Abstract
Introduction
A fundamental challenge facing machine learning (ML) model development is the acquisition and processing of large, high-quality datasets of representative real-world data necessary for training. 1 In medical imaging projects, much of this labor-intensive work has been done manually by radiologists or trained researchers and is often the rate limiting step in the development pipeline. 2 In order to expand the scale, streamline dataset curation, and accelerate model development, automating aspects of dataset curation are becoming increasingly important.
Medical imaging studies that meet inclusion criteria are downloaded from a Picture Archiving and Communication Systems (PACS) in Digital Imaging and Communications in Medicine (DICOM) format for subsequent processing. Each study consists of one or more series of images, which can differ in the anatomical region imaged, imaging axis, convolutional kernel, slice thickness, and presence and timing of intravenous (IV) contrast. Developers have an explicit definition of the imaging that needs to be included in terms of these basic parameters based on the target input of the final model. In principle, data extraction should be a straightforward process of using DICOM metadata to select desired imaging data but in practice this is very often a time consuming, frustrating, and highly manual selection process due to the heterogeneity and inconsistency of metadata. 3 This process is only worsened when the imaging data is derived from equipment of multiple vendors or different institutions.
A system that operates independently of DICOM metadata and accurately categorizes imaging along key parameters would be valuable in both model development and deployment. To our knowledge, there has been limited prior investigation of ML mediated CT series classification models. Several ML models for classifying body parts and contrast enhancement have been developed but these handled a limited number of body parts and phases of post-contrast imaging.4-8
The aim of this study was the development and evaluation of ML models that automatically classify CT series by the body part(s) imaged, axis of imaging, and IV contrast enhancement.
Methods
This retrospective study was approved by the local Hospital Research Ethics Review Board with a waiver for informed consent.
Dataset Curation
The radiology information system (syngo, Siemens Medical Solutions) was searched using Nuance mPower (Nuance Communications) to determine the names and frequency of CT protocols performed between January 1, 2010 and September 30, 2021. These protocols are indication-based and define the anatomical region to be imaged and specific technical parameters used for image acquisition. The most common and representative protocols for neuro, thoracic, and abdominopelvic imaging were identified. These 83 protocols accounted for 79.7% of all CT imaging performed during this period. A random sample of 15 studies from each of these protocols was performed. Images in DICOM format were downloaded from Philips Vue PACS (Philips) using RSNA Anonymizer and underwent de-identification using SapienSecure (Sapien). Scout images, scanned documents, dose reports, and post-processed images were removed. A custom Python (Python Software Foundation) script further limited DICOM metadata fields to a “white-list”. Manual review of images ensured no private health information remained. The CT studies were performed on a multi-detector CT scanner (Revolution, LightSpeed 64, or Optima 64; General Electric Medical Systems).
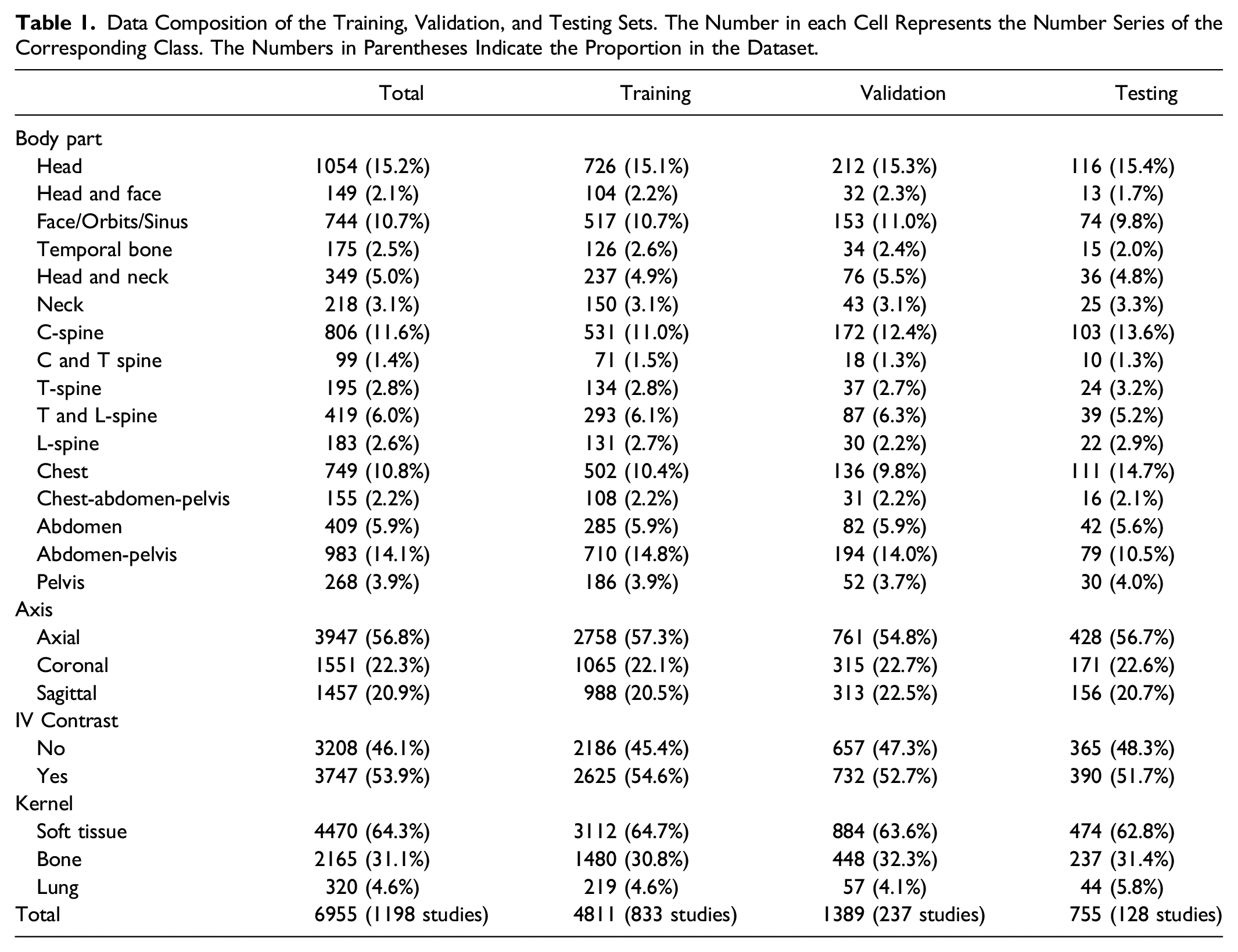
Data Composition of the Training, Validation, and Testing Sets. The Number in each Cell Represents the Number Series of the Corresponding Class. The Numbers in Parentheses Indicate the Proportion in the Dataset.
