Abstract
Objectives
Human endogenous retroviruses (HERVs) are integral components of the human genome, and their reactivation has been implicated in the pathogenesis of some malignancies. External viral co-infections are suspected to play a role in HERV transactivation. This study aimed to investigate the expression of HERV-K np9 elements and HERV-R env gene in pediatric acute lymphoblastic leukemia (ALL) patients. Additionally, we explored potential correlations between HERV expression and common viral infections prevalent in this group of patients.
Methods
Blood samples were collected from 43 pediatric ALL patients and 48 age- and sex-matched healthy controls. Quantitative real-time PCR (qRT-PCR) was used to assess the expression of HERV-K np9 and HERV-R env, along with herpes simplex virus (HSV), parvovirus B19, and polyomavirus BK.
Results
HERV-K np9 and HERV-R env showed significantly higher expression in the peripheral blood of ALL patients compared to healthy controls (p < .001 and p = .003, respectively). HSV positivity was associated with significantly increased HERV-K np9 expression. No significant correlations were observed between other investigated viruses and HERV gene expression.
Conclusion
The overexpression of HERV-K np9 and HERV-R env in pediatric ALL patients suggest their potential role in leukemogenesis. Our findings also suggest a possible link between HSV infection and HERV reactivation in this population. Future investigations are needed to understand the precise roles of these genes and viral infections in the development of ALL.
Introduction
Human endogenous retroviruses (HERVs), which constitute roughly 8% of the human genome, represent remnants of ancient retroviral infections that integrated into human germline millions of years ago. 1 Although HERVs remain silent for many years, they retain the potential to reactivate and contribute to various diseases. Numerous studies have implicated HERV reactivation in autoimmune disorders such as rheumatoid arthritis 2 and malignancies such as breast cancer 3 and acute and chronic leukemias.4,5 Within the diverse HERV families, HERV-K stands out for its unique characteristics. Several factors including its recent arrival, exhibiting minimal corruption and the high levels of transcriptional activity, make it a prime suspect for potential oncogenic activity. 6
Acute lymphoblastic leukemia (ALL) is the most common pediatric leukemia, with the highest incidence occurring in children under 5 years old.7,8 This disease itself, or its treatment with chemotherapy, can lead to significant immunodeficiency in children. 9 This immunocompromised state increases susceptibility to the reactivation of latent infections, such as herpes simplex virus (HSV) and polyomavirus BK (BKV).10,11 Additionally, immunocompromised patients are more likely to experience persistent parvovirus B19 (B19 V) infection. 12
Given the established link between HERV-K np9 and HERV-R env and malignancies,4,5,13 this study investigated the expression patterns of specific HERV genes/elements in pediatric ALL patients. We focused on HERV-K np9, a spliced product of the env gene implicated in cancer development,14,15 and HERV-R env, known for its high expression in many cancer types such as colon, liver, stomach, esophageal, cervical, and ovarian cancers. 1 In addition, we aimed to explore potential associations between the reactivation of these HERV-K and R genes and infections with prevalent viruses in ALL patients, such as herpes simplex virus, human parvovirus B19, and polyomavirus BK.
Subjects and methods
Subjects
This cross sectional study recruited pediatric patients diagnosed with acute lymphoblastic leukemia (ALL) at the Pediatric Oncology Department of South Egypt Cancer Institute (SECI), Assiut University, Egypt. Patients were age- and sex-matched with healthy controls. To ensure a homogenous study population, only patients prior to chemotherapy initiation were included, including both newly diagnosed and relapsed cases. Patients under active chemotherapy were excluded. Written informed consent was obtained from legal guardians in accordance with the ethical approval granted by the local ethics committee of SECI.
Methods
Two-milliliter blood samples were collected from each participant. Plasma, separated from the first EDTA-anticoagulated sample, was frozen for subsequent viral nucleic acid detection. Peripheral blood mononuclear cells (PBMCs) were isolated from the second EDTA blood sample for HERV-K np9 and HERV-R env expression analysis. PBMCs were enriched via centrifugation on a Ficoll-Paque (GE Healthcare) density gradient. Total RNA was subsequently extracted from the isolated PBMCs using the RNAeasy Mini Kit (Qiagen) according to the manufacturer’s protocol. cDNA synthesis was then performed using the RevertAid First Strand cDNA Synthesis Kit (ThermoFisher Scientific), adhering to the provided instructions.
