Abstract
Objective:
Atypical antipychotics are linked to a higher incidence of metabolic side effects, including weight gain, dyslipidemia, and diabetes. In this study, we examined the prevalence and potential genetic predictors of metabolic side effects in 60 adult patients on clozapine.
Method:
Genetic variants of relevance to clozapine metabolism, clearance, and response were assessed through targeted genotyping of cytochrome P450 enzymes CYP1A2 and CYP2C19, the efflux transporter ABCB1, the serotonin receptor (HTR2C), leptin (LEP), and leptin receptor (LEPR). Clozapine levels and other potential confounders, including concurrent medications, were also included in the analysis.
Results:
More than half of the patients were obese (51%), had metabolic syndrome (52.5%), and 30.5% were overweight. There was a high prevalence of antipsychotic polypharmacy (61.9%). With multivariable linear regression analysis, LEP –2548G>A, LEPR c.668A>G, and HTR2C c.551-3008 C>G were identified as genetic predictors of body mass index (BMI) after considering effects of clozapine dose, blood level, and concurrent medications (adjusted R2 = 0.305). Metabolic syndrome was found to be significantly associated with clozapine level and CYP2C19*2 and LEPR c.668 G alleles. Clozapine levels in patients with metabolic syndrome were significantly higher compared to those without metabolic syndrome (1886 ± 895 vs. 1283 ± 985 ng/mL, P < 0.01) and were associated with the CYP2C19*2 genotype. No association was found between the genetic variants studied and lipid or glucose levels.
Conclusion:
This study confirms a high prevalence of metabolic side effects with clozapine and suggests higher clozapine level and pharmacogenetic markers in CYP2C19, LEP, LEPR, and HTR2C receptors as important predictors of BMI and metabolic syndrome.
Clinical Implications
Pharmacogenetic markers in LEP, LEPR, and HTR2C receptors are important predictors of body mass index in patients on clozapine. Education is needed to reduce antipsychotic polypharmacy, particularly with clozapine. Clinicians should use clozapine levels as a tool to monitor for risk of metabolic syndrome.
Limitations
The study was cross-sectional in nature and had a relatively small sample size. Larger controlled prospective studies are warranted to confirm clinical implications of the genetic risk factors identified in this study.
Introduction
Schizophrenia is a severe mental illness and is associated with mortality rates 2 to 3 times higher than the general population. 1 Schizophrenia has been reported as an independent risk factor for metabolic syndrome, including insulin resistance and increased visceral adiposity. 2,3 Several studies now confirm that increased risk of mortality in schizophrenia is attributable to unhealthy lifestyle and physical illnesses, with cardiovascular disease (CVD) being the major contributor. 4,5
Atypical antipsychotics are currently the first-line treatment options for schizophrenia and other psychotic illnesses. These are preferred over the typical antipsychotics due to the decreased incidence of extrapyramidal side effects. However, atypical antipsychotics such as clozapine, olanzapine, risperidone, and quetiapine are associated with varying degrees of metabolic side effects, including weight gain, diabetes mellitus, and dyslipidemia, thereby further increasing the metabolic and cardiovascular (CV) risks in patients treated with such agents. 6,7 Among those, clozapine and olanzapine are reported to carry the highest risk for metabolic side effects. 8,9 Clozapine, not preferred as a first-line agent due to the risk of a serious though rare side effect of neutropenia, is the only antipsychotic reserved for treatment-resistant schizophrenia. Significant improvement has been observed in 30% to 60% of such patients when switched to clozapine treatment. 10 Unfortunately, almost half of the individuals on clozapine develop metabolic side effects over the course of therapy. 11
Considerable variability has been reported in the incidence of metabolic side effects among patients prescribed atypical antipsychotics. This may result from differences in the drugs’ pharmacokinetics (drug metabolism and transport affecting systemic drug exposure) and pharmacodynamics (pharmacological action) that are influenced by genetic factors. 12 Clozapine acts as antagonist of dopamine (D2 and D4) and serotonin (5HT2 and 3) receptors, as well as adrenergic, cholinergic, and histamine (H) 1 receptors. Antagonistic drug effects at the H1 and serotonin 2C subtype receptor (HTR2C) have been implicated in weight gain as well as obesity-independent diabetes. 13 Two meta-analyses reported an association of HTR2C genotype (rs3813929) with antipsychotic-induced weight gain. 14,15 There is also evidence that the metabolic effects of atypical antipsychotics may be mediated by peptide hormone regulators of metabolic control, including leptin and ghrelin. 16 To date, associations of gene polymorphisms in 5HTR2C (rs3813929, rs1414334) as well as leptin (LEP; rs7799039) and leptin receptor (LEPR; rs1137101) with antipsychotic-induced weight gain and metabolic syndrome have been widely reported yet with varying results, and sex-specific effects have been postulated by some. 14,17 –22 Furthermore, leptin has also been suggested to increase blood pressure through sympathetic activation in the circulatory system or at the renal level, 23 and LEP gene polymorphisms as well as leptin plasma levels were shown to correlate with blood pressure, particularly in women. 24,25
Clozapine is predominantly metabolized by cytochrome P450 (CYP) 1A2 to its active metabolite norclozapine (also N-desmethylclozapine), whereas CYP2C19, CYP2D6, and CYP3A4 are thought to represent minor deactivating pathways. 26,27 Norclozapine exposure is reported to be about 50% to 70% of that of the parent compound with similar affinity to D2 and 5HT2 receptors. 27 In a recent study, impaired-function polymorphisms in CYP1A2 (*1C; *1D) were associated with higher serum clozapine concentration but also increased insulin and lipid levels in a small set of patients, whereas no effect was observed for the CYP2D6 genotype. 28 Furthermore, a potential role of gene variants in CYP2C19 (*1; *2; *17) and P-glycoprotein (ABCB1; rs1128503), an efflux transporter of the ATP binding cassette (ABC) family, on clozapine exposure has been reported, but metabolic side effects were not explored. 29 There are no studies in a Caucasian population that specifically investigate both the pharmacokinetic and pharmacodynamic genetic factors contributing to the metabolic side effects of clozapine in addition to clozapine trough concentration as well as important clinical confounders. Therefore, this exploratory study was conducted in a cohort of patients prescribed clozapine as a part of standard patient care, to more fully address the above gap in knowledge.
