Abstract
Objectives:
Gout is a common inflammatory arthritis, which is caused by hyperuricemia. Limited efforts have been paid to systematically explore the relationships between gout and common psychiatric disorders.
Methods:
Genome-wide association study summary data of gout were obtained from the GeneATLAS, which contained 452,264 participants including 3,528 gout cases. Linkage disequilibrium score regression (LDSC) was first conducted to evaluate the genetic relationships between gout and 5 common psychiatric disorders. Transcriptome-wide association studies (TWAS) was then conducted to explore the potential biological mechanism underlying the observed genetic correlation between gout and attention-deficit hyperactivity disorder (ADHD). The Database for Annotation, Visualization and Integrated Discovery online functional annotation system was applied for pathway enrichment analysis and gene ontology enrichment analysis.
Results:
LDSC analysis observed significant genetic correlation between gout and ADHD (genetic correlation coefficients = 0.29, standard error = 0.09 and P value = 0.0015). Further TWAS of gout identified 105 genes with P value < 0.05 in muscle skeleton and 228 genes with P value < 0.05 in blood. TWAS of ADHD also detected 300 genes with P value < 0.05 in blood. Further comparing the TWAS results identified 9 common candidate genes shared by gout and ADHD, such as CD300C (P gout = 0.0040; P ADHD = 0.0226), KDM6B (P gout = 0.0074; P ADHD = 0.0460), and BST1 (P gout = 0.0349; P ADHD = 0.03560).
Conclusion:
We observed genetic correlation between gout and ADHD and identified multiple candidate genes for gout and ADHD.
Background
Gout is a common inflammatory arthritis, which is caused by abnormal metabolism of uric acid. 1 Gout is characterized by the elevation of serum uric acid levels and deposition of uric acid crystals in the joint and kidneys. 2 The main clinical manifestations of gout include joint pain, chronic joint damage, renal stone formation and potential renal insufficiency. The worldwide prevalence rate of gout ranges from 0.1% to approximately 10% and the incidence rate from 0.3 to 6 cases per 1,000 person per year. 3 The risk of gout is affected by multiple environmental factors such as diets, lead exposure, and alcohol consumption. 4 Additionally, genetic factors contribute greatly to the development of gout. It was reported that 60% of variation of uric acid level can be explained by genetic factors. 5 Genetic studies identified multiple susceptibility genes for gout, for instance SLC2A9, SLC22A12, and ABCG2. 6,7
Uric acid is an organic acid, which is the end product of purine metabolism and is recognized as a marker of oxidative stress in human body. 8 Recent studies provided evidence supporting the impact of uric acid on brain function. For instance, it was reported that patients with bipolar and depressive disorder were at a higher risk of gout than healthy controls. 9 Previous studies have suggested that uric acid plays a causal role in neurodegenerative diseases. 10 Uric acid has neuroprotective effects due to its anti-oxidative action. 10 Because oxidative stress plays a critical role in neurodegeneration, the low concentration of uric acid may adversely lead to increased oxidative stress and damage to neural cells. 10 Impulsivity is an essential component of various psychiatric disorders. Some evidence suggested that the role of uric acid in impulsivity comes from Lesch–Nyhan syndrome. Lesch–Nyhan syndrome’s behavioral manifestations include impulsivity, and even individuals with less mutation still exhibit attention deficits. 11 However, limited efforts have been paid to systematically explore the relationships between gout and common psychiatric disorders. Metabolic abnormalities, as well as hyperlipidemia, central obesity, hyperglycemia, hyperuricemia, and increased body mass index (BMI), are closely associated with psychiatric disorders. 9 Several studies suggested that gout is associated with depression symptoms. For instance, in the adults over 65 years of age, gout is independently associated with a 42% higher risk of developing depression. 12 In a study of 90 French children (aged 3.5 to 4.5 years), serum uric acid levels were correlated positively with hyperactivity, short attention span, impulsivity, and anger control. 13 However, the potential relationships between gout and psychiatric disorders remain elusive now. In this study, we aimed to systematically explore the relationship between gout and common psychiatric disorders in this study.
