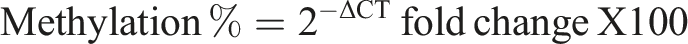Abstract
Objectives: Gliobalstoma is the most common primary brain tumor in adults with an extensive genetic and transcriptional heterogeneity, still identification of the role of DNA methylation, as one of epigenetic alterations, is emerged. Authors aimed to study the clinical role of N-myc downstream-regulated gene 2 (NDRG2) –based methylation among GBM patients versus benign neurological diseases (BND), investigate its prognostic role and its relation with survival outcomes. Methods: A total of 78 FFPE specimens were recruited as follows: GBM (n = 58) and BND (n = 20) then analyzed for NDRG2 methylation using Methyl II quantitative PCR system. The sensitivity and specificity of methylation was detected using receiver operating characteristic (ROC) curve and the relation with clinicopathological criteria for GBM and response to treatment were studied. Survival patterns; progression free survival (PFS) and overall survival (OS) were analyzed using Kaplan-Meier analyses. Results: Mean methylation NDRG2 level was significantly increased in GBM patients as compared to BND and its sensitivity and specificity were 96.55% and 95%, respectively with area under curve (AUC) equals 0.973. Among the clinical characteristic factors, mean methylation level reported significant difference with ECOG and tumor site. Survival out comes revealed that NDRG2 methylation increased with worse PFS and OS at significant level (long rank test X 2 = 13.3, p < .0001; and X 2 = 7.1, p = .008, respectively). Conclusion: Current findings highlight the importance of studying DNA methylation of NDRG2 as a key factor to understand the role of epigenetic alterations in GBM.
Introduction
The most reciprocal and aggressive primary tumor in adults is glioblastoma (GBM). Although multimodal treatment strategies i.e. surgery, radiotherapy and chemotherapy, its prognosis remains tremendously poor. 1 Due to the latest advances in genomic 2 and proteomic levels 3 the advantageous for controlling robust signatures and clinically appropriate molecular classifiers of GBM have now been emerged. 4
Impact of epigenetic mechanisms in tumorgenesis and it aggressiveness is well recognized, since hypermethylation of CpG island leads to silencing of tumor suppressor genes, while its hypomethylation encourages the transcriptional activation of oncogenes and hence promotes chromosomal instability. 5 These alterations in epigenetic are potentially reversible and hence can be counted as promising targets for epigenetic anti-cancer treatments. 6
In the current study, authors have investigated the methylation pattern for N-myc downstream-regulated gene 2 (NDRG2), one of the members of NDRG family, 7 that act as tumor suppressor gene and regulate several processes as differentiation, proliferation, metastasis and apoptosis. Hence its overexpression inhibits tumor metastasis and progression, through regulating the malignant performances of tumors by modifying the expression of c-Myc, CD24, Fas, and MSP58.8–10 The inactivation of its role as tumor suppressor gene has been proved as being due to DNA methylation. 11 Accordingly, authors aimed to assess the role of NDRG2 methylation among GBM patients and study its relation with clinical criteria, treatment strategies and survival outcomes, also to investigate its methylation level between GBM patients and others with benign neurological diseases to address its role in carcinogenesity in GBM. In addition authors utilize public databases to investigate the relationship between NDRG2 gene methylation and NDRG2 gene expression, as well as survival analysis. Moreover use public available datasets to analyze the pathways influenced by the NDRG2 gene.
Materials and methods
Tissue specimens and patient samples
A total of 78 specimens were retrieved from formalin – fixed paraffin embedded (FFPE) tissue blocks from the Department of Neuropathology - Faculty of Medicine, Ain Shams University, Egypt. Twenty of them were benign neurological disease specimens serve as tumor-free brain tissue samples. These specimens were surgically resected before they received any treatments, investigations were done in accordance with the principles of Declaration of Helsinki and approved by the Medical Ethics Committee from National Research Centre (ID#20110), and all patients gave written informed consent. Their clinic-pathological data were collected from their medical records.
Histopathologic examination
In this prospective study, FFPE specimens were recruited from adult GBM patients (age >18 years) with viable tumor tissue for GBM more than 80%, then cutting at full slices with 4 microm in thickness and stained with hematoxylin and eosin H&E stain, finally examined by pathologist according to 2016 CNS Tumors WHO classification. Another fresh FFPE sections (around 8 sections) with 10 micro-thickness were a liquated for molecular biology examinations.
