Abstract
Clock gene expression is not only confined to the master circadian clock in the suprachiasmatic nucleus (SCN) but is also found in many other brain regions. The phase relationship between SCN and extra-SCN oscillators may contribute to known differences in chronotypes. The Octodon degus is a diurnal rodent that can shift its activity-phase preference from diurnal to nocturnal when running wheels become available. To understand better the relationship between brain clock gene activity and chronotype, we studied the day-night expression of the Period genes, Per1 and Per2, in the SCN and extra-SCN brain areas in diurnal and nocturnal degus. Since negative masking to light and entrainment to the dark phase are involved in the nocturnalism of this species, we also compare, for the first time, Per expression between entrained (EN) and masked nocturnal (MN) degus. The brains of diurnal, MN, and EN degus housed with wheels were collected during the light (ZT4) and dark (ZT16) phases. Per1 and Per2 mRNA levels were analyzed by in situ hybridization. Within the SCN, signals for Per1 and Per2 were higher at ZT4 irrespective of chronotype. However, outside of the SCN, Per1 expression in the hippocampus of EN degus was out of phase (higher values at ZT16) with SCN values. Although a similar trend was seen in MN animals, this day-night difference in Per1 expression was not significant. Interestingly, daily differences in Per1 expression were not seen in the hippocampus of diurnal degus. For other putative brain areas analyzed (cortices, striatum, arcuate, ventromedial hypothalamus), no differences in Per1 levels were found between chronotypes. Both in diurnal and nocturnal degus, Per2 levels in the hippocampus and in the cingulate and piriform cortices were in phase with their activity rhythms. Thus, diurnal degus showed higher Per2 levels at ZT4, whereas in both types of nocturnal degus, Per2 expression was reversed, peaking at ZT16. Together, the present study supports the hypothesis that the mechanisms underlying activity-phase preference in diurnal and nocturnal mammals reside downstream from the SCN, but our data also indicate that there are fundamental differences between nocturnal masked and entrained degus.
The circadian system generates near 24-h rhythms in physiology, metabolism, and behavior and allows living organisms to anticipate and prepare in advance for cyclic changes in environmental conditions. The mammalian master circadian clock is located in the suprachiasmatic nucleus (SCN) of the anterior hypothalamus (Moore and Eichler, 1972; Stephan and Zucker, 1972). Here, the coordinated activity of cells containing an internal molecular clock generates and orchestrates endogenous circadian rhythms in the body to ensure an optimal temporal organization in physiology and behavior with respect to time cues in the environment (Liu et al., 2007; Welsh et al., 2010). To maintain a stable phase relationship with the external world, the daily activity of the SCN is reset by recurrent environmental synchronizers (zeitgebers) of which the light/dark cycle is the most important (Lowrey and Takahashi, 2000).
In mammals, the core molecular circadian clock is an autoregulatory oscillator integrating transcriptional/translational feedback loops that drive recurrent rhythms in mRNA and protein levels of key clock components, including the Period genes (Per1/2) and their protein products (PER1/2; Reppert and Weaver, 2001, 2002; Hastings and Herzog, 2004; Mohawk and Takahashi, 2011).
Despite the variety of activity patterns across animal species, some characteristics of SCN function, such as daily neural and metabolic patterns, are fundamentally conserved (Schwartz et al., 1983). Moreover, except for the blind rat (Oster et al., 2002), the phases of core-clock gene expression and their corresponding protein rhythms are similar in diurnal and nocturnal animals studied so far, despite major differences in the temporal patterns of their physiology and behavior (e.g., Vosko et al., 2009; Ramanathan et al., 2010). Together, this suggests that the mechanisms underpinning the different temporal niche preferences in most diurnal or nocturnal species are located downstream from the master pacemaker (but see Oster et al., 2002). Indeed, it is suggested that the circadian signals coming from the SCN are interpreted in an opposite manner in diurnal and nocturnal mammals (Smale et al., 2003, 2008).
Clock gene expression and its protein products are not only confined to the SCN but are also found in many other brain regions (Yamamoto et al., 2001; Shieh, 2003). Daily rhythms in these genes/proteins are reported in extra-SCN brain sites and in most peripheral tissues and are believed to form part of a network of “slave” oscillators that, despite having their own endogenous rhythms, obtain essential time-of-day cues from the SCN (Guilding and Piggins, 2007; Dibner et al., 2010). Collectively, these extra-SCN oscillators are believed to be important for daily tissue-specific timing in physiology and/or behavior (Mendoza and Challet, 2009). Thus, the circadian system is an organized hierarchy of multiple circadian oscillators, with the SCN as the main circadian pacemaker receiving external light/dark information, which allows synchronizing/entraining of other oscillators residing in other regions of the nervous system and peripheral tissues.
The phase relationship between SCN and extra-SCN oscillators may contribute to differences in chronotype (Lambert and Weaver, 2006; Vosko et al., 2009; Ramanathan et al., 2010). Although oscillations in Per1 and Per2 expression and in the protein products (PER1 and PER2) in the SCN do not differ with chronotype, their expression outside the SCN in nocturnal animals is inverted when compared with diurnal animals (Mrosovsky et al., 2001; Vosko et al., 2009; Ramanathan et al., 2010). That is, in extra-SCN brain regions, Per1 and Per2 rhythms in nocturnal rodents peak at night, which is nearly 180° out of phase with the peaks reported in diurnal species (Masubuchi et al., 2000; Abe et al., 2001; Mrosovsky et al., 2001; Wakamatsu et al., 2001; Shieh et al., 2005; Vosko et al., 2009).
The Octodon degus is a diurnal rodent with major episodes of activity at dawn and at dusk (Fulk, 1976). Interestingly, in laboratory conditions, the degus can shift their activity phase preference from diurnal to nocturnal when running wheels become available (Kas and Edgar, 1999). Moreover, nocturnal chronotype can be achieved through 2 different mechanisms: negative masking to light and a true entrainment to the dark phase (Vivanco et al., 2009, 2010b). This species, therefore, offers an excellent opportunity to investigate how shifts in activity pattern affect the internal temporal organization of the circadian system.
Previous multivariate analysis in the degus shows that when diurnal degus become nocturnal, a complete inversion in the temporal order in physiology, hematology, and biochemistry does not occur in parallel to the shift seen in their wheel-running activity (WRA) and body temperature patterns (Otalora et al., 2010). This raises the tantalizing question as to whether the phasing of clock gene expression rhythms in extra-SCN brain regions in degus changes in animals that have transitioned from their diurnal to a nocturnal chronotype. Our objective was to study Per1 and Per2 day-night expression in the SCN and in extra-SCN brain areas in diurnal and nocturnal degus to understand better the relationship between brain clock gene activity and chronotype. Following the diurnal to nocturnal shift, some of the nocturnal degus become entrained to the night phase, while others become nocturnal by negative masking (Vivanco et al., 2009). We also compared, for the first time, Per1 and Per2 expression between nocturnal degus with entrained and masked activity patterns.