Data Preprocessing
Hounsfield unit (HU) values were extracted from each DICOM file and transformed from 512 × 512 into 224 × 224 8-bit grayscale images. Hounsfield unit values ranging between −1000 and 400 were converted to a [0, 255] scale while values below −1000 HU were assigned 0 and those above 400 HU were assigned 255. Image pixel values were normalized to the range of [0, 1]. For each series, images were stacked to construct a 3D matrix with dimension N x 224 × 224, where N is the total number of images in the series. This matrix was resized to 128 × 224 × 224 by down-sampling and up-sampling operations to match the model’s input.
Model Development
The 3D residual neural network (3D ResNet)
9
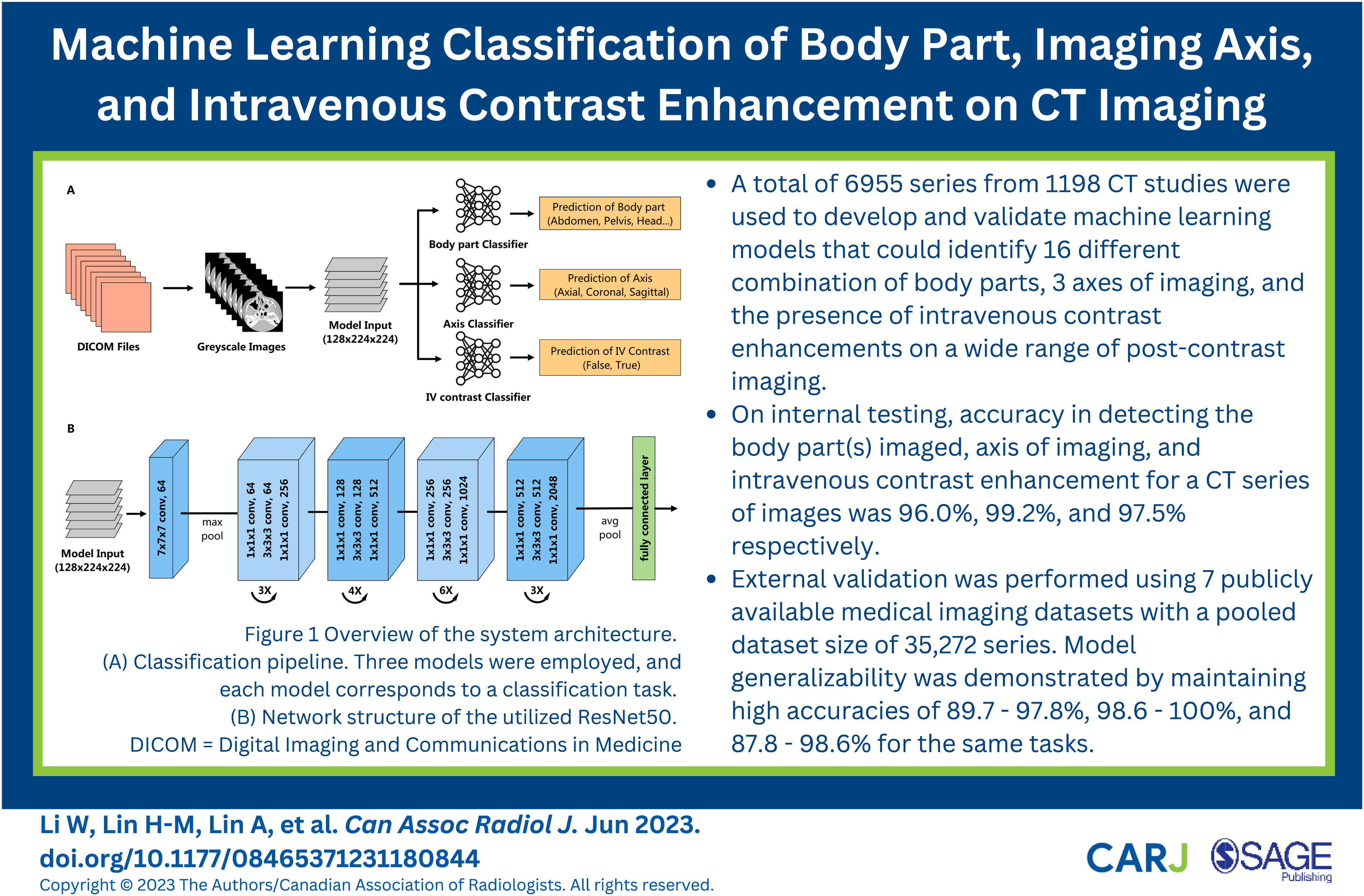
was utilized to perform the CT series-level classification tasks in our study. The 3D ResNet is composed of 3D convolution blocks and pooling layers. Shortcut connections were included to link early to late layers and skip layers in between, making the networks easier to train, especially for very deep neural networks. The 3D ResNet was adjusted to accept inputs of size 128 × 224 × 224 with 50 layers. The model output was controlled by a fully connected layer that transformed the extracted features into classification scores, normalized into positive values