Quantitative real-time PCR
Quantitative real-time PCR (qRT-PCR) analysis was used to quantify messenger RNA (mRNA) expression levels. Assays for HERV-K np9 and HERV-R env were run on an Applied Biosystems 7500 real-time PCR system using Maxima SYBR Green qPCR Master Mix (ThermoFisher Scientific). Gene expression levels were normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as a housekeeping gene. Primer sequences for all targets are provided in Supplemental Table 1. The ΔΔCt method was utilized for relative quantification and data analysis, with results expressed as fold change. 16 Following data acquisition, the control samples were automatically incorporated into the analysis, and their mean CT value was calculated for the entire run. Subsequently, the CT value of each individual sample was compared to the mean CT value of the controls to determine the relative expression level of each target gene.
Viral DNA extraction
Viral DNA extraction was performed on previously frozen plasma samples following thawing, using The GeneJET Viral DNA and RNA Purification Kit (ThermoFisher Scientific).
Detection of herpes simplex virus and Parvo B19 virus and second step of semi-nested PCR of Polyoma virus
Qualitative real-time PCR was employed for viral detection, utilizing Maxima SYBR Green PCR Master Mix (ThermoFisher Scientific) on an Applied Biosystems 7500 real-time system. Primer sequences used for these reactions are detailed in Supplemental Table 1. Each run incorporated positive controls obtained from previously confirmed patients, alongside a non-template control to verify the absence of spurious amplification.
Polyomavirus detection was performed through semi-nested PCR using DreamTaq Green PCR Master Mix (ThermoFisher Scientific) on an Applied Biosystems ARKTIK thermal cycler (ThermoFisher Scientific).
Statistical analysis
Data entry and analysis were conducted using the IBM SPSS 20.0 statistical software package (IBM; Armonk, New York, USA) and GraphPad Prism version 8.0.2 (GraphPad, San Diego, CA). Data normality was assessed with Shapiro-Wilk and Kolmogorov-Smirnov tests. Quantitative non-parametric data were reported as median, minimum, and maximum range, while categorized data were presented as both number and percentage. The non-parametric Mann-Whitney U test and Kruskal-Wallis test were used for comparisons between independent groups, while the Chi-square test and Fisher’s exact test were employed for comparisons of categorical variables. Binary and multiple logistic regression analyses were performed to evaluate the combined influence of independent variables on the target (dependent) variable. Statistical significance was established at a p-value less than 0.05.
Results
General patients’ characteristics
The study included 43 pediatric patients with ALL and 48 age- and sex-matched healthy controls. The median age of the ALL patients was 7 years, with 60.5% being male. Among the ALL cases, 33 were classified as B-ALL, including 10 newly diagnosed patients. Additionally, 12 patients reported a family history of cancer, and five exhibited BCR-ABL translocations confirmed by conventional cytogenetic and/or molecular techniques. Supplemental Table 2 provides further details about the clinical and demographic characteristics of the participants.
Gene expression analysis of HERV-R env and HERV- K np9 in patients and controls
Expression of HERV-R env gene and HERV- K np9 elements in ALL cases and controls.
