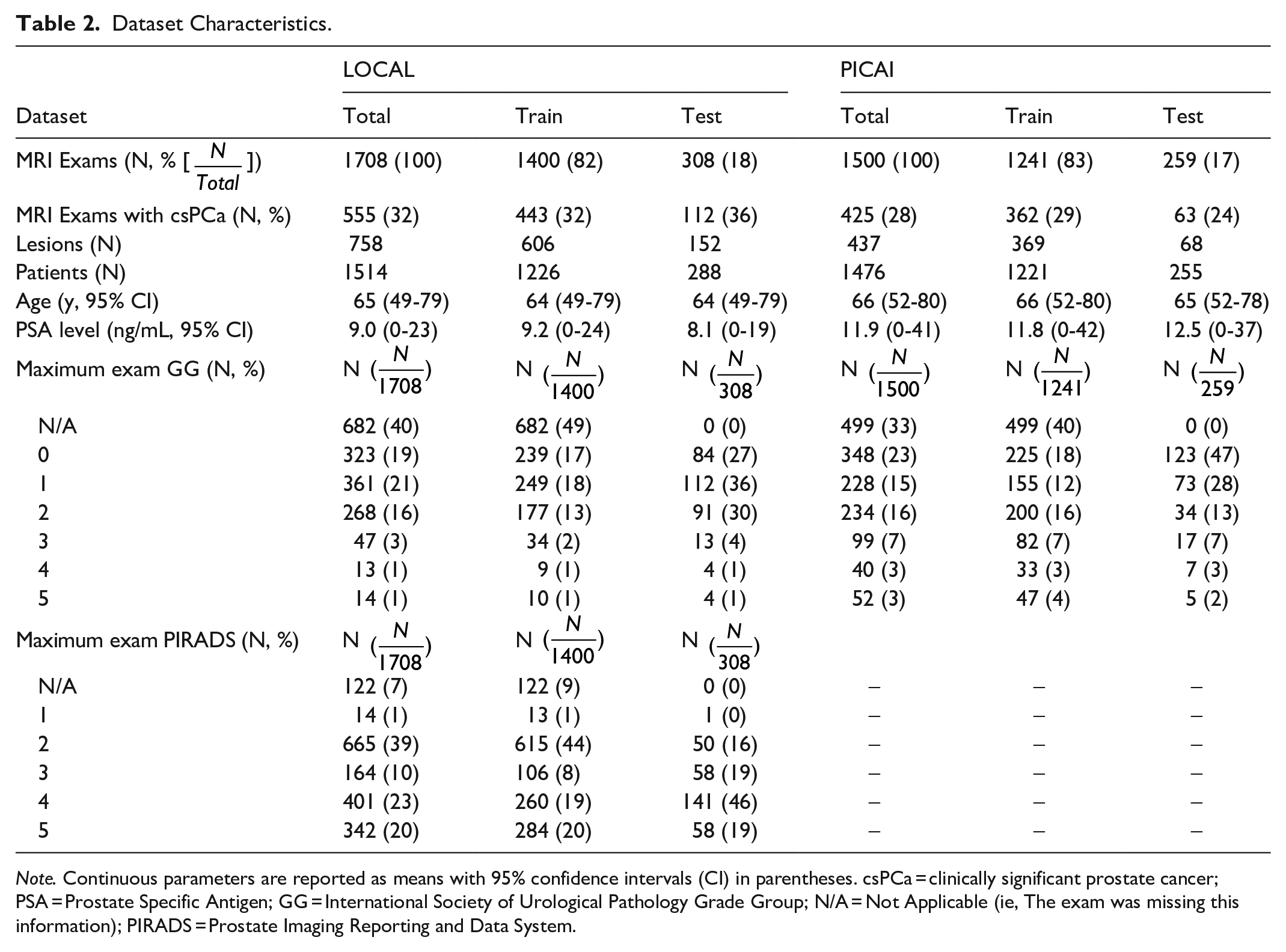
Dataset Characteristics.
