Abstract
The circadian clock controls 24-h biological rhythms in our body, influencing many time-related activities such as sleep and wake. The simplest circadian clock is found in cyanobacteria, with the proteins KaiA, KaiB, and KaiC generating a self-sustained circadian oscillation of KaiC phosphorylation and dephosphorylation. KaiA activates KaiC phosphorylation by binding the A-loop of KaiC, while KaiB attenuates the phosphorylation by sequestering KaiA from the A-loop. Structural analysis revealed that magnesium regulates the phosphorylation and dephosphorylation of KaiC by dissociating from and associating with catalytic Glu residues that activate phosphorylation and dephosphorylation, respectively. High magnesium causes KaiC to dephosphorylate, whereas low magnesium causes KaiC to phosphorylate. KaiC alone behaves as an hourglass timekeeper when the magnesium concentration is alternated between low and high levels in vitro. We suggest that a magnesium-based hourglass timekeeping system may have been used by ancient cyanobacteria before magnesium homeostasis was established.
Introduction
Almost all organisms on earth have adapted to environmental cycles by developing their own timekeeping system, a circadian clock, to predict daily changes. The simplest circadian model system is found in cyanobacteria, a single-celled organism. The central oscillator of the cyanobacterial circadian clock is composed of only 3 proteins, KaiA, KaiB, and KaiC (Golden and Canales, 2003). Because oscillations continue without transcriptional-translational feedback loops (Tomita et al., 2005), the cyanobacterial circadian clock can be reconstituted in vitro by mixing those 3 proteins and adenosine 5′-triphosphate (ATP) with magnesium ions (Nakajima et al., 2005). KaiC undergoes rhythmic autophosphorylation and autodephosphorylation with a 24-h period in the presence of both KaiA and KaiB (Iwasaki and Kondo, 2004). These autokinase and autophosphatase activities in KaiC are regulated by the conformation of the A-loop, which is known as a day/night switch located in the C-terminus of KaiC (residues 487-519; Kim et al., 2008). Without KaiA and KaiB, KaiC alone undergoes dephosphorylation in the in vitro reaction because the default conformation of the A-loop is the “buried” state (Kim et al., 2008). The A-loop changes its conformation from the buried to the “exposed” state by binding KaiA (Kim et al., 2008; Vakonakis and LiWang, 2004). When the A-loop is in the exposed conformation, kinase activity is predominant and KaiC is phosphorylated. KaiA keeps the A-loop in the exposed conformation by binding the A-loop with differential affinity, while KaiB returns the A-loop to a buried conformation by sequestering KaiA far from the A-loop (Snijder et al., 2017; Tseng et al., 2017). At the active site of KaiC, S431 and T432 residues are phosphorylated and dephosphorylated sequentially over the daily cycle (Nishiwaki et al., 2007; Rust et al., 2007). Because the active site is distal from the A-loop, the kinase and phosphatase activity of KaiC may be regulated by an allosteric effect that changes the conformation of the active site. Currently, the mechanism acting downstream of the A-loop is unknown. In this study, we found that magnesium is a key element regulating the kinase and phosphatase activity of KaiC as a downstream regulatory element of the A-loop. Based on our findings, we suggest a possible direction of the evolution of the cyanobacterial circadian oscillator in cyanobacteria.
Materials And Methods
Cloning, Protein Purification, and Expression
Cloning and purifications were performed essentially as described previously (Kim et al., 2015; Kim et al., 2012). The KaiC E318D protein expression plasmid was generated using the site-directed mutagenesis method.
Structure Analysis of KaiC Hexamer
Every structure analysis was performed with UCSF Chimera (Pettersen et al., 2004). Distance measurements between OGlu and OThr were performed with the “Structure Analysis” function in UCSF Chimera.
In Vitro Assay of KaiC Phosphorylation
All assays were performed essentially as described previously (Kim et al., 2015) with the following modification: ethylenediaminetetraacetic acid (EDTA) was completely removed for all reactions that were performed without KaiA. The magnesium concentration was modified as indicated in the main text. Disodium ATP (initial concentration: 1 mM), rather than MgATP, was used for all in vitro reactions to avoid the addition of magnesium ions from another source.
Molecular Dynamics Simulation of KaiC Hexamer
All-atom molecular dynamics simulations were performed using GROMACS version 5.1. The Charmm36 force field was used to account for the different interactions between the atoms of the system that comprised the KaiC protein in the hexameric state solvated with 60,000 to 70,000 TIP3P water molecules. For the initial conformation of the KaiC hexamer, we used its experimentally resolved crystal structure (i.e., PDB ID: 1U9I), wherein positions of each atom of residues 14-497 of KaiC were given. We also studied a variant of this experimental structure in which the A-loop (residue 489-497) was deleted from the PDB entry. The latter mimics the exposed conformation of the A-loop, in which KaiC becomes phosphorylated. Moreover, to study the effects of magnesium, 2 sets of simulations were performed for these 2 initial conformations: one with magnesium ions and the other without them. The initial location of the magnesium atoms was indicated in the PDB structure. The net charge of the system was neutralized by adding Na and Cl ions to the solvent.
The 4 systems examined in this study underwent energy minimization as well as equilibration in the NVT ensemble for 10 ns at 300 K, with heavy atoms of the protein restrained to their initial positions via 1000 kcal mol−1Å−2 springs. The leap-frog algorithm with a time step of 2 fs was used to integrate the equations of motion, and the neighbor list was updated every 10 steps. A Lennard-Jones cutoff of 1.0 nm was used. Electrostatics were treated using the smooth particle mesh Ewald method with a grid spacing of 0.13 nm and a 1.0 nm real-space cutoff. The production run was performed in the NPT ensemble (1 atm and 300 K), in which the temperature was controlled using the v-rescale thermostat (τT = 0.1 ps) and the pressure was controlled using the Parrinello-Rahman barostat (τP = 1 ps). The total simulation time of the production runs was 50 ns.