Materials and Methods
Animals and Housing
A total of 23 male degus between 23 and 25 mo of age were obtained from a breeding colony at the Animal Service at the University of Alicante (Spain). Animals were individually housed in Plexiglas cages (52 × 15 × 27 cm) equipped with running wheels kept in an isolated room with controlled temperature (28 ± 1 °C), light intensity (300 lux), and photoperiod (12:12 h light-dark [LD] cycle; lights-on at 0800 h). Food (A04 rat-mouse maintenance; Panlab, Barcelona, Spain) and drinking water were available ad libitum.
Another batch of 7 male degus (15-20 mo) was obtained from a breeding colony at the University of Michigan (Ann Arbor, MI). Animals were also individually housed in plastic cages (48 × 26.8 × 20.3 cm) equipped with running wheels (9 cm width × 34.5 cm diameter) maintained under a 12:12 LD cycle (100 lux) and with free access to food (5001 Rodent Diet; PMI Nutrition, Henderson, CO) and water. Room temperature was maintained at 19 ± 1.5 °C.
All experimental procedures were in compliance with the Principles of Animal Care and were approved both by the University Committee for the Care and Use of Animals at the University of Michigan and by the Bioethical Committee of the University of Murcia, Spain.
Chronotype Characterization
WRA was continuously recorded as wheel revolutions/10-min intervals for at least 15 days using a data acquisition system (Electronic Service at the University of Murcia, Spain) at the University of Murcia and using Vitalview and Actiview software (Minimitter Company, Inc., Bend, OR) at the University of Michigan.
WRA actograms were analyzed using El Temps (version 1.228, copyright Diez-Noguera, University of Barcelona). Degus were classified as diurnal or nocturnal based on the percentage of daily activity they exhibited during the photophase. Animals were considered diurnal if their activity during the day was >50%. When the percentage was <30%, animals were included in the nocturnal chronotype group. Considering that some nocturnal degus are entrained to the scotophase while others become nocturnal by negative masking (Vivanco et al., 2009), we assessed negative masking by transferring nocturnal degus from LD conditions to continuous darkness (DD) for 5 days. Nocturnal degus who started to free run in DD from their former LD phase were considered to be “entrained nocturnal,” whereas those who showed an immediate phase advance (by 2 to 6 h) from the first days under DD so that they ran during their subjective day were defined as “masked nocturnal.” When these criteria were applied, 8 degus exhibited a diurnal chronotype, 9 were classified as entrained nocturnal animals, and the 13 remaining animals were classified as masked nocturnal degus.
Tissue Collection/Sampling
Degus from all chronotypes were rapidly anesthetized with isoflurane and were sacrificed by decapitation within 30 min of the zeitgeber times (ZT) ZT4 and ZT16 (ZT0 corresponds to lights-on). Their brains were removed and immediately flash-frozen by immersion for 20 to 30 sec in isopentane cooled with dry ice. Tissue was stored at −80 °C until use. Brain collection at ZT16 (dark phase) was performed under dim red light. Unfortunately, practical considerations (availability of animals) and design requirements (selection of animals for desired groups) severely limited the number of time points that could be studied. Previous work (Yamamoto et al., 2001; Vosko et al., 2009) indicated that the 2 points examined would be the most informative.
In Situ Hybridization
The in situ hybridization methodology was performed using a protocol similar to that used by Vosko et al. (2009) and Koch et al. (2009). During the in situ hybridization procedure, brain tissue from all chronotypes at both time points (ZT4 and ZT16) was processed in parallel and under identical conditions. Vectors (pGEM-T Vector) containing fragments from Degu Per1 (892-bp, GenBank accession no.EU715821) and Per2 (532-bp, GenBank accession no. EU590918), courtesy of Drs. Jeremiah Shepard and Steven McKnight (Southwest Medical School, Dallas, TX), were purified using the QIAGEN Plasmid Midi Kit. Then, the plasmids were linearized with restriction enzymes, and the completeness of the digest was assessed using gel electrophoresis.
Radioactive Riboprobe Generation
To produce a radioactive RNA probe that was complementary (antisense) to the RNA of interest (Per1 and Per2), we prepared a solution containing the linearized plasmid DNA template, RNA polymerase (T7 for Per1 or SP6 for Per2), nucleotides (GTP, ATP), radioactive labeled 35S-UTP and 35S-CTP nucleotides (Perkin Elmer, Boston, MA), and transcription buffer. To initiate transcription, this solution was incubated in a water bath at 37 °C for 2 h, followed by a treatment with DNase to stop the reaction. Finally, the probes were purified using Micro Bio-Spin 6 Chromatography Columns (Bio-Rad Laboratories, Hercules, CA).
Tissue Preparation
Brain tissue was cut into 16-µm coronal sections using a cryostat (Leica CM1850) and collected on 4 series of glass slides. This tissue was then stored at −80 °C until the day of the in situ hybridization protocol. Four to 6 frozen slides (4 sections/slide) per animal were prepared for in situ hybridization for Per1 and Per2 using previously described methods (Vosko et al., 2009). Briefly, slides containing the brain sections were fixed in 4% paraformaldehyde for 1 h and rinsed 3 times in 2× saline sodium citrate (SSC). Then, the sections were incubated in freshly prepared 0.1 M triethanolamine containing 0.25% acetic anhydride for 10 min and afterward washed in distilled water. Finally, slides were dehydrated through a graded ethanol series (50%, 70%, 90%, 95%, and 100%) and left to air dry completely for 1 to 3 h.
Hybridization
Hybridization solution containing the radioactive antisense Per1 and Per2 mRNA probes was prepared in hybridization buffer as previously described by Badiani et al. (1998), yielding a concentration of 2 to 3 million cpm per 100 µL. This solution was applied onto coverslips, which were then placed onto the slides containing the brain sections. The slides were incubated overnight in an oven at 55 °C in boxes containing filter paper soaked in 50 mL of a 50% formamide solution. After hybridization, the slides were immersed in 2× SSC to aid in coverslip removal, washed 3 times in 2× SSC, and then incubated in an RNase A solution (200 µg/mL) at 37 °C for 1 h to destroy unbound probe. The slides were sequentially washed in 2× SSC, 1× SSC, and 0.5× SSC for 1 min in each solution, placed in 0.1× SSC at 65 °C for 1 h to reduce nonspecific binding, and then quickly rinsed in distilled water. Finally, the brain sections were dehydrated in graded alcohols and, after air drying, placed in a cassette with Kodak BioMax film (Kodak, Rochester, NY) for 9 days (Per1) or 14 days (Per2). Our laboratory has previously verified the absence of nonspecific labeling when using this protocol using sense Per1 and Per2 probes (Vosko et al., 2009).
Analysis
Autoradiographic films were digitized with a Microtek ScanMaker 1000XL and ScanWizard Pro software (Microtek, Cerritos, CA). The digitized images were analyzed for the optical density of the hybridization signal using NIH ImageJ software (National Institutes of Health, Bethesda, MD) by an investigator blind to the images’ ZT and chronotype group identity.