that sum to 1 using a SoftMax function. The size of the fully connected layer was adjusted in each task to map to the number of classes. Three models were constructed and separately trained, where each model corresponded to a classification task (Figure 1). Model outputs for each CT series were compared to ground truth labels to evaluate comprehensive predictions. Overview of the system architecture. (A) Classification pipeline. Three models were employed, and each model corresponds to a classification task. (B) Network structure of the utilized ResNet50. DICOM = Digital Imaging and Communications in Medicine.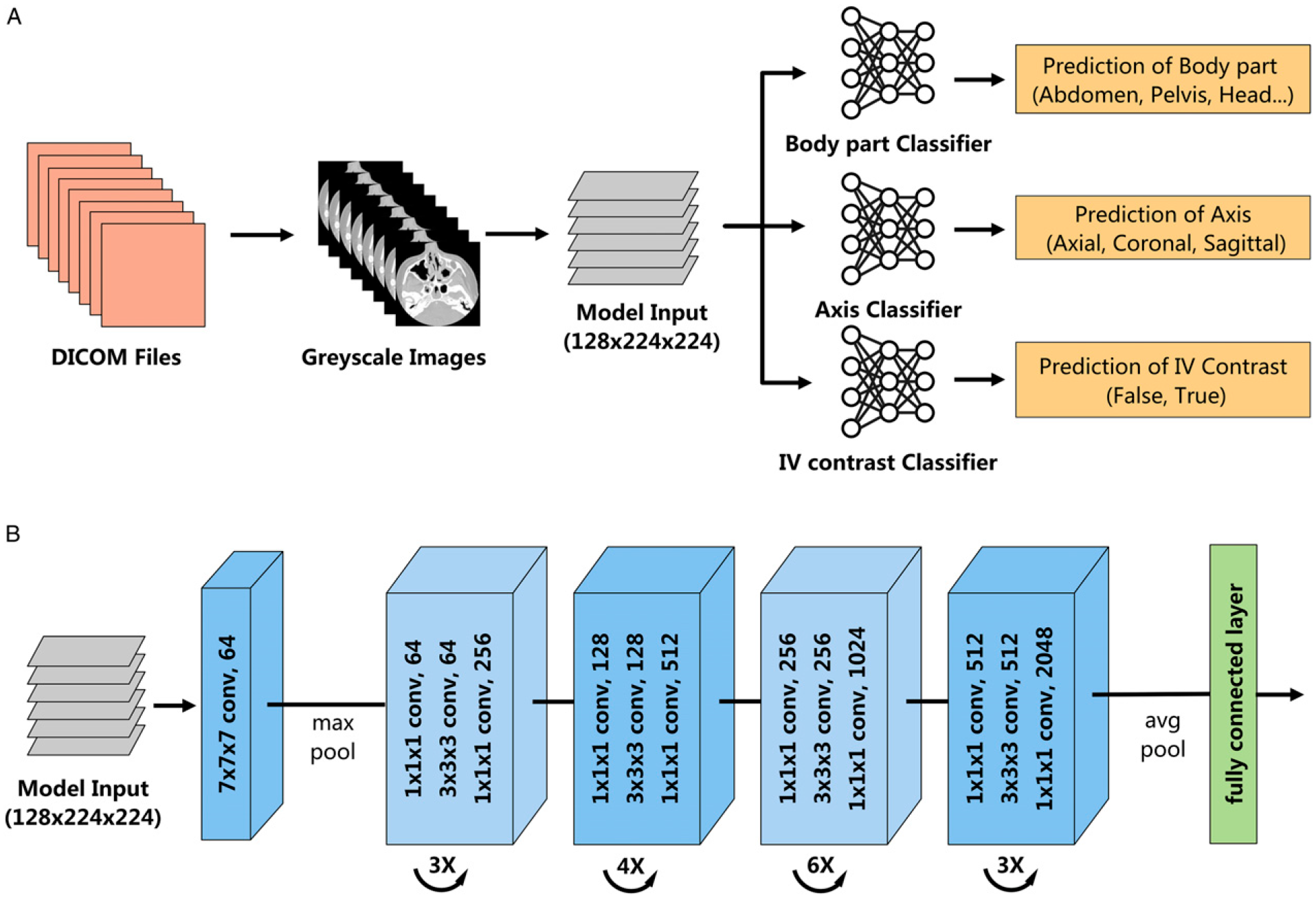
During model development, data was divided at the study-level with a proportion of 70%, 20% and 10%, corresponding to the training, validation, and testing sets, respectively. The models were trained from scratch with a stochastic gradient descent optimizer and initial learning rate of .001. The learning rate was decayed every 7 epochs by a factor of .1. Validation loss (cross-entropy) was evaluated at the end of each epoch, and an early stopping strategy was utilized with the patience of ten epochs. The training and testing were constructed in the PyTorch framework (The Linux Foundation) on an Nvidia Quadro RTX 8000 48G GPU (NVIDIA Corporation).