Hourglass Timer In Vitro
The KaiC (3.4 µM, 2 mL)–alone reaction mixture was prepared with the modified in vitro reaction buffer (150 mM NaCl, 20 mM Tris-HCl, 5 mM MgCl2, pH = 8.0). The reaction mixture was incubated at 30 °C for 16 h to completely dephosphorylate KaiC. The incubated reaction mixture was passed through a spin desalting column (Zeba spin desalting column, Fisher Scientific, Waltham, MA) at 100 ×
Mathematical Modeling of Hourglass Timer
Our mathematical model of magnesium-dependent phosphorylation and dephosphorylation cycles of KaiC in the absence of KaiA and KaiB is based on the Rust model of ordered multisite phosphorylation for the KaiABC oscillator (Rust et al., 2007). The model is a 3-dimensional system of ordinary differential equations that captures the temporal dynamics of KaiC in 4 different forms: unphosphorylated (
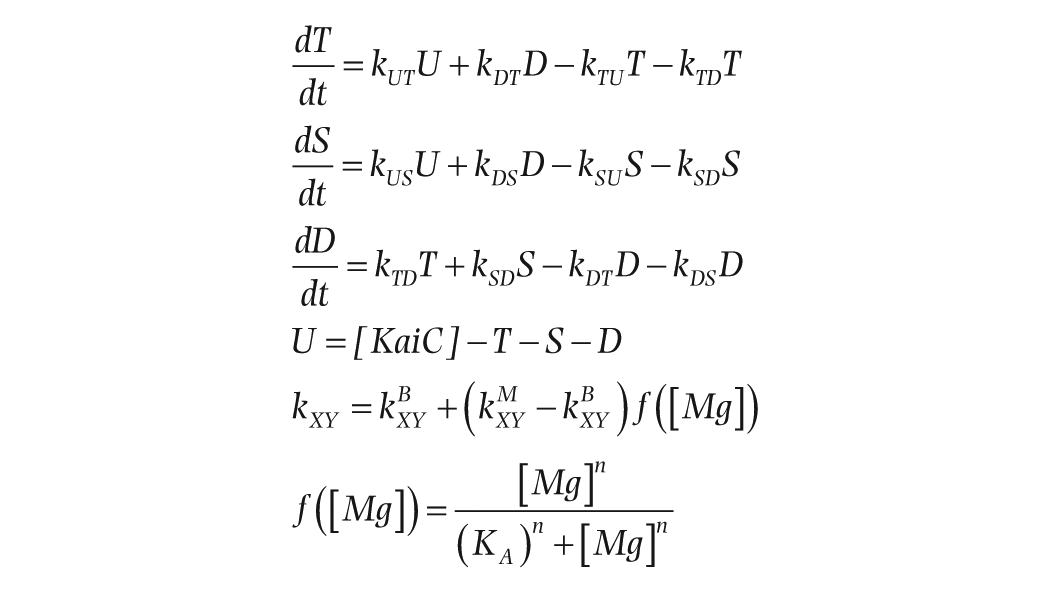
where
To determine the remaining parameters
Adenosine diphosphate (ADP) inhibits the kinase reaction involved in KaiC phosphorylation, and periodic changes in the ATP/ADP ratio can entrain the KaiABC oscillator (Rust et al., 2011). To incorporate the effect of the ATP/ADP ratio into our model, we followed Rust’s model (Rust et al., 2011) and added a multiplicative term to the transition rates for the phosphorylation (but not the dephosphorylation) reactions:
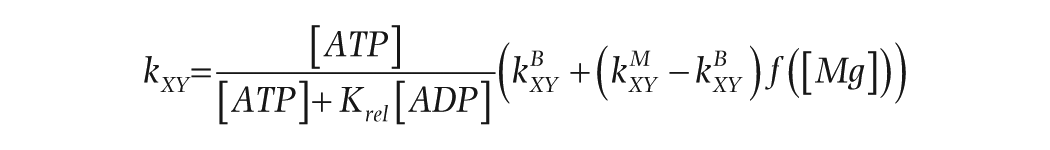
where
Results
Magnesium Ions Are Necessary for the Dephosphorylation of KaiC
To obtain insight into how the allosteric effect of the A-loop induces a conformational change in the active site of KaiC phosphorylation, we analyzed the crystal structure of the KaiC hexamer from a cyanobacterium,
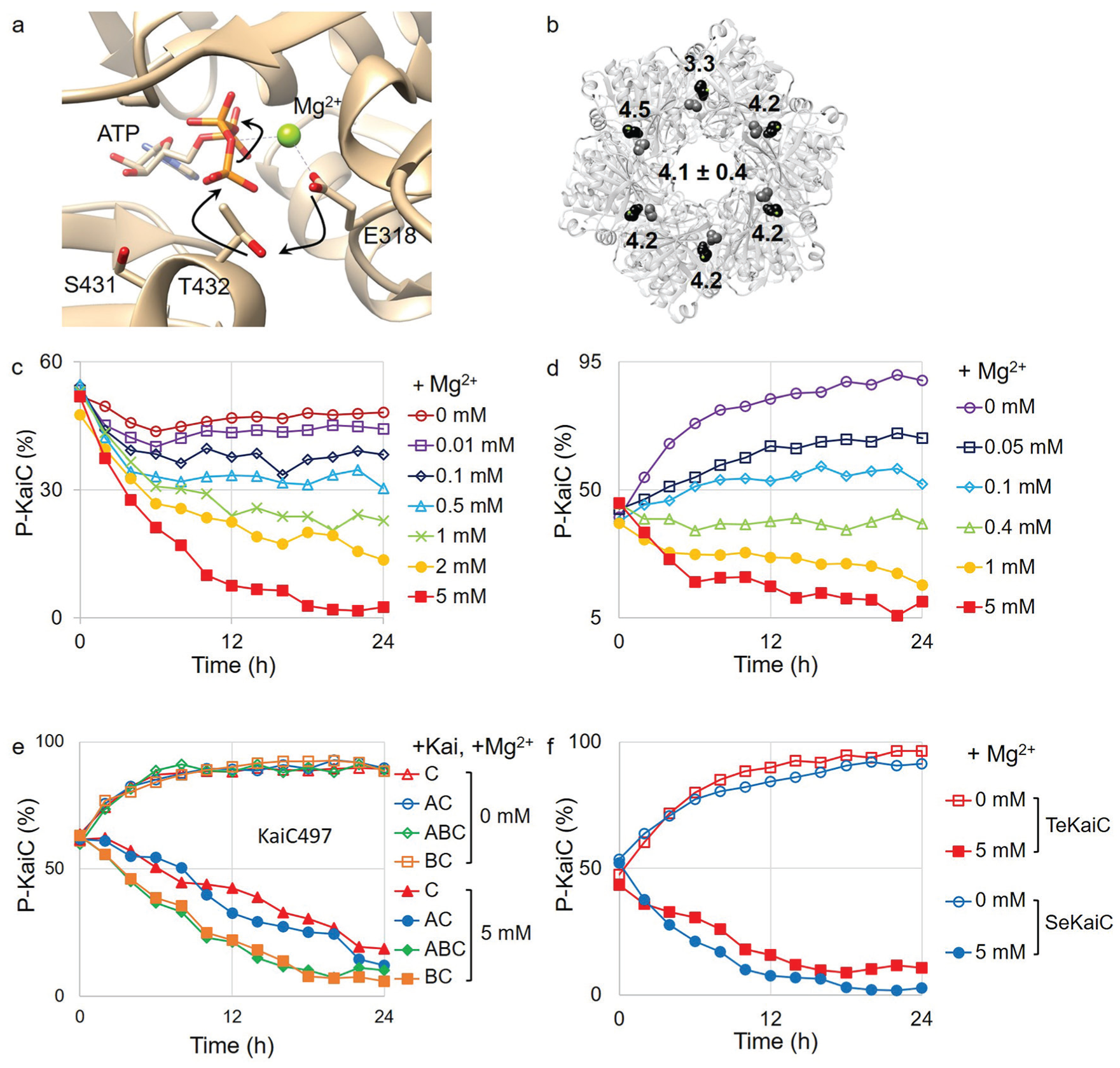
Magnesium regulates the phosphorylation and dephosphorylation of KaiC. (a) Crystal structure of an active site in the KaiC hexamer (PDB ID: 1U9I). The phosphate group on T432 was removed for clearance. Black arrows indicate the electron movements in the phosphoryl transfer reaction. (b) Distances (in angstroms, Å) between the oxygen atom in E318 and the oxygen atom in T432 are labeled on the crystal structure of the KaiC hexamer. Black represents E318, and gray represents T432. (c) Phosphorylation state of KaiC in the in vitro reaction with many different magnesium concentrations. Magnesium concentrations are labeled on the right of the graph. (d) Phosphorylation state of KaiC in the absence of ethylenediaminetetraacetic acid with many different magnesium