Materials and Methods
Patients and Study Design
This was a cross-sectional study of adult patients (aged 18-65 years) with a psychotic disorder (schizophrenia, schizoaffective disorder, or delusional disorder), under the care of the Prevention and Early Intervention Program for Psychoses (PEPP), London, Canada, who were on stable clozapine therapy for at least 6 months. The study was conducted between June 2012 and May 2014. Exclusion criteria were serious uncontrolled illness, nonadherence with medication (confirmed by the patient or blood clozapine levels after recruitment), and/or organic psychosis. Meeting the above inclusion criteria, 60 patients were recruited for this study after providing written informed consent. Information on diagnosis, medications prescribed in the previous year, age, sex, ethnicity, waist circumference, body mass index (BMI), blood pressure (BP), blood sugar, glycosylated haemoglobin (HbA1c) if diabetic, lipid profile (including total cholesterol, high-density lipoprotein [HDL] cholesterol, low-density lipoprotein [LDL] cholesterol, and triglycerides), smoking status, and family history of diabetes and CVD was collected. Patients on clozapine under the care of the PEPP routinely undergo 6-monthly assessment of physical health, including the above parameters, as well as estimation of plasma levels of clozapine and norclozapine. For purposes of this study, at the time of their routine blood work, an extra sample of blood was collected for genetic testing. This study was approved by the Research Ethics Board of the Western University (London, Ontario).
Drug-Level Analysis
Drug-level analysis was performed as part of the therapeutic drug monitoring for patients by a clinical laboratory. Time of last dose and time of blood withdrawal are not routinely recorded for patients, and thus hours after the last clozapine dose are unknown, but the majority of patients (87%) were reported to take clozapine once daily at bedtime, and blood was taken the following day at their clinic visit, suggesting >12 hours postdose. Clozapine and norclozapine plasma concentrations were measured by liquid chromatography–mass spectrometry (LC-MS). Plasma aliquots were precipitated with phosphate buffer (pH 7) and organic solvent containing the internal standard (d8-clozapine), then evaporated to dryness and reconstituted in injection buffer (15% methanol in ammonium formate buffer, pH 3) for analysis by LC-MS (Agilent 1100 Series; Agilent, Santa Clara, CA, USA). The single quadruple LC-MS was equipped with a Phenomenex Kinetex (Torrance, CA, USA) column (50 × 3 mm, 5 micron). Analytes were separated using the mobile phase (38% methanol in 25 mM ammonium formate buffer, pH3) at 0.9 mL/min. Typical between-day coefficients of variation for the analysis were in the range of 3% for clozapine and 6% for norclozapine.
Genotyping
The single-nucleotide polymorphisms (SNPs) of the most functionally relevant genes were selected on the basis of their potential linkage to clozapine metabolism and its metabolic side effects shown in previous studies as well as reported allelic frequency >5% (see Table 1 and www.cypalleles.ki.se). Genetic analysis was conducted to determine SNPs of the drug-metabolizing enzymes CYP1A2 (rs762551, rs2069514, rs35694136) and CYP2C19 (rs4244285, rs12248560), the drug transporter P-glycoprotein/ABCB1 (rs1128503), central serotonin receptor HTR2C (rs3813929, rs3813928, rs1414334), and LEP (rs7799039) and LEPR (rs1137101). A CYP2C19 intermediate metabolizer (IM) was defined by a CYP2C19*1/*2 or *17/*2 genotype, a poor metabolizer (PM) by a CYP2C19*2/*2 genotype, an ultrarapid metabolizer (UM) by a CYP2C19*17/*17 genotype, and an extensive metabolizer (EM) in the absence of *2 and *3 variant alleles (*1/*1 genotype or wildtype) or a *1/*17 genotype. Genomic DNA was isolated from venous blood samples with the Gentra Puregene extraction kit (Qiagen, Alameda, CA, USA) and concentration determined by PicoGreen (Invitrogen, Carlsbad, CA, USA). Genotyping was performed by TaqMan Assays (Applied Biosystems, Carlsbad, CA, USA) using a 7500 Real-time PCR system (Applied Biosystems). Hardy-Weinberg equilibrium (HWE) was tested using the χ2 method. All genotypes were in Hardy-Weinberg equilibrium with the exception of HTR2C rs3813929 (P < 0.001), rs3813928 (P < 0.001), and rs1414334 (P < 0.001).
Allelic Frequency of Single-Nucleotide Polymorphisms (SNPs) of Candidate Genes Analysed (n = 60).
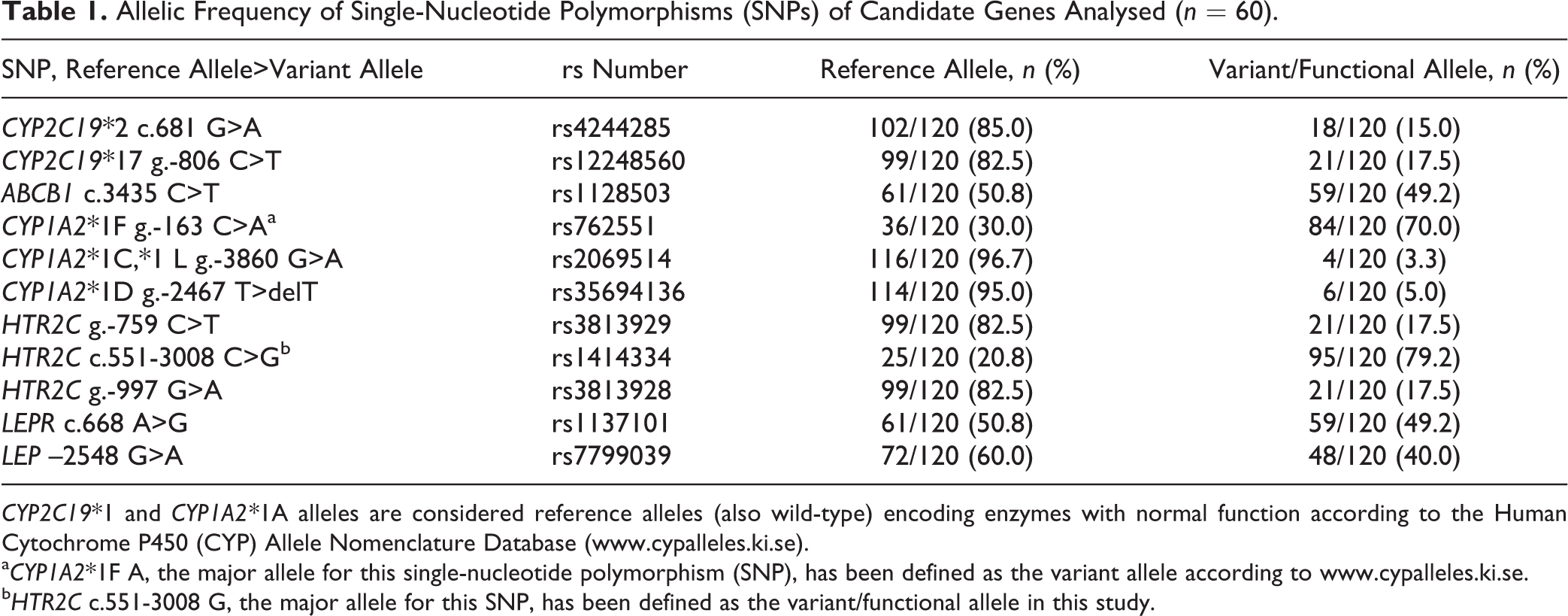
CYP2C19*1 and CYP1A2*1A alleles are considered reference alleles (also wild-type) encoding enzymes with normal function according to the Human Cytochrome P450 (CYP) Allele Nomenclature Database (www.cypalleles.ki.se).
aCYP1A2*1F A, the major allele for this single-nucleotide polymorphism (SNP), has been defined as the variant allele according to www.cypalleles.ki.se.
bHTR2C c.551-3008 G, the major allele for this SNP, has been defined as the variant/functional allele in this study.
