Abstract
Introduction
Type 1 diabetes mellitus (T1DM) is characterized by autoimmune destruction of insulin-producing pancreatic beta (β-) cells. Previous studies suggested an imbalance between and pro- and anti-inflammatory cytokines exacerbates T1DM development.
Objectives
We aimed to test the hypothesis that patients with T1DM carry a higher frequency of regulatory genes associated with low levels of the anti-inflammatory cytokines interleukin-4 (IL-4), its receptor (IL-4R), and interleukin-10 (IL-10).
Methods
Accordingly, we compared frequencies of five different single nucleotide polymorphisms (SNPs) in T1DM patients and healthy controls who had been typed for HLA-DRB1, HLA-DQA1, and HLA-DQB1 genes.
Results
The frequencies of rs2070874 (IL-4) alleles C and T differed between T1DM patients and controls (cp = 0.0065), as did their codominant (cp = 0.026) and recessive (cp = 0.015) models. Increased frequencies were observed in T1DM patients for HLA alleles: DRB1*03 (pc < 0.0013), DRB1*04 (cp = 0.0169), DQA1*03 (cp = 0.0222), DQA1*05 (cp < 0.0006), DQB1*02 (cp = 0.0005), and DQB1*06 (cp < 0.0005). And lower frequencies were observed for: DRB1*07 (cp = 0.0078), DRB1*11 (cp = 0.0013), DRB1*13 (cp < 0.0364), DRB1*15 (cp < 0.0013), DQA1*01 (cp < 0.0006), and DQA1*02 (cp = 0.0348). Certain DRB1: DQA1: DQB1 haplotypes showed greater frequencies, including, 03:05:02 (p < 0.0001) and 04:03:03 (p = 0.0017), whereas others showed lower frequencies, including, 07:02:02 (p = 0.0032), 11:05:03 (p = 0.0007), and 15:01:06 (p = 0.0002). Stratification for the above HLA haplotypes with rs2070874 C/C exhibited no significant differences between T1DM patients overall and controls. However, when stratified for the vulnerable HLA haplotype (03:05:02/04:03:03), young patients in whom T1DM began at ≤13 years had a higher frequency of the SNP (rs2070874 C/C); a gene associated with low IL-4 production (p < 0.024).
Conclusion
This study suggests that possession of the rs2070874 C/C genotype, which is associated with low production of IL-4, increases the risk of T1DM in young individuals carrying vulnerable HLA alleles/haplotypes.
Keywords
Introduction
Type 1 diabetes mellitus (T1DM) is a chronic disease characterized by autoimmune destruction of insulin-producing pancreatic beta cells (β-cells), leading to insulin deficiency. 1 Between 5% and 10% of all diabetes patients worldwide have T1DM, and the incidence is estimated to be approximately 3% annually with significant geographical variations 2 ; the highest incidence being in Finland (40.2/100,000), 3 and the lowest in China (0.1/100,000). 4 In 2017, Saudi Arabia was ranked eighth worldwide regarding the numbers of TIDM patients and fourth in terms of the incidence rate of the disease, which was estimated to be (33.5 per 100,000 individuals) of TIDM. 5
T1DM occurs more frequently in genetically susceptible individuals who are subject to various environmental and epigenetic factors, as indicated by the concordance rate of T1DM in monozygotic twins, ranging between 13% and 65%.6,7 Importantly, in twins of patients with early onset of T1DM (<24 years) the probability of progression to T1DM was 38% compared to late-onset patients (>24 years) where the probability was 6% (Redondo et al. 2001) 6 ; suggesting an inverse association between the burden of genetic risk and age of onset. This is accounted for by multiple genes, including those associated with peptide processing and regulation of inflammatory reactions being implicated in pathogenesis.
