Abstract
Background
Nasopharyngeal carcinoma has unique epidemiological characteristics. Screening for this currently lacks a highly efficient, non-invasive, and inexpensive method. Serum microRNA (miRNA), which is stable and commonly present, has the potential to serve as a novel marker for nasopharyngeal carcinoma diagnosis.
Objectives
This study aims to find a highly efficient, non-invasive, and inexpensive biomarker for nasopharyngeal carcinoma diagnosis.
Methods
This study, involving 52 patients with nasopharyngeal carcinoma and 56 healthy controls, was conducted in two phases to identify miRNAs in the serum suitable for nasopharyngeal carcinoma diagnosis using quantitative reverse transcription polymerase chain reaction. Stepwise logistic regression analysis was then used to identify a miRNA panel with high diagnostic efficiency. Additionally, we used bioinformatic analysis to explore the potential biological functions of the crucial miRNAs.
Results
A three-miRNA panel (miR-148b-3p, miR-10b-5p, and miR-18a-5p) has a high diagnostic value for nasopharyngeal carcinoma (area under the curve = 0.872; 95% confidence interval: 0.793–0.928; sensitivity = 78.57%; specificity = 86.54%). Through bioinformatics analysis we found that CC2D2B, PCDH9, and FOXP1 may be potential target genes of these three miRNAs.
Conclusion
This three-miRNA panel (miR-148b-3p, miR-10b-5p, and miR-18a-5p) represents a highly efficient, non-invasive, and inexpensive biomarker for diagnosing nasopharyngeal carcinoma.
Introduction
Nasopharyngeal carcinoma (NPC) has unique epidemiological characteristics, with its incidence varying significantly by race, age, and gender. Among these factors, age has a significant impact on the prognosis of NPC. For the youngest group (ages 15–45), the 5-year survival rate is 72%, while for the oldest group (ages 65–74), it drops to just 36%. 1 NPC prognosis is also influenced by gender; generally, men have a worse prognosis than women.2,3 Between 2009 and 2015, the five-year survival rate for NPC patients in the USA was 60%. 1 The current diagnostic method involves the biopsy of the primary nasopharyngeal tumor through endoscopy for a definitive diagnosis. For early-stage nasopharyngeal cancer, when endoscopically guided tumors are not detected but magnetic resonance imaging or positron emission tomography scans show positive signals in the nasopharyngeal tissues, a biopsy is required to make a definitive diagnosis. 4 However, this diagnostic method is invasive, costly, and time-consuming, which often results in poor patient compliance. Therefore, there is an urgent need to develop a non-invasive, low-cost, and convenient method for diagnosing NPC.
Human microRNAs (miRNAs) are highly conserved, containing 19–23 nucleotides, and are among the most abundant molecules involved in gene regulation. They may regulate the expression of protein-coding genes. 5 The miRNA profiles contain a large amount of information that can reflect the differentiation status and type of tumors. 6 Numerous studies have shown that extracellular miRNAs in plasma and serum are associated with various cancers, making it possible to diagnose cancer by detecting and monitoring miRNA concentrations. 7 In summary, miRNAs hold significant potential for tumor diagnosis.
The objective of this research is to identify diagnostic serum miRNAs as biomarkers for screening NPC. The study was divided into four phases to evaluate the diagnostic capabilities of serum miRNAs. We amplify serum miRNAs using quantitative reverse transcription-polymerase chain reaction (RT-qPCR) to identify a set of miRNAs as novel biomarkers for NPC. Additionally, we analyzed the potential target genes of these miRNAs and their clinical significance.
Material and methods
Ethical approval and informed consent
This study involved the random selection of 108 volunteers from Peking University Shenzhen Hospital between 2017 and 2023, including 52 individuals with NPC and 56 healthy controls (HCs). For the diagnosis of nasopharyngeal cancer, we strictly followed the classification criteria for epithelial tumors of the nasopharynx established by the World Health Organization in 2016. On the other hand, for tumor node metastasis (TNM) staging and clinical staging, we strictly followed the TNM staging and clinical staging criteria for NPC developed by the American Joint Committee on Cancer (AJCC) in 2017 to ensure the accuracy of diagnosis and rationality of treatment. None of the NPC patients had metastatic tumors or cardiac insufficiency; all cases were primary tumors. All HCs were from the physical examination department of the hospital, and no disease was detected in them after thorough examination. None of the HCs had a history of malignant tumors or other acute or chronic diseases. The study was approved by the Ethics Committee of Peking University Shenzhen Hospital, and all volunteers signed informed consent forms, fully understanding and supporting the study.
Study design
Supplementary Figure 1 shows four stages of this study. Initially, we selected candidate miRNAs that were differentially expressed in NPC tissues from a database published in the Gene Expression Omnibus as potential biomarkers. We then identified 10 differentially expressed miRNAs associated with NPC or tumor-related tissues through a literature search. The training and validation phases were conducted to evaluate these candidate biomarkers. In the training phase, we used qRT-PCR to analyze 10 candidate miRNAs in serum samples from NPC patients and HCs, selecting miRNAs with differential expression for further study. In the validation phase, we expanded the sample size and remeasured the expression levels of the miRNAs identified in the previous phase. Finally, we constructed a miRNA panel with the highest diagnostic value for NPC using reverse stepwise logistic regression analysis. Finally, we performed bioinformatics analysis of the miRNAs.