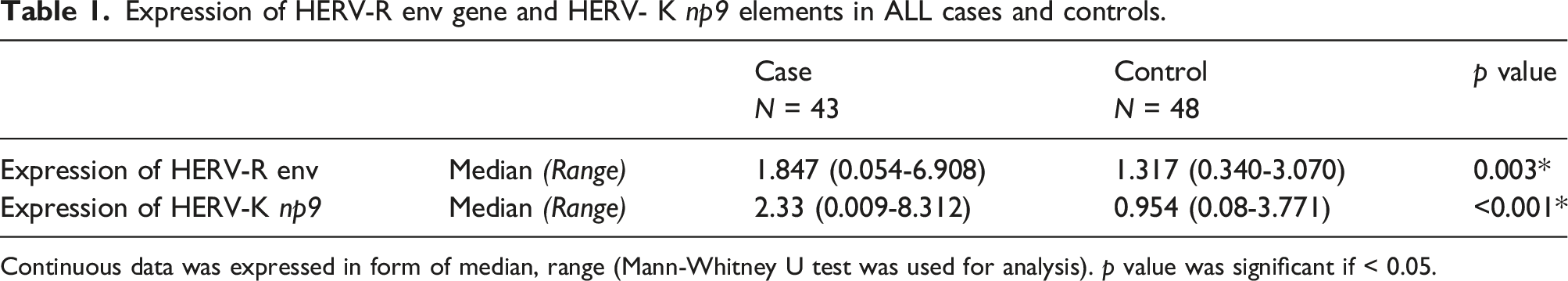
Continuous data was expressed in form of median, range (Mann-Whitney U test was used for analysis). p value was significant if < 0.05.
Analysis revealed no significant influence of gender or age group (less than 5 years vs greater than 5 years) on the expression of HERV-K np9 elements or HERV-R env gene in ALL patients. Similarly, no differences were observed across ALL subtypes (between B and T ALL). However, in the B-ALL subgroup, expression patterns of both genes were significantly higher than the control group (p = .001 for HERV-R env and p < .001 for HERV-K np9) as detailed in Supplemental Table 3. This expression profile was not mirrored in the T-ALL subgroup, where only HERV-K np9 showed a statistically significant difference (p < .001) as shown in Figure 1. Box plot showing expression of HERV-R env gene and HERV- K np9 elements in ALL cases and controls.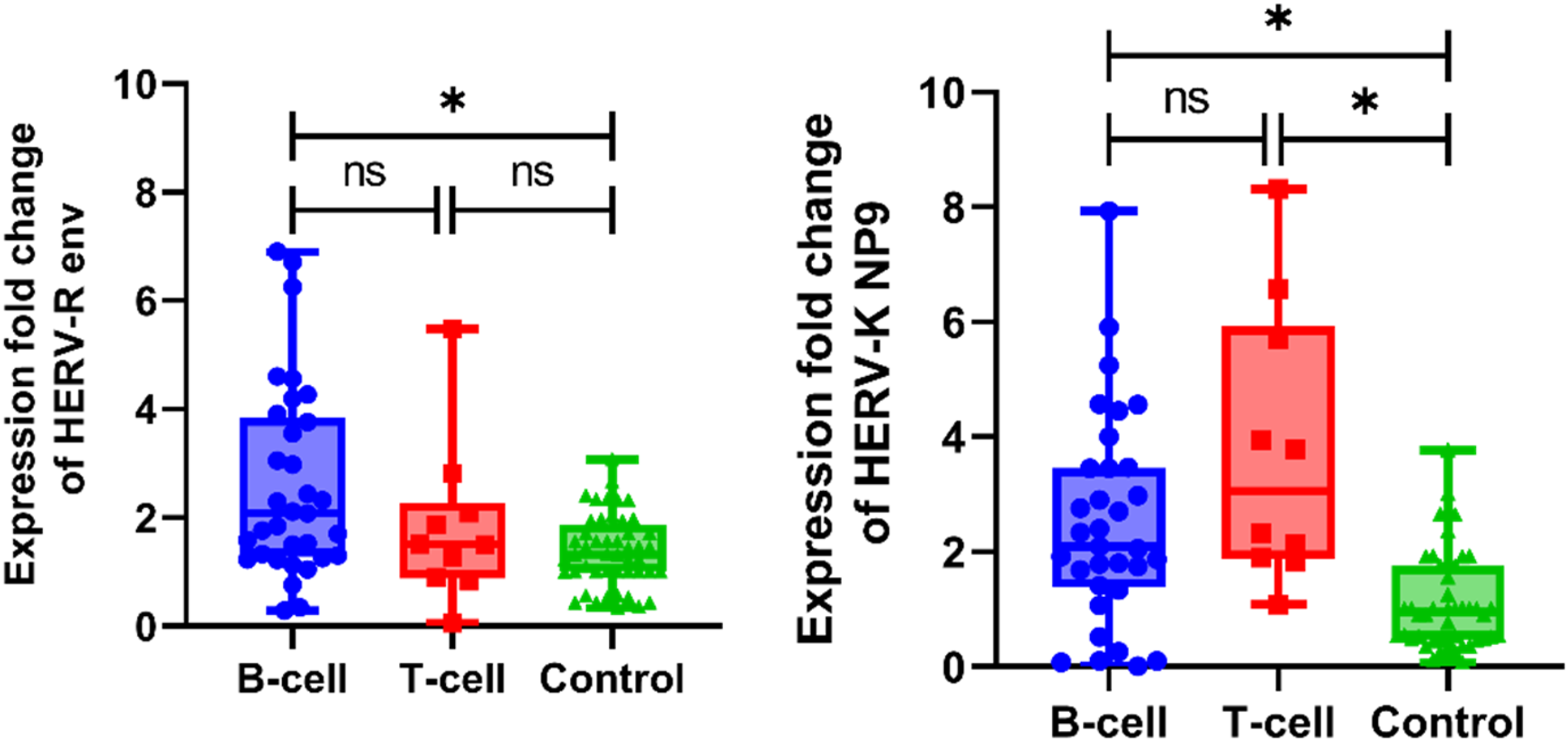