Self-reactive T cells are normally heavily suppressed to prevent autoimmunity. However, in T1DM, genetic predisposition carried by alleles within the human leukocyte antigen (HLA) region leads to a breakdown in peripheral tolerance and the progressive destruction of insulin-producing β-cells. 8 In addition, genetic regulation of expression, function, and production of both pro-and anti-inflammatory cytokines plays a role in triggering the inflammatory β-cell destruction.9-11 While the HLA-DRB1/DQA1/DQB1 genes on chromosome six encode cell surface molecules that determine which self-peptides are presented via T-cell receptors (TCRs), cytokines regulate the expansion or suppression of T-cell clones.12,13 The interleukin-4 (IL-4) gene on chromosome five and the interleukin 10 (IL-10) gene on chromosome one encode cytokines that down-regulate inflammatory responses. However, decreased levels have been observed in both newly diagnosed T1DM patients and non-obese diabetic (NOD) mice thereby facilitating insulitis.14,15 Conversely, genetic overexpression of IL-4 prevents insulitis and decreases the incidence of diabetes. 16 Notwithstanding these findings, others have claimed to have refuted a role for IL-4 regulator genes (Riemsnider et al. 2000). 17 IL-10 also generates feedback regulation of autoimmunity by binding to the IL-10 receptor (IL-10R) expressed on the surface of many immune cells. 15
Multiple research studies, including our own, have investigated associations between single nucleotide polymorphisms (SNPs) linked to regulatory or structural regions of cytokine genes and T1DM.18-21 Depending on their location within the genome, SNPs are flags highlighting possible differences in the expression, function, and/or production of cytokines within T1DM phenotypes. For example, the IL-4 SNP rs2070874 is located in the 5′-untranslated region of exon one of the IL-4 gene and, as such, could regulate the levels of cytokine production. In addition, rs1800871 and rs1800872 are located in the promoter region of the IL-10 gene, influencing messenger RNA transcription and expression. 22
In this study, we hypothesized that in T1DM “Self-peptides” associated with β-cell are targeted by autoimmune T-cell clones, whose evolution is facilitated by an excess of pro-inflammatory cytokines (e.g., TNF-α) and/or a dearth of anti-inflammatory cytokines (e.g., IL-4 or IL-10); both scenarios facilitating β-cell destruction. Based on this hypothesis, we studied SNPs located within the IL-10 (rs1800896, rs1800871, and rs1800972), IL-4 (rs2070874), and IL-4R (rs1801275) genes in T1DM patients and controls typed for HLA - DRB1, DRA1, and DQB1 loci. Our objective was to identify significant differences in gene frequencies between patients, with age of onset above and below 13, and controls.
Materials and methods
This case-controlled cohort study of 371 individuals included 180 patients diagnosed with T1DM and 191 healthy controls. The sample size calculation was performed based on a descriptive epidemiological study on the Saudi population by Robert et al. utilizing Fisher’s exact test at a level of 5% and power of 80. Patients were diagnosed according to the American Diabetes Association criteria. 23 We included individuals of Saudi origin (patients and controls) and only patients who were less than 30 years of age at the time of blood collection. Exclusion criteria included non-Saudi patients, patients with type 2 diabetes, young patients with maturity-onset diabetes, patients with other forms of secondary diabetes, and immunocompromised patients. The healthy unrelated control group was selected randomly from the list of bone marrow transplant (BMT) donors of King Fahad Medical City (KFMC) Hospital who had no history of diabetes or other autoimmune diseases.
Guidelines of the Helsinki Declaration on Human Experimentation were implemented, and the Institutional Review Board (IRB) at KFMC approved the study. An exemption from obtaining informed consent from patients was also approved by the IRB because we used clinical samples collected from patients for hemoglobin A1c or archived DNA of BMT donors. Peripheral blood sample were collected into EDTA tubes between January 2016 and December 2019 for T1DM patients and from February 2015 to July 2020 for healthy controls. Blood samples integrity was assessed for quality purposes at the time of collection. All procedures adhered to rules and regulations of the Saudi government, the KFMC/IRB policies and procedures, and the IHC Good Clinical Practice guidelines.