Extraction of miRNAs and collection of serum samples
Serum samples were collected from untreated NPC patients, with no prior interventions, such as surgery, radiotherapy, or chemotherapy. The collection process adhered to the regulations of the Ethics Committee of Peking University Shenzhen Hospital. Initially, total RNA was extracted from the serum samples using the TRIzol LS isolation kit. To minimize variability in total RNA extraction, each serum sample was treated with 2 μL of miR-54 (10 nm/L, Ribobio). The concentration and purity of the extracted RNA were then assessed using a NanoDrop 2000c spectrophotometer.
RT-qPCR
The miRNAs were amplified using Bulge-Loop miRNA qRT-PCR primers specific to reverse transcription. The LightCycler 480 Real-Time PCR system, along with TaqMan probes, was used to detect miRNA expression levels. The RT-qPCR process involved 40 cycles at 95°C for 20 s, followed by denaturation at 95°C for 10 s, annealing at 60°C for 20 s, and extension at 70°C for 10 s.
Statistical analysis
During the testing and validation phases of the study, we used the 2−△△Cq method to determine miRNA expression levels in serum samples. 8 Additionally, we compared miRNA expression levels between two independent cohorts using the Student's t-test and the Chi-square test. Clinicopathological parameters between different cohorts were compared using the Kruskal–Wallis test and the Mann–Whitney test. To evaluate the specificity, sensitivity, and diagnostic capability of miRNAs in NPC, we used receiver operating characteristic (ROC) curves and calculated the area under the ROC curve (AUC). The optimal diagnostic group was identified using the Youden index. A P-value of less than 0.05 was considered statistically significant. All data analyses were conducted using MedCalc (Version 19), SPSS Statistics 23.0, and GraphPad Prism 8.3.0. Data were presented as numbers or percentages, along with means and standard deviations.
Bioinformatics analysis
Using miRWalk, we could predict and validate the interactions between genetically significant miRNAs and their disease-associated targets. 9 Therefore, we used miRWalk 2.0 to predict the targets of miRNAs with high diagnostic value. Additionally, these miRNAs were predicted to play a role in the onset and progression of NPC. Enrichr (http://amp.pharm.mssm.edu/Enrichr/), a comprehensive gene search engine, was used to enrich these miRNAs and annotate their target genes. 10 For functional annotation, we utilized the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) databases. OncoLnc (http://www.oncolnc.org/), an interactive tool for exploring survival relevance, was used to link the diagnostic potential of candidate miRNAs with survival data from The Cancer Genome Atlas (TCGA), resulting in the generation of Kaplan–Meier survival curves. 11
Results
Clinical and demographic characteristics of the volunteers
The study included a total of 108 participants: 52 with NPC and 56 HCs. All NPC cases were primary tumors, with no metastatic tumors or cardiac insufficiency observed. The HCs had no history of malignant tumors or other acute or chronic diseases. NPC diagnoses were based on pathological analysis of nasopharyngeal biopsy samples. The diagnosis of NPC in all patients was based on TNM staging, and clinical staging adhered to AJCC criteria. During the study's testing and validation phases, the clinical and demographic characteristics of NPC patients and HCs were compared, as shown in Table 1. There were no significant differences in gender and age between the two groups. The statistical analysis was conducted using the Wilcoxon–Mann–Whitney test, and parameters are presented as numbers (percentages).
Demographic manifestation and clinical features of 164 participants (NPC patients and HCs).
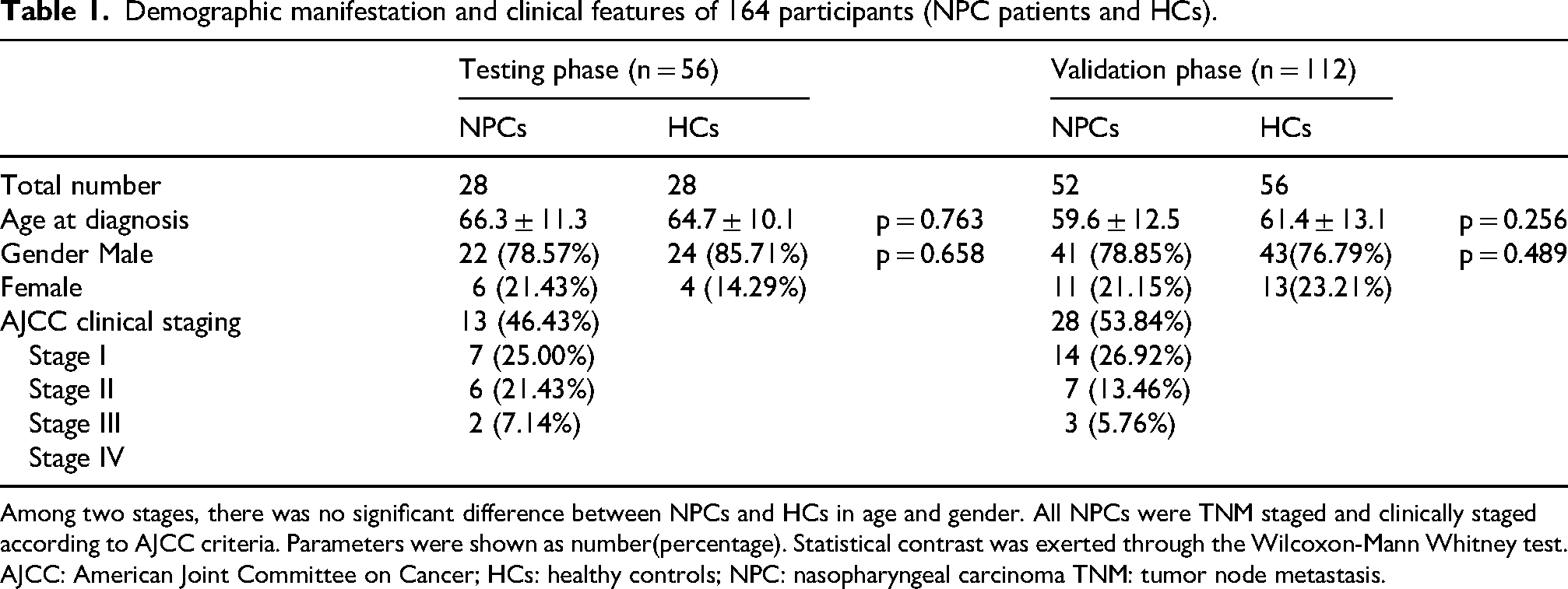
Among two stages, there was no significant difference between NPCs and HCs in age and gender. All NPCs were TNM staged and clinically staged according to AJCC criteria. Parameters were shown as number(percentage). Statistical contrast was exerted through the Wilcoxon-Mann Whitney test.