Evaluation of Model Generalizability
To demonstrate generalizability of model performance, external validation was performed using 7 publicly available medical imaging datasets. These datasets include the Radiological Society of North America (RSNA) Intracranial Hemorrhage (ICH) Detection, 10 RSNA 2022 Cervical Spine Fracture Detection, 11 RSNA Pulmonary Embolism CT, 12 SPIE-AAPM Lung CT Challenge, 13 StageII-Colorectal-CT, 14 CPTAC-LSCC, 15 and C4KC-KiTS 16 datasets. The axis of imaging, body part, kernel, and presence of IV contrast is homogeneous for the RSNA datasets. The non-RSNA datasets were annotated by a board-certified radiologist.
Statistical Analysis
For each input series, the final prediction aligned with the class that had the maximum score. In each task, model performance was evaluated by determining accuracy, top-2 accuracy, micro averaged F1 score and confusion matrices. Furthermore, area under the receiver operating characteristic curve (AUC), accuracy, balanced accuracy, F1 score, positive and negative predictive values, sensitivity, specificity for each class were also calculated. Bootstrapping experiments were conducted with 1000 iterations and 95% CIs were calculated with .025 and .975 percentiles. Analyses were performed with metrics API from Scikit-learn version 1.1.1.
Results
Performance of Body Part Classification
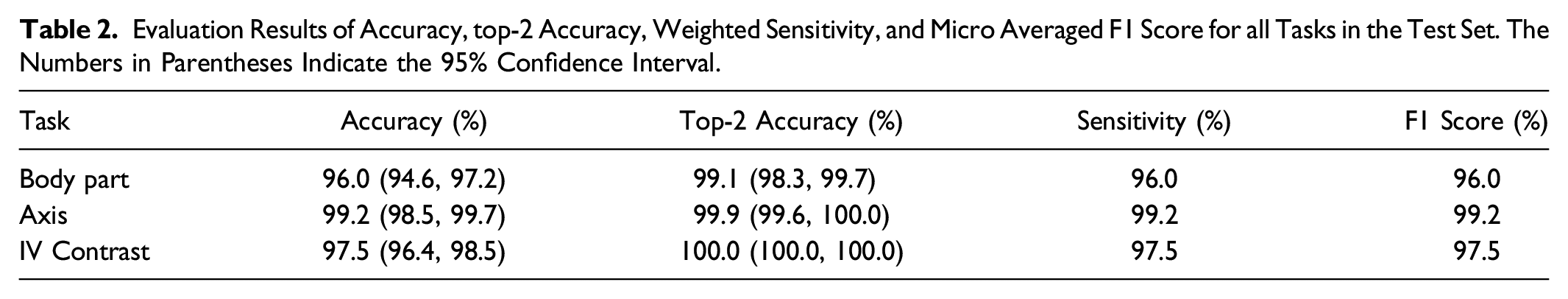
Evaluation Results of Accuracy, top-2 Accuracy, Weighted Sensitivity, and Micro Averaged F1 Score for all Tasks in the Test Set. The Numbers in Parentheses Indicate the 95% Confidence Interval.
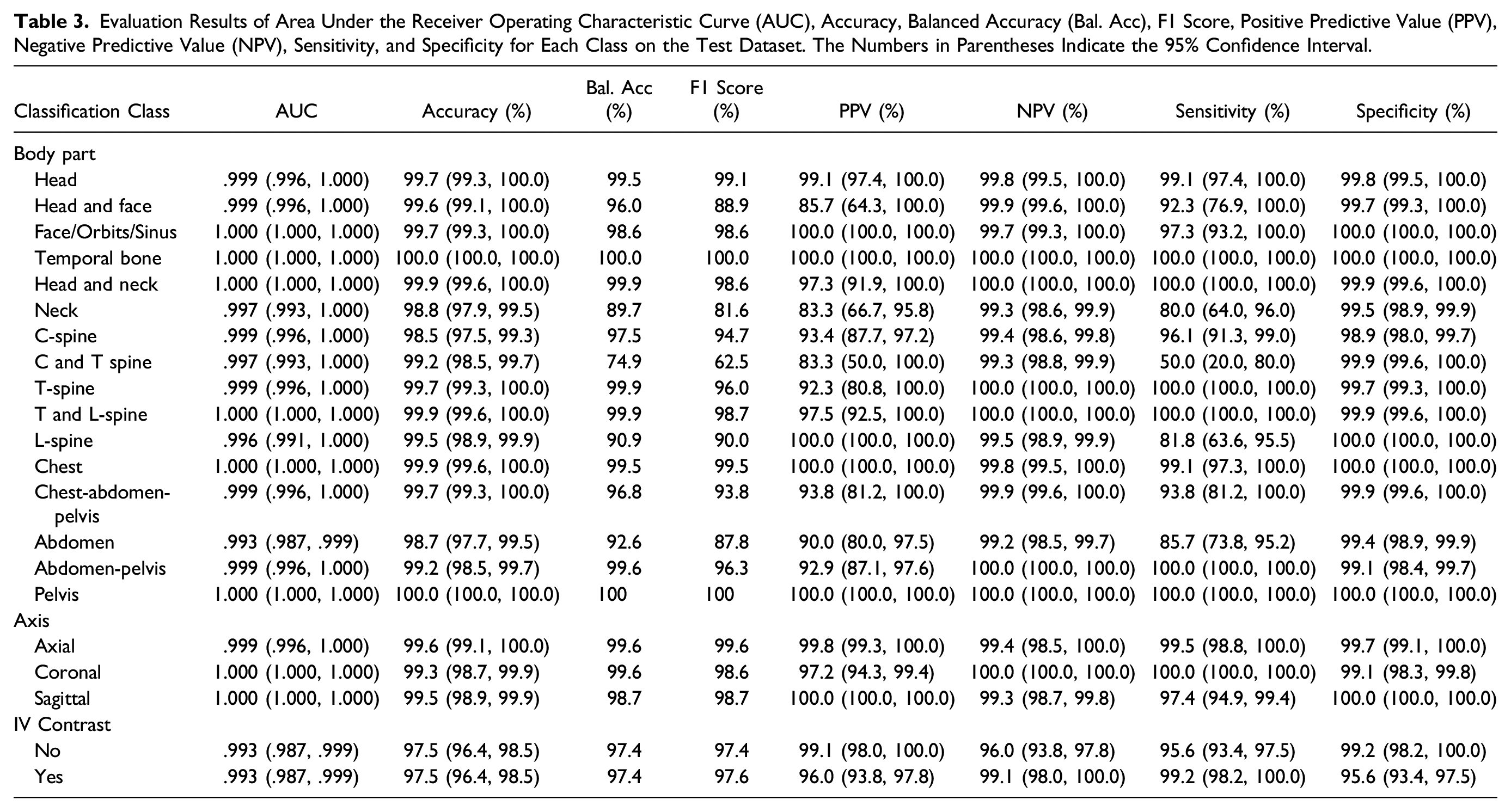
Evaluation Results of Area Under the Receiver Operating Characteristic Curve (AUC), Accuracy, Balanced Accuracy (Bal. Acc), F1 Score, Positive Predictive Value (PPV), Negative Predictive Value (NPV), Sensitivity, and Specificity for Each Class on the Test Dataset. The Numbers in Parentheses Indicate the 95% Confidence Interval.
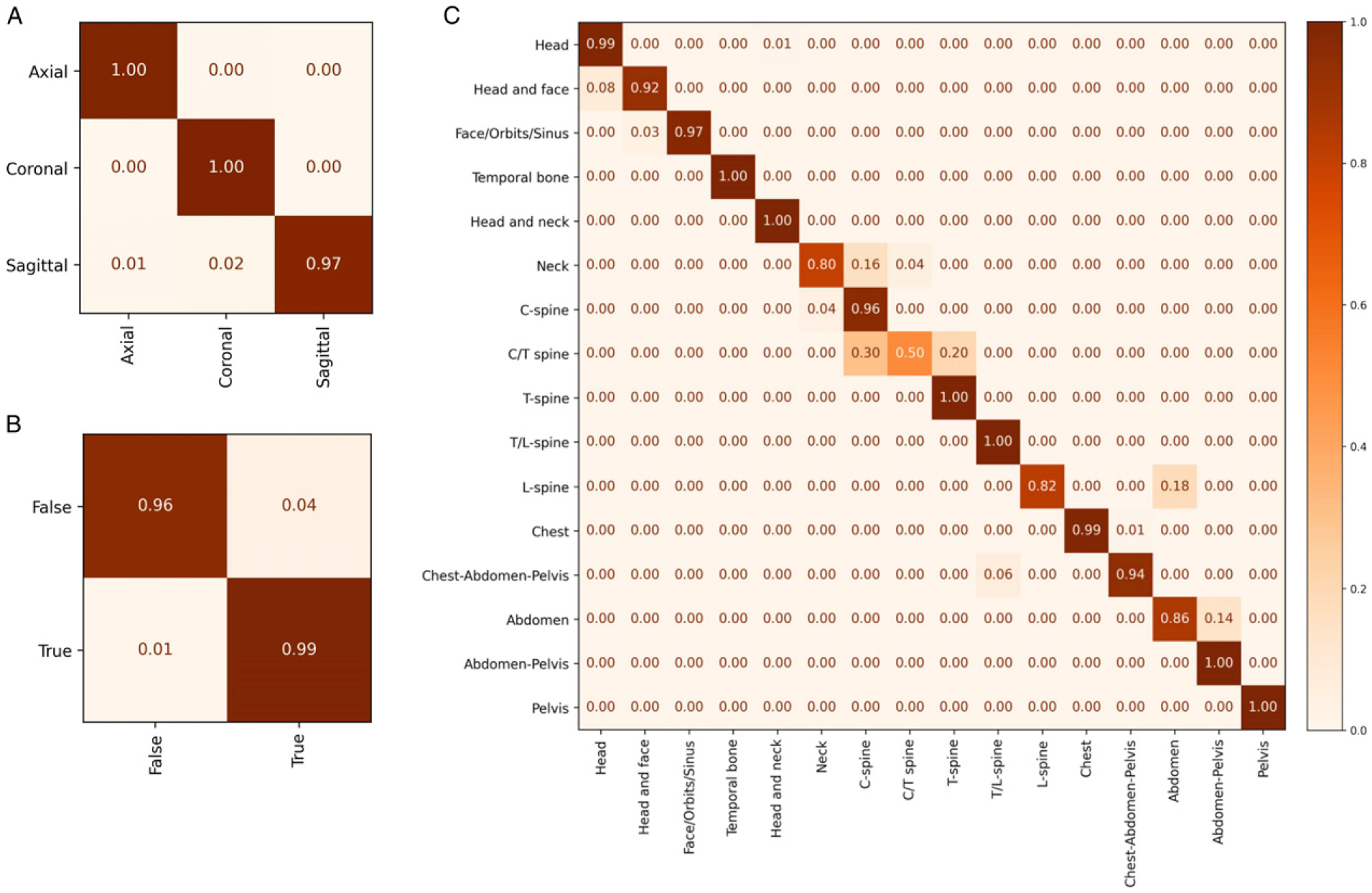
Confusion matrices for all tasks in the test dataset. (A) axis of imaging, (B) presence of intravenous contrast, and (C) body part. The x-axis represents the predicted labels while the y-axis indicates the ground truth.