The brain areas that we analyzed were identified using previous anatomical verification (SCN: Goel et al., 1999) or using local landmarks (e.g., corpus callosum, optic chiasm, third ventricle) and a rat brain atlas (Paxinos and Watson, 2007). Since in the degus, brain anatomy outside the SCN has not been rigorously mapped, we refer to all other brain sites mentioned here as putative (see also Vosko et al., 2009). Measurements for Per1 signal were performed for the SCN, putative subparaventricular zone (SPVZ), striatum, cingulate, parietal and piriform cortices (in coronal sections anatomically similar to bregma = −0.70 mm in the rat), hippocampal CA1 and CA3, the hilus, and the dorsal blade of the dentate gyrus (DGd), arcuate nucleus (ARC), and ventromedial hypothalamus (VMH; in coronal sections anatomically similar to bregma = −3.20 mm in the rat). Except for SPVZ, ARC, VMH, striatum, and parietal cortex, these brain regions were also analyzed for Per2 signals. In these latter brain sites, the signal for Per2 was low and could not be rigorously measured. The hybridization signal for each brain area was expressed as the intensity of the labeling (optical density of gray-scale units) relative to the background signal measured from the corpus callosum (mean ± 2.0 SD). Measurements were taken from 2 sections per animal for each area sampled, and the individual mean value was calculated. In ImageJ, the area analyzed for each region was defined by boxes/circles as shown in Fig. 1. We chose these extra-SCN areas for analysis because previous reports indicated that they contain rhythmic clock gene expression that might be associated with chronotype (Wakamatsu et al., 2001; Shieh et al., 2005; Guilding and Piggins, 2007; Vosko et al., 2009; Ramanathan et al., 2010; Hagenauer et al., unpublished data). Since the activity patterns of animals from both colonies are the same (Labyak et al., 1997; Hagenauer and Lee, 2008; Vivanco et al., 2009; Otalora et al., 2010) and the variance in the combined groups is small, there was no reason to suspect differences between the 2 colonies in what is being measured. So, data from the 2 colonies were appropriately pooled.
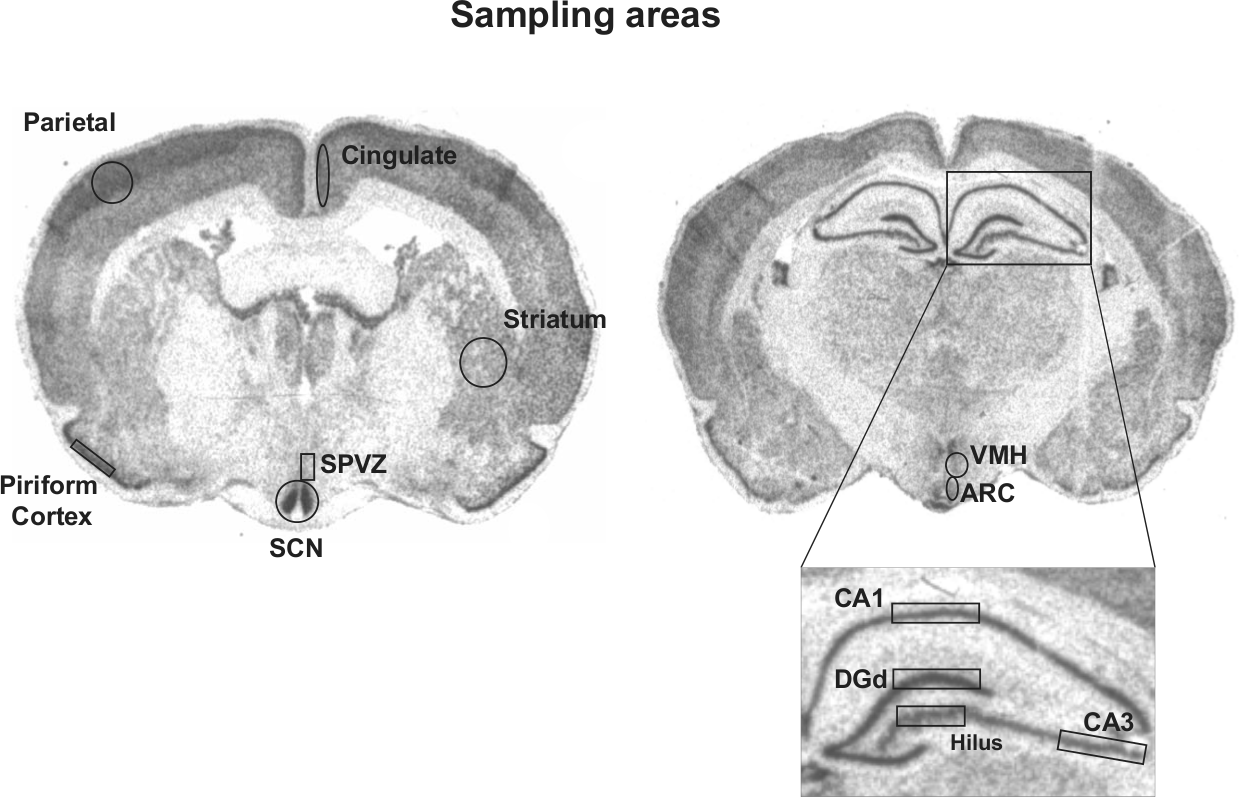
Autoradiographs illustrating the specific regions that were analyzed in each of the brain areas (suprachiasmatic nucleus [SCN], subparaventricular zone [SPVZ], putative striatum, piriform, parietal and cingulate cortices, arcuate nucleus [ARC], the dorsal blade of the dentate gyrus [DGd], hippocampus CA1 and CA3, ventromedial hypothalamus [VMH]). Coronal brain sections corresponding to diurnal degus sacrificed at ZT4 (left panel) and at ZT16 (right panel) and labeled for Per1 mRNA.
Statistics
For each brain region and each probe used, statistical differences between chronotypes (diurnal, masked, and entrained nocturnal) and ZT (ZT4 and ZT16) were assessed by 2-way factorial analysis of variance. Significant interactions were followed by an evaluation of the simple main effects of chronotype within each ZT, then by Sidak post hoc pairwise comparisons when appropriate. Simple main effects of ZT for each chronotype were analyzed by independent Student t tests. All data showed normal distributions and homogeneity of variance. Values of p < 0.05 were considered to be statistically significant. All data in texts and figures are expressed as mean ± SEM. All statistical tests were performed using SPSS 15.0 (Statistical Package for Social Sciences, Chicago, IL).