Statistical Analysis
Statistical analysis was performed using the statistical software R 30 and GraphPad Prism (V5.02; GraphPad Software, La Jolla, CA, USA). Differences in clozapine concentration, dose-normalized concentration, parent-to-metabolite ratio, and clinical variables were analyzed by Kruskal-Wallis test and Dunn’s multiple-comparisons test or Mann-Whitney test, as appropriate. Descriptive statistics were used to summarize patient characteristics. Allelic, additive, dominant, and recessive models were considered for each SNP and the genetic model that best describes the outcomes in linear regression models chosen. Multivariable linear regression models were used to evaluate the association of SNPs with drug concentration and metabolic side effects (BMI, blood sugar, and lipids) after log transformation, controlling for the confounding factors age, sex, concurrent medication with known effects on weight increase (quetiapine, valproate, lithium, risperidone, and olanzapine), smoking status, personal and family history of diabetes, and CVD. Ethnicity was not a significant covariate in any model and thus has been omitted in our analyses. We also tested for associations with metabolic syndrome (diagnosed when individuals have high waist circumference with 2 or more abnormal parameters including triglycerides, HDL cholesterol, blood pressure, and fasting glucose) defined by Adult Treatment Panel (ATP) III criteria (American Heart Association) using multivariable logistic regression models. 31 Adjustment of the significance threshold for multiple testing of the 11 SNPs genotyped was not applied due to the exploratory character of this pilot study. We used a statistical significance level of 0.05 for testing; however, we thought it was worth reporting P values close to 0.05 as these are marginally significant.
Results
The sample included patients with schizophrenia (n = 46) and schizoaffective (n = 13) and delusional disorders (n = 1). The majority of subjects were Caucasian (93%); the remainder were Asians (n = 2) and First Nations (n = 2). Thirty percent of patients were on clozapine monotherapy. Antipsychotic medications were co-prescribed in 61.9% of patients (olanzapine, risperidone, aripiprazole, ziprasidone, quetiapine, lurasidone, or asenapine), 21.4% were on a mood stabilizer (valproate, lithium, carbamazepine), and 14.3% were taking antidepressants (sertraline, paroxetine, venlafaxine, fluoxetine, citalopram, and escitalopram). Table 2 summarizes the clinical characteristics of patients and their distribution among those with and without metabolic syndrome; in particular, 52% of the patients had metabolic syndrome, 51% were obese (BMI >30), and an additional 30% were overweight (BMI 25-30). The observed allelic frequencies for genetic markers as presented in Table 1 are similar to those previously reported in white populations.
Clinical Characteristics of Patients and Their Distribution among Those with and without Metabolic Syndrome.
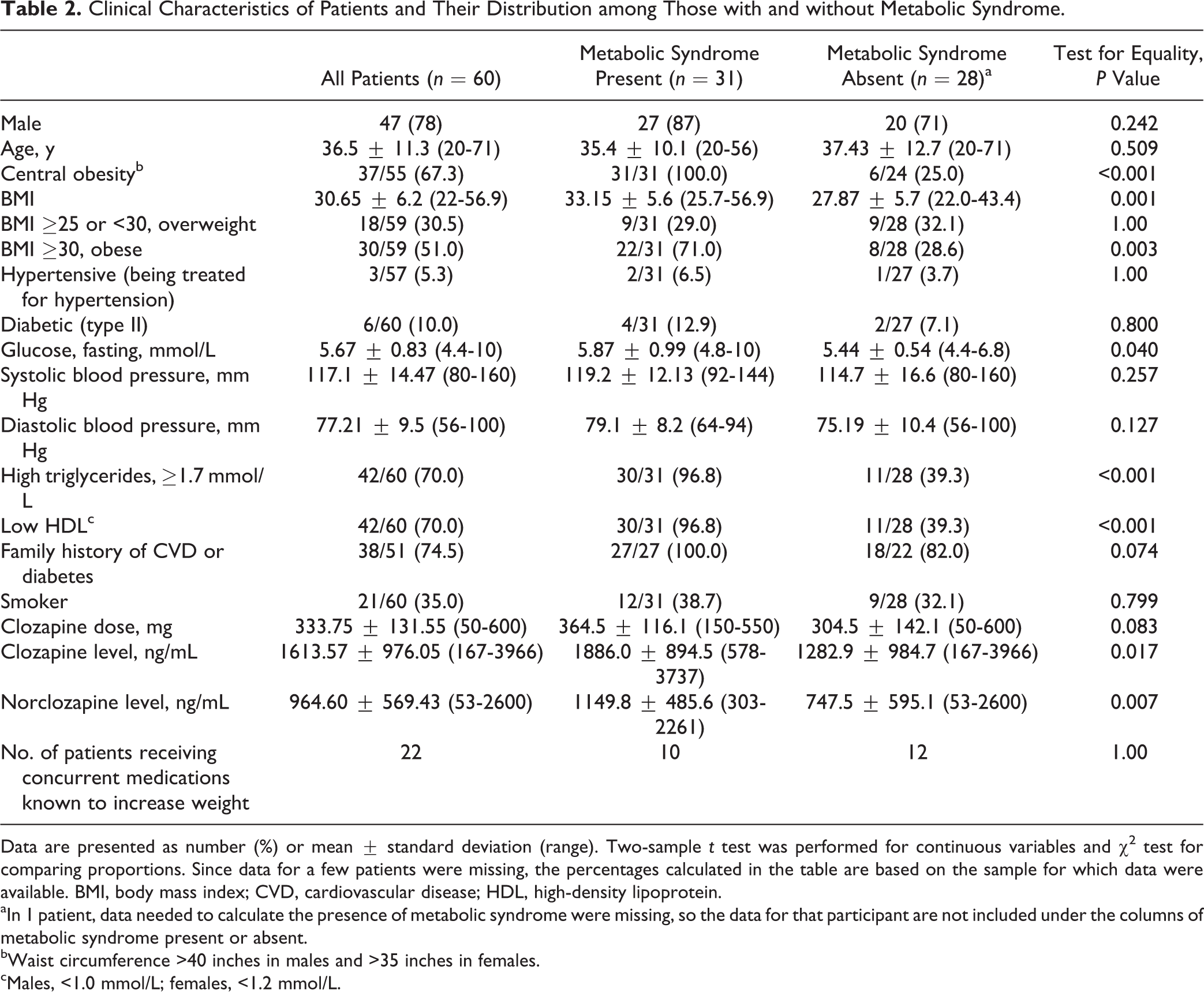
Data are presented as number (%) or mean ± standard deviation (range). Two-sample t test was performed for continuous variables and χ2 test for comparing proportions. Since data for a few patients were missing, the percentages calculated in the table are based on the sample for which data were available. BMI, body mass index; CVD, cardiovascular disease; HDL, high-density lipoprotein.
aIn 1 patient, data needed to calculate the presence of metabolic syndrome were missing, so the data for that participant are not included under the columns of metabolic syndrome present or absent.
bWaist circumference >40 inches in males and >35 inches in females.
cMales, <1.0 mmol/L; females, <1.2 mmol/L.