Association between HERV-K np9 and HERV-R env expression and ALL diagnosis
Regression analysis for the association between HERV-K np9 and HERV-R env expression and ALL diagnosis.
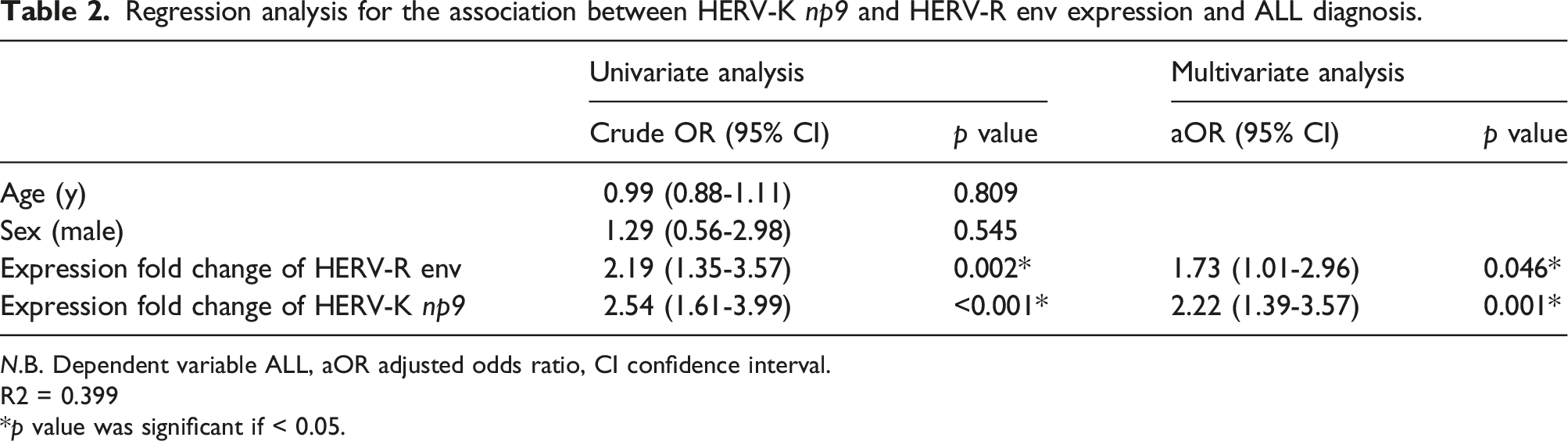
N.B. Dependent variable ALL, aOR adjusted odds ratio, CI confidence interval.
R2 = 0.399
*p value was significant if < 0.05.
Viruses detected in cases and controls
Figure 2 and supplemental table 4 show that ALL patients had a significantly higher prevalence of all three investigated viruses; herpes simplex virus, human parvovirus B19, and polyomavirus BK compared to controls (p = .001 for HSV, HPV B19, and p < .001 for BKV). Bar chart showing Viruses detected in cases and controls.