Model
We used nnUNet-v2 to provide a well understood baseline model for comparison. 11 This choice promotes reproducibility and aligns with tools commonly used in clinical research settings. The “3d_fullres” configuration was used, with 3 volumes (T2W, BHIGH, ADC) and outputting a continuous csPCa heat-map. For each exam, the BHIGH image and ADC map were resampled to match the resolution of the T2W image as per PICAI preprocessing methods. 12 We used the nnUNet-v2 plans file generated from LOCAL-TRAIN, and hence the same model architecture and training settings, for all experiments, updating only the stored dataset statistics used by nnUNet-v2 for normalization prior to training. Training followed nnUNet-v2 defaults, except we replaced the loss function with an equal-weighted sum of Focal Loss and Binary Cross Entropy Loss, as this was used in training the PICAI-PROVIDED model. 12 Additional details on data preprocessing, model architecture, and model training are included in the Supplemental Material.
Evaluation Metrics and Statistics
Previous studies have used various voxel and exam level metrics to evaluate performance,5
-10 but inconsistent post-processing methods limit comparability.
18
To address this, we adopted the standardized evaluation pipeline from the PICAI competition,
12
which emphasizes lesion-level assessment over voxel-level accuracy (see Supplemental Material). As noted in the competition guidelines, the PICAI Score “mandates coupling the tasks of lesion detection and patient diagnosis to promote interpretability and disincentivize AI solutions that produce inconsistent outputs.”
19
The PICAI Score was the primary metric and is the mean of 2 submetrics, an area under the receiver operating characteristic (AUROC) based on the model’s overall confidence that an exam contains csPCa and an average precision (AP) based on the model’s csPCa confidence for individual predicted lesions (for which there may be multiple in a single exam). We report both of these submetrics in our results. For evaluation, we selected the best-performing model checkpoint for each fold based on validation performance and used these checkpoints to generate predictions on the test sets. For each test sample, an ensembled prediction was generated by averaging the outputs from all folds. These ensembled predictions were used to compute the PICAI Score, exam-level AUROC, and lesion-level AP. Reported values for these metrics represent the mean performance over 1000 bootstrap iterations. We used bootstrapping to estimate 95% confidence intervals and compute
Results
The 1500 publicly available exams from the PICAI dataset were included in this study. For the LOCAL dataset, 1063 exams were not used in this study based on the exclusion criteria with 1708 exams from 1514 patients remaining. Both the PICAI and LOCAL datasets were partitioned into independent training and testing sets (PICAI-TRAIN, PICAI-TEST, LOCAL-TRAIN, LOCAL-TEST). Dataset characteristics for all partitions are presented in Table 2.
Model Performance on Full Datasets
We evaluated the LOCAL-TRAIN, PICAI-TRAIN, CENTRAL-TRAIN, and the PICAI-PROVIDED models on the LOCAL-TEST cohort (Table 3). Both LOCAL-TRAIN (66,
PICAI Scores for Each Full Dataset Trained Model on LOCAL-TEST Dataset.
Model Performance on Partial Datasets
To estimate how many local exams are needed to match the performance of a model trained on the full PICAI training set and explore the effect of dataset size on model performance, we retrained the LOCAL-TRAIN model on progressively smaller subsets of its data. The PICAI-TRAIN and CENTRAL-TRAIN models were similarly retrained on subsets of their data to assess how performance scales with training volume. PICAI Scores for all 12 models (3 models × 4 dataset sizes) on the LOCAL-TEST dataset are shown in Figure 3. Exam-Level AUROC and Lesion-Level AP are shown in Figures 4 and 5, respectively, with a summary of all metrics across all subsets in Table 4. Results for the PICAI-TEST set are provided in the Supplemental Material.