Determinants of Metabolic Side Effects Associated with Clozapine Therapy
Metabolic syndrome as well as known predictors of metabolic syndrome such as BMI, lipids, and fasting glucose levels were evaluated as clinically relevant metabolic side effects. Genetic and clinical determinants of BMI are summarized in Table 3. LEP –2548 G>A, LEPR c.668 A>G, and –HTR2C c.551-3008 C>G were identified as genetic predictors of BMI and explained, together with concurrent medications known to increase BMI, 30.5% of the observed variability (Table 3). BMI decreased with increasing number of variant alleles in these 3 genes, LEP –2548 A, LEPR c.668 G, and HTR2C c.551-3008 G (P = 0.0479; Figure 1A). Concurrent medications known to increase BMI resulted in a 1.11-fold difference in BMI compared to patients who did not take these medications (32.7 vs. 29.4, respectively; Figure 1B). Clozapine dose and blood level were not associated with BMI (Table 3).
Genetic and Clinical Determinants of BMI: Linear Regression Model Coefficients from Prospective Cohort of 60 Patients Prescribed Clozapine.
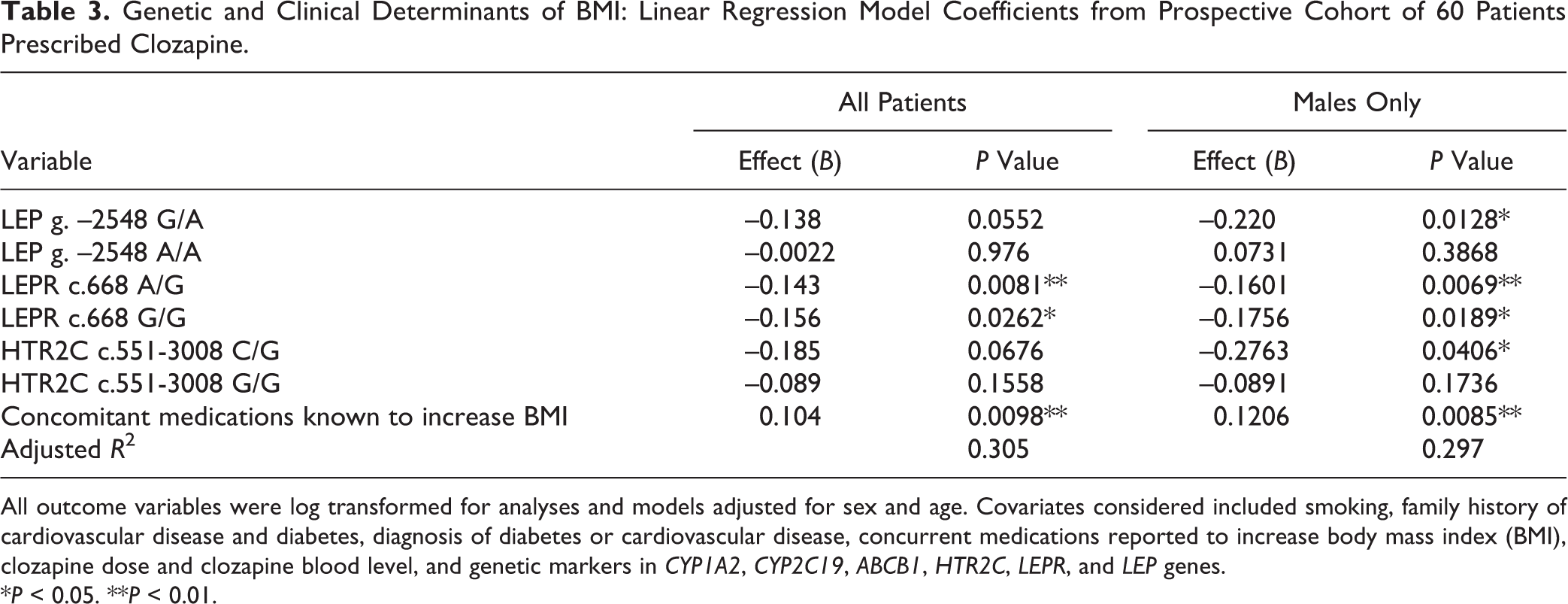
All outcome variables were log transformed for analyses and models adjusted for sex and age. Covariates considered included smoking, family history of cardiovascular disease and diabetes, diagnosis of diabetes or cardiovascular disease, concurrent medications reported to increase body mass index (BMI), clozapine dose and clozapine blood level, and genetic markers in CYP1A2, CYP2C19, ABCB1, HTR2C, LEPR, and LEP genes.
*P < 0.05. **P < 0.01.