A MagNa Pure Compact instrument (Roche Diagnostics GmbH, Roche Applied Science, Mannheim, Germany) was used to extract genomic DNA according to the manufacturer’s instructions, and a minimum of 20 ng/μl of DNA with 260/280 nm values between 1.6 and 2.0 was set as the standard for performing the genotype assays.
Selected SNPs for IL-10, IL-4, and IL-4R genes (https://www.snpedia.com).
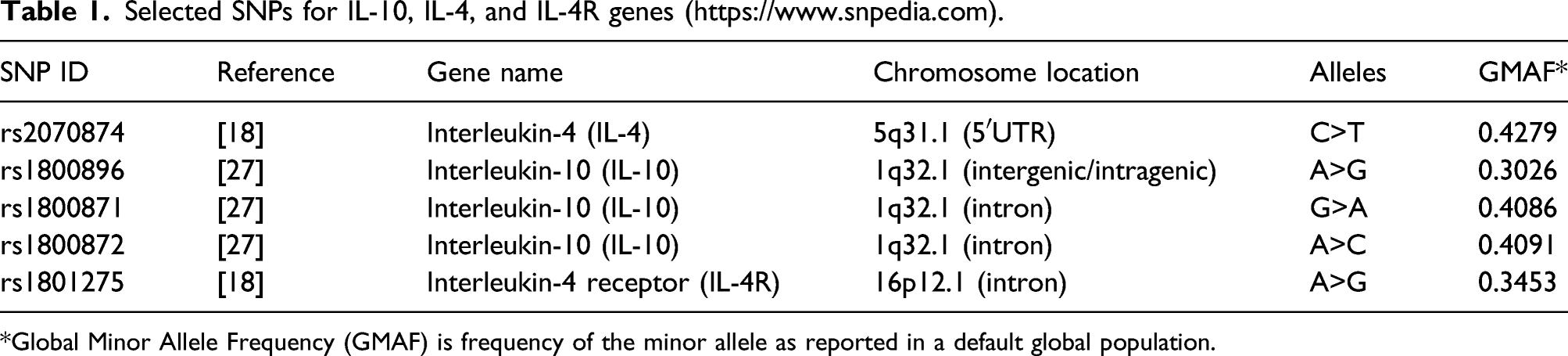

*Global Minor Allele Frequency (GMAF) is frequency of the minor allele as reported in a default global population.
For HLA genotyping, we used a sequence-specific oligonucleotide probe (SSOP) utilizing a Luminex-based method (One Lambda, San Diego, USA) to detect the HLA-DRB1, HLA-DQA1, and HLA-DQB1 genes according to the manufacturer’s instructions (http://www.onelambda.com). The HLA genotyping procedure performed in our laboratory is continuously monitored for quality assurance by the College of American Pathologists (CAP).
Statistical analysis
Hardy-Weinberg equilibrium (HWE) for expected and observed allele frequencies for control data was assessed based on the χ2 distribution (degree of freedom = 1). 24 For each SNP allele, allele frequencies and genotype models (codominant, dominant, and recessive) were derived by an algorithm based on the direct counting method utilizing SNPStats software (https://www.snpstats.net/start.htm).
Dominance is defined as the relationship between the two alleles of one gene, where the effect of the first allele (dominant) in the phenotype masks the contribution of the second allele (recessive). 25
Codominance is defined as a form of dominance in which two different heterozygous alleles of one gene are fully expressed, and the offspring’s phenotype is neither dominant nor recessive. 26
D′ statistic and p-value tests were implemented to analyze the linkage disequilibrium (LD) between the SNPs. The D′ LD measurement ranged from 0 (no LD) to 1 (strong LD), and significant differences were indicated by a p-value less than 5%. 27 Fisher’s exact test and the log of the odds (logit) for the presence of each SNP were used to specify the significant differences between patients and controls at an overall level of p < 0.05. HLA alleles, haplotype frequencies, and stratification with SNPs were conducted using SPSS software version 22.0. The Bonferroni correction test was used whenever it was necessary to correct multiple comparisons, and the results are presented as corrected p (cp) values.