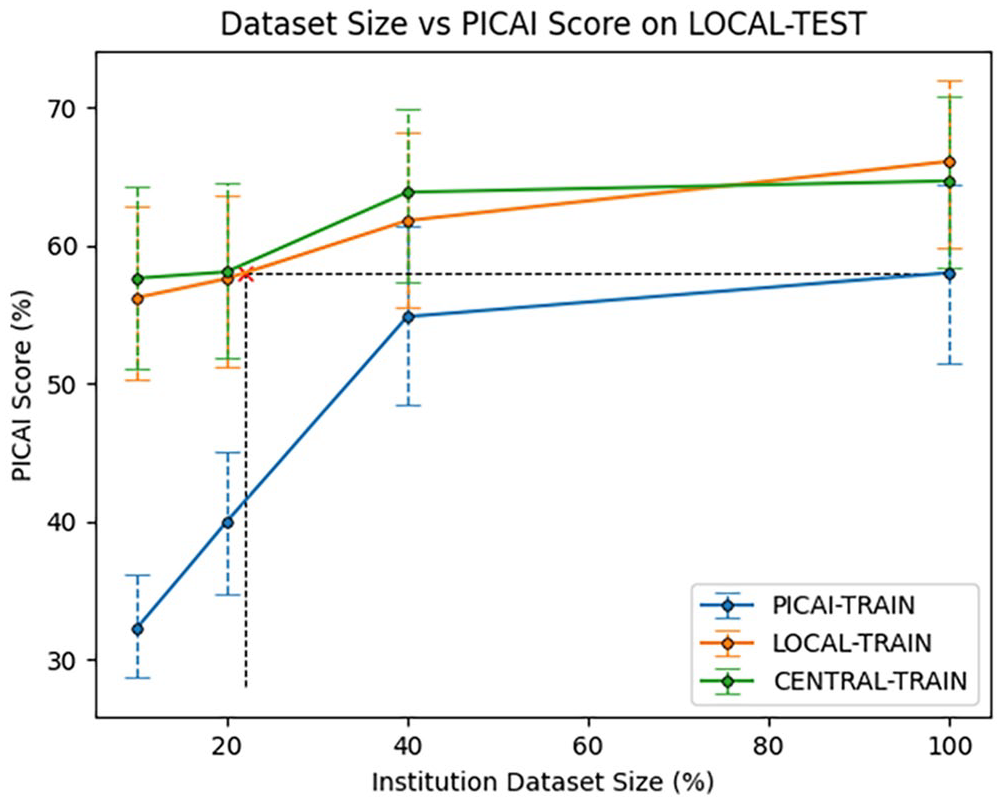
Institutional Dataset Size versus PICAI Score on LOCAL-TEST Dataset. The
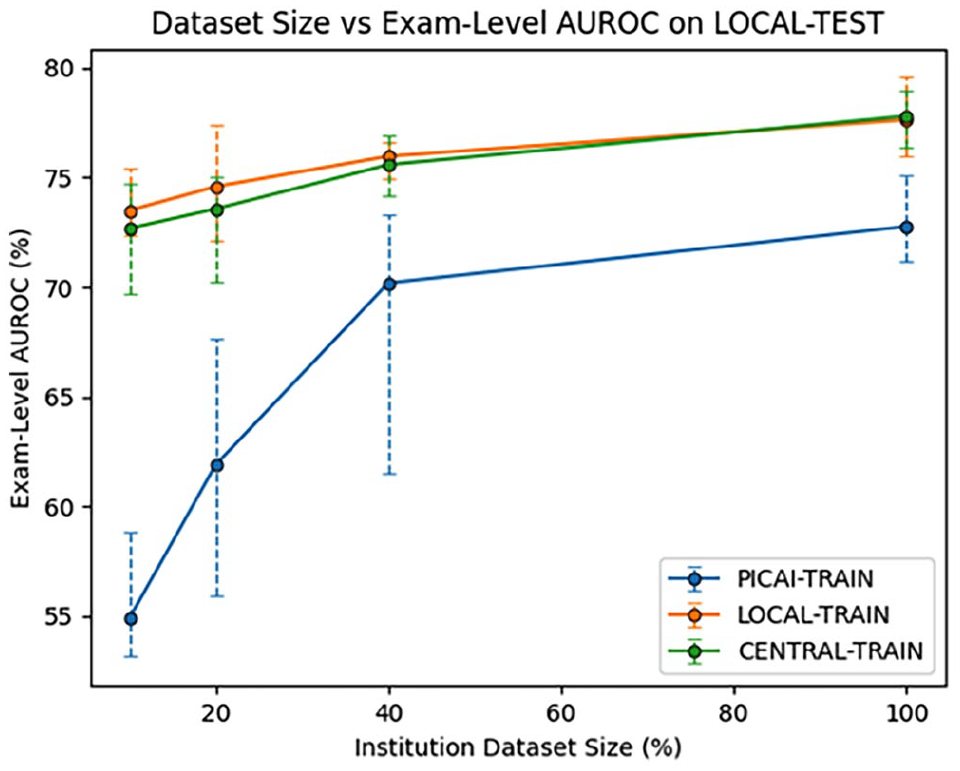
Institutional Dataset Size versus Exam-Level AUROC on LOCAL-TEST Dataset. The