Mental/psychiatric disorders are clinical manifestations of behavioral, psychological, and biological dysfunction in individuals with major distress or disability. 14 The common psychiatric disorders mainly include attention-deficit hyperactivity disorder (ADHD), anxiety disorder (AD), autism (AUT), bipolar disorder (BD), schizophrenia (SCZ), and major depressive disorder (MDD). ADHD is a neurodevelopment disorder of childhood characterized by persistent impairment of attention, hyperactivity, and/or impulsivity. 15 AUT is a developmental mental disorder characterized by difficulties with social interaction and communication and restricted repetitive behavior. SCZ is a highly heritable and neuropsychiatric disorder characterized by hallucination, delusions, and extremely disordered thinking and behavior. 16 MDD is characterized by a persistent feeling of sadness or a lack of interest in stimuli. 17 AD mainly manifests as extreme fear or worry, panic disorder and panic attack, selective mutism, and specific phobias. 18
Recent studies demonstrated the generality of genetic correlations among complex diseases and traits. Linkage disequilibrium score regression (LDSC) is an efficient method for estimating genetic correlation and heritability from genome-wide association study’s (GWAS) summary statistics. 19 Utilizing GWAS summary data, LDSC provides an easy and reliable way to simultaneously screen thousands of traits and find out the real genetic correlations among them. 20 For instance, Duncan et al. suggested that LDSC was a powerful approach for assessing the genetic relationships among SCZ, BD, and MDD. 21 The transcriptome-wide association study (TWAS) have been recently advocated as a powerful approach to prioritize candidate risk gene underlying complex traits, and this approach mostly utilized as a test of association to identify risk region where eQTLs that involved in disease risk. 22
In this study, LDSC was first conducted to evaluate the genetic relationships between gout and common psychiatric disorders including ADHD, AD, AUT, BD, SCZ, and MDD. Significant genetic correlation was detected between gout and ADHD. TWAS of gout and ADHD were then conducted and compared to detect the common genes shared by gout and ADHD.
Methods and Materials
GWAS Data of Gout and Uric Acid Level
The GWAS summary data of gout were obtained from the GeneATLAS website (http://geneatlas.roslin.ed.ac.uk/). Briefly, a total of 452,264 participants including 3,528 gout cases of European ancestry were included in this study. 23 DNA extracting and genotyping were performed by Affymetrix Research Services Laboratory, using Applied Biosystems UK Biobank Axiom Array. Genotype imputation was performed by IMPUTE4 against the Haplotype Reference Consortium as reference panel. GWAS of gout were performed using a linear mixed model. Detailed information of the subjects, genotyping, imputation, and quality control can be found in the published study. 23
Since elevated concentrations of serum urate—hyperuricemia can cause gout. We also use the GWAS summary data of uric acid levels to detect its genetic correlation with gout and ADHD. The GWAS meta-analysis of uric acid level was derived from >140,000 individuals of European ancestry within the Global Urate Genetics Consortium. 24 In each study of the discovery GWAS, genotyping was performed on genome-wide chips, and imputation was conducted using HapMap 2 data as the reference. Detailed information of the subjects, genotyping, imputation, and quality control can be found in the published study. 24
GWAS Datasets of ADHD, AD, AUT, BD, SCZ, and MDD
The latest GWAS summary data of ADHD, AD, AUT, BD, SCZ, and MDD were driven from the Psychiatric Genomic Consortium (PGC; http://pgc.unc.edu/). Briefly, these GWAS data sets contained 20,183 ADHD cases and 35,191 controls for ADHD 25 ; 7,016 AD cases and 14,745 controls for AD; 5,305 AUT cases and 5,305 controls for AUT; and 33,426 cases and 54,065 controls for SCZ. 26 All study subjects were European whites. Genotyping was conducted using commercial platforms such as Illumina 610 K and Affymetrix SNP 6.0 chips. Genotype imputation was conducted against the public reference panels, for instance, the HapMap and 1,000 Genomes Project reference panels. GWAS including 59,851 MDD cases and 113,154 controls were collected from deCODE, generation Scotland, UK biobank (not including 23 and Me). 27 Imputation was performed using the 1,000 genomes project multi-ancestry reference panel. 27 Genotype data for all PGC29 samples, GERA, and iPSYCH were performed by the PGC ricopili pipeline for standardized quality control, imputation, and analysis. 27 Detailed information of the subjects, genotyping, imputation, and quality control can be found in the published study.