Results
WRA patterns from diurnal and nocturnal degus are shown in Fig. 2. Although both chronotypes showed 2 crepuscular peaks of activity, the diurnal type exhibited the majority of its WRA in the photophase, whereas the nocturnal degus were mostly active during the scotophase (Fig. 2A, 2B). When nocturnal degus (n = 22) were released into constant darkness (DD), 9 of them (entrained nocturnal) continued to exhibit the majority of their activity during the subjective night. By contrast, the activity pattern of masked nocturnal degus (n = 13) showed an immediate phase advance in the first few days under DD, eventually resulting in the majority of the animals’ WRA to be confined to the subjective day (Fig. 2B).
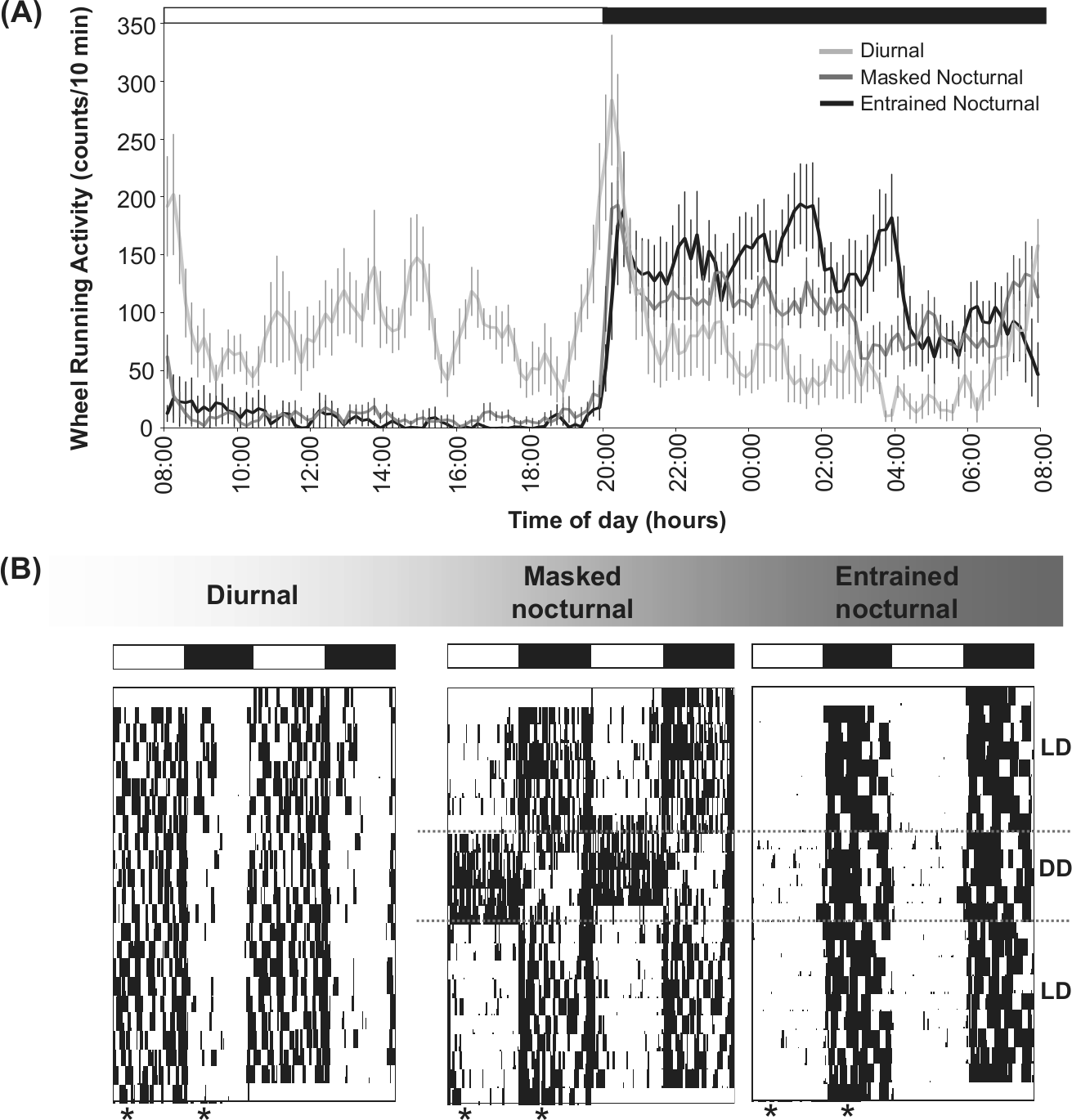
Chronotype characterization. (A) Wheel-running activity waveforms (WRA) for diurnal (n = 8), masked nocturnal (n = 13), and entrained nocturnal degus (n = 9) exposed to a 12:12 light/dark (LD) cycle. Values are expressed as mean ± SEM. (B) Double-plotted WRA actograms for representative diurnal, masked nocturnal, and entrained nocturnal degus exposed to a 12:12 LD cycle. Eventually, nocturnal degus were placed into constant darkness (DD), as indicated on the right side of the actograms. Asterisks indicate the time of group brain collection (zeitgeber times ZT4 and ZT16, with ZT0 correspond to the daily time of lights-on). The LD cycle is represented at the top of each of the graphs by white and dark bars.
In situ hybridization analysis from all animals showed strong day-night differences in Per1 and Per2 expression in the SCN (Fig. 3A). A quantitative analysis of the autoradiographs revealed that the expression of these 2 genes in the SCN was significantly higher at ZT4 than at ZT16, F(1, 24)= 277.31, p < 0.001, for Per1; F(1, 24) = 157.01, p < 0.001 for Per2, with no main effect of chronotype (p = 0.484 for Per1; p = 0.579 for Per2) or significant ZT × chronotype interaction (p = 0.689 for Per1; p = 0.478 for Per2; Fig. 3B). As in the SCN, Per1 expression in the SPVZ also showed a significant change over time with higher levels during the light phase than at night, F(1, 24) = 32.209, p < 0.001, with no main effect of chronotype (p = 0.851) or significant interaction (p = 0.543; Suppl. Fig. S1).