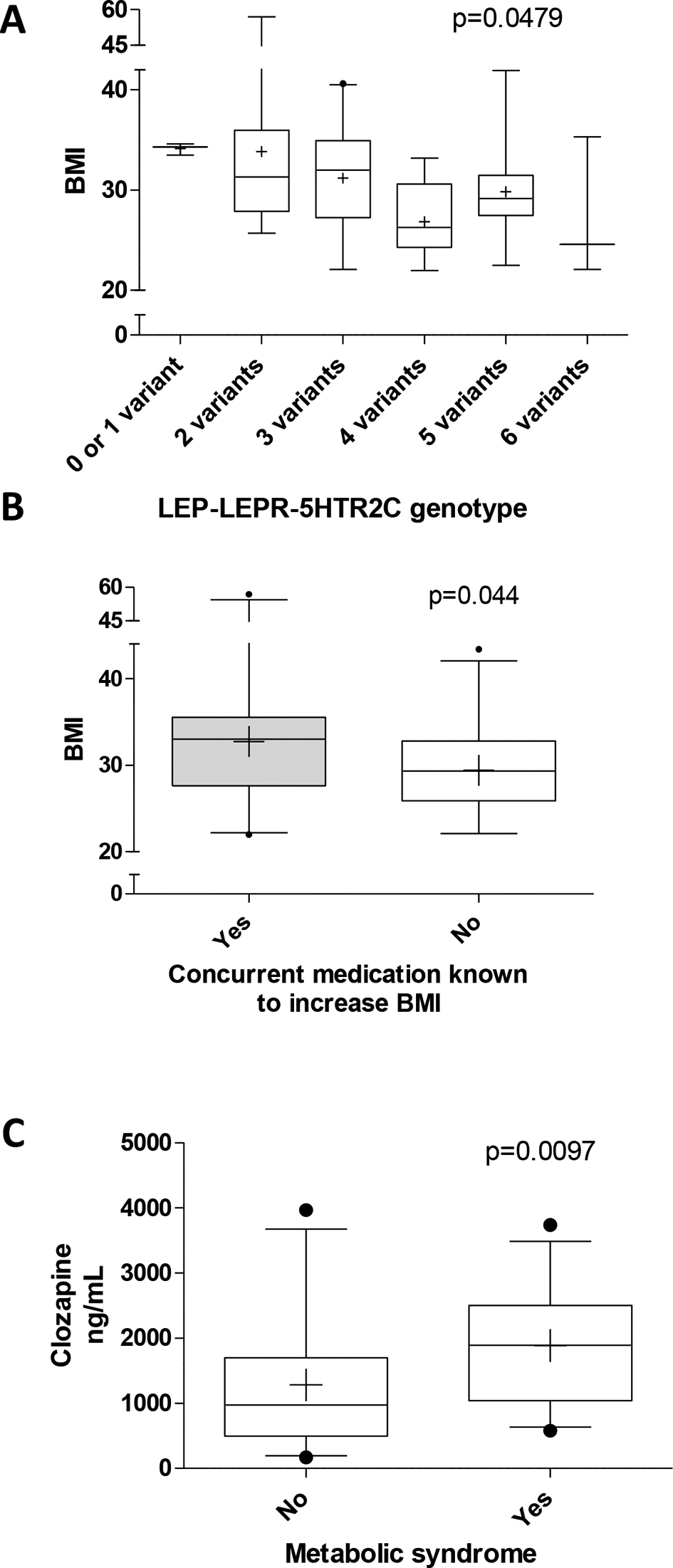
Effect of genetic variation, clinical factors, and clozapine exposure on BMI and metabolic syndrome. (A) Cumulative effect of variant alleles, including LEP –2548 A, LEPR c.668 G, and HTR2C c.551-3008 G on BMI. (B) Effect of concurrent medications known to increase BMI. (C) Clozapine blood levels in patients with or without metabolic syndrome. Statistical analyses were performed with Kruskal-Wallis test with Dunn’s multiple-comparison or Mann-Whitney test as appropriate. BMI, body mass index; HTR2C, serotonin receptor; LEP, leptin; LEPR, leptin receptor.
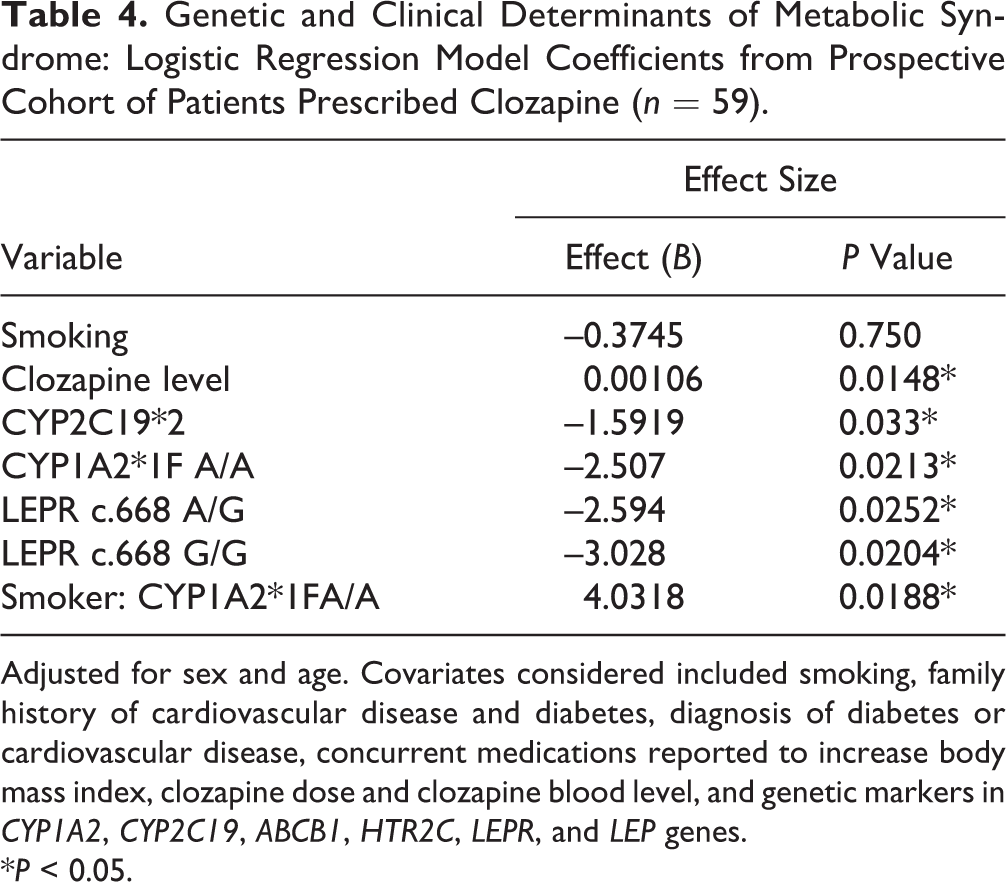
Metabolic syndrome was found to be associated with the presence of at least 1 CYP2C19*2 allele (P = 0.033) and the LEPR c.668 A/G (P = 0.0252) and G/G (P = 0.0204) genotypes, which resulted in a 80%, 92.5%, and 95.2% lower odds, respectively, compared to their reference genotypes as indicated by logistic regression analysis (Table 4). Also, a significant interaction was observed between smoking and CYP1A2 *1F A/A carrier status (P = 0.028). Our findings suggest that clozapine-treated smokers with the CYP1A2*1F A/A genotype have a 4.6 times higher odds for metabolic syndrome, whereas nonsmokers with CYP1A2*1F A/A have an about 92% lower odds compared to CYP1A2 C/C carriers. Importantly, clozapine blood level was significantly associated with a higher risk for metabolic syndrome, with an 11% increase in the odds per 100 ng/mL (Table 4). Clozapine blood levels in patients with metabolic syndrome were about 1.5 times higher compared to those without metabolic syndrome (1886.0 ± 894.5 vs. 1282.9 ± 984.7, P = 0.0097; Figure 1C). Aside from known demographic and clinical predictors of lipids or fasting glucose, no association was found for the genetic variants studied here (data not shown).
Genetic and Clinical Determinants of Metabolic Syndrome: Logistic Regression Model Coefficients from Prospective Cohort of Patients Prescribed Clozapine (n = 59).
Adjusted for sex and age. Covariates considered included smoking, family history of cardiovascular disease and diabetes, diagnosis of diabetes or cardiovascular disease, concurrent medications reported to increase body mass index, clozapine dose and clozapine blood level, and genetic markers in CYP1A2, CYP2C19, ABCB1, HTR2C, LEPR, and LEP genes.
*P < 0.05.