DNA extraction
DNA was extracted from FFPE specimens through using QIAamp FFPE kit (Cat no. 56404) as per manufacturer instructions. Then purity and concentration for each sample were detected using nono-drop spectrophotometer (Quawell, Q-500, Scribner, USA) through measuring the absorbance at 260 and 280 nm, then checked on 1%agarose gel, finally extracted DNA samples were stored in −20 for further processing to analyze NDRG2 methylation.
Detection of NDRG2 methylation pattern using methyl II quantitative PCR system
NDRG2 methylation pattern was assessed in DNA extracted samples using EpiTect fast DNA bisulfite kit (Qiagen, Germany, cat. No. 59824) according to manufacture user guide converted DNA was tested using specific designed methylation specific primers and unmethylation specific primers that flanks a promoter region of interest by real-time PCR. by using real-time PCR (Max3005P QPCR system; Stratagene, Agilent Technologies, CA, USA) 5 μL from the bisulfite treated DNA was mixed directly with qPCR master mix (RT2 qPCR SYBR Green/ROX Master Mix, Cat number 330520) and were dispensed into a PCR plate comprising pre-aliquoted NDRG2 methylations specific PCR primers which will amplify the NDRG2 gene if it was methylated primer and un-methylated primers which will amplify the un-methylated NDRG2 gene (Left methylated primer GTTTTTGGAGTTTTAGTTTTTGTGC Right methylated primer GAACGATATAATTAATCCGCGTC) (Left un-methylated primer TTTTGGAGTTTTAGTTTTTGTGTGT Right un-methylated primer CAAACAATATAATTAATCCACATC). Real-time PCR is carried out by listed cycling conditions: one cycle [95°C for 10 min], then three cycles [99°C for 30 and 72°C for 1 min], finally 40 cycles [97°C for 15 sec and 72°C for 1 min]. Calculations were done by collecting, the raw ΔCT values for each PCR reaction tube (UM and M) for every sample. Because in the qPCR reaction the M was used hence the DNA in which all CpG sites are methylated will be identified by real-time PCR
12
by following equations:
Treatment strategies
Recruited patients were undergo multimodality approach as the standard primary method for GBM as they were evaluated clinically (through complete history, both clinical and neurologic examinations and neurologic examination) then by brain imaging in order to receive their standardized treatment protocol, that include maximum safe resection, followed by radiotherapy conventional fractions (total dose of 60 Gy, given 2 Gy per fraction for 30 fractions over 6 weeks) or hypofractionation (45 Gy in 15 fractions over 3 weeks) by concomitant temozolomide
(TMZ) chemotherapy (75 mg/m2 every day till end of radiotherapy) by systematic follow up, after wards re-evaluated clinically and radiologically, followed by adjuvant 6 cycles of TMZ treatment at a dose of 150 mg/m2 body surface area from days 1 to 5 every 28 days with clinical monitoring. Throughout regular clinical follow up, patient were evaluated by gadolinium-enhanced magnetic resonance imaging (Gd-MRI) 45 days after RT and then completed every 3 months or at time of clinical signs of neurologic progression. Tumor response was calculated on the basis of radiological RANO response criteria (2010). 13 Complete response (CR) defined as disappearance of all recognized brain lesion. Partial response (PR) is known as 50% or greater decrease in assessable brain lesion or an actual enhancement in evaluable brain lesion. Stable disease (SD): brain lesion unchanged (<50% decrease or <25% increase in the size of measurable lesions). Progressive disease (PD): ≥25% increase in size of some or all of brain lesions and/or the appearance of any new brain lesions.
Survival outcome assessment
Survival outcome for recruited GBM patients were categorized into progression free survival (PFS) which is defined as the time from start enrollment of the patients in the study till progression of the disease confirmed by MRI imaging or other clinical deteriorations. And the other category is overall survival (OS) and this is defined as the period from the date of random assignment of the patient till the date of death or lost follow-up.
Statistical analysis
Statistical analysis was performed using SPSS 16 (IBM software). Association between the clinico-pathological and demographic factors with NDRG2 methylation was calculated by ANOVA analysis. Comparing between responses to treatment was calculated by Independent samples t test was used for continuous variables and Fisher exact test for proportions. Receiver operating characteristic (ROC) curve was plotted between GBM patients and BND to detect sensitivity, specificity, positive and negative productive values and accuracy and their clinical efficacy. 14 Survival analyses were performed by the Kaplan–Meier method and log-rank test was used to compare difference of survival curve between high and low methylation levels. To show the reliability of the survival estimate, the confidence interval (CI) with 95% confidence level was reported.