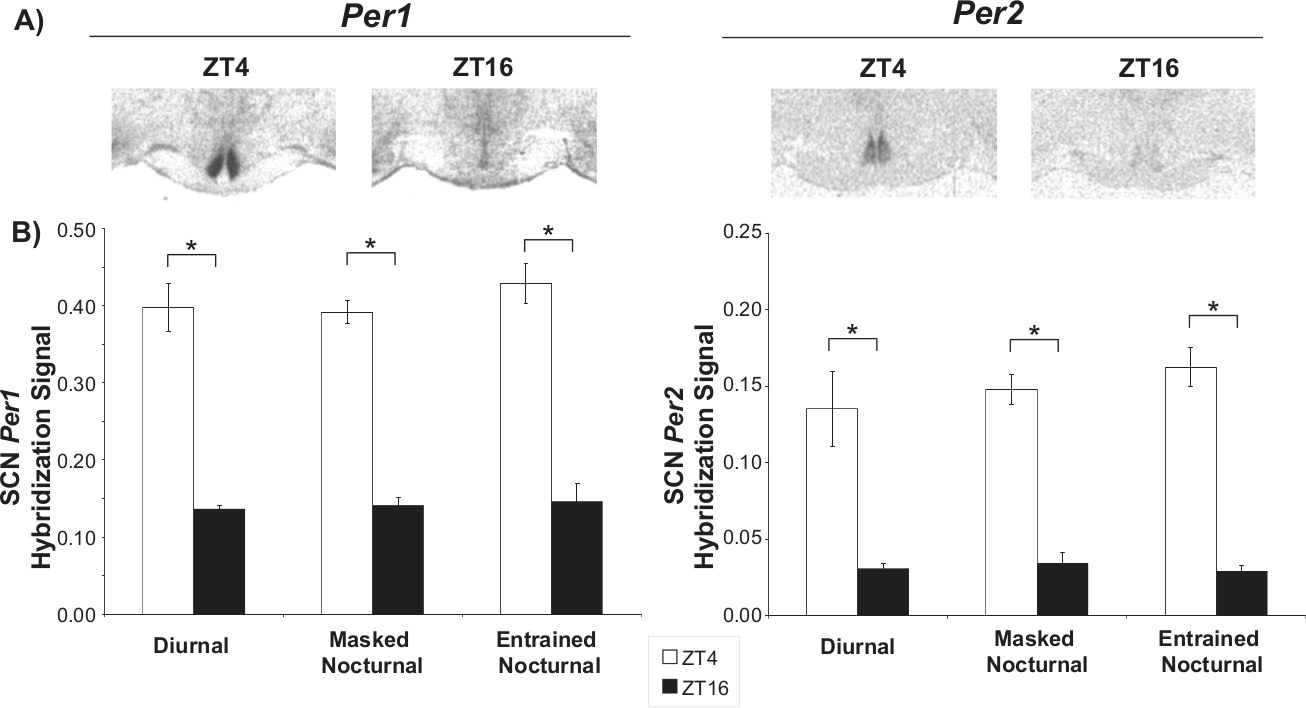
Per1 and Per2 signal in the suprachiasmatic nucleus (SCN) of the degu at ZT4 and ZT16. (A) Representative autoradiographs showing Per1 and Per2 hybridization signal at the 2 time points. (B) Comparison of Per1 and Per2 gene mRNA levels in the SCN of diurnal, masked nocturnal, and entrained nocturnal animals at ZT4 and ZT16 (ZT0 corresponds to lights-on). Values are expressed as mean ± SEM (n = 4-6; *p < 0.05). The hybridization signal for both clock genes was higher at ZT4 irrespective of chronotype.
Strong hybridization signals for Per1 and Per2 were also observed in many brain areas outside the SCN, including the CA1, CA3, hilus, and dentate gyrus regions of the hippocampus and in the cortex (Figs. 1 and 4A). Analysis revealed that Per1 expression showed a significant ZT × chronotype interaction in the CA1, F(2, 23) = 12.705, p < 0.001; CA3, F(2, 23) = 9.165, p < 0.01; and the hilus, F(2, 23) = 6.473, p < 0.01, indicating that in these brain areas, the effect of ZT on Per1 expression was chronotype dependent. Follow-up analyses of the simple main effects of chronotype within each ZT are summarized in Supplementary Table S1. The results showed a significant effect of chronotype at ZT4 in these hippocampal areas, with lower Per1 expression levels in entrained nocturnal animals than in diurnal ones. Per1 expression at this time point in masked nocturnal degus, however, was at an intermediate level when compared with these 2 chronotypes (Fig. 4B).
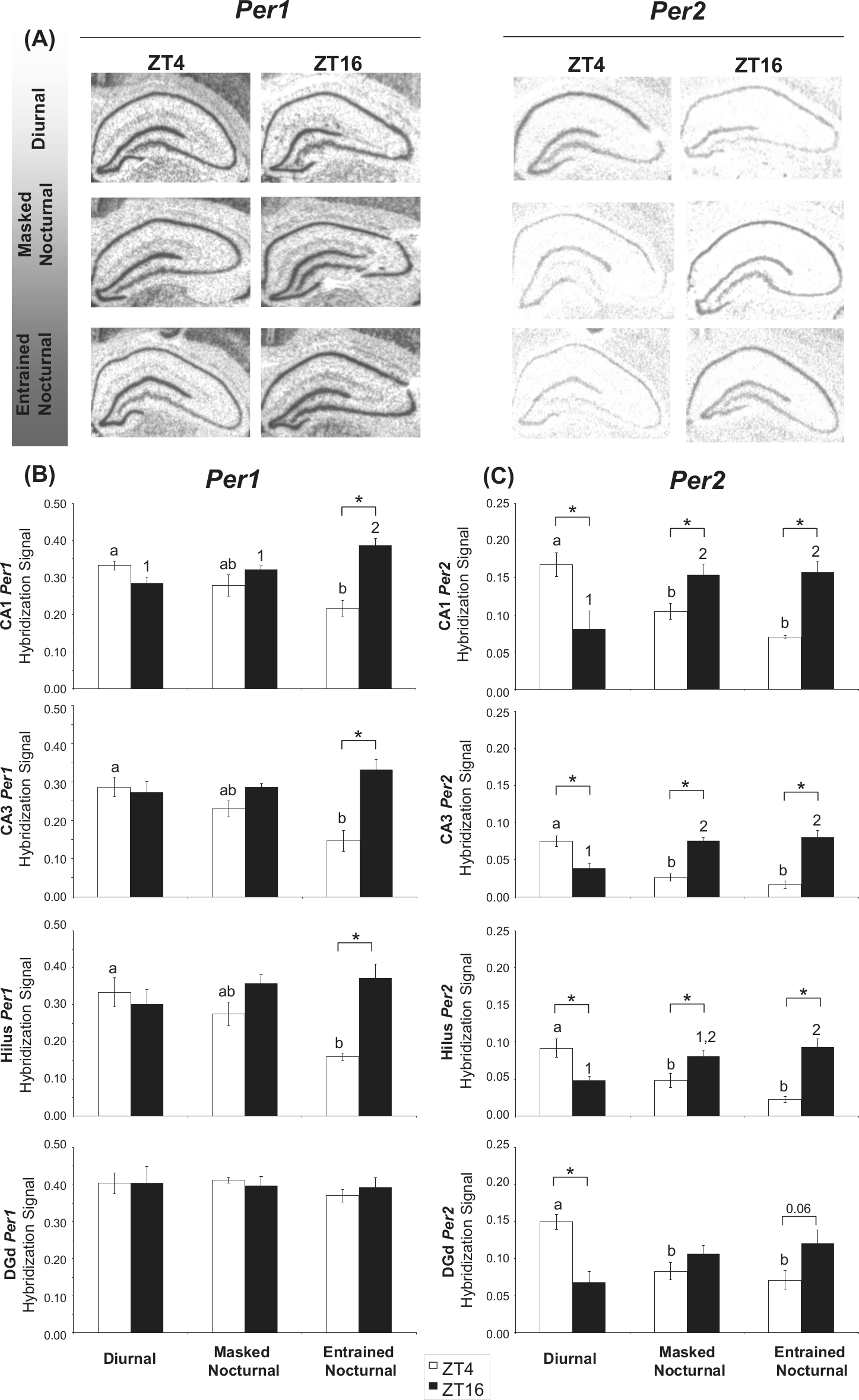
(A) Representative autoradiographs showing Per1 and Per2 signal in CA1, CA3, hilus, and the dorsal blade of the dentate gyrus (DGd) regions of the hippocampal formation of diurnal, masked nocturnal, and entrained nocturnal degus sacrificed at ZT4 and ZT16 (ZT0 corresponds to lights-on). Comparison of Per1 (B) and Per2 (C) signal in the CA1, CA3, hilus, and DGd regions of the hippocampus from diurnal, masked nocturnal, and entrained nocturnal degus at ZT4 and ZT16. Values are expressed as mean ± SEM (n = 4-6). Significant differences between chronotypes are noted by different letters (at ZT4) or different numbers (at ZT16; *p < 0.05).