Determinants of Clozapine Exposure
Clozapine plasma level was highly correlated with norclozapine level (r = 0.873) (Figure 2A). Patients with 2 CYP2C19*2 alleles (poor metabolizer [PM] phenotype; Figure 2B) showed a trend to higher dose-normalized clozapine concentration (P = 0.1056) and a significantly increased clozapine to norclozapine ratio (P = 0.0237), suggesting enhanced systemic exposure due to impaired clozapine metabolism. No significant differences were observed for CYP1A2 or ABCB1 genotypes (data not shown). In addition, smokers had a lower clozapine concentration (P = 0.064, marginally significant) and metabolic ratio (P = 0.044) compared to nonsmokers (Figure 2C), suggesting induction of CYP1A2 metabolism by smoking.
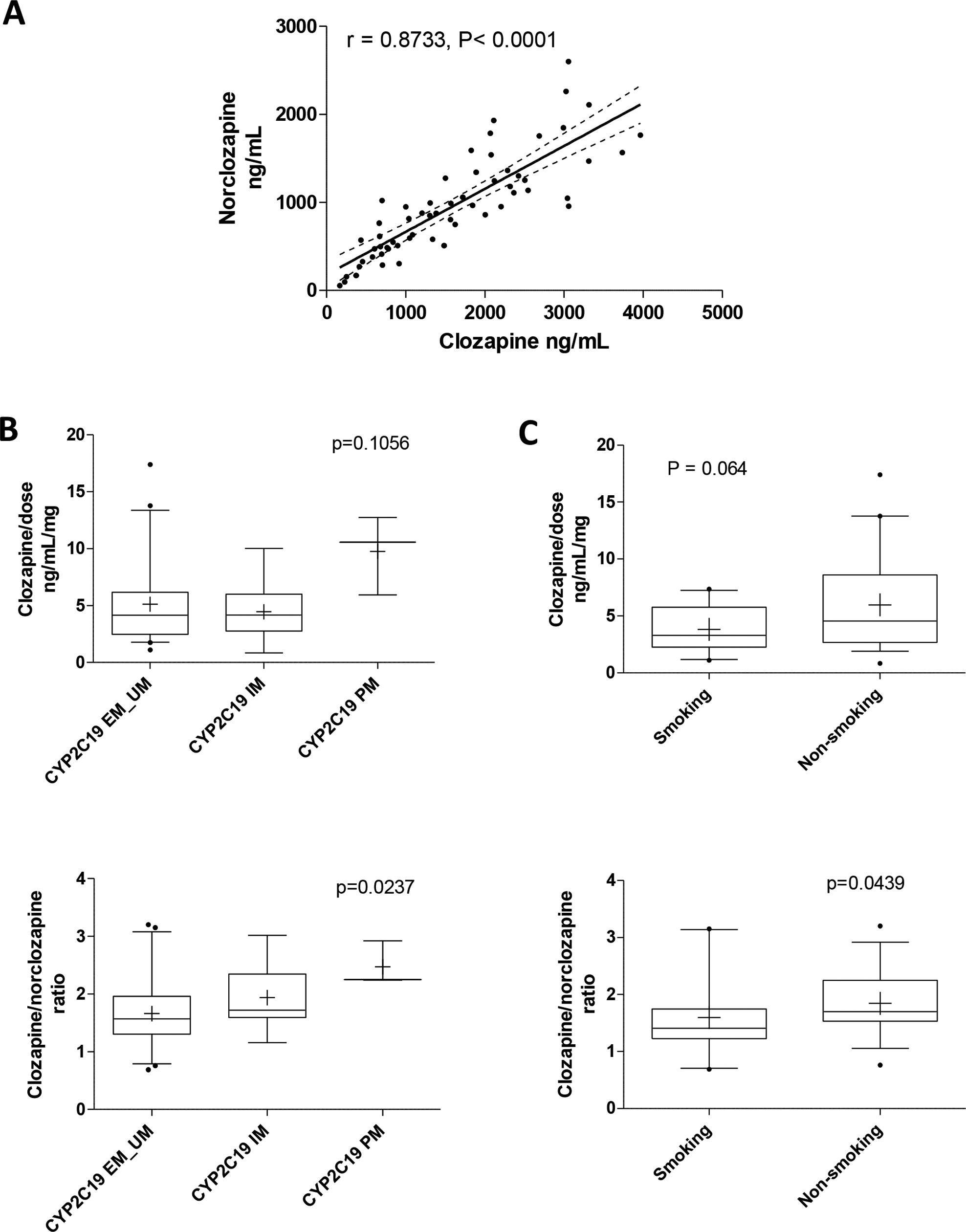
Characteristics of systemic drug exposure in patients prescribed clozapine (n = 60). (A) Spearman correlation between clozapine and norclozapine drug level. (B) Effect of CYP2C19 phenotype and (C) smoking status on dose-normalized clozapine levels and clozapine to norclozapine ratio. Statistical analyses were performed with Kruskal-Wallis test with Dunn’s multiple-comparison or Mann-Whitney test as appropriate. EM, extensive metabolizer; IM, intermediate metabolizer; PM, poor metabolizer; UM, ultrarapid metabolizer.
Multivariable linear regression analysis confirmed CYP2C19 genotype as a major determinant of clozapine level after adjustment of other covariates, indicating that CYP2C19 PM status is associated with higher blood level (Table 5). Furthermore, a significant interaction was observed between smoking and CYP1A2*1F g.-163 C/A carrier status (P = 0.0499). Compared to non-CYP1A2*1F carriers (C/C genotype), the presence of 1 CYP1A2*1F A risk allele resulted in a 0.71 times lower clozapine level among smokers while enhancing exposure among nonsmokers 2.8-fold. The interaction between smoking and CYP1A2 g.-163 A/A carrier status was not significant (Table 5). After adjustment for age and sex, CYP2C19 genotype, CYP1A2*1F-smoking interaction, clozapine dose, and BMI explained 35.2% of the observed variability in clozapine blood level in our final model. Previously reported predictors of clozapine exposure such as CYP1A2 and ABCB1 genotype were not significantly associated with clozapine level in this model (Table 5). Assessment of clozapine to norclozapine ratio by multiple linear regression further supported a role of CYP2C19*2 genotype (P = 0.027) and smoking (P = 0.052, marginally significant).
Genetic and Clinical Determinants of Blood Clozapine Concentration: Linear Regression Model Coefficients from Prospective Cohort of Patients Prescribed Clozapine.