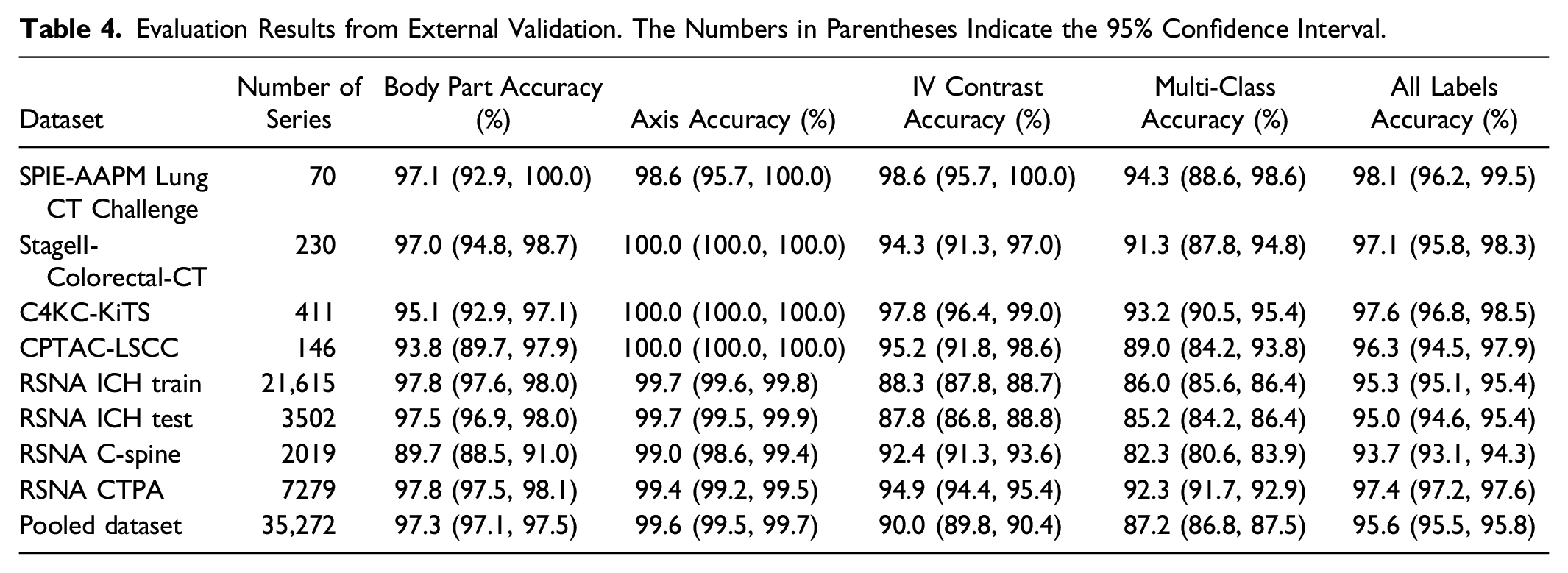
Evaluation Results from External Validation. The Numbers in Parentheses Indicate the 95% Confidence Interval.