Results
Demographic and clinical data for T1DM patients and Controls.
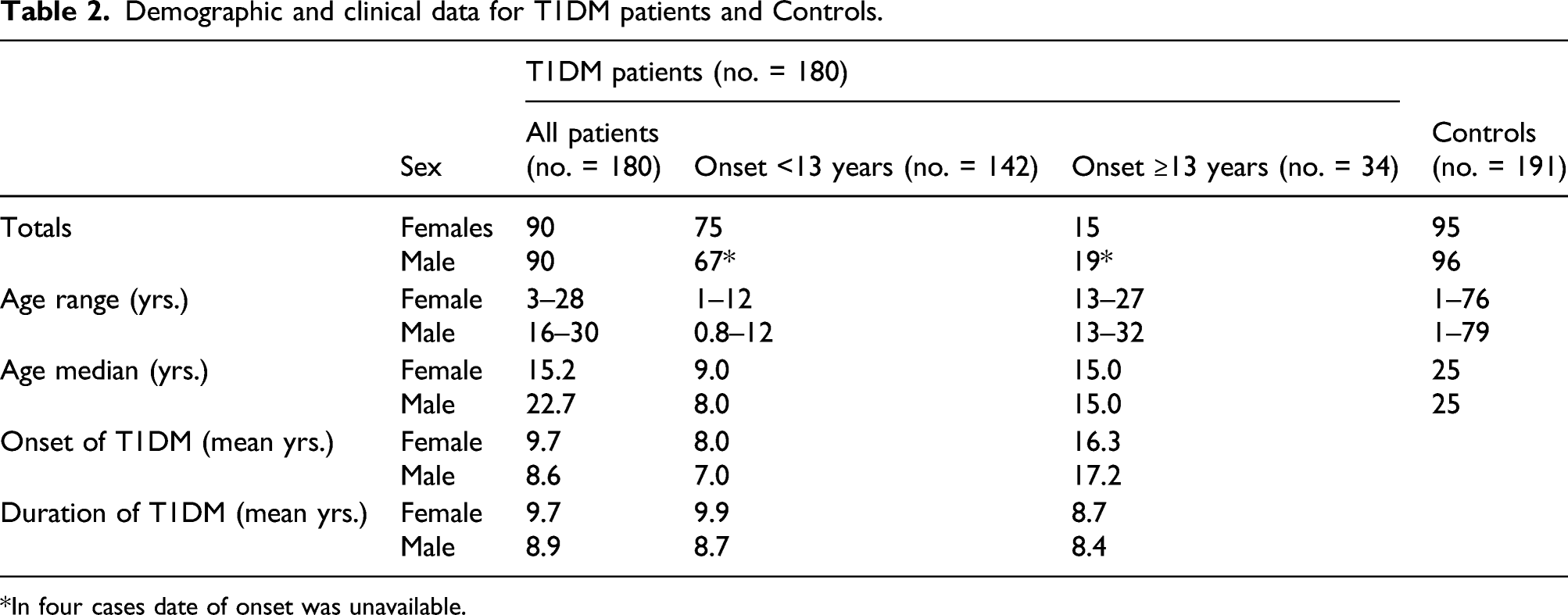
*In four cases date of onset was unavailable.
Single nucleotide polymorphisms
Conformation to HWE was observed for the three SNPs, rs1800871, rs1800872, and rs1801275 with expected statistical values (p > 0.05); however, two SNPs, rs2070874 and rs1800896, did not conform to HWE (p < 0.05). Among the five SNPs tested for possible LD, only rs1800896 and rs1800872 showed significant statistical values (D′ = 0.9993 and p < 0.001) and were considered to exhibit high LD. No other significant values for LD were detected.
Association between SNPs and susceptibility to T1DM.

*The second most common allele occurring in the population.
The allele frequency for rs1800871 (IL-10) exhibited significant variation that did not remain after Bonferroni correction. No other differences were found, and interaction analysis for these SNPs revealed no difference between males and females.