AJCC: American Joint Committee on Cancer; HCs: healthy controls; NPC: nasopharyngeal carcinoma TNM: tumor node metastasis.
Screening of candidate miRNAs
We selected 10 candidate miRNAs from the Encyclopedia of miRNA Interactomes database, including miR-196a-5p, miR-106b-5p, miR-10b-5p, miR-126-3p, miR-150-5p, miR-148b-3p, miR-129-1-3p, miR-18a-5p, miR-143-3p, and miR-29b-3p (Figure 1).
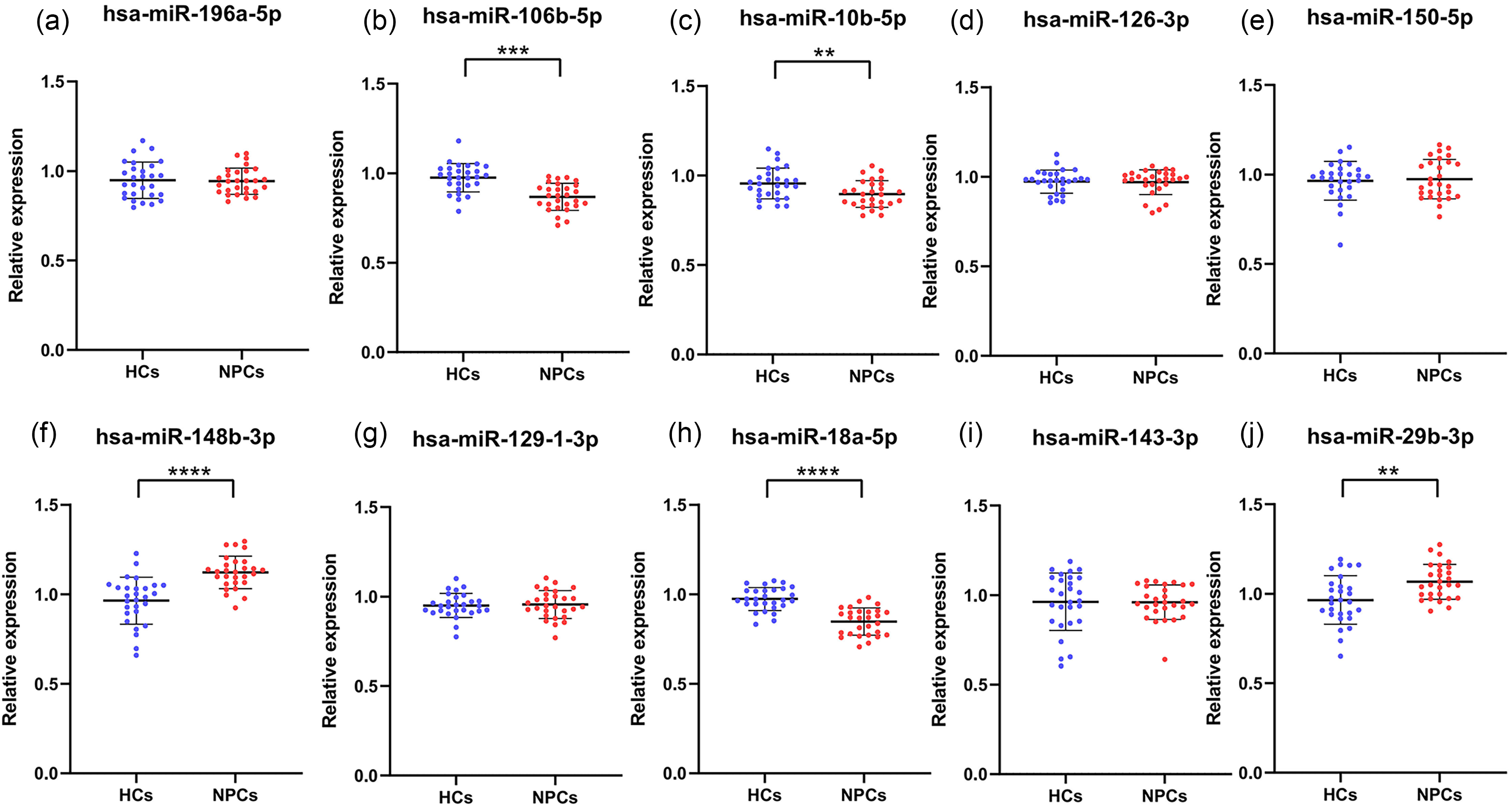
Ten candidate miRNAs were tested for their expression levels in the training phase. Five miRNAs were chosen with differential expression as candidate miRNAs for the validation phase. Conditions for screening: **P < 10−2, ***P < 10−3, ****P < 10−4. In the training phase, serum from 28 NPCs and 28 HCs was used. The blue dots denote HCs, while the red dots denote NPCs. The ordinate denotes the relative expression levels of serum miRNAs.