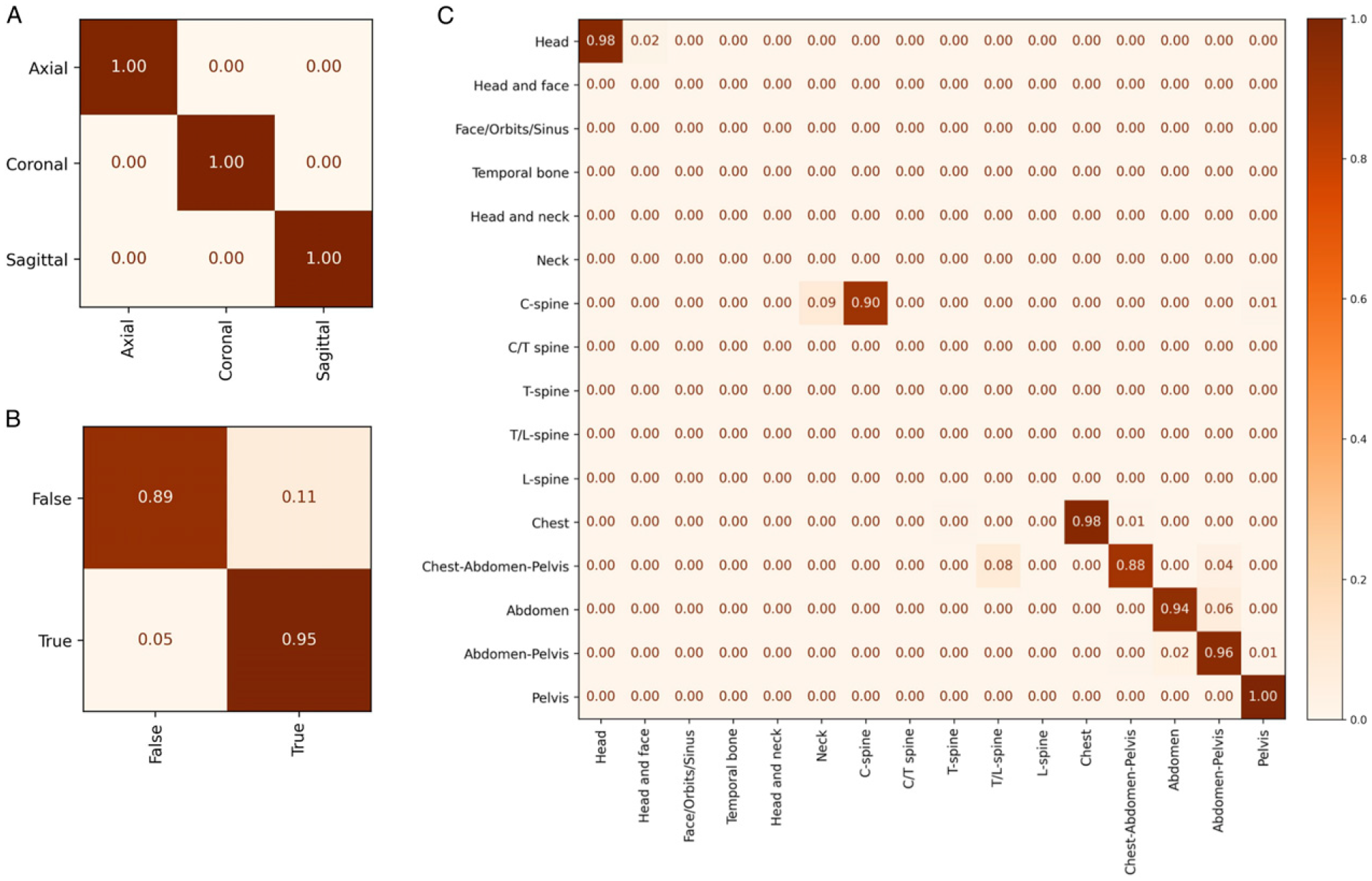
Confusion Matrices from the pooled external validation datasets. (A) axis of imaging, (B) presence of intravenous contrast, and (C) body part. The x-axis represents the predicted labels while the y-axis indicates the ground truth.
Performance of Axis of Imaging Classification
Three standard CT axes of imaging (axial, coronal and sagittal) were used as classification model outputs. Overall accuracy on the test set was 99.2% (95% CI: 98.5%, 99.7%) with a top-2 accuracy of 99.9% (95% CI: 99.6%, 100%) (Table 2). Only the sagittal orientation was ever misclassified while the other orientations were always correctly identified (Figure 2A). Model performance was maintained on external validation with classification accuracy ranging between 98.6% (95% CI: 95.7%, 100%) and 100% (95% CI: 100%, 100%) (Table 4).
Performance of Intravenous Contrast Classification
CT imaging can be performed with or without an IV contrast agent. Classification accuracy for IV contrast was 97.5% (95% CI: 96.4%, 98.5%) (Table 2). On external validation, model accuracy ranges between 87.8% (95% CI: 86.8%, 88.8%) and 98.6% (95% CI: 95.7%, 100.0%) (Table 4). A notable drop in performance was observed when external validation was performed with the RSNA ICH datasets. This is attributable to CT studies with intracranial hemorrhage (ICH) that were misinterpreted as being performed with IV contrast agent. This suggests under-representation in our original training set of cases with ICH which can be a mimic of IV contrast.
Performance of Multi-Class Classification
The results of the 3 prior tasks were compiled to determine simultaneous overall performance of the models on a complete classification task. On the test set, the model successfully classified a CT series in all 3 tasks correctly in 700 out of 755 (92.7%), incorrect on only one of the parameters on 55 series (7.3%), and all series had at least 2 labels correctly classified. There was similar performance on the pooled external validation dataset with accuracy ranging between 93.7% (95% CI: 93.1%, 94.3%) and 98.1% (95% CI: 96.2%, 99.5%) (Table 4).
Discussion
A key challenge in model development and deployment is ensuring that the correct studies are identified for use in training or as input to the final system. The use of DICOM metadata to filter imaging studies for these tasks is often employed, which should work in theory but is frequently problematic in practice due to erroneous or inconsistent data.3,17 Reliance on metadata can result in the need for a resource intensive manual review of the dataset prior to training or model failure in clinical practice when there is a mismatch between expected and actual studies used as model inputs. To help address both problems, we have developed ML models that can classify the body part(s) imaged, imaging axis, and IV contrast enhancement for a series of CT images based on images alone.