Genetic Correlation Scan between Gout, Uric Acid Level, and Psychiatric Disorders
Following the document recommended by the developers, 19 LDSC software (v1.0.0; https://github.com/bulik/ldsc) was applied to the GWAS summary data for evaluating the genetic relationships between gout, uric acid level, and each of ADHD, AD, AUT, BD, SCZ, and MDD, respectively. The principle of the LD score regression is based on the fact that the genetic variants tagging more of genome (e.g., having high LD scores) are more likely to tag causal variants and therefore have higher statistics on average than the genetic variants with low LD scores. LDSC only requires GWAS summary-level data and can distinguish genuine polygenicity from the bias caused by population stratification and cryptic relatedness. 19 Bonferroni correction has become the main method to solve the problems of multiple statistical tests in ecological research. We made multiple comparisons according to Bonferroni correction for the number of analyzed psychiatric disorders (0.05/5).
TWAS of Gout and ADHD
It is well known that gene expression is under genetic control and a large part of the candidate loci identified by GWAS affects diseases by regulating gene expression. 28 Another limitations of GWAS is that it cannot consider tissue specificity in association analysis. 29 This motivates the development of TWAS, which take tissue specificity into account by imputes transcriptome levels of genes with models trained in measured transcriptome datasets (e.g., GTEx). 30 Specific for this study, the expression weights of brain RNA-seq (DLPFC), peripheral blood RNA array, and whole blood RNA array were downloaded (http://gusevlab.org/projects/fusion/) and used as reference data in the TWAS of ADHD, and muscle skeletal peripheral, blood RNA array, and whole blood RNA array were used as reference data in the TWAS of gout. The gene expression weights were first calculated using the prediction models of FUSION. Let w denotes the weights. Z denotes the scores of diseases (gout and ADHD). L denotes the single nucleotide polymorphism (SNP) correlation (LD) matrix. The association testing statistics between predicted gene expression and disease was calculated as Z TWAS = w′Z/(w′Lw)1/2. 30 The imputed gene expression data can be viewed as a linear model of genotypes with weights based on the correlation between SNPs and gene expression in the training data while accounting for LD among SNPs. 30 FUSION by default uses a range of models to derive the SNP weights for each gene, such as “blup”, “bslmm,” and “lasso.” We conducted TWAS analysis by using the default setting of FUSION. For TWAS analysis, a permutation-based correction for multiple testing was implemented in FUSION software. We conducted 2,000 times permutation for the genes with TWAS P < 0.05 and genes with TWAS permutation-based empirical P value <0.05 was considered as significant. We compared the TWAS results of gout and ADHD to identify the common candidate genes shared by gout and ADHD. To avoid bias due to the expression correlation between genes, lme4qtl 31 package in R software was used to fit a mixed model regression of TWAS Z-score, accounting for the correlation in Z-scores between genes due to LD. In brief, predicted expression levels for genes from all SNP-weight sets were imputed into the 1,000 genomes reference dataset used by FUSION using the FUSION protocol which involves PLINK (https://github.com/opain/Predicting-TWAS-features). 32 The correlation between genes that were more than 5 Mb apart or on separate chromosomes was set to zero. Any gene–gene correlations with an R 2 less than 0.0001 were set to zero. The software used for this analysis (TWAS-GSEA) is publically available (https://github.com/opain/TWAS-GSEA). 33 We used the procedure to detect the correlation between genes of gout and ADHD due to LD.