Training phase: identification of candidate miRNAs
During the training phase, we randomly selected 28 NPC patients and 28 HCs from the 108 participants. We used qRT-PCR analysis to determine whether there were significant differences in the expression of the 10 candidate miRNAs between NPC patients and HCs. The results are shown in Figure 1. After analysis, we identified five miRNAs with significant differences in serum samples between NPC patients and HCs: miR-106b-5p, miR-10b-5p, miR-148b-3p, miR-18a-5p, and miR-29b-3p. We then proceeded to further validate these five miRNAs.
Validation of candidate miRNAs as diagnostic biomarkers
To further verify the diagnostic capability of the five miRNAs and their potential as novel biomarkers for NPC screening in the validation phase, we selected 52 NPC patients and 56 HCs. According to the results of qRT-PCR analysis, Figure 2 shows that the relative expression levels of miR-106b-5p (Figure 2(a)), miR-10b-5p (Figure 2(c)), miR-148b-3p (Figure 2(e)), miR-18a-5p (Figure 2(g)), and miR-29b-3p (Figure 2(i)) in serum samples were significantly decreased in NPC patients.
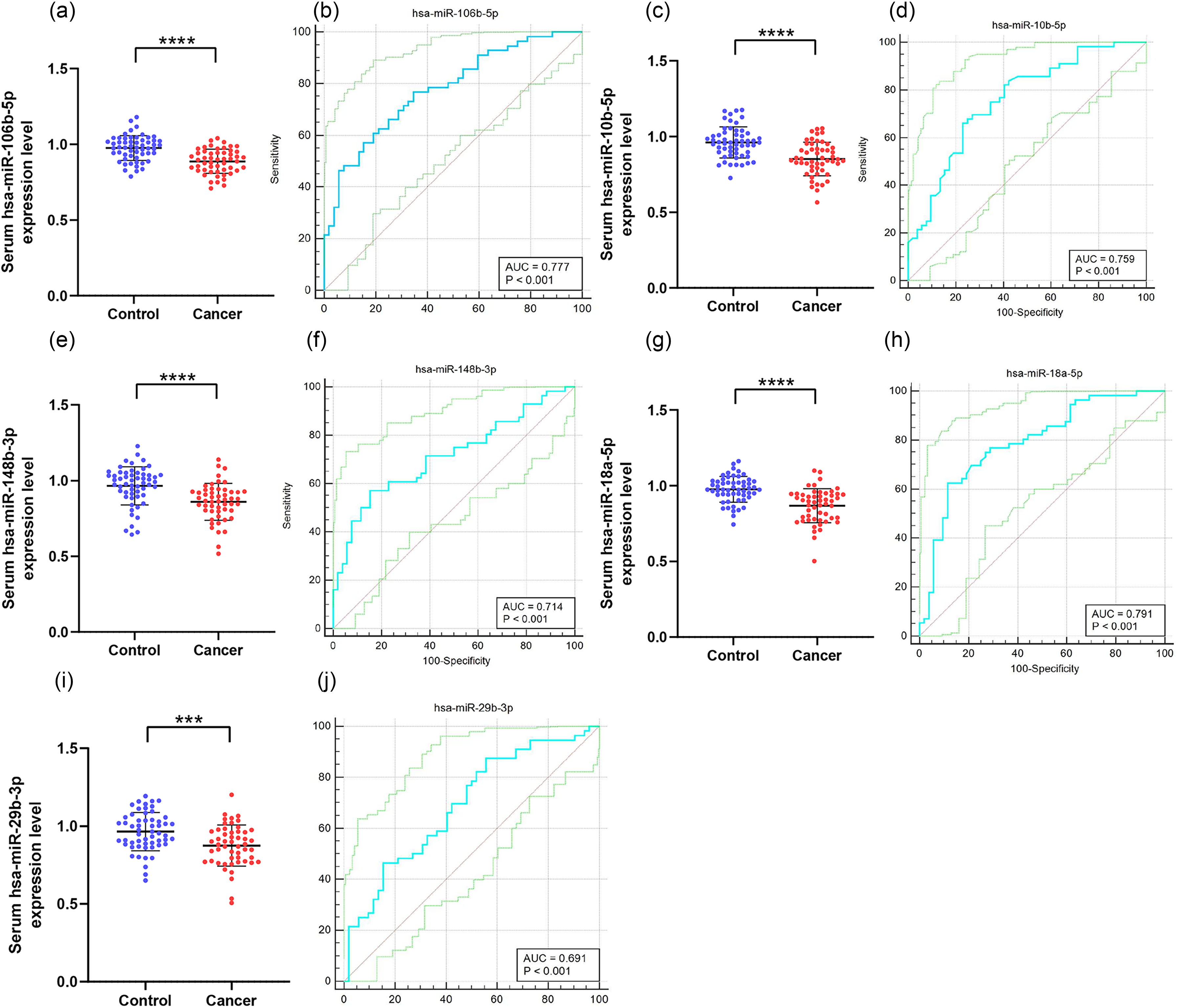
The image shows the variances in the levels of expression for five miRNAs detected in serum samples collected from 52 NPCs as well as 56 HCs, along with the corresponding ROC curve for these miRNAs. The control group is denoted by blue dots, while the cancer group is denoted by red dots. *** P < 10−3, **** P < 10−4. The ROC curve is represented by the blue line, while the 95% confidence interval for the ROC curve is indicated by the green line. The red line corresponds to the diagonal. The ordinate of (a), (c), (e), (g), and (i) denotes the relative expression levels of serum miRNAs. The ordinate of (b), (d), (f), (h) and (j) denotes the sensitivity of this miRNAs to screen for NPCs, while the ordinate denotes the specificity. HCs: healthy controls; miRNA: micro RNA; NPC: nasopharyngeal carcinoma; ROC: receiver operating characteristic.