concentrations. Magnesium concentrations are labeled on the right of the graph. (e, f) Phosphorylation state of the KaiC mutant and other species of cyanobacteria. Open markers indicate the absence of magnesium. Closed markers indicate the presence of magnesium (5 mM). KaiC497 was mixed with KaiC497 alone (▲), KaiA (•), KaiB (■), and KaiA and KaiB (♦) (e). Phosphorylation state of KaiC from
EDTA Inhibits the Phosphorylation of KaiC in the Absence of KaiA and Magnesium
Because KaiC alone did not show phosphorylation in the absence of magnesium, we hypothesized that at least one of the components in the reaction mixture possibly inhibits the kinase activity of KaiC. For the original in vitro oscillator developed by Kondo (Nakajima et al., 2005), EDTA is used as a metal chelator; however, EDTA is not present as a natural component in cyanobacteria. We removed EDTA from the in vitro reaction to determine whether the kinase activity was recovered. In the absence of magnesium and EDTA, KaiC was phosphorylated to the same level as that observed when KaiC was incubated with KaiA in vitro (Fig. 1d; Supplemental Fig. S2b). KaiC was still dephosphorylated with higher magnesium concentrations in the absence of EDTA. The removal of EDTA did not affect the dephosphorylation of KaiC. To confirm the inhibition of kinase activity by EDTA, we added different concentrations of EDTA to the KaiC-alone in vitro reactions in the absence of magnesium. The intensity of the phosphorylation was decreased by increasing the EDTA concentration in the reaction mixture (Supplemental Fig. S3). Therefore, the deficiency of magnesium turns on the kinase activity of KaiC, while EDTA inhibits its kinase activity.
Magnesium Regulation of KaiC Phosphorylation Is Universal
KaiC497 is a KaiC mutant with 23 residues (residues 497-519) deleted from the A-loop, maintaining a dephosphorylation state even in the presence of KaiA due to the lack of the KaiA binding site (Kim et al., 2008). We checked the phosphorylation level of KaiC497 to test the magnesium regulation of KaiC phosphorylation in this mutant. KaiC497 alone spontaneously dephosphorylates with a high magnesium concentration while phosphorylating without magnesium (Fig. 1e; Supplemental Fig. S4). The addition of KaiA or KaiB did not affect KaiC phosphorylation at either concentration. Therefore, the magnesium regulation of KaiC phosphorylation is a downstream regulatory element of the A-loop for the phosphorylation and dephosphorylation of KaiC.