Correlation between viruses detected in the blood of ALL patients and relative gene expression of HERV-R env gene and HERV- k np9 elements
Correlation between viruses detected in the blood of ALL cases and relative gene expression of HERV-R env gene and HERV- k np9 elements (n = 43).

Data are presented as median (range).
*p value was significant if < 0.05.
Predictors of HERV-R env and HERV-K np9 expression in patients
Multiple linear regression analysis of HERV-R env and HERV-K np9 expression among all participants.

Dependent Variables: Expression fold change of HERV-R env and HERV-K np9.
The complete model, including all tested variables, and the final model after stepwise exclusion of all insignificant variables is given for each gene.
HERV-R env: R2 = 0.163
HERV-K np9: R2 = 0.301
*p value was significant if < 0.05.
Discussion
Among the various factors believed to contribute to HERV reactivation, external viral infection stands out as the most well-documented. 17 Studies showed a significant increase in HERV transcription in response to various viruses including, including herpes simplex virus (HSV), 18 hepatitis B virus (HBV), 19 human herpesvirus (HHV) 6, 20 Epstein-Barr virus (EBV), 21 dengue virus, 22 and Coxackie B virus. 23 Cancer is also a recognized trigger for HERV reactivation and expression, although the precise mechanism of this phenomenon in oncogenesis remains unclear. 24 While the underlying mechanisms linking exogenous viral infection and HERV expression in cancer are still under investigation, its potential involvement in the cell transformation process has been proposed. 25 Our findings add to the evidence about this connection, by demonstrating a strong association between ALL diagnosis, elevated expression of HERV-K np9 and HERV-R env, and the presence of herpes simplex virus infection.
Multiple factors contribute to the increased HERV transcription observed in cancer. These factors include the immunosuppressive environment associated with malignancies, alterations in the tumor microenvironment leading to activation of diverse signaling pathways and cytokine secretion, and potential secondary viral infections triggered by the cancerous state. 26 A general upregulation trend of HERV families across various cancer types is now well-established. 27
HERV-K is expressed in several solid and hematological malignancies. 28 For instance, studies have reported significantly elevated expression of specific HERV-K genes in various leukemias; such as env gene in acute myeloid leukemia, 29 pol gene in acute lymphoblastic leukemia, 30 gag gene in leukemia samples, 31 and np9 elements in chronic lymphocytic leukemia. 5 Furthermore, the HERV-K Env protein has been implicated in the oncogenesis of some solid malignancies, including melanoma 32 and pancreatic cancer. 33
We observed a significantly elevated expression of the HERV-K np9 in ALL patients. This finding aligns with previous research suggesting a significant role of np9 in tumorigenesis. 34 High expression of np9 has been documented across various malignancies, including teratocarcinoma cell lines, 34 melanoma cell lines and biopsies, 35 colon and breast tissues,36,37 and approximately 70% of chronic lymphocytic leukemia cases. 5
To the best of our knowledge, our study is the first to investigate np9 expression in ALL. The literature on its role in leukemia offers contrasting perspectives. Chen et al. demonstrated a critical role for np9 in the growth and survival of lymphoblastic and myeloid leukemia cells in both laboratory models and living organisms. 13 Conversely, Engel et al. did not observe upregulation of HERV-K in leukemia or lymphoma cell lines. 38 Adding to this complexity, Armbruester et al. reported minimal to undetectable np9 protein levels in normal tissues, 39 while Schmitt et al. identified it as a component of the human proteome, expressed in various healthy tissues, including PBMCs. 40
While research on HERV-R expression in leukemias is limited, our study contributes valuable new data. We observed a statistically significant upregulation of the HERV-R env gene in peripheral blood mononuclear cells (PBMCs) of ALL patients compared to controls. This finding aligns with Bergallo et al.'s work, where bone marrow analysis revealed higher HERV-R levels in both ALL and myeloid leukemia patients. 4 Notably, their study did not detect significant differences between lymphoid and myeloid leukemia subtypes in terms of HERV-R expression. 4
Upregulation of endogenous retrovirus Group 3 member 1, envelope (ERV3-1) was observed in the myeloid lineage compared to the monocytic lineage in both blood and bone marrow. 41 Conversely, using a different approach, Engel et al. reported downregulation of ERV3 in specific subtypes of acute leukemia. 38 These contrasting findings highlight the need for further investigation into HERV-R expression patterns across various leukemia subtypes and the underlying mechanisms involved.