To assess the diagnostic capability of these 5 miRNAs in NPC, we evaluated them using ROC curves. The AUC was 0.777 (95% confidence interval (CI): 0.687–0.852; Figure 2(b)) for miR-106b-5p, 0.759 (95% CI: 0.667–0.836; Figure 2(d)) for miR-10b-5p, 0.714 (95% CI: 0.619–0.796; Figure 2(f)) for miR-148b-3p, 0.791 (95% CI: 0.702–0.863; Figure 2(h)) for miR-18a-5p, and 0.691 (95% CI: 0.595–0.776; Figure 2(j)) for miR-29b-3p. The optimal sensitivity and specificity for NPC diagnosis are provided in Supplementary Table 1. Based on the ROC curve analysis, miR-10b-5p, miR-148b-3p, and miR-18a-5p demonstrated moderate diagnostic capability for NPC, with AUC values of 0.759, 0.714, and 0.791, respectively.
Searching for the optimal miRNA diagnostic panels
The accuracy of NPC diagnosis using a diagnostic panel formed by multiple miRNAs may be greater than that of a single miRNA. During the validation phase, miR-148b-3p, miR-10b-5p, and miR-18a-5p were identified as moderately capable of diagnosing NPCs. Consequently, we constructed a diagnostic panel consisting of these three miRNAs. The optimal combination for screening NPC was determined using a stepwise logistic regression model, which evaluated all possible combinations of the three miRNAs. The model's calculation formula is derived as follows: Logit(P) = 20.651 + (−8.459 × miR-10b-5p) + (−5.227 × miR-148b-3p) + (−8.991 × miR-18a-5p). Supplementary Figure 2 presents the ROC curve of this diagnostic panel. The AUC of this diagnostic panel was 0.872 (95% CI: 0.793–0.928), with a sensitivity of 78.57% and specificity of 86.54%, as shown in Supplementary Table 1.
Candidate miRNAs: biological analysis
Potential target genes for miR-10b-5p, miR-148b-3p, and miR-18a-5p were predicted using MiRWalk 2.0. We identified a set of 31 genes predicted to be targeted by these miRNAs, with three genes (CC2D2B, PCDH9, and FOXP1) showing differential expression in NPCs. Survival analysis was conducted on these three target genes, as illustrated in Figure 3. Each survival analysis included two groups: high expression and low expression. The P-values from the Log rank test for the three groups were 0.014, 0.018, and 0.025, respectively—all below 0.05—indicating a significant correlation between long-term survival and gene expression levels. Furthermore, the hazard ratio for the high-expression groups of CC2D2B and FOXP1 were 0.71 and 0.74, respectively—both below 1—suggesting a protective effect against NPC. The P-values for HR were 0.015 and 0.026, respectively, both below 0.05, indicating statistical significance. Conversely, the HR for the high-expression group of PCDH9 was 1.4, indicating that this gene was a risk factor for NPC. Its P-value for HR was 0.019, also below 0.05, indicating significant statistical relevance. These findings suggest that these three genes might have the potential to be targeted by the miRNA screening panel.