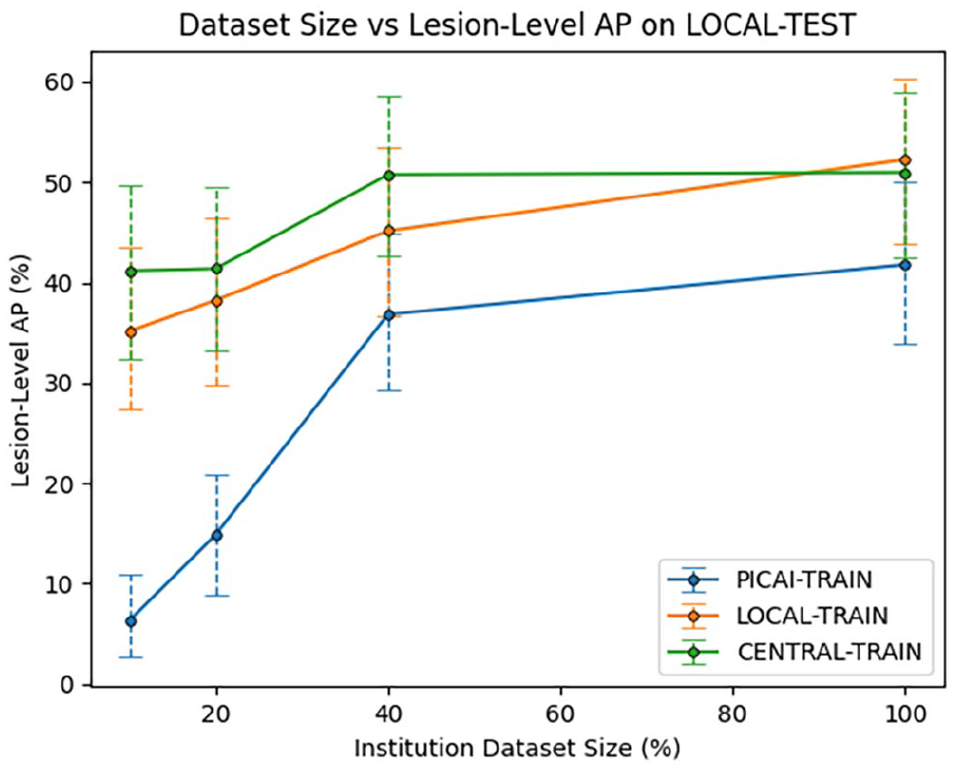
Institutional Dataset Size versus Lesion-Level AP on LOCAL-TEST Dataset. The
Results on the LOCAL-TEST Dataset With Partial Dataset Sizes.