Analysis of the simple main effects of time for each chronotype revealed that Per1 expression in CA1, CA3, and the hilus in entrained nocturnal degus was out of phase (higher levels at ZT16 than at ZT4) when compared with SCN values. Interestingly, in masked nocturnal animals, although a similar relationship in Per1 expression between the SCN and these hippocampal regions was seen, this inversion in Per1 expression was not significant (p = 0.214 for CA1; p = 0.053 for CA3; p = 0.063 for the hilus; Fig. 4B).
The Per1 hybridization signal in the DGd, striatum, ARC, VMH, and cortical areas (parietal and piriform cortices) did not show any significant ZT × chronotype interaction or main effects of chronotype or ZT (Fig. 4B, Fig. 5A, Suppl. Fig. S1). The only exception was in the cingulate cortex, where a main effect of chronotype was found, F(2, 23) = 8.544, p = 0.002, with the overall expression of Per1 elevated in the nocturnal animals compared with the diurnal degus (Fig. 5A).
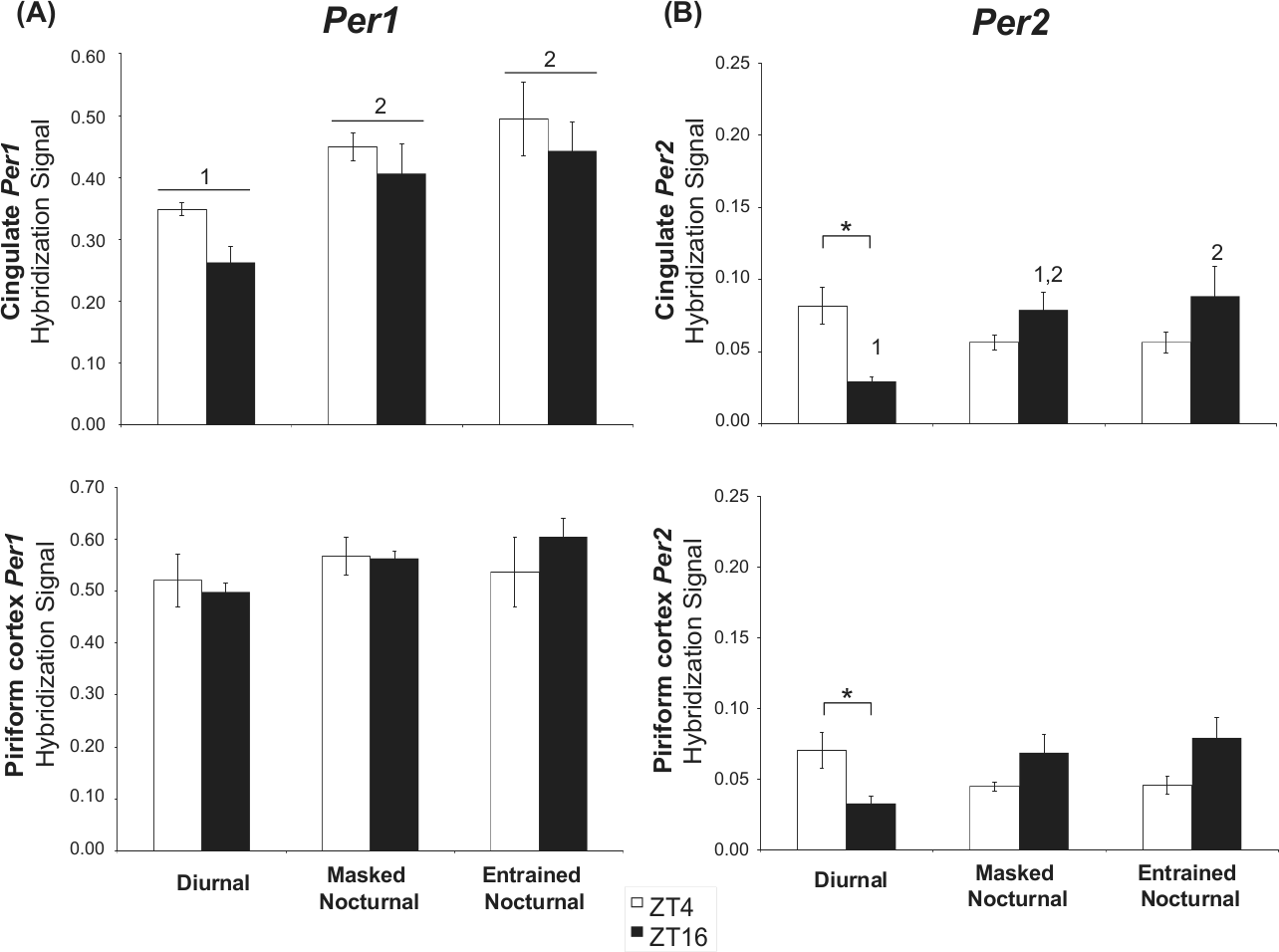
Comparison of Per1 (A) and Per2 (B) signal in the putative cingulate and in the piriform cortex from diurnal, masked nocturnal, and entrained nocturnal animals at ZT4 and ZT16. Values are expressed as mean ± SEM (n = 4-6). Significant differences between chronotypes are noted by different numbers (*p < 0.05).