Result
Clinical and demographic data for GBM cases.

NDRG2 methylation level among investigated groups
Significant difference was reported between the two groups regarding methylation level as plotted in Figure 1, as the median level (mean ± SD) was 19 (20 ± 9) and 67 (63 ± 16) for BND and GBM, respectively. To investigate the sensitivity and specificity for NDRG2 methylation among recruited patients, ROC curve was plotted (Figure 2) with area under curve equals 0.9, the sensitivity and specificity recorded 96.55% and 95%, respectively. The best cutoff point recorded to be 32, by considering this cutoff point 19 out of 20 (96.6%) BND were below this point and only one patient (5%) reported methylation level above this point. For GBM, 56 out of 58 (95%) were above the cutoff point and 2 cases (3.4%) were below cutoff point at significant level (X
2
= 63.4, p < .0001). Mean level of NDRG2 methylation among GBM cases and benign neurological diseases. ROC curve for methylated NDRG2. Arrow donates for the best cut off point.

Association between clinical criteria and methylation level
Association between NDRG2 methylation and clinical pathological criteria.

Relation between response to treatment and methylation level
GBM received standard of care treatment and accordingly their response was recorded as responders (n = 30) or non-responders (n = 28). Levels of methylation (median, mean ± SD) were reported as (56.5, 57.5 ± 14) and (74, 69 ± 15), respectively at significance of (F = 8.8, p = .004) (Figure 3(a)). Among the entire group of responders (CR, PR and SD) and non-responders also significant level was recorded as level of methylation was for CR (n = 15) (55, 58.3 ± 14), PR (n = 8) (57, 53 ± 15), SD (n = 7) (65, 60 ± 15.5), and non-responders as (n = 28) (74, 69 ± 15) at significant level (F = 3.2, p = .029) (Figure 3(b)). Frequency of NDRG2 methylatioon with response was reported in Table 3 Relation between NDRG2 methylation and response to treatment. (a) Relation between NDRG2 methylation and both responders vs non-responders. (b) Relation between NDRG2 methylation and entire groups representing different responses levels versus non responders. Frequency of NDRG2 methylation with survival outcome criteria.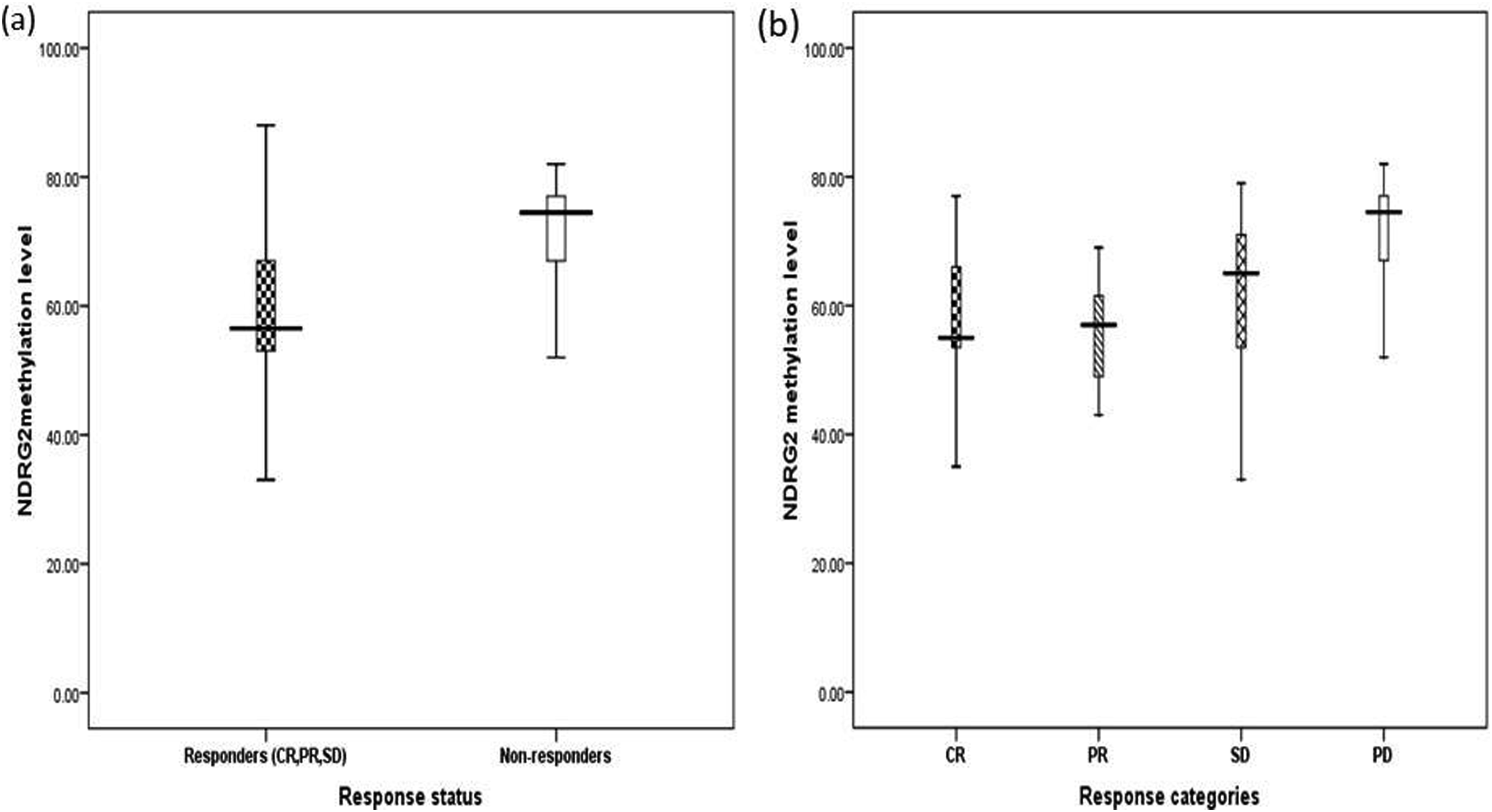

Impact of methylation level and survival outcome