Gene Set Enrichment Analysis
The Database for Annotation, Visualization and Integrated Discovery (DAVID), a full knowledgebase online functional annotation system, was applied for pathway gene enrichment analysis and gene ontology (GO) enrichment analysis of gout and ADHD associated genes identified by TWAS. In this study, the genes with permutation-based empirical P value < 0.05 identified by TWAS of ADHD and gout were selected for GO and pathway enrichment analysis by DAVID (https://david.ncifcrf.gov/.). DAVID now provides researchers a comprehensive set of functional annotation tools to understand the biological meaning behind a large number of genes. DAVID tools are able to identify enriched biological themes, particularly GO terms and discover enriched functional related gene group. We used the multiple testing corrected Benjamini P values <0.05 as significant threshold.
Results
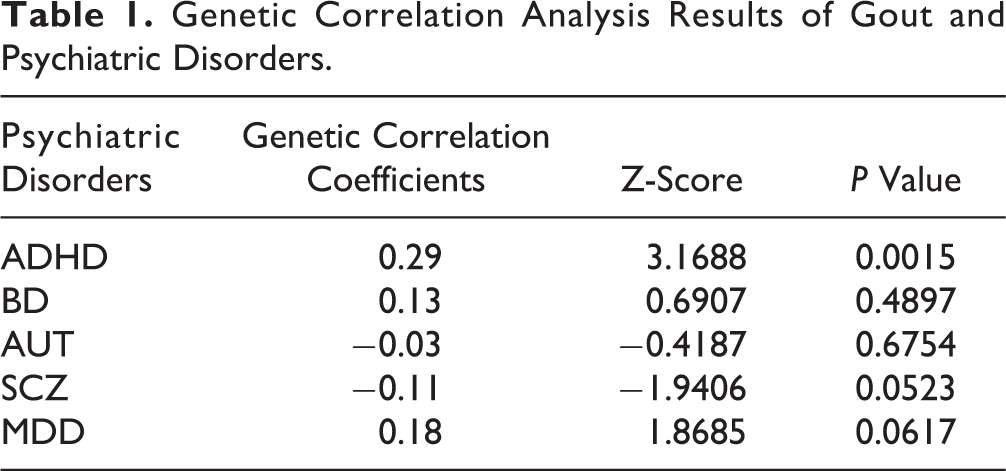
LDSC analysis observed significant genetic correlation between gout and ADHD (P value = 0.0015, genetic correlation coefficients = 0.29, and Z-score = 3.1688; Table 1). LDSC analysis also observed significant genetic correlation between uric acid level and ADHD (P value = 0.0057, genetic correlation coefficients = 0.14, and Z-score = 2.7632) and gout (P value = 4.88×10-33, genetic correlation coefficients = 0.9184, and Z-score = 11.9737). Table 2 summarized the liability scale heritability of each phenotype included in this study.
Genetic Correlation Analysis Results of Gout and Psychiatric Disorders.
The Liability Scale Heritability of Each Phenotype.

Further TWAS of gout identified 105 genes with permutation-based empirical P value < 0.05 in muscle skeleton, such as PCOLCE2 (P = 0.0005) and ZNF669 (P = 0.0139; Supplemental Table S1). TWAS of gout also identified 228 genes with permutation-based empirical P value < 0.05 in blood, such as ADAM metallopeptidase domain 17 (ADAM17; P = 0.0247) and CYP4F3 (P = 0.0005; Supplemental Tables S2 and S3). For ADHD, TWAS identified 258 genes with permutation P value <0.05 in brain tissue, such as VSIG10 (P = 0.0005) and KIAA1671 (P = 0.0084; Supplemental Table S4). Additionally, TWAS of ADHD detected 300 genes with permutation-based empirical P value < 0.05 in blood, such as SDF2 (P = 0.0015) and oligodendrocyte transcription factor 2 (OLIG2; oligodendrocyte transcription factor 2P = 0.0359; Supplemental Tables S5 and S6). GO enrichment analysis of the significant genes identified by TWAS detected 1 GO terms, membrane (DAVID P = 0.0002) for gout.