When in situ hybridization analysis for Per2 expression was carried out, a significant interaction of ZT × chronotype in the CA1, F(2, 23) = 16.758, p < 0.001; CA3, F(2, 23) = 34.250, p < 0.001; hilus, F(2, 23) = 17.568, p < 0.001; DGd, F(2, 23) = 11.835, p < 0.001; cingulate cortex, F(2, 23) = 6.925, p < 0.01; and piriform cortex, F(2, 24) = 6.401, p < 0.01, was found. Follow-up analyses in the hippocampus revealed significant simple main effects of chronotype within each ZT (Suppl. Table S1). Interestingly, in both diurnal and nocturnal degus, Per2 levels in these areas of the hippocampus were in phase with their WRA rhythms. Thus, diurnal degus showed higher Per2 levels at ZT4 than at ZT16, whereas in both types of nocturnal degus (entrained and masked), Per2 expression was reversed, showing greater values at ZT16 than at ZT4 (Fig. 4C). It is noteworthy that although the differences between these 2 nocturnal chronotypes for Per2 levels in the hippocampus did not reach statistical significance, the day-night differences in its expression levels in this brain region tended to be higher in the entrained than in the masked nocturnal animals (Fig. 4C).
As in the hippocampus, Per2 levels in the cingulate and piriform cortices of diurnal degus were significantly higher during the day than at night, coinciding with their active phase, whereas in nocturnal degus, a trend to higher values at night was seen (Fig. 5B).
Discussion
To understand the relationships between daily brain clock gene activity and chronotype, the present study compares Period gene (Per1 and Per2) expression within the SCN and in key extra-SCN brain areas in both diurnal and nocturnal degus that had access to running wheels. Since some nocturnal degus were entrained to the dark phase (entrained nocturnal), while others became nocturnal by negative masking (masked nocturnal; see also Vivanco et al., 2009), we also discriminate, for the first time, Per1 and Per2 expression patterns in the SCN and other brain areas in both types of nocturnal animals to determine any temporal differences in transcription of these clock genes between entrained and masked nocturnal degus.
Within the SCN, the expression patterns of Per1 and Per2 did not differ between the 3 chronotypes (diurnal, masked, and entrained nocturnal), with high levels during the light phase (ZT4) and low levels at night (ZT16). This temporal pattern of Per1 and Per2 expression is similar to the profiles previously reported in diurnal degus that had no running wheel access (Vosko et al., 2009; Koch et al., 2009) and to other diurnal rodents, including ground squirrel, grass rat ansorgei, and grass rat niloticus (Caldelas et al., 2003; Mrosovsky et al., 2001; Novak et al., 2006). Similarly, this temporal pattern of Per1 and Per2 expression in the SCN was typical of nocturnal species, such as rats, mice, and hamsters (Asai et al., 2001; Nielsen et al., 2001; Shieh et al., 2005; Yan and Silver, 2002; Yamamoto et al., 2001; Wang et al., 2009). Recent observation in the SCN of the grass rats also supports our finding given that no differences in the phases for PER1 and PER2 protein rhythms were found when day- and nighttime active animals were compared (Ramanathan et al., 2010).
The day-night differences in Per1 expression in the putative SPVZ remained similar in all chronotypes. This concurs with previous results in the Nile grass rats (Ramanathan et al., 2010), which also show no phase differences in PER expression in this area between day- and night-active animals. A small caveat here is that although it appears that Per1 expression at this anatomical level is similar in both species, to date no evidence in the degu suggests an area that functions in the same way as the lower SPVZ in the Nile grass rats (Hagenauer and Lee, 2008).
Outside of the SCN, entrained nocturnal degus showed a pattern of hippocampal Per expression that resembled that of previously characterized nocturnal species (Wakamatsu et al., 2001), with levels peaking during the night at ZT16 and significantly lower levels at ZT4. For Per1 expression, this was found only in entrained nocturnal degus, and although a similar trend was seen in nocturnal masked animals, this day-night difference in Per1 expression did not reach statistical significance. Interestingly, day-night differences in Per1 gene expression were not seen in diurnal degus, and with the exception of the hippocampus of entrained nocturnal degus, we did not find day-night differences in Per1 expression in any of the putative brain areas (cortices, striatum, arcuate, the dorsal blade of the dentate gyrus, ventromedial hypothalamus) analyzed across the chronotypes. Together, these observations suggest that in analyzed extra-SCN brain regions, Per1 gene expression is rhythmic only in entrained nocturnal degus, with masked nocturnal degus representing an intermediate pattern between entrained diurnal and nocturnal degus. We must stress, however, that since we evaluated only 2 time points, we can neither completely exclude the possibility that Per1 expression in these extra-SCN brain areas analyzed here is rhythmic nor discount that there may be differences between the chronotypes. In favor of our conclusion, however, previous studies in our lab have shown that Per1 gene expression in the brain cortices of the degu is not rhythmic (Vosko et al., 2009, Hagenauer and Lee, unpublished data).
The daily pattern of Per2 expression outside the SCN differed between diurnal and nocturnal degus. In areas in the hippocampal formation examined (CA1, CA3, hilus, and DGd), and in putative cingulate and piriform cortices, Per2 expression levels were in phase with the animals’ behavioral rhythms. That is, Per2 levels were highest at ZT4 (light period) in diurnal animals and at ZT16 (dark period) in nocturnal degus. We reasoned, therefore, that while SCN peak levels of Per2 expression remained unaltered in diurnal and nocturnal degus, in extra-SCN brain regions, this peak in Per2 gene expression shifted in phase with the animal’s behavioral patterns and thus showed elevated levels at night in nocturnal degus. This pattern of Per2 expression seen in the hippocampus and the day-night trend observed in the cortex of nocturnal degus are consistent with previous studies in other nocturnal species (mice and rats) whereby Per rhythm also peaked at night (hippocampus: Wakamatsu et al., 2001; Wang et al., 2009; Jilg et al., 2010; and cortex: Wakamatsu et al., 2001; Masubuchi et al., 2000; Vosko et al., 2009; Abe et al., 2001). Further, in the diurnal grass rats, PER1 and PER2 rhythms in extra-SCN oscillators shift to a nocturnal phase when the animals change from day- to night-time activity (Ramanathan et al., 2010). Interestingly, in extra-SCN brain areas, a similar relationship between activity and Per gene expression is seen in nocturnal species that are driven to become active during the light phase (for example, by food restriction: Wakamatsu et al., 2001; or chronic exposure of methamphetamine: Masubuchi et al., 2000). Here, a phase reversal in Per rhythm is reported with peak expression shifted from night to light period, while SCN expression pattern remained unaffected.
Together, our results suggest that while there are no fundamental temporal differences in Period gene (Per1 and Per2) expression in the SCN of diurnal and nocturnal degus, in extra-SCN brain sites, the pattern for Per2 is significantly different between the chronotypes; peak expression occurs when the animals are active. In the CA1 and CA3 of the hippocampus and in the hilus, the timing of the Per1 peak was also in phase with the animal’s activity pattern.
The causal relationship between the expression patterns of Per and locomotor activity in extra-SCN brain regions remains unclear. For example, we are still unsure whether the phasing of Per in these extra-SCN brain regions is set by changes in the animal’s phase preference (diurnal to nocturnal) or vice versa. In the current study, clock gene analyses were carried out after the behavioral patterns were firmly established in these animals. Much light would be shed onto these issues in future studies assessing clock gene levels from brain tissues collected immediately after the animals shift their activity-phase preference.