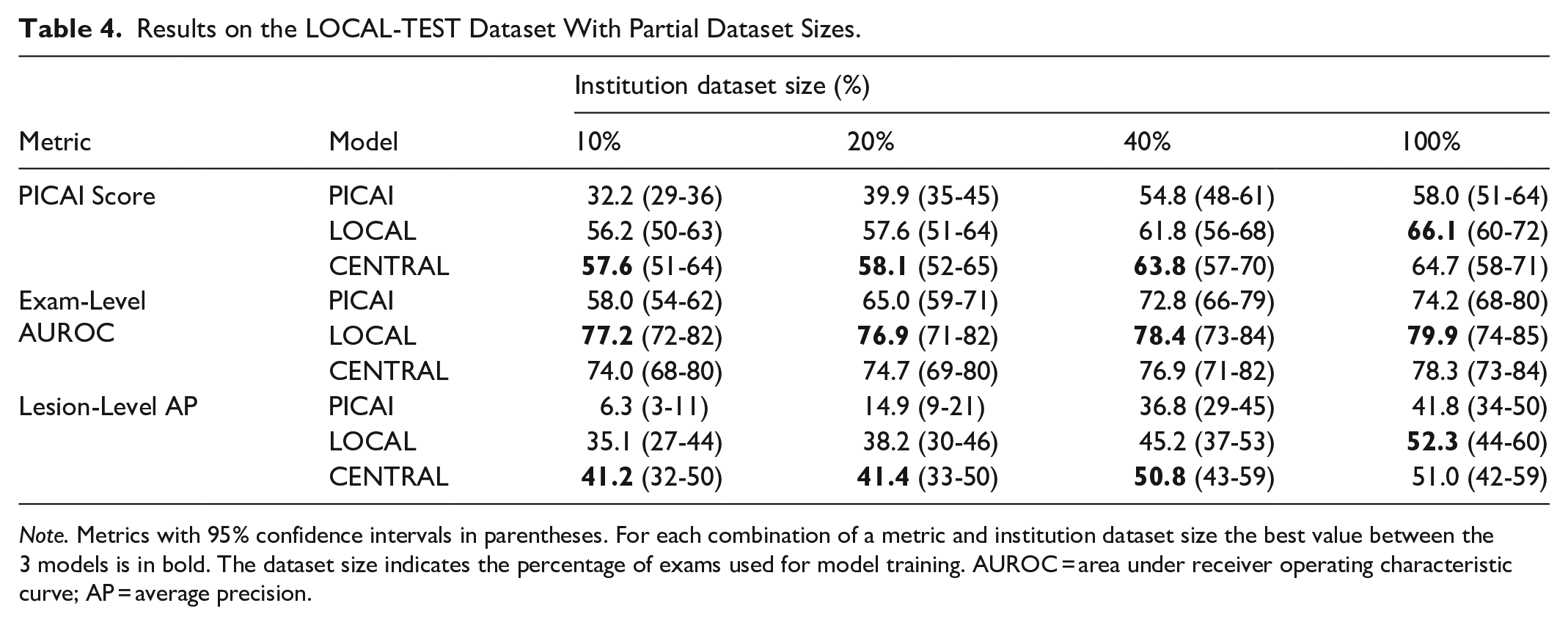
CENTRAL-TRAIN and LOCAL-TRAIN performance were consistently better than PICAI-TRAIN performance on the LOCAL-TEST dataset across all dataset sizes (
Discussion
Using a nnUNet-v2 architecture trained on varying amounts and combinations of local and external data, we investigated the relative value of local versus external exams for training AI models to segment and detect csPCa in bpMRI. Models trained with local data consistently outperformed those tested on an external cohort, even when using substantially less data. We found that just 309 local exams would be sufficient to match the local performance of training with a much larger multi-institutional dataset of 1241 external exams. The model provided by PICAI also underperformed on our local test set, suggesting that in-house implementation biases were not a factor. Even combining all available data into a centralized model yielded only minimal, statistically insignificant gains over local training alone.
While previous studies have explored the impact of dataset composition on model generalization,6 -9 our work uniquely assesses these impacts on a much larger scale using established public-domain model architectures and metrics. Importantly, we conduct our analysis using standardized, publicly available models and metrics tailored for csPCa by the PICAI group, 12 facilitating reproducibility and cross-study comparison. In contrast, many prior studies employ a wide range of models and evaluation metrics. The latter are often inconsistently applied making it difficult to determine which metric should serve as the primary measure of performance.7,10,20 Existing metrics frequently lack either clinical relevance or the ability to comprehensively assess segmentation performance.21,22 Moreover, prior studies are typically based on relatively small cohorts. For example, Netzer et al, Rodrigues et al, and Provenzano et al used only 640, 733, and 175 exams, respectively.6,7,9 To our knowledge, only 3 related studies have exceeded 1000 exams. Rajagopal et al used 1959 exams, however nearly half were acquired with an endorectal coil, making the domain shift overt and undermining its relevance. 8 Hosseinzadeh et al used 2734 exams 5 but their only domain shift-related analysis involved testing their local model on 296 external exams, which showed no significant drop in performance. Furthermore, their data lacked biopsy confirmation, instead using PIRADS scores as the reference standard and PIRADS 3 lesions were treated as csPCa-negative.