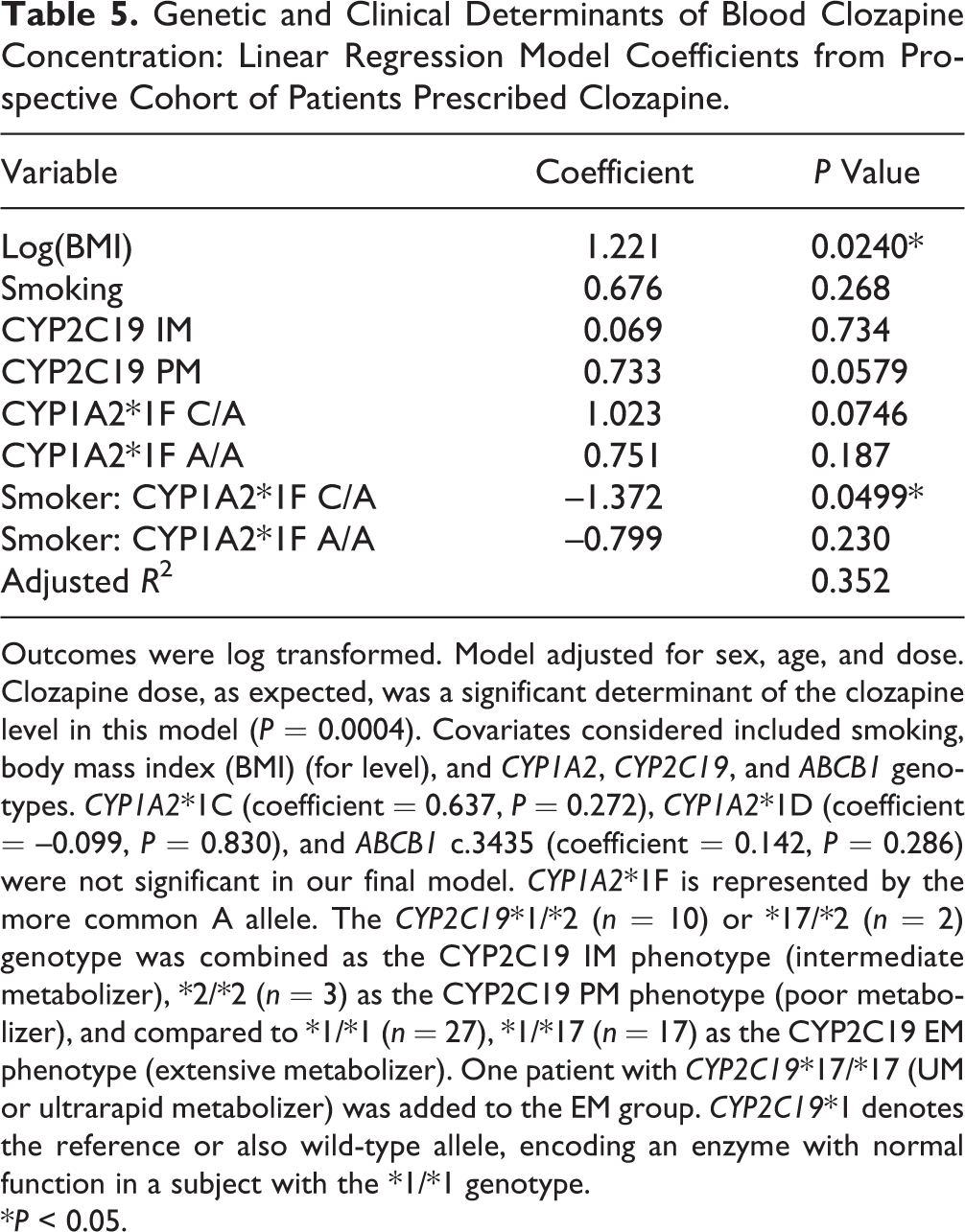
Outcomes were log transformed. Model adjusted for sex, age, and dose. Clozapine dose, as expected, was a significant determinant of the clozapine level in this model (P = 0.0004). Covariates considered included smoking, body mass index (BMI) (for level), and CYP1A2, CYP2C19, and ABCB1 genotypes. CYP1A2*1C (coefficient = 0.637, P = 0.272), CYP1A2*1D (coefficient = –0.099, P = 0.830), and ABCB1 c.3435 (coefficient = 0.142, P = 0.286) were not significant in our final model. CYP1A2*1F is represented by the more common A allele. The CYP2C19*1/*2 (n = 10) or *17/*2 (n = 2) genotype was combined as the CYP2C19 IM phenotype (intermediate metabolizer), *2/*2 (n = 3) as the CYP2C19 PM phenotype (poor metabolizer), and compared to *1/*1 (n = 27), *1/*17 (n = 17) as the CYP2C19 EM phenotype (extensive metabolizer). One patient with CYP2C19*17/*17 (UM or ultrarapid metabolizer) was added to the EM group. CYP2C19*1 denotes the reference or also wild-type allele, encoding an enzyme with normal function in a subject with the *1/*1 genotype.
*P < 0.05.
Discussion
This study examined the prevalence and pharmacokinetic and pharmacodynamic genetic risk factors associated with clozapine-induced metabolic side effects in a cohort of 60 patients who were prescribed clozapine for a minimum of 6 months, taking into account drug exposure as well as other potential confounders such as concurrent medications known to increase body weight as well as smoking. Our findings show that about two-thirds of the patients on clozapine were overweight or obese and had abnormal triglycerides and HDL cholesterol, with about half of the patients having metabolic syndrome. These results are similar to those reported by others. 11 One important finding that merits discussion is the high prevalence of antipsychotic polypharmacy (61.9%) with clozapine. The evidence-based report of the Canadian Agency for Drugs and Technologies in Health (CADTH) clearly indicated that high-dose or combination antipsychotic strategies are not known to be more effective and may be more harmful. 32 Our study confirms that concurrent use of medications known to increase BMI significantly contributed to the increased BMI in patients on clozapine. The ‘Choosing Wisely Canada’ guidelines for psychiatry encourages physicians to not routinely prescribe high-dose or combination antipsychotic treatment strategies in the treatment of schizophrenia. 33
With respect to genetic markers, in our study, 3 SNPs related to leptin and serotonin signalling were identified as key genetic predictors of BMI: LEP –2548 G>A, LEPR c.668 A>G, and HTR2C c.551-3008 C>G. An increasing number of variant alleles in these 3 genes were associated with decreased BMI, suggesting a protective effect. Previous cross-sectional and longitudinal studies have assessed the relationship of HTR2C, LEP, and LEPR polymorphisms with metabolic outcomes in psychiatric patients receiving antipsychotic medications, but the evidence remains equivocal, 12,34 likely due to differences in study design, treatment duration, sex, and assessed confounders. Our findings are consistent with previous studies in patients on olanzapine or clozapine that support associations between the LEP –2548 G allele and increase in weight or BMI. 35 –40 Only 2 other studies, 41,42 one in an Asian population and the other in children and adolescents, reported that the LEP-2548 A allele was associated with weight gain, but participants in those studies received risperidone or antipsychotic medications other than olanzapine or clozapine, suggesting that clozapine and olanzapine may share a unique mechanism of causing weight gain compared to other atypical antipsychotic medications. 43,44 Previously, a significant interaction between HTR2C and LEP variants with respect to obesity and metabolic syndrome has been reported. 35,40 Yevtushenko et al. 35 showed that among patients (n = 134, on olanzapine, clozapine, and risperidone) with the HTR2C –759C/CC genotype, those with the LEP –2548G allele had a higher BMI, larger waist circumference, and presence of metabolic syndrome compared to AA carriers. Another study showed that the combined absence of the HTR2C T allele and presence of the LEP-2548 G allele was associated with obesity in patients on atypical antipsychotic medications. 40
Previously, associations of LEPR c.668A>G (Q223 R; rs1137101) with weight gain or BMI also have been shown but only when sex or LEP variants are taken into consideration. 19,36 Gregoor et al. 19 observed that female carriers of LEPR 223QQ had higher rates of obesity compared to R carriers in a cross-sectional study of 200 patients (67% male) on different antipsychotic medications, including clozapine, olanzapine, and risperidone. A small prospective study reported a significant interaction between the G alleles of LEPR and LEP genes and greater weight gain at higher serum concentrations of olanzapine. 36 In contrast, another cross-sectional study in patients prescribed clozapine (n = 56, 78% males) did not find an association between LEP and LEPR variants and frequency of metabolic syndrome or obesity when comparing LEP-2548 G/G versus A allele carriers and LEPR 223 Q/Q versus R allele carriers, but LEPR 223Q/Q predicted lower triglyceride levels in males after controlling for dose. 45 Similarly, 2 other studies 46,47 found no associations between LEPR variants and change in BMI or weight gain or waist circumference, but the assessed populations were on multiple antipsychotic medications, including clozapine 46 or excluding clozapine. 47 Since the sample in our study consisted of predominantly male patients, it is difficult to draw any conclusions regarding the effect of sex on the association of LEP or LEPR polymorphism with BMI (Table 3).