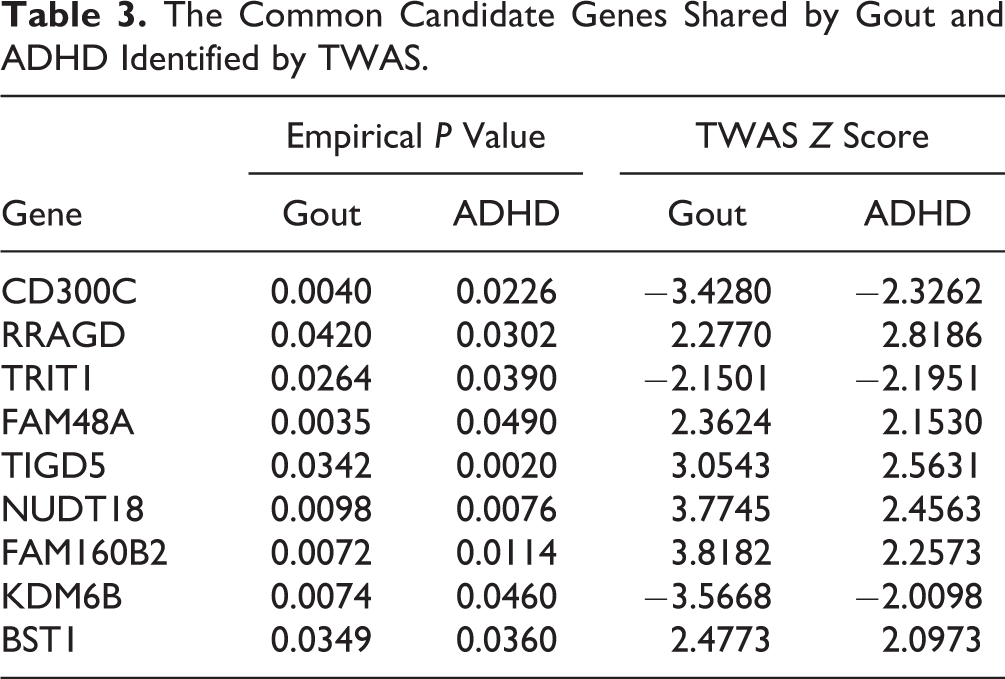
Further comparing the TWAS results identified 9 common candidate genes shared by gout and ADHD, such as CD300 antigen-like family member C (CD300C; P gout = 0.0040; P ADHD = 0.0226), KDM6B (P gout = 0.0074; P ADHD = 0.0460), and bone marrow stromal cell antigen-1 (BST1; P gout = 0.0349; P ADHD = 0.03560; Table 3).
The Common Candidate Genes Shared by Gout and ADHD Identified by TWAS.
Discussion
The biological mechanism of the observed correlation between gout and psychiatric disorders remains elusive now. In this study, we conducted a genetic correlation scan between gout and 5 common psychiatric disorders using LDSC and large-scale GWAS summary data. Significant correlation was observed between gout, uric acid level, and ADHD. TWAS of gout and ADHD were then conducted and identified some common candidate genes shared by gout and ADHD.
There are some evidences supporting a possible role of uric acid in the development ADHD. In a study of 90 French children (aged 3.5 to 4.5 years), serum uric acid levels were correlated positively with hyperactivity, short attention span, impulsivity, and anger control. 13 It has been reported that low-level lead intoxication is associated with increased risk of ADHD, and lead intoxication is a mechanism for increasing uric acid levels. 34,35 Recent studies have also provided evidence supporting the correlation between gout and other common psychiatric disorders. 9 –11 A dose–response relationship was uncovered between serum uric acid and brain infraction among women, which notice that the higher serum uric acid the faster rate of Parkinson disease progression in women. 36 Uric acid may influence emotion by modifying the function of brain region that associated with emotion-related psychopathology such as anxiety and depression. 37 Uric acid could also affect the central nervous system. For example, increased uric acid levels could accelerate purine transformation thus leading to alterations of neural transmission. 37 Uric acid is considered a neuroprotective factor against oxidative stress, which is specifically, suppresses the damaging effects of reactive oxygen species on neurons by preserving the integrity of the plasma membrane. 37 Our results were consistent with previous study results, supporting the biological effects of uric acid on brain function as well as the development of ADHD.