We also tested the magnesium effect with another cyanobacterial species,
Molecular Dynamics Simulation Reveals That E318 Activates T432 for the Phosphorylation of KaiC in the Absence of Magnesium
To provide structural insight into the effect of magnesium on the regulation of KaiC phosphorylation, we performed molecular dynamics simulations of the crystal structure of the KaiC hexamer, which has the A-loop in the buried conformation. The overall backbone conformation of KaiC did not significantly change in the simulations performed with or without magnesium (Fig. 2a; Supplemental Fig. S6). However, the distances between OGlu and OThr were significantly longer than 3 Å when the simulations were performed with magnesium (Fig. 2b; Supplemental Fig. S7a). In the simulations performed without magnesium, 1 of the 6 pairs of OGlu-OThr distances was found to be sufficiently short (less than ~3 Å) to allow OGlu to interact with HThr (Fig. 2c; Supplemental Fig. S7b). This interaction enables T432 to be activated for phosphorylation. Thus, these molecular dynamics simulations suggest that magnesium interacts with OGlu and keeps it away from HThr, which leads to the suppression of KaiC phosphorylation. In contrast, in the absence of magnesium, OGlu can move and/or rotate more freely toward HThr, which enables residue T432 to be activated for phosphorylation.
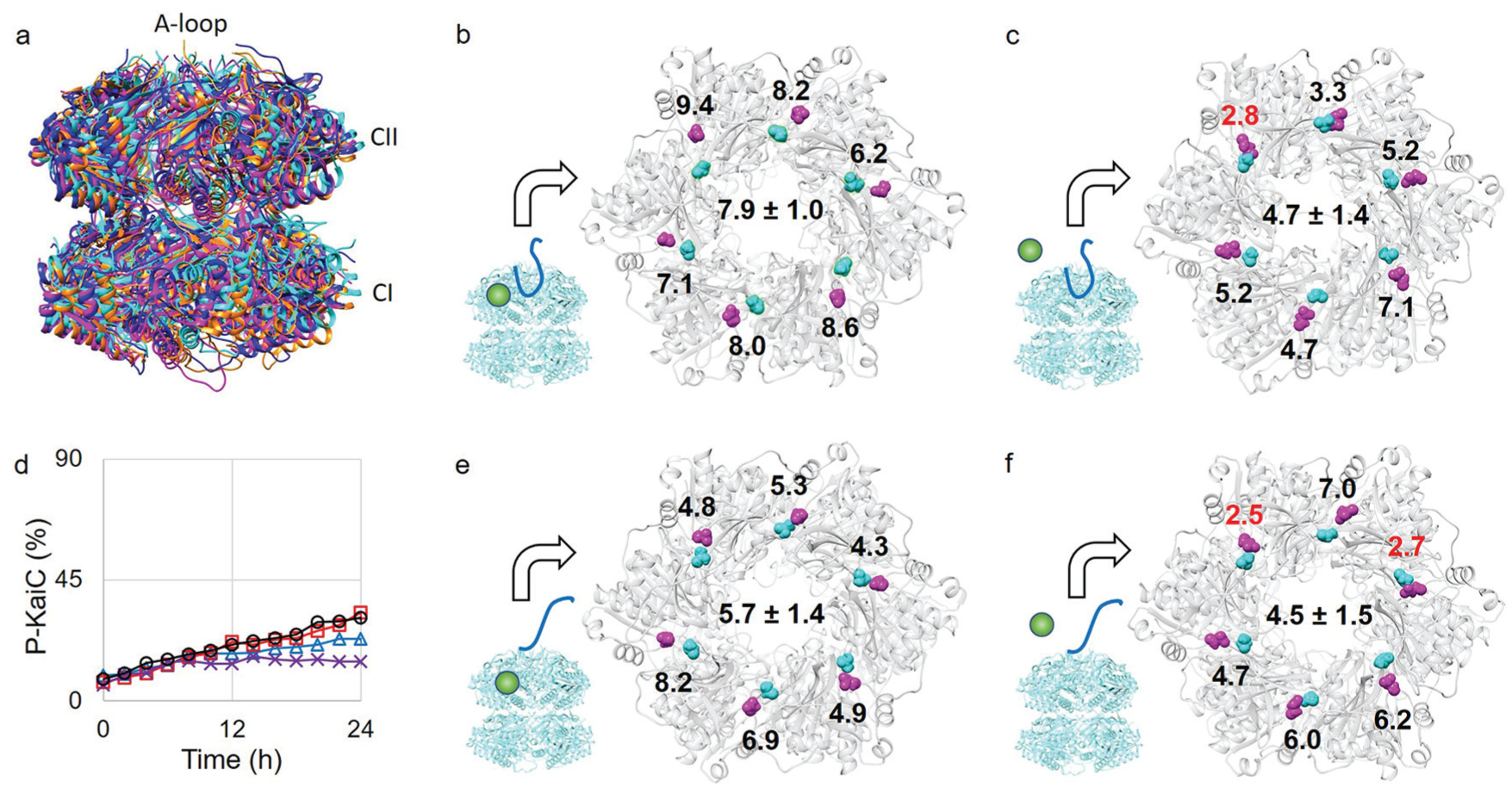
Molecular dynamics (MD) simulations of the KaiC hexamer. (a) Four MD simulation structures are superimposed. The “buried” conformations are shown in magenta (with magnesium) and cyan (without magnesium). The “exposed” conformations are shown in orange (with magnesium) and blue (without magnesium). The 3 domains of KaiC are labeled at the position. (b,c,e,f) MD simulation structures of the KaiC hexamer. Magenta represents E318, and cyan represents T432. The distances (in angstroms, Å) between the oxygen atom in E318 and the oxygen atom in T432 are labeled on the structure. Distances shorter than 3 Å are labeled in red. The average distance of all 6 subunits is labeled at the center of the structure. The combinations of the A-loop conformation and magnesium (see the cartoon representations) were the buried conformation with magnesium (b), the buried conformation without magnesium (c), the exposed conformation with magnesium (e), and the exposed conformation without magnesium (f). (d) Phosphorylation state of KaiC E318D. Four different reaction mixtures were prepared to measure the phosphorylation state: KaiC alone (Δ), KaiC and KaiA (□), KaiC and KaiB (×), KaiC, KaiA, and KaiB (○).