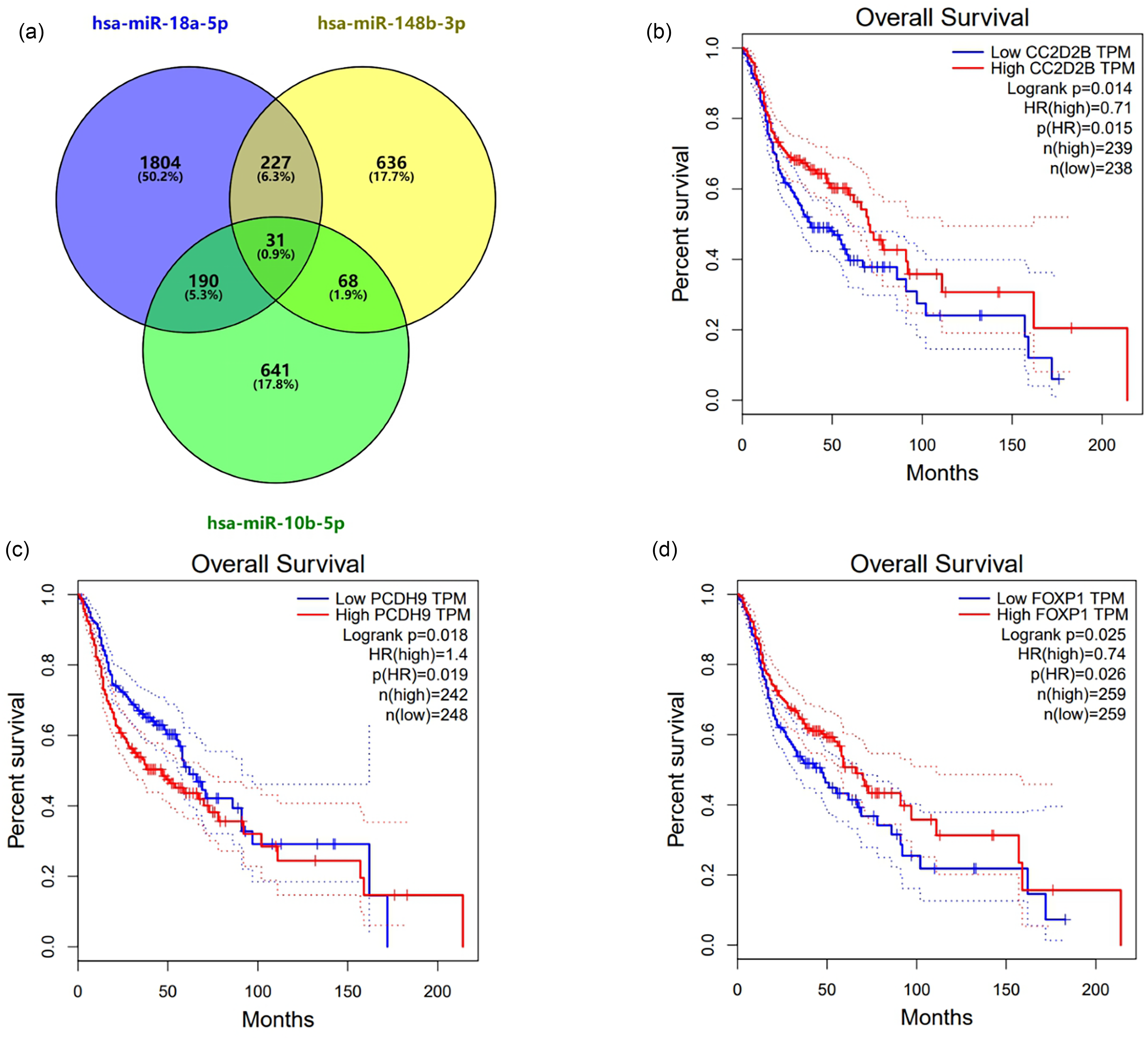
(a): Utilizing miRWalk target gene prediction and Enrichr annotation to identify 31 relevant genes. Relevant genes were selected as targets based on the prediction of three shared genes. (b) to (d): Out of the 31 genes that were predicted as potential targets, basing on the GEPIA database, there are three genes having diversities in the survival analysis: CC2D2B (b), PCDH9 (c), FOXP1 (d).
Subsequently, we conducted further research using the Enrichr database for KEGG pathway enrichment analysis and GO function annotation. The analysis revealed that relevant genes were expressed in pathways such as cell adhesion molecules, cholinergic synapse, ubiquitin mediated proteolysis, and focal adhesion (Figure 4(d)). GO functional annotations covered three categories: biological processes (BP), cellular components (CC), and molecular functions (MF), as shown in Figure 4(a) to (c). Significant annotations included Regulation of mRNA Splicing, Via Spliceosome (GO:0048024), Positive Regulation of Sodium Ion Transport (GO:0010765), and Modulation of Chemical Synaptic Transmission (GO:0050804) in the BP category; Neuron Projection (GO:0043005), Glutamatergic Synapse (GO:0098978), and Dendrite (GO:0030425) in the CC category; and Amino Acid: Sodium Symporter Activity (GO:0005283), Histone H3 Methyltransferase Activity (GO:0140938), and SH3 Domain Binding (GO:0017124) in the MF category.
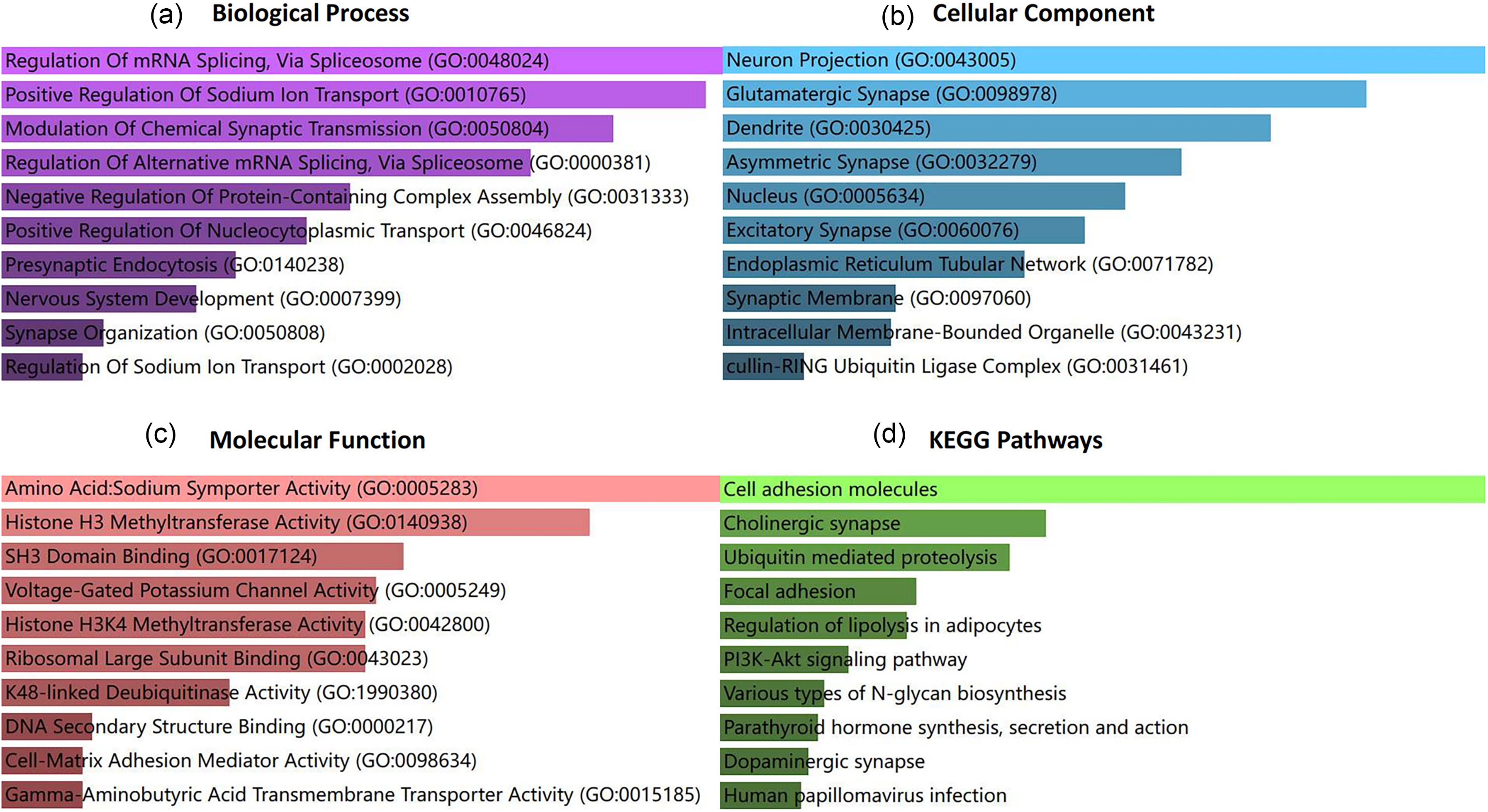

Go functional annotation (a) to (c) and enrichment analysis of KEGG pathways (d) were performed for the target genes regulated by miR-148b-3p, miR-10b-5p, and miR-18a-5p.
Relationship between relative expression of serum miRNAs and clinical characteristics
The Kruskal–Wallis test was used to assess whether the expression levels of the three selected miRNAs (miR-10b-5p, miR-148b-3p, miR-18a-5p) showed statistically significant differences across clinical characteristics of the disease. The analysis included subjects from the validation cohort. When NPC patients in the validation cohort were grouped according to primary tumor extent (T-stage), lymph node metastasis (N-stage), and distant metastasis (M-stage), no significant associations were observed between the expression levels of the three miRNAs and these clinical characteristics (Supplementary Table 2).
Discussion