Some life-style factors may underlie the relationship between gout and ADHD, such as alcohol intake and obesity. Some studies examine whether individuals with a high genetic risk for both ADHD and BMI benefit in particular from interventions targeting one or both of these conditions. 38 The reduction in ADHD symptoms can associate with decreased BMI, which is especially for overweight adolescents. Few studies suggested that BMI and excessive consumption of alcoholic beverages, meat, soft drinks, and fruit juices increase the risk of developing gout. 39 In addition, it has been suggested that use of alcohol temporarily destroys cognitive functioning in a manner similar to persistent cognitive impairment that is characteristic of disorders such as ADHD. 40 Johnson et al. reviewed preclinical and clinical data suggested overlaps among ADHD, sugar and obesity, and modifiable risk factors for gout include obesity and consumption of purine-rich alcoholic beverages. 41
ADAM17 is a nominal significant gene identified by TWAS for gout. It was known as Tumor necrosis factor-α-converting enzyme and has been implicated in various biological processes involving cell–cell and cell–matrix interactions, including fertilization, muscle development, and neurogenesis. 42 ADAM17 is associated with various diseases including polycystic kidney disease, Alzheimer disease and autoimmune diseases such as rheumatoid arthritis (RA). 43 ADAM17 is expressed on RA synovial tissues and fibroblast-like synoviocytes and is associated with disease activity in RA. 43 ADAM17 substrate glycoprotein VI is critical for platelet microparticle formation involved in the inflammation that is important for hemostasis, thrombosis, cancer, and inflammation. Platelet microparticles are found in arthritis as well as in other joint inflammatory disorders like gout and juvenile arthritis. 44 In 2007, a epidemiology study reported that a 25-year cumulative gout prevalence of more than 5% in patients with RA. Few studies suggested that a significant proportion of RA patient’s exhibit monosodium urate deposition consistent with gout, challenging the previous perceptions that gout and RA are mutually exclusive conditions. 45
OLIG2 is nominal significant gene identified by TWAS for ADHD. OLIG2 is a candidate gene for SCZ, which expressed points to an etiological mechanism in SCZ. 46 OLIG2 gene is essential regulator in the development of white matter, which influences risk for psychosis in people with Alzheimer disease. 47 The OLIG2 gene is highly expressed in amygdala, thalamus, and caudate nuclei–brain regions associated with obsessive compulsive disorder. 48 OLIG2 is necessary for the development of oligodendrocytes, which are responsible for the production of white matter (myelin), and obsessive compulsive disorder is associated with white matter abnormalities. 48 A recent study suggested that OLOG2 is significantly associated with SCZ in the British population. 49 ADHD is associated with many psychiatric disorders such as MDDs, obsessive compulsive disorder, and SCZ. 50,51
TWAS detected several common genes shown nominal significance shared by gout and ADHD, such as BST1, CD300, and KDM6B. BST1 plays an important role for B-lymphocyte development. 52 The BST-1 is expressed on human bone marrow stromal cells, myeloid lineage cells, and lymphoid progenitors. BST-1 was involved in both ADP-ribosyl cyclase and cyclic ADP-ribose hydrolase activities. 53 BST-1 expression is raised in bone marrow stromal cell lines derived from patients with RA. 54 SNPs in the BST-1 gene were associated with the increased risk of Parkinson disease due to modified ADP-ribosyl cyclase activity. 55
CD300c is a protein that belongs to the CD300 family. The CD300 family is functional in modulating a large range of immune cell processes via their activating and inhibitory receptors. 56 The expression of CD300c on primary monocytes is regulated upon stimulation with TLR4 and TLR5 ligands, which is a costimulatory effect with lipopolysaccharides in the secretion of pro-inflammatory cytokines. 56 CD300c is an activating receptor expressed on monocytes and is important for inflammatory responses. 56 CD300 family is emerging as novel immune regulators in health and disease due to the interaction between their lipid-nature ligands. 57
KDM6B, also known as JMJD3, is essential for normal organ development and can affect bone formation and the cartilage maturation during endochondral bone formation. 58 JMJD3 can enhance pro-inflammatory and anti-inflammatory responses by targeting different transcription factors in a context-sensitive manner in gene promoters. 59 JMJD3 affects the peripheral nervous system, indicating that JMJD3 plays a role in microglia polarization and may change the immune pathogenesis of Parkinson disease. 58 JMJD3 is also involved in the neural development and inflammation that plays a protective role in Alzheimer disease. 58
There are 2 advantages of this study. First, we applied the TWAS approach that took tissue specificity genetic effect of gene expression into account. TWAS can help us to identify the candidate genes, whose predicted expression levels were associated with ADHD and gout. Additionally, other powerful approach can be used, such as MAGMA. 60 Second, large-scale GWAS and powerful LDSC approach were used here, which ensure the accuracy of our study results. LDSC is an effective and reliable method by using GWAS summary data to estimate the SNP heritability of complex traits and diseases. LDSC contributes to an accurate estimate of the genetic correlation between 2 traits, even if neither trait’s heritability was well estimated nor it was also important for using of LDSC, which was the estimation, and correction of confounding factors.