In our study, metabolic syndrome was found to be associated with the presence of at least 1 CYP2C19*2 allele and the LEPR c.668 A/G and G/G genotypes, which resulted in a 80%, 92.5%, and 95.2% lower odds, respectively, compared to their reference genotypes as indicated by logistic regression analysis. In addition, clozapine blood level significantly predicted a higher risk for metabolic syndrome. Since there was a clear association of clozapine level with increased risk, the negative correlation of CYP2C19 PM status with a lower odds for metabolic syndrome remains difficult to explain. However, CYP1A2 has been reported as the main metabolic pathway for clozapine, and thus additional metabolic effects mediated by CYP1A2 that were not captured with the evaluated genotypes may play a role. 27,29
An association of clozapine level with metabolic side effects, including BMI, insulin resistance, and glucose and lipid levels, has been reported in 2 previous studies. In 17 clozapine-treated patients, Melkersson et al. 28 found that higher serum clozapine concentration was linked with a higher risk of developing insulin and lipid elevation on a given dose of clozapine. Similarly, in a retrospective survey of 100 patients on clozapine, increased drug exposure in women was associated with a higher BMI and glucose concentration. 48 Taken together, these finding suggest that clozapine plasma level, rather than dose, may be a suitable tool to monitor the risk of metabolic side effects associated with clozapine.
Although we observed a linear relationship between clozapine level and prescribed dose (r = 0.4055, P = 0.0013), clozapine level was highly variable among patients, supporting previous reports. 49 Clinical and genetic factors identified in our study explained 35% of the observed variability. Our findings confirm the role of CYP2C19 polymorphism in the pharmacokinetics of clozapine as previously shown by in vitro and in vivo studies. 26,29 After controlling for other covariates, CYP2C19 PM status was found to be associated with higher blood level of clozapine. In contrast to previous reports, CYP1A2 and ABCB1 genotype as well as smoking were not found to be significantly associated with clozapine level in our study. 12,28,50 However, a significant interaction was observed between smoking and CYP1A2*1F g.-163 CA carrier status. CYP1A2 AA genotype is thought to enhance inducibility and therefore increased enzymatic activity, 51,52 and smoking is a known inducer of CYP1A2. 29,53,54 This may explain the observed lower clozapine exposure among smokers carrying the CYP1A2*1F polymorphism. 51,52 However, there are conflicting reports on the influence of CYP1A2*1F polymorphism on clozapine blood levels. 51,52,55,56
To our knowledge, this is the first study in a primarily Caucasian population that specifically targeted clozapine to investigate the genetic factors in both drug metabolism and response contributing to its metabolic side effects, taking into account clozapine blood concentration as well as important confounders. Many previously reported studies lack information regarding treatment duration, dose, or concomitant medications or simultaneously assess multiple atypical antipsychotics, thus making it difficult to interpret results. Patients in our study were enrolled after at least 6 months of clozapine therapy, and prescribed dose and blood level were obtained. The detailed data collected here helped control for the majority of the clinical confounding factors affecting metabolic outcomes, including age, sex, concurrent medications with known effects on weight increase, smoking status, personal and family history of diabetes, and CVD. We also examined the effects of genetic polymorphisms on other metabolic parameters, including glucose and lipid profile.
Limitations of our study include its cross-sectional nature and a relatively small sample size, which made it difficult to analyze sex differences or effects of more rare genotypes such as CYP1A2*1C and *1D. Many previous studies have similar patient numbers ranging from 37 to 74 subjects, with more than two-thirds being males. 36,37,39,42,45 Absence of a patient control group not prescribed clozapine made it difficult to account for the contributory effects of other concomitant atypical antipsychotics on BMI and metabolic syndrome, but this was in part overcome by including concurrent medications with known effects on weight gain in the applied regression model. We did not collect data on severity of symptoms, which may influence medication adherence and hence plasma levels of clozapine. Genotype distribution in a disease population (i.e., schizophrenia) may differ from the general population, leading to deviation of HWE as observed for the 3 assessed HTR2C SNPs. Although we have to use caution if SNPs violating HWE are included in the analyses because they might result in spurious results, they can be truly important SNPs hovering in the disease population and associating with outcome of interest.
In conclusion, this exploratory study confirms the high prevalence of metabolic side effects in patients on clozapine, with higher clozapine plasma levels being an important risk factor for metabolic syndrome. Pharmacogenetic markers in genes encoding CYP2C19, leptin, leptin receptor, and HTR2C receptor were identified as main predictors of metabolic syndrome as well as BMI. Clinicians should be aware of potentially increasing a patient’s risk when co-prescribing antipsychotic medications with clozapine and use clozapine plasma level to minimize the risk of metabolic syndrome while titrating the dose to the desired therapeutic effect. Larger controlled prospective studies are warranted to confirm clinical implications of these findings.
Footnotes
Acknowledgements
We thank Norman B. Smith, PhD, FCACB, Section Head, Toxicology & Therapeutic Drug Monitoring, Biochemistry, Pathology & Laboratory Medicine London Health Sciences Centre & St. Joseph’s Health Care London, for analyzing the blood clozapine level and providing analytical method details. We also thank Deborah Windell, research associate with PEPP, for helping with data collection and analysis, and Dr. Rommel Tirona for helpful suggestions for the manuscript.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: KV was supported for this project by the Academic Medical Organisation of Southwestern Ontario (AMOSO) Opportunities Fund. RBK is supported by the Wolfe Medical Research Chair in Pharmacogenomics.