The molecular dynamics simulations reveal that certain OGlu-OThr distances are shorter than 3 Å but longer than 2 Å in the absence of magnesium. We hypothesized that if E318 is mutated to D318, this mutant would not be able to activate T432 for phosphorylation because of the shortness of the side chain. Indeed, KaiC E318D cannot phosphorylate in the absence of magnesium (Fig. 2d; Supplemental Fig. S8). Interestingly, the addition of KaiA cannot activate the kinase activity of KaiC E318D (Fig. 2d; Supplemental Fig. S8). Therefore, the OGlu−OThr distance is an important factor for KaiC phosphorylation and is regulated by magnesium.
Magnesium Is a Downstream Regulator of the A-Loop Conformation in the Circadian Oscillation of KaiC Phosphorylation
We hypothesized that magnesium regulation of KaiC phosphorylation is correlated with the conformation of the A-loop. A molecular dynamics simulation of KaiC was previously reported with an exposed A-loop conformation that is mimicked by removing the entire A-loop (residue 487 to 518) from KaiC (Egli et al., 2013). When the A-loop is in the exposed conformation, the flexibility of the 422-loop in KaiC is increased in the presence of magnesium, but the distance information between OGlu and OThr has not been reported (Egli et al., 2013). To understand the relationship between the A-loop conformation and magnesium regulation, we performed molecular dynamics simulations with an A-loop deletion mutant KaiC in both the presence and absence of magnesium. As in the simulations performed with the A-loop, the overall backbone conformation of KaiC did not significantly change in the absence of the A-loop (Fig. 2a; Supplemental Fig. S6). All OGlu−OThr distances were longer than 3 Å in the presence of magnesium (Fig. 2e; Supplemental Fig. S7c). In contrast, the distances in the 2 subunits were shorter than 3 Å in the absence of magnesium (Fig. 2f; Supplemental Fig. S7d). We suggest that this rare chance for activation helps explain why phosphorylation in the cyanobacterial circadian oscillator is so slow. These results also suggest that the distances between OGlu and OThr were significantly affected not by the conformation of the A-loop but by the presence or absence of magnesium (Fig. 2b, c, e, and f). However, the conformation of the A-loop does enhance the probability of phosphorylation in the absence of magnesium, as we found more subunits that were within the hydrogen bonding distance when the A-loop was in the exposed conformation (Fig. 2f) than in the buried conformation (Fig. 2e). The average OGlu−OThr distance of all subunits was the shortest when the A-loop was exposed in the absence of magnesium (Fig. 2f) and the longest when the A-loop was buried in the presence of magnesium (Fig. 2b). Therefore, the phosphorylation and dephosphorylation of KaiC is regulated by the presence or absence of magnesium, which is an effect downstream of the A-loop conformation.
We tested this finding using the KaiC E487A mutant, which shows constitutive phosphorylation without KaiA (Kim et al., 2008). The A-loop conformation in the mutant KaiC seems to stay exposed by breaking the H-bond network formed between the A-loops in the KaiC hexamer (Kim et al., 2008). The binding affinity of the regulatory magnesium ion should be significantly lower in this mutant, keeping KaiC in the hyperphosphorylated state even in a high magnesium concentration. To test this hypothesis, we constructed the in vitro reaction with KaiC E487A in a high magnesium concentration. Indeed, KaiC E487A was able to stay hyperphosphorylated in 20 mM magnesium (Supplemental Fig. S9).
The possible molecular mechanism of KaiC phosphorylation is proposed in Figure 3. When the A-loop in KaiC is exposed by binding KaiA, magnesium is released from the active site of KaiC, and OGlu moves and rotates toward HThr to activate T432 for the phosphoryl transfer reaction (Fig. 3). While the A-loop is buried after sequestration of KaiA by KaiB, magnesium is tightly bound to OGlu, making it unable to activate T432 for phosphorylation, and KaiC undergoes dephosphorylation (Fig. 3).