A small number of published studies have investigated the use of ML for the classification of body parts imaged by CT.4,5 Na et al 4 reported a model that could classify a CT into 5 anatomical locations with an accuracy of 100% on internal and 99.8% on external validation sets. Recently, Raffy et al 18 proposed a model that could classify 17 body parts with a sensitivity of 92.5% at the image-level and 1.1% improvement at the series-level. Our study differs from previous work by providing more detailed body part classification by including 16 labels, many of which cover multiple anatomical regions. This granular label schema better reflects real-world clinical practice in which CT studies may have overlapping coverage and different scanning parameters such as the field-of-view, radiation dose, slice thickness, and phase of post-contrast imaging. Instead of processing each image sequentially in a CT series, we used 3D models to capture the features of an entire CT series simultaneously for the classification tasks. As a result, our models demonstrated high accuracy in both internal and external testing despite the greater number of body parts and variety of imaging axes, kernels, and phases of post-contrast imaging.
When the model failed at predicting the body part it typically occurred under 2 conditions. The first is when the ground truth and predicted body part anatomy overlapped such as “C spine” and “C and T spine”. This misclassification is likely accounted for by variations in Z-axis scan coverage which is manually defined by a CT technologist at the time of imaging. For example, a technologist may extend the field of view of a cervical spine CT to the mid-thoracic spine, or a substantial portion of the chest may be included in an abdominal CT. The second condition is when the same anatomy was imaged but with different acquisition parameters. For example, a CT of the cervical spine is optimized to assess bony structures and uses different scanning parameters than a CT of the neck which is optimized for soft tissue structures even though they cover mostly the same anatomical area. High performance may be possible if related labels were collapsed but we felt granular labels would better filter data for use by dataset curators.
The model demonstrated high performance in imaging axis classification on both internal and external validation. The model for IV contrast detection showed an AUC of .99 on the internal test set and ranged between .96 and 1.0 on external validation. This performance is slightly higher than a recent study that reported models with AUCs ranging between .95 and 1.0. 5 Moreover, that study included studies from 2 body parts each with a single phase of contrast enhancement, whereas our dataset included 15 phases and 16 body part labels.
External validation was performed on 7 publicly available datasets with representation of multiple body parts, kernels, and non/post-contrast imaging. External datasets ranged in size from 70 to 21,615 series with a pooled dataset size of 35,272 series. Models generalized well with high performance in all 3 tasks. Like the internal validation, incorrect body part predictions typically occurred when the ground truth label and prediction were similar. Examples include “Head” and “Head and face”, “C-spine” and “Neck”, and “Abdomen” and “Abdomen-Pelvis”. The lower accuracy in detecting IV contrast on the RSNA ICH dataset is accounted for by the high prevalence of studies with ICH which can resemble IV contrast within cerebral blood vessels.
The automated classification of CT series has uses beyond dataset curation. Similar problems can exist during clinical deployment in correctly identifying appropriate imaging studies to serve as model input. With deployed models, there is no longer any realistic possibility for manual study assignment and there are significant consequences of failing to apply the model to a valid imaging study or applying it to an invalid one. ML models are inherently fragile when exposed to data dissimilar to their training data, so ensuring this does not occur is a necessary practical step before safe deployment can occur. In addition, the automated classification of CT series could help optimize image display organization in PACS workstations. Hanging protocols based on DICOM metadata are unreliable and radiologists are forced to constantly spend unproductive time organizing their work environments.19,20 ML models hold promise in improving the reliability and robustness of hanging protocols when compared to using DICOM metadata.21,22 In addition, automatic series classification could potentially map similar series and protocols to a standardized schema and help facilitate the curation of multi-institutional datasets. Accurate series classification may also result in more accurate clinical ML outputs by preventing suboptimal or incorrect series from being sent to ML models, thereby potentially improving patient care.
This study has several limitations. The dataset spanned over a 10-year period during which CT technology consistently improved. It is possible that more remote CT scans may be less relevant for model development given the trend towards thinner acquisitions and recent innovations such as dual energy CT. While the data used to train our models was generated from 3 different CT scanners, they were from a single manufacturer. A broader representation of scanners and manufacturers may further improve model generalizability.
Conclusion
ML models can accurately classify a CT series along 3 important parameters (body part, imaging axis, and IV contrast enhancement) commonly used in model development and deployment. The models demonstrated efficacy and generalizability on a large-scale pooled dataset of numerous well-curated external datasets covering all commonly used study types and are ready for deployment in data processing pipelines.
Supplemental Material
Supplemental Material - Machine Learning Classification of Body Part, Imaging Axis, and Intravenous Contrast Enhancement on CT Imaging
Supplemental Material for Machine Learning Classification of Body Part, Imaging Axis, and Intravenous Contrast Enhancement on CT Imaging by Wuqi Li, Hui-Ming Lin, Amy Lin, Marc Napoleone, Robert Moreland, Alexis Murari, Maxim Stepanov, Eric Ivanov, Abhinav Sanjeeva Prasad, George Shih, Zixuan Hu, Suvd Zulbayar, Ervin Sejdić, and Errol Colak in Canadian Association of Radiologists Journal
Footnotes
Acknowledgments
The authors would like to acknowledge the contributions and support of Derek Beaton, Blair Jones, Kate MacGregor, William Parker, and Colin Purcell.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Errol Colak received funding from the Odette Professorship in Artificial Intelligence for Medical Imaging, St. Michael’s Hospital, Unity Health Toronto.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