The literature extensively emphasizes the pivotal role of miRNA expression in regulating cell proliferation, differentiation, and survival. In addition, miRNAs are significant in the development of malignancies by influencing cellular metabolic pathways and cell–cell communication. 12 Because miRNA expression is important for distinguishing normal cells from tumor cells, it has emerged as a new biological marker for non-invasive cancer diagnosis. 13 In our study, we aimed to identify a panel of miRNAs for the accurate screening of NPC. Our findings revealed significant differences in the levels of three miRNAs (miR-148b-3p, miR-10b-5p, and miR-18a-5p) between serum samples from individuals with NPC and HCs, as confirmed through a comprehensive screening, training, and validation process. Therefore, we developed a diagnostic panel consisting of these three miRNAs (AUC = 0.872) to screen for NPC and to explore the potential roles and clinical importance of each miRNA.
MiR-148b-3p expression (AUC = 0.714; 95% CI: 0.619–0.796) was significantly reduced in NPCs within the diagnostic group. The literature consistently highlights the significant under-expression of miR-148b-3p in osteosarcoma (OS) cells. However, upregulation of miR-148b-3p in OS cells has been shown to inhibit cellular proliferation, migration, and invasion while simultaneously promoting apoptotic responses. Therefore, targeting miR-148b-3p could be a potential therapeutic strategy for OS. 14 Further research has also suggested that miR-148b-3p may serve as a potential biomarker for prostate cancer. 15 Comparing our findings with previous studies, it was evident that miR-148b-3p played a significant role in the pathogenesis of various cancers.
MiR-10b-5p (AUC = 0.759; 95% CI: 0.667–0.836) was significantly reduced in NPCs within the diagnostic group. Several studies have demonstrated that miR-10b-5p inhibits the occurrence and progression of gastric cancer by downregulating Tiam1. 16 Other research highlighted the inhibitory effects of miR-10b-5p on the proliferation and invasion of primary liver cancer cells, as well as its role in promoting cell apoptosis by downregulating hepatocyte receptor A2 (EphA2) expression induced by erythropoietin. 17 Our research also indicated that miR-10b-5p might have significant screening potential for NPC.
MiR-18a-5p expression (AUC = 0.791; 95% CI: 0.702–0.863) was significantly reduced in NPCs within the diagnostic group. A substantial body of literature highlights the crucial role of miR-18a-5p in suppressing the incidence and progression of HER2-positive breast cancer by modulating the PI3 K/AKT signaling pathway. 18 Additionally, miR-18a-5p is significant in the growth of esophageal squamous cell carcinoma (ESCC), suggesting that it could improve future diagnosis and treatment of ESCC. 19 Our study indicated that miR-18a-5p could be effectively used to screen for NPC.