HLA-DRB1
Frequency distribution and association of HLA-DRB1 alleles in T1DM and controls.
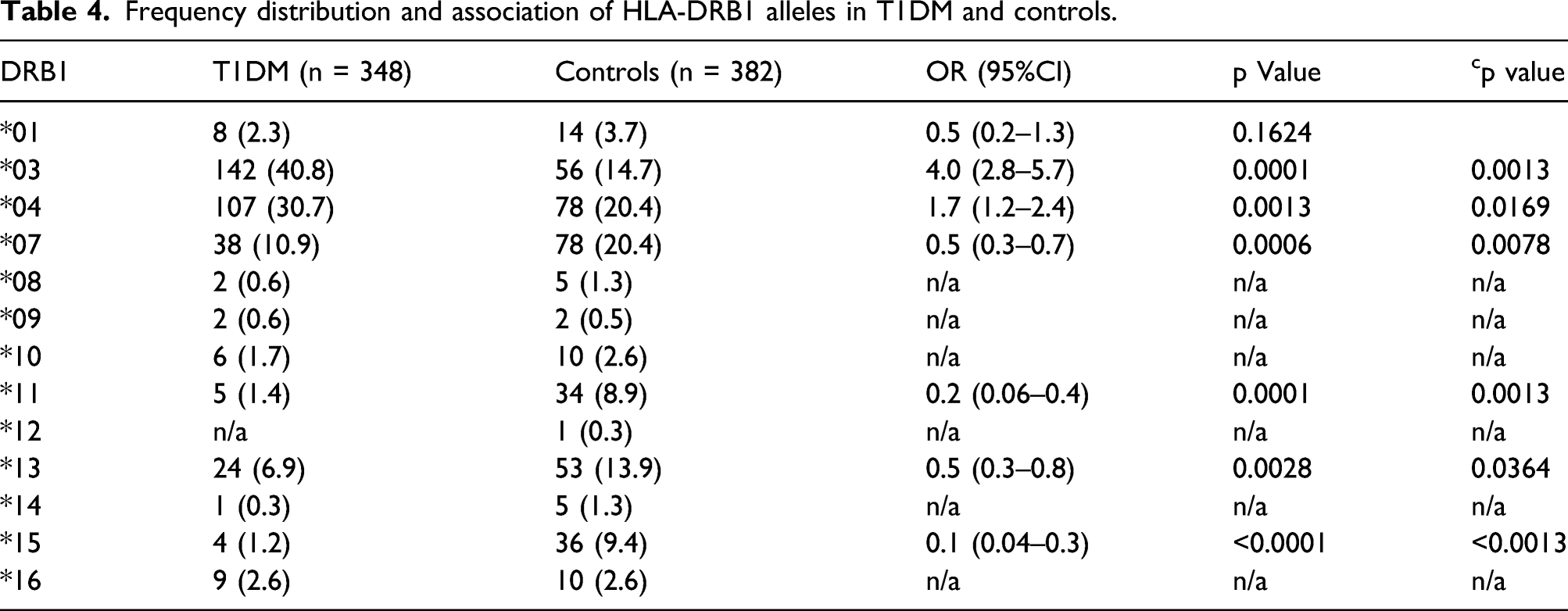
HLA-DQA1
Frequency distribution and association of HLA-DQA1 alleles in T1DM and controls.