The possible mechanisms underlying the shifts seen in extra-SCN oscillators may involve assertion of otherwise self-sustained circadian oscillators that reside outside of the SCN. These oscillators may switch their phases and then communicate their new phase information to other semiautonomous and/or slave extra-SCN oscillators. These self-sustained oscillators include the main olfactory bulb (OB) and the SCN-independent food-entrainable oscillator (FEO; reviewed in Guilding and Piggins, 2007). It is known that the OB controls daily changes in olfaction sensitivity and drives daily circadian rhythms in one of its primary targets, the piriform cortex, which is lost following bulbectomy (Granados-Fuentes et al., 2006). Since the OB also sends direct and indirect projections to other brain areas known to contain rhythmic clock gene expression (e.g., amygdala: McDonald, 1998; hippocampus: Insausti et al., 2002), the possibility that these extra-SCN brain areas may also be under OB’s circadian drive cannot be discounted. Future experiments addressing the contribution of the OB to daily clock gene expression in extra-SCN oscillators will provide important information regarding this issue.
Restricted feeding schedules that engage the FEO can also alter behavioral and physiological rhythms independently of the SCN clock (Stephan, 2002) and adjust the phase of Per oscillations in some extra-SCN regions (Wakamatsu et al., 2001; Angeles-Castellanos et al., 2007; Feillet et al., 2008). Although the involvement of the FEO cannot be discounted here, it is unlikely that a FEO is involved since food was available ad libitum and not restricted to a specific schedule over the circadian cycle. Indeed, although in ad libitum feeding conditions, diurnal degus concentrate their feeding activity during the light period (in phase with their activity pattern), and food restriction to 12 h at night does not induce nocturnalism in these animals (Vivanco et al., 2010a), the feeding patterns of nocturnal degus are less understood. Future studies addressing the role of FEO in extra-SCN clock gene expression both in masked and entrained nocturnal animals need to combine ad libitum and feeding schedules with measurements of brain clock gene activity. Analysis of clock gene expression in extra-SCN areas of diurnal degus with food availability confined to 12 h at night will also shed light on the involvement of the FEO in extra-SCN Per gene rhythm reversal.
Studies simulating body temperature rhythms in cultured cells and tissue explants have shown that temperature can sustain and provide important resetting cues for mammalian peripheral oscillators in vitro (Brown et al., 2002; Burh et al., 2010; Saini et al., 2012). Although this raises the possibility that body temperature rhythms associated with the active phases of diurnal and nocturnal degus (Vivanco et al., 2009; Otalora et al., 2010) may provide an underlying drive in Per expression in extra-SCN oscillators seen here, further in vivo experiments are needed to confirm this. Indeed, the day-night fluctuation of environmental temperature in our study was between 1 and 1.5 °C. Previous work (Kas and Edgar, 1999; Vivanco et al., 2009, 2010c) indicated that this variation in environmental temperature does not induce nocturnal chronotypes. Therefore, it is unlikely that the behavioral and Per expression patterns seen here were due to day-night changes in environmental temperature.
The physiological functions of clock genes in areas outside the SCN are just beginning to be scrutinized. The brain areas studied here are involved in learning and memory, neurogenesis, activity, neuroendocrine system, feeding, reward, depression, and addiction. It is suggested that daily intrinsic oscillations in clock gene activity in these extra-SCN brain areas are important in the normal physiological function of these brain regions (Menet and Rosbash, 2011), with the SCN acting as the master global time giver coordinating daily circadian gene activity in these brain areas (Abe et al., 2002). Indeed, several studies have highlighted the importance of the circadian system in hippocampal-dependent memory formation and neurogenesis and the importance of the SCN in coordinating these events (reviewed in Eckel-Mahan and Storm, 2009; Gerstner and Yin, 2010; Menet and Rosbash, 2011). One might therefore argue that the phase reversal seen in Per2 gene expression in the hippocampus of the nocturnal degus aligns the optimal time of cognitive functioning with their daily activity patterns. To address this hypothesis, future studies need to perform simultaneous behavioral test and clock gene activity measurements in diurnal and nocturnal degus or animals that have recently shifted from a diurnal to a nocturnal phase of activity.
The physiological roles for Per1 and/or Per2, however, may be brain region and species specific. In some brain structures, rhythmic expression is seen only in 1 of the 2 protein products of these genes, depending on the species being studied (Zheng et al., 2001). Our results add another interesting layer of complexity showing that the day-night changes in Per2 expression were present in all extra-SCN brain regions investigated, whereas the day-night differences in Per1 gene activity were present only in hippocampal CA1, CA3, and hilus regions of entrained but not in masked nocturnal degus. This indicates that in certain brain regions, fundamental differences in clock gene activity exist between entrained and masked nocturnal animals, raising the possibility that underlying circadian system processes may be fundamentally different in these 2 types of nocturnal degus. This finding may provide a golden opportunity to study the mechanisms involved and perhaps identify any advantages/disadvantages for full entrainment versus masking. Indeed, results emerging from our lab suggest that entrained degus are more efficient at thermoregulation than masked animals. We noted that although the overall amount of WRA at night is significantly higher in entrained than in masked nocturnal degus, no differences were found in their nighttime body temperature values (see Suppl. Table S2). This suggests that entrained animals may run more because they can regulate their body temperature more effectively than those that are masked. This higher WRA in entrained nocturnal degus may provide stronger activity-dependent feedback onto the master circadian clock and extra-SCN oscillators of these animals than in their masked counterparts, potentially causing an increase in the amplitude of clock gene expression outside the SCN.
In conclusion, our data indicate that daily Period gene expression in extra-SCN brain regions in diurnal and nocturnal degus was in antiphase while their pattern of expression in the SCN in all chronotypes remained similar, regardless of their activity pattern. Taken together, the results of the present study suggest that the mechanisms underlying activity phase preference in the degus reside downstream from the SCN. These data also indicate that there are fundamental differences between entrained and masked nocturnal degus. It is now important to address how coupling between the SCN and extra-SCN oscillators is established in mammals and how the phase reversals of gene expression in extra-SCN brain regions affect physiology. This will be of particular relevance to understand better the consequences associated with conditions causing phase reversal of activity pattern in humans, such as during night-shift work.
Footnotes
Acknowledgements
This project was funded to J.A.M. by the Instituto de Salud Carlos III (RETICEF, RD06/0013/0019 and RD12/0043/ 0011), the Ministry of Education and Science, and the Ministry of Economy and Competitiveness (BFU2010-21945-C02-01 and IPT-2011-0833-900000 co-financed by the FEDER Fund). B.B.O. was supported by a PhD scholarship (AP2006-04117). We wish to thank Ana Kantorowski, Jin Yan, Brittany Ajegba, Doug Orzel, Andy Williams, Jim Mossner, Li Zheng, and Alejandro Sempere for their help with data collection. We would also like to thank Drs. Stan Watson and Huda Akil for the use of their dark room.
Conflict of Interest Statement
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