In contrast to prior work, our study explicitly compares models trained on local versus external data across 3208 prostate MRI exams, more than any previous study. This allows the assumption that training with a sufficiently large and diverse external dataset can match or exceed the performance of local training, to be tested at an unprecedented scale and level of data diversity. Despite the growing availability of large, multi-institutional datasets,12,23 our findings emphasize the continued importance of local data. Even when trained on over 1200 external exams from 3 institutions, the PICAI-TRAIN model still underperformed on data from a new independent institution. Training with less than one quarter the number of cases using local data matched the external-data trained model performance. Even when combining all the available data to train on a total of 2641 exams, the additional 1241 external exams included in training (representing a 47% increase in data volume) did not result in any significant performance improvements. This suggests that training or fine-tuning with local data remains essential in real-world deployment, even when leveraging external multi-institutional datasets on the order of 1000 exams.
It is noteworthy that when studying the PICAI-TEST dataset the LOCAL-TRAIN model was only 1.6% worse than the PICAI-TRAIN model while the spread for the LOCAL-TEST dataset was 8.1% (Table 3) suggesting our locally trained model was more generalizable. This requires further investigation. We did not insist on pathology confirmation for every case in our local training (just testing) while all PICAI cases were pathology confirmed. We can speculate that perhaps this improves training diversity and generalizability by including more cases that did not go to biopsy.
While some hypothesize that domain shift can be mitigated with sufficiently large and diverse datasets6 -9 our findings suggest that 1200 exams across 3 sites are insufficient. The true threshold may be much higher, and maintaining such a model over time would be costly given evolving local practices. Our study shows that even small amounts of local data significantly improve local performance. This suggests frameworks of deployment should support ongoing data sharing or fine-tuning to allow clients to update the central model. How to do this is an important direction for future research in domains such as federated learning, quality assurance, and models that can correct for domain shift with minimal local data.
Limitations of our study include the absence of pathological confirmation for all LOCAL-TRAIN cases, whereas PICAI-TRAIN was completely biopsy-confirmed. However, this would bias against local training, the opposite of what we observed. Additionally, all local test cases were biopsy confirmed. Another limitation is that the publicly available PICAI dataset is only a subset of the full PICAI data, which remains private. Notably, we have not studied the performance of much larger external data sets trained over larger numbers of institutions and encompassing all major MRI manufacturers and field strengths as, to our knowledge, such a dataset is not publicly-available. It remains possible that an order of magnitude larger centralized data may obviate the need for local training.
In conclusion, domain shift limits MRI prostate cancer segmentation performance even when training with over 1000 exams from 3 external institutions. Use of local data is paramount at these scales.
Supplemental Material
sj-docx-1-caj-10.1177_08465371251367620 – Supplemental material for Training With Local Data Remains Important for Deep Learning MRI Prostate Cancer Detection
Supplemental material, sj-docx-1-caj-10.1177_08465371251367620 for Training With Local Data Remains Important for Deep Learning MRI Prostate Cancer Detection by Shawn G. Carere, John Jewell, Paola V. Nasute Fauerbach, David B. Emerson, Antonio Finelli, Sangeet Ghai and Masoom A. Haider in Canadian Association of Radiologists Journal
Footnotes
Abbreviations
ADC apparent diffusion coefficient
AP average precision
AUROC area under the receiver operating characteristic
BHIGH high b-value
bpMRI biparametric MRI
csPCa clinically significant prostate cancer
GG International Society of Urological Pathology grade group
PIRADS Prostate Imaging Reporting and Data System
PSA prostate specific antigen
T2W T2 weighted image
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially funded by the Sinai Health Foundation, Toronto, Canada.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