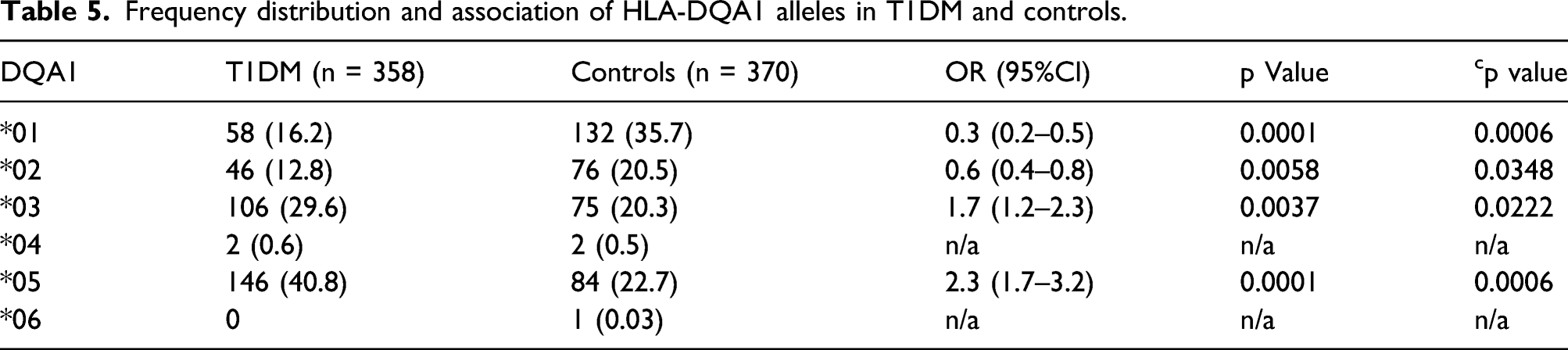
HLA-DQB1
Frequency distribution and association of HLA-DQB1 alleles in T1DM and controls.

HLA-DRB1:DQA1: DQB1 haplotypes
Frequency distribution and association of DRB1: DQA1: DQB1 haplotypes in T1DM and controls.
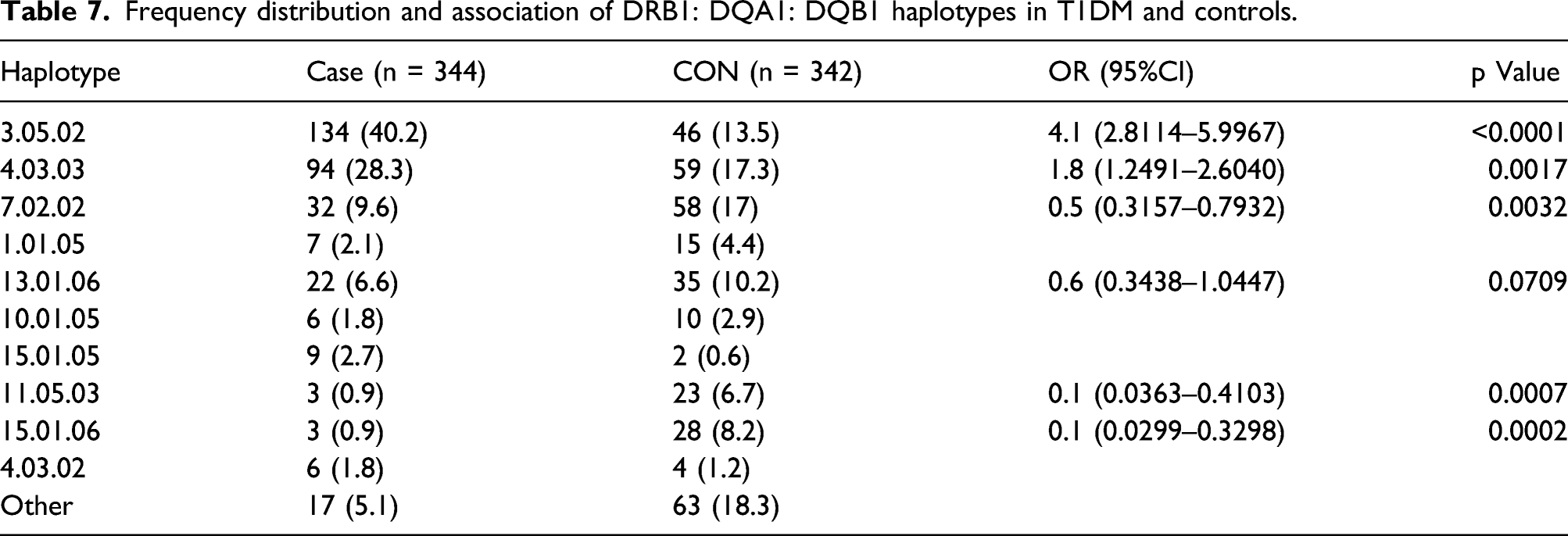
Stratification by SNP and HLA-DRB1 genotype
Stratification rs2070874 C/C genotype with HLA- DRB1.