GBM patients were followed up for a median of 10 months and their progression free survival was recorded at median follow-up of 13 months (Figure 4(a)), by considering the mean level of methylation among GBM patients as 63, those with high methylation status (above median level) reported worse PFS as compared to those with low methylation level at significant level (Figure 4(b)). Similarly, OS was recorded at median of 16 months (Figure 5(a)), were significantly related with NDRG2 methylation as those with high methylation level reported shorter OS compared to those with low methylation level as plotted in (Figure 5(b), Tables 4 and 5). Progression free survival for GBM patients. (a) PFS concerning all recruited GBM patients, (b) PFS of GBM patients concerning the different methylation level of NDRG2. Overall survival for GBM patients. (a) OS concerning all recruited GBM patients, (b) OS of GBM patients concerning the different methylation level of NDRG2.

In silico analysis of the relation between NDRG2 gene methylation and its gene expression
By using cBioportal for cancer genomics (TCGA, PanCancer Atlas) dataset at https://www.cbioportal.org/study/summary?id=gbm_tcga_pan_can_atlas_2018, the relation between NDRG2 methylation and NDRG2 gene expression was examined and demonstrated significant difference as shown in (Figure 6(a)) and their levels were examined in relation to survival analysis and revealed significant difference as shown in Figure 6(b) and (c). To analyze the pathways influenced by the NDRG2 gene, authors used STRING online available database was used to analyze pathways influenced by NDRG2 as protein. Acordingly, NDRG2; contributes to the regulation of the Wnt signaling pathway, down-regulates CTNNB1-mediated transcriptional activation of target genes, such as CCND1, and may thereby act as tumor suppressor, also it may be involved in dendritic cell and neuron differentiation, (371 aa), as shown in (Figure 7). Scattered graph using data set (TCGA, PanCancer Atlas) in GBM to show: (a) correlation between the NDRG2 methylation status and NDRG2 gene expression level, (b) NDRG2 methylation status and progression free survival (PFS), (c) NDRG gene expression level and progression free survival (PFS). NDRG2 protein - protein interaction network.

Discussion
Number of events, mean levels of methylated NDRG2 with progression free survival (PFS).

aMean level of methylation level among GBM cases.