Our studies also have 3 limitations. First, TWAS detected some common candidate genes between gout and ADHD. But it is difficult to clarify the potential genetic mechanism of identified candidate genes contributing to both gout and ADHD only using GWAS summary data in this study. Functional studies are warranted to investigate whether the significant associations of the identified genes are driven by the same causal variants between gout and ADHD. Second, the genes near one another are likely to correlate in their predicted expression due to LD, which can lead to multiple significant TWAS hits within a single locus, thus bias the results of gene set enrichment analysis. Further studies are needed to validate our findings and reveal related biological mechanism. Last, we have not accounted for multiple testing in TWAS results, thus the genes identified for ADHD and gout are only suggestive evidence for the associations. Further functional studies are warranted to explore the underlying mechanism.
Conclusion
In summary, we conducted a genetic correlation scan between gout and 5 common psychiatric disorders and observed significant genetic correlation between gout and ADHD. Further TWAS identified multiple common candidate genes between gout and ADHD.
Supplemental Material
Supplemental Material, Table_S1 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S1 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Supplemental Material
Supplemental Material, Table_S2 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S2 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Supplemental Material
Supplemental Material, Table_S3 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S3 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Supplemental Material
Supplemental Material, Table_S4 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S4 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Supplemental Material
Supplemental Material, Table_S5 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S5 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Supplemental Material
Supplemental Material, Table_S6 - Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité
Supplemental Material, Table_S6 for Genetic Correlation Analysis and Transcriptome-wide Association Study Suggest the Overlapped Genetic Mechanism between Gout and Attention-deficit Hyperactivity Disorder: L’analyse de corrélation génétique et l’étude d’association à l’échelle du transcriptome suggèrent un mécanisme génétique superposé entre la goutte et le trouble de déficit de l’attention avec hyperactivité by Om Prakash Kafle, Xi Wang, Shiqiang Cheng, Miao Ding, Ping Li, Bolun Cheng, Xiao Liang, Li Liu, Yanan Du, Mei Ma, Lu Zhang, Yan Zhao, Yan Wen and Feng Zhang in The Canadian Journal of Psychiatry
Footnotes
Authors’ Note
OK and XW contributed equally to this work. OK and FZ conceived and designed the study, and wrote the manuscript; OK and FZ collected the data and carried out the statistical analyses; MD, SC, YD, YW, YZ, PL, LL, BC, LZ, MM and XL made preparations for the manuscript at first. All authors reviewed and approved the final manuscript.
Acknowledgments
The authors would like to acknowledge the funding received from National Natural Scientific Foundation of China (81472925 and 81673112); Key Projects of International Cooperation among Governments in Scientific and Technological Innovation (2016YFE0119100); Natural Science Basic Research Plan in Shaanxi Province of China (2017JZ024); Fundamental Research Funds for the Central Universities.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research and/or authorship of this article: This study received funding from National Natural Scientific Foundation of China (81472925 and 81673112); Key Projects of International Cooperation among Governments in Scientific and Technological Innovation (2016YFE0119100); Natural Science Basic Research Plan in Shaanxi Province of China (2017JZ024); Fundamental Research Funds for the Central Universities.
Supplemental Material
The supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