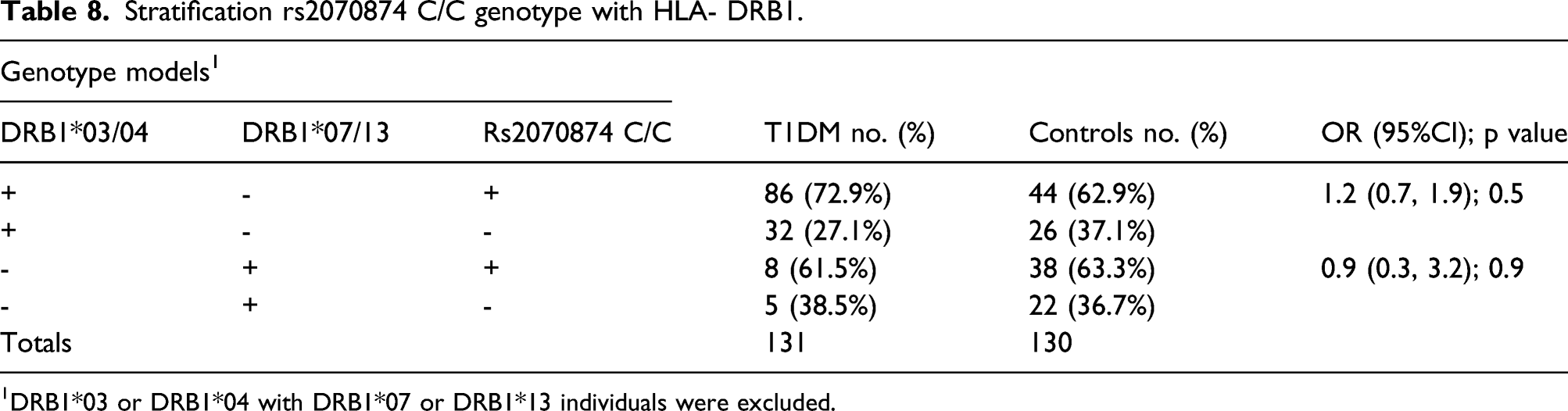
1DRB1*03 or DRB1*04 with DRB1*07 or DRB1*13 individuals were excluded.
Stratification of rs2070874 C/C genotypes with HLA - DRB1 by age of onset of T1DM.

1DRB1*03 or DRB1*04 with DRB1*07 or DRB1*13 individuals were excluded.
Stratification by SNP and HLA-DRB: DQA1:DQB1 haplotype
Stratification of rs2070874 C/C genotype with HLA - DRB1: DQA1; DQB1 haplotypes.

1DRB103:05:02 or 04:03:03 with 07:02:02 or 11:05:03 DRB1 individuals were excluded.
Stratification of rs2070874 C/C with HLA - DRB1: DQA1; DQB1 haplotypes, according to age of onset of T1DM.

1DRB103:05:02 or 04:03:03 with 07:02:02 or 11:05:03 DRB1 individuals were excluded.