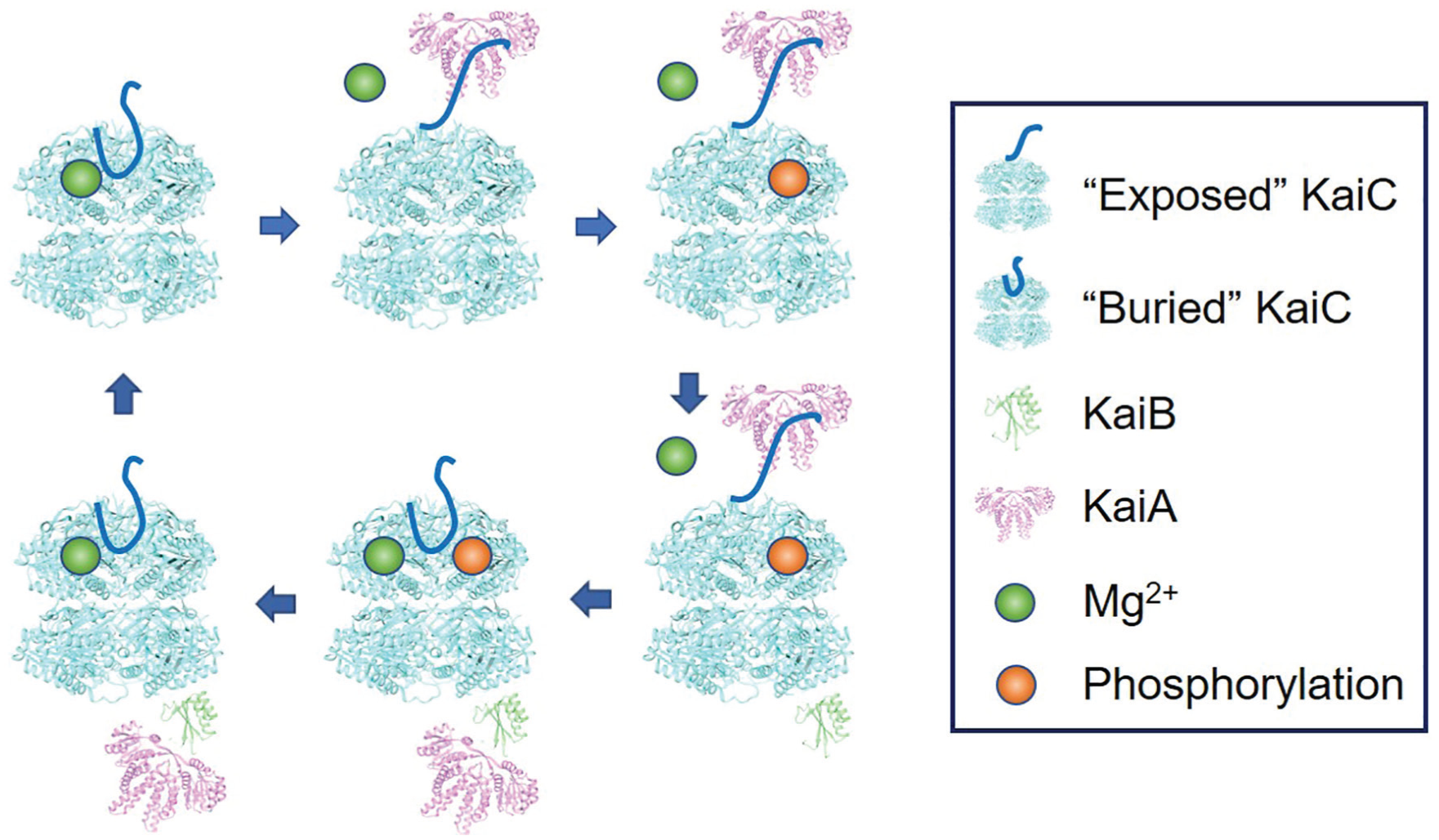
Molecular mechanism of the circadian oscillator in cyanobacteria. KaiC is fully dephosphorylated, the A-loop (solid line) is “buried,” and the magnesium ion is bound on E318 in the active site (upper left). KaiA is bound to the A-loop to keep it in the “exposed” conformation, and magnesium is released from the active site (upper center). KaiC is fully phosphorylated (upper right). KaiB is bound on the CI domain of KaiC to sequester KaiA from the A-loop (lower right). A-loop turns to the buried conformation by dissociation of KaiA, and magnesium is bound on E318 in the active site (lower center). KaiC is fully dephosphorylated (lower left), and KaiB and KaiA are dissociated from KaiC (upper left).
The Circadian Oscillator Has Possibly Evolved from an Hourglass to a Self-sustained Oscillator in Cyanobacteria
This magnesium regulation of KaiC phosphorylation enables the construction of an hourglass-type in vitro reaction with KaiC alone that is suggestive of the evolutionary trajectory of the circadian clock in cyanobacteria. Previous genetic analysis indicated that KaiC is the oldest protein among the oscillator components and that KaiB and KaiA were added later, respectively (Dvornyk et al., 2003; Tauber et al., 2004). Based on the evolutionary information of the oscillator components, a reasonable hypothesis was proposed: the prokaryotic timekeeping system evolved from an hourglass timer to a self-sustained oscillator in response to selective pressures (Johnson et al., 2017; Ma et al., 2016). Before the appearance of KaiB or KaiA, day/night alterations of the environment caused the phosphorylation state of KaiC to oscillate as an hourglass. In other organisms, magnesium concentration is altered by the natural light/dark cycle and provides feedback to the circadian clock (Feeney et al., 2016). Thus, we hypothesized that the KaiC-alone timekeeping system may operate as an hourglass timer driven by the influx and efflux of magnesium, assuming the magnesium concentration was altered by an environmental signal such as the light/dark cycle in the ancient cyanobacteria. If we assume that the environmental magnesium concentration in the habitat of cyanobacteria oscillated daily as a result of evaporation during the day and condensation during the night, then the KaiC-alone hourglass could have been used as a timekeeping system in the ancient cyanobacteria before establishing the current magnesium homeostasis system.