To the best of our knowledge, our study also represents the first to directly compare HERV gene expression levels between B-ALL and T-ALL subtypes. We did not observe any significant difference in the expression levels of either HERV-K np9 elements or HERV-R env gene between these subtypes. This finding provides some support for the observations of Chen et al., who reported high np9 expression specifically in leukemia progenitor/stem cells, but not in normal CD34+ hematopoietic stem cells, which are precursors to all lymphocyte lineages. 13
Recognizing the potential for common viral infections to act as cofactors for HERV transactivation in the immunosuppressed environment of ALL, we explored the relationship between HERV gene expression and specific viral infections. Notably, HSV infection emerged as the only significant factor associated with elevated HERV-K np9 expression. This finding could represent the first in vivo evidence linking HSV infection to increased HERV-K np9 expression in ALL patients. While previous studies have established the ability of HSV to transactivate HERVs in cell lines, our findings represent the first demonstration of this association in leukemia patients.
The observed significant upregulation of the np9 in response to various viral infections has led some researchers to propose it as a potential link to the transformative abilities of certain oncogenic viruses. This hypothesis is further supported by documented findings of elevated np9 expression in EBV-transformed lymphocytes, HHV8-infected Kaposi sarcoma cells, and AIDS-associated Kaposi sarcoma tumor biopsies. 42
Our study has two main limitations: first, we did not perform a sample size/power analysis, limiting the robustness of our results. Second, the number of participants with positive herpes virus tests restricts the generalizability of our findings regarding the association with HERV-K np9 expression. Future studies would benefit from recruiting a larger and more diverse control group, including cancer-free children with documented herpes virus infections. Additionally, incorporating microarray technology and sequencing with qPCR could offer a better analysis of HERV family expression in all participants, especially considering the diverse splicing variants within HERV-K (including np9 and rec) along with related gag, pol, etc. sequences. Sequencing data would be valuable to confirm our findings and potentially identify other HERVs involved in ALL development.
Conclusions
This study contributes evidence to the growing body of research on HERV involvement in ALL. We observed a significant overexpression of HERV-K np9 and HERV-R env in the peripheral blood PBMCs of pediatric ALL patients compared to controls. This upregulation warrants further investigation to determine the potential role of these HERV genes in ALL development or progression. Additionally, our analysis revealed a novel association between HSV infection and increased expression of HERV-K np9 in leukemia patients. This finding shed light on a potential mechanism by which viral infections might contribute to oncogenesis. A deeper understanding of the relationship between HERV transactivation and viral infections holds promise for advancing our knowledge of cancer pathogenesis. Such insights could pave the way for the development of innovative therapeutic strategies targeting either HERVs themselves or the virally induced pathways they activate.
Supplemental Material
Supplemental Material - HERV-K np9 and HERV-R env: Two endogenous retrovirus products with potential Pathogenic roles in pediatric acute lymphoblastic leukemia
Supplemental Material for HERV-K np9 and HERV-R env: Two endogenous retrovirus products with potential Pathogenic roles in pediatric acute lymphoblastic leukemia by Noha Gaber, Ahmed Samir Abdelhafiz, Nivin Abdel-Azim, Azza Shibl, Lamiaa Fadel, Zeinab Korany Hassan and Eman Hassan Ahmed in International Journal of Immunopathology and Pharmacology
Footnotes
Author contributions
Noha Gaber, Eman Hassan Ahmed, and Zeinab Korany Hassan: Conceptualization, Methodology. Ahmed Samir Abdelhafiz: Writing original draft. Nivin Abdel-Azim, Azza Shibl, and Lamiaa Fadel: Investigation and data collection
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical statement
Study approval statement
This study was conducted with ethical approval obtained from the South Egypt Cancer Institute (SECI) ethics committee (approval no. 18-2023-662) on 13/11/2023.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