Discussion
Our findings indicate genes associated with low production of IL-4 facilitate the early onset of T1DM in individuals with vulnerable HLA genotypes. The association between HLA and T1DM has been extensively studied, but the involvement of IL-4 has been unclear; and whereas low IL-4 was implicated in NOD mice (Shoda et al. 2004) 33 and patients with T1DM (Hagar and Zohreh 2016), 34 others claimed to have disproved a role for IL-4 regulatory genes (Reimsnider et al. 2000). 17 But no account was taken of the age of onset of T1DM in their analysis. Our data are consistent with and add to, the observation that early-onset T1DM is associated with a higher twin-concordance rate in younger patients due to an increased burden of non-HLA genes controlling B and T-cell development (Inshaw et al. 2020, Redondo et al., 2001and 2020).35,36
The well-established role of HLA Class II molecules is to process peptides external to the cell surface and present them to T-helper cells to generate an immune response, or to T-regulator cells to generate tolerance. In this context, our data imply the HLA haplotype most strongly associated with T1DM (03:05:02/04:03:03) presents a unique array of β-cell derived self-peptides to T-helper cells triggering autoimmunity. Conversely, the HLA haplotype most strongly associated with protection against T1DM (07:02:02/11:05:03) produces β-cell self-peptides triggering tolerance. Thus immunity, or tolerance, to β-cell peptides is dictated initially by the HLA haplotype, and subsequent evolution of responses is directed by pro or anti-inflammatory cytokines.
Within the context of the simplistic paradigm whereby autoimmune diseases are facilitated by an imbalance between anti-inflammatory (IL-4, IL-10, IL-13) and pro-inflammatory cytokines (TNF- α, IFN- γ) low levels of the former and high levels of the latter facilitate T1DM onset (Moudgil and Chouby 2011)39. Both our former finding (Osman et al. 2021), 37 that T1DM was associated with high TNF producer genes (rs361525), and current findings of low IL-4 producer genes are consistent with this model.
As demonstrated by next-generation genotyping array technology, several other genes are associated with aggressive early-onset T1DM in children under 7 years. 36 These include genes expressed in β-cells (GLIS3) and others affecting immune function via B-cell, T-cell, and Thymus development (IL2-RA, IL-10, IKZF3, THEMIS, CTSH). In Inshaw et al.’s study SNPs detecting IL-10 differed from those in ours and may account for our negative finding of an association between T1DM and IL-10. Furthermore, no SNPs linked to IL-4 and IL-4R, in particular the rs2070874, were used in their study. Hence, our results are concordant with their observation that the number of susceptibility genes is inversely correlated with the age of onset of T1DM.
Selection of controls in our study deserves comment in so far as bone marrow volunteer donors are selected for health and absence of autoimmune diseases; a feature that may have increased the differences between them and patients. Furthermore, they were not strictly matched for age range; the T1DM patients being 3–30 and controls ranging from 1 to 79 years. These differences may have introduced artefacts into the comparison between the groups and be addressed in future studies with larger numbers designed to take account of the entire age range of disease onset. Such studies will allow cumulative effects of susceptibility genes to be compared between the young and elderly. The ultimate application of this work is to develop early prophylactic screening of vulnerable individuals allowing intervention designed to arrest disease progression.
Conclusions
In this study, young T1DM patients (age of onset ≤13 years old) carrying vulnerable HLA genes had a higher frequency of the IL-4 “low producer” linked SNP rs2070874 C/C, consistent with the view that low levels of anti-inflammatory cytokines facilitate autoimmunity to β-cells.
Footnotes
Acknowledgements
This study is fully supported by King Fahad Medical City, Riyadh, Saudi Arabia. The authors would like to thank the patients and BMT donors who participated in this study.
Author’s Contribution
AO and MH designed the study. IB provided clinical information, and A AlQurashi and A Al-Jurayyan acquired information. AO and A Al-Jurayyan supervised laboratory work. AO and BB interpreted the data. AO drafted the manuscript. AO, BB, and IB revised the manuscript. BB supervised the manuscript and critically added comments.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was completely funded by King Fahad Medical City.
Ethics approval
Ethical approval for this study was obtained from Institutional Review Board (IRB) at King Fahad Medical City, Riyadh, Saudi Arabia (IRB Log No: 16–054)
Informed consent
Informed consent was not sought for the present study because we used leftover clinical blood specimens in this project and approval from our IRB was obtained.