To test the possibility of an hourglass timer in vitro, we constructed an in vitro reaction with KaiC alone. By altering the magnesium concentration every 12 h, KaiC showed a robust phosphorylation and dephosphorylation cycle with a 24-h period, although the variation of magnesium concentrations (0.5-5 mM) was somewhat greater than that observed under the physiological condition (Fig. 4a; Supplemental Fig. S10). We still observed the phosphorylation and dephosphorylation cycle in the low magnesium concentration range (0.05-0.1 mM), although the amplitude was significantly decreased compared with that of the KaiABC oscillator. Simulations of a mathematical model of KaiC alone (see the Materials and Methods section) can reproduce the hourglass timer (Fig. 4b). In cyanobacteria, light alters ADP concentration (Rust et al., 2011; Simons, 2009) in addition to magnesium concentration. Increasing ADP can induce dephosphorylation of KaiC in cyanobacteria and entrain the self-sustained KaiABC circadian oscillator (Rust et al., 2011). This ADP effect may enhance the amplitude of the phosphorylation rhythm in the hourglass timer by inducing robust dephosphorylation at night. When we applied the ADP effect in simulations of the mathematical model of the KaiC-alone hourglass timer, robust oscillations were observed across the entire range of magnesium concentrations (Fig. 4c). The existence of the hourglass timer provides evidence for the hypothesis that the molecular evolution of the cyanobacterial circadian clock started from a KaiC-alone hourglass timer. To further test steps along an evolutionary path to an oscillator, we constructed an in vitro reaction with KaiB and KaiC, which was proposed to be a damped oscillator (Johnson et al., 2017). Many different magnesium concentrations were screened for the signature of a damped oscillator, but all the phosphorylation patterns were similar to the KaiC-alone reaction (Supplemental Fig. S11). The in vitro reaction with KaiB and KaiC also behaved as an hourglass timer (Fig. 4d; Supplemental Fig. S12). It is possible that the KaiBC timekeeping system in each species may be at a different stage of evolution. Thus, whether the mechanism is an hourglass timer, a damped oscillator, or a self-sustained oscillator is species-dependent. In this study, we found an hourglass KaiBC timer using both KaiB and KaiC from
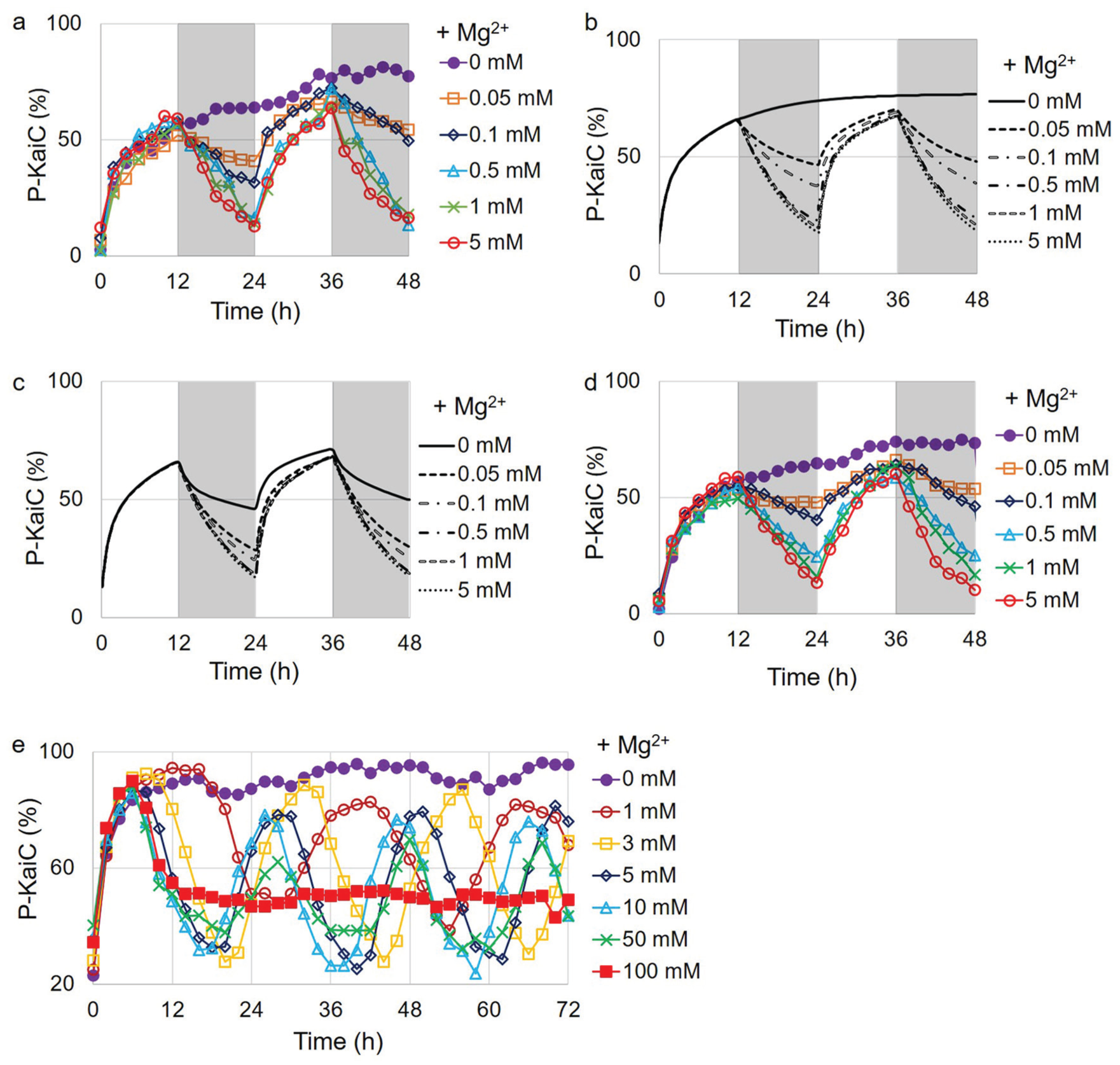
Molecular evolution of the circadian clock in cyanobacteria. (a) Hourglass timer of KaiC alone in vitro. The magnesium concentration was altered every 12 h (low in white and high in gray). The magnesium concentrations in gray are labeled on the right of the graph. (b) Simulations of a mathematical model of the hourglass KaiC timer. All other features are the same as those in (a). (c) Simulations of a mathematical model of the hourglass KaiC timer with ATP/ADP effect. All other features are the same as those in (b). (d) Hourglass-type oscillation of KaiBC in vitro oscillator. All other features are the same as those in (a). (e) Magnesium effect on the KaiABC self-sustained oscillator. Magnesium concentrations are labeled on the right of the graph.