Number of events, mean levels of methylated NDRG2 with overall survival (OS).

aMean level of methylation level among GBM cases.
Relation between NDRG2 methylation with demographic and clinical characteristic factors for GBM reported, accordingly there was an elevation in mean level of methylation in GBM patients with age >60 as compared to those below 60, this finding agrees with previous study 15 but ours did not reached significant difference which reveal that increased methylation pattern in older ages tend to correlate with the aggressiveness of GBM. 16 Median methylation levels was significantly increased in men versus women and this is inconsistent with previous findings that GBM incidence is ranged 1.6 times higher in men than women 1 which may be attributed to biological and environmental variables. Some studies have reported a link between sex hormones and susceptibility to GBM17,18 as a strong link between estrogen and their neuroprotective effects, while in men a relationship between testosterone and GBM tumorgenesis since this sex hormone has been recommended to have responsibility in the male predominance of the disease, 1 hence mechanistic studies are urged regarding the potential reasons of gender differences in GBM and they should assimilate observations at metabolomic, genomic and proteomic levels. NDRG2 methylation was elevated among those GBM patients with a family history as 2 of the enrolled GBM patients reported their 1st degree relatives with primary brain tumors, although the difference was not significant but still an evidence that agree with a previous study 19 that a great concern should be addressed to persons with a family history of glioma.
In the current study 29 GBM patients reported to have tumor on the left side, 23 patients on the right side and the remaining 6 patients were having the tumor at both sides, when authors investigate the level of methylation it was significantly increased in patients having tumor in multiple sites indicating that NDRG2 methylation is linked to GBM tumorgenesity. Moreover, performance status (PS) was estimated according to ECOG at both diagnosis and treatment and accordingly GBM patients were categorized as below or above 2. Significant difference was detected in NDRG2 methylation regarding ECOG criteria as those reported to have worse PS showing high methylation indicating that methylation of NDRG2 is linked to pathogenicity of GBM. 5 In addition our results showed that NDRG2 methylation increased in GBM patients with multiple tumor sites as compared to those having either left or right side tumor which points out that NDRG2 methylation is related to the aggressiveness of the tumor. 5
Survival outcomes for GBM patients were recorded and the impact of NDRG2 methylation on both PFS and OS reported significant association. GBM patients with high NDRG2 methylation level reported worse PFS and lower OS and vice versa. These results were in concordance with previous findings 15 and can point out the importance for detection of gene methylation (as NDRG2 methylation) since its level can aid in GBM progression monitoring and function as a prognostic marker to characterize patient’s survival outcomes. 20
DNA methylation is among epigenetic alterations which has been reported to play an important role in gene regulation. 21 In cancer, the role of DNA methylation in cancer initiation and progression is being clarified as extensive – scale data become accessible, among these data bases is The Cancer Genome Atlas (TCGA) which offers abundance of knowledge for the investigation of many molecular aspects of cancer genetics. 22 Thus in the current study authors have used the cBiPortal Cancer Genomics databases to analyze the relation between levels of NDRG2 methylation and its gene expression as well as its impact on survival and significant difference was reported.
In conclusion, current study reports the clinical impact role of NDRG2 methylation in GBM as useful diagnostic marker and its importance to predict their prognosis and survival outcomes.
Limitations of the study
Although the current study revealed an evidence for the role of NDRG2 gene methylation in GBM, but several limitation of the study can be reported as: small sample size, because GBM is somehow a rare disease among Egyptians and there was no calculation for samples enrolled in the study. Also detection of NDRG2 gene expression or NDRG2 protein level in GBM samples were not detected which could emphasize the relation between both NDRG2 gene methylation and its gene expression or protein level, hence future study is in progress to overcome these limitations.
Footnotes
Author contribution
Study conception and design: MS, AMN and MSM. Provision of samples and clinical follow-up: LRE, KE and ME, Acquisition of data: MK and AMN. Analysis and interpretation of data: MS; MK, AMN and MS. Drafting of manuscript: MS; LRE; MK and AMN. Critical revision: all authors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This paper is based upon work supported by Science, Technology & Innovation Funding Authority (STDF) Basic and Applied Research Support Grant Project (BARG) [No.25562]. The instruments listed in the current study were purchased through a grant from Science Technology & Innovation Funding Authority (STDF) through Capacity Building Grant Fund (CBG) [No. 4940], Egypt.