Discussion
We successfully demonstrate the phosphorylation of wild-type KaiC without KaiA. Until now, it was believed that KaiC needed KaiA or a mutation on the A-loop to undergo phosphorylation (Kim et al., 2008; Nishiwaki et al., 2004). Because of this limitation, although the KaiC-alone hourglass was proposed before as a major step in the evolutionary pathway of the circadian oscillator, no experimental proof was reported (Axmann et al., 2009; Ma et al., 2016). We present a possible pathway for the evolution by removing EDTA, an unnatural reagent commonly used in reactions, from the hourglass reaction mixture. The inhibition of the phosphorylation of KaiC by EDTA was not observed in the presence of KaiA. Currently, the detailed mechanism of the inhibition of kinase activity by EDTA is unknown. We thought that the examination of the A-loop conformation could serve as a basis for elucidating the effect of EDTA on KaiC phosphorylation.
The general consensus on the mechanism of the phosphoryl-transfer reaction is that the magnesium ion acts as a cofactor to neutralize the negative charges on the γ-phosphate of ATP to facilitate the nucleophilic phosphoryl-transfer reaction. Our findings on the activation of KaiC kinase activity run counter to this general behavior. However, there are other examples of bacterial autokinases that respond in a similarly unusual way to changes in magnesium concentration. In PhoQ, which is involved in magnesium homeostasis in bacteria, the autokinase and the autophosphatase activities are predominant in low and high magnesium concentrations, respectively (Montagne et al., 2001). Although the detailed reaction mechanism at the atomic level is not yet elucidated, the magnesium ion regulates the autokinase activity of PhoQ in the same way as KaiC. In either case, the addition of EDTA inhibits the autophosphorylation in low magnesium concentration (Castelli et al., 2000). Therefore, we cannot exclude the possibility that a trace amount of magnesium ions from the impurities of the reagents may act as a cofactor for the phosphoryl-transfer reaction. EDTA may strip magnesium ions from the active sites, resulting in the inhibition of kinase activity. Indeed, 2 magnesium ions are found within some KaiC crystal structures. One is located at the same position that we are claiming here as a “regulatory magnesium,” and the other is located near the γ-phosphate of ATP (Supplemental Fig. S14). However, even if the kinase activity is inhibited in absolute zero magnesium concentration, our findings here are still valid because absolute zero magnesium is impossible in cyanobacteria.
The magnesium concentration in cyanobacteria may be affected by fluctuating environmental conditions such as evaporation and condensation, because cyanobacteria can live even in the small amount of moisture found in a desert. Although magnesium concentration did not have a large effect on the self-sustained period, it is still possible that day/night magnesium alternations play a role in entraining the KaiABC oscillator because small changes in the self-sustained period can have a large effect on the entrained phase (Granada et al., 2013). We also hypothesize that signaling pathways other than magnesium, such as quinone signaling through KaiA (Kim et al., 2012), evolved to enable effective entrainment of the self-sustained oscillator. For example, CikA, a major input pathway protein (Kaur et al., 2019; Schmitz et al., 2000), directly interacts with the KaiABC oscillator to entrain circadian rhythms in cyanobacteria (Tseng et al., 2017). The step-by-step evolution from an hourglass timer permitted the development of a robust and entrainable circadian clock system in this single-celled organism.
Conclusion
Until recently, adding KaiA was the only way to activate the kinase activity of wild-type KaiC. We found that magnesium regulates KaiC phosphorylation downstream of the A-loop conformation, which is the master regulator of KaiC phosphorylation. By modulating magnesium concentration, the phosphorylation state of KaiC can be regulated without KaiA and KaiB. In ancient cyanobacteria, the KaiC-alone hourglass timekeeping system, which evolved before the current homeostasis system was established, may be used for the regulation of gene expression in the fluctuating magnesium environment. The current self-sustained circadian oscillator possibly evolved from the KaiC-alone hourglass controlled by magnesium.
Supplemental Material
JBR_20190412_SI_1 – Supplemental material for Magnesium Regulates the Circadian Oscillator in Cyanobacteria
Supplemental material, JBR_20190412_SI_1 for Magnesium Regulates the Circadian Oscillator in Cyanobacteria by Young M. Jeong, Cristiano Dias, Casey Diekman, Helene Brochon, Pyonghwa Kim, Manpreet Kaur, Yong-Sung Kim, Hye-In Jang and Yong-Ick Kim in Journal of Biological Rhythms
Footnotes
Acknowledgements
We thank Hana Nim for insightful discussions, Dr. Carl Johnson and Daniel Kim for critical comments, and Alejandra Lopez-Diaz, Brianna Porr, and Neha Thati for technical assistance. This work was supported by a New Jersey Institute of Technology start-up grant and a faculty seed grant awarded to Y. I. Kim and the National Science Foundation grants DMS-1412877 and DMS-1555237 and the U.S. Army Research Office grant W911NF-16-1-0584 awarded to C. Diekman.
Conflict Of Interest Statement
The authors have no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Notes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