Our findings suggest that CC2D2B, PCDH9, and FOXP1 are potential target genes for the miRNA diagnostic panel. Literature has highlighted that CC2D2B is a significant shared probe between brain metastatic papillary thyroid carcinoma and primary brain tumors. 20 Overexpression of PCDH9 inhibits the migration of melanoma cells and may be a potential target for melanoma therapy. 21 PCDH9 is poorly expressed in melanoma cells, and its overexpression can induce cell cycle arrest in the G0/G1 phase, thereby inhibiting tumor progression. 22 Increased FOXP1 expression is associated with various malignant tumors. FOXP1 acts as a suppressor of P21 and retinoblastoma protein transcription and interacts directly with the tumor suppressor p53 to inhibit OS activity. 23 Given these findings and our research results, CC2D2B, PCDH9, and FOXP1 are likely important in the development and progression of NPC. Further investigation into their specific mechanisms is warranted.
Although our research presented clinically significant findings, there were some limitations. Firstly, the number of NPC patients in our study was relatively small. Due to the low incidence of NPC in our region, and the strict enrollment criteria, we were unable to collect as many samples as possible over a long period of time. However, the results from the experimental data are satisfactory, but if the sample size is larger, the experiment will be more convincing. Secondly, there are many potential confounding factors in the experiment. Although we excluded the influence of gender, age, and other diseases on the experiment, we did not evaluate the influence of tumor grade and stage. The group of patients selected for this study was not homogeneous in terms of staging, with the predominant proportion of patients being in Stage I and Stage II. This distribution of stages may have introduced a source of heterogeneity that could potentially affect the validity of our findings. In future studies, we will consider using stratified statistical methods to better understand the relationship between miRNA panels and different stages of nasopharyngeal cancer. Meanwhile, we will try to recruit a more balanced sample population of NPC of all stages to minimize the impact of stage heterogeneity on the study results. Thirdly, as serum samples were collected before treatment, we could not assess whether miRNA levels changed post-treatment. Future studies should focus on tracking changes in miRNA levels after treatment. Fourthly, the study is currently in an exploratory phase, where we conclude that a three-miRNA panel can be used to diagnose nasopharyngeal cancer, but there is no explicit cut-off when comparing healthy samples with NPC samples. More studies are needed before it can be used in the clinic. Fifthly, miR-148b-3p, miR-10b-5p, andmiR-18a-5p are expressed in a variety of solid tumors and belong to pan-target genes. The specific studies on the roles in NPC are relatively limited, therefore more experimental studies and clinical data are needed in the future to reveal the specific mechanisms and specificity of these miRNAs in laryngeal cancer. Meanwhile, due to the complexity and diversity of miRNAs, their expression and role may differ in different types of tumors, so further research is needed for specific tumor types.
Conclusion
The three-miRNA diagnostic panel (miR-148b-3p, miR-10b-5p, and miR-18a-5p) has high efficiency in NPC diagnosis (AUC = 0.872; 95% CI: 0.793–0.928; sensitivity = 78.57%; specificity = 86.54%). It represented a highly efficient, non-invasive, and inexpensive biomarker for NPC diagnosis.
Supplemental Material
sj-tif-1-jbm-10.1177_03936155251329041 - Supplemental material for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma
Supplemental material, sj-tif-1-jbm-10.1177_03936155251329041 for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma by Zhenjian Ge, Pengwu Zhang, Yong Xia, Chong Lu, Chen Sun, Zhenyu Wen, Wenkang Chen, Yingqi Li, Shengjie Lin, Yutong Wu, Xutai Li, Huimei Zhou, Wuping Wang, Siwei Chen, Ling Ji and Yongqing Lai in The International Journal of Biological Markers
Supplemental Material
sj-tif-2-jbm-10.1177_03936155251329041 - Supplemental material for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma
Supplemental material, sj-tif-2-jbm-10.1177_03936155251329041 for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma by Zhenjian Ge, Pengwu Zhang, Yong Xia, Chong Lu, Chen Sun, Zhenyu Wen, Wenkang Chen, Yingqi Li, Shengjie Lin, Yutong Wu, Xutai Li, Huimei Zhou, Wuping Wang, Siwei Chen, Ling Ji and Yongqing Lai in The International Journal of Biological Markers
Supplemental Material
sj-docx-3-jbm-10.1177_03936155251329041 - Supplemental material for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma
Supplemental material, sj-docx-3-jbm-10.1177_03936155251329041 for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma by Zhenjian Ge, Pengwu Zhang, Yong Xia, Chong Lu, Chen Sun, Zhenyu Wen, Wenkang Chen, Yingqi Li, Shengjie Lin, Yutong Wu, Xutai Li, Huimei Zhou, Wuping Wang, Siwei Chen, Ling Ji and Yongqing Lai in The International Journal of Biological Markers
Supplemental Material
sj-docx-4-jbm-10.1177_03936155251329041 - Supplemental material for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma
Supplemental material, sj-docx-4-jbm-10.1177_03936155251329041 for A three-miRNA panel in serum: Serving as a novel diagnostic method for nasopharyngeal carcinoma by Zhenjian Ge, Pengwu Zhang, Yong Xia, Chong Lu, Chen Sun, Zhenyu Wen, Wenkang Chen, Yingqi Li, Shengjie Lin, Yutong Wu, Xutai Li, Huimei Zhou, Wuping Wang, Siwei Chen, Ling Ji and Yongqing Lai in The International Journal of Biological Markers
Footnotes
Acknowledgments
Not applicable.
Ethical considerations
The research protocol received approval from the Ethics Committee of Peking University Shenzhen Hospital (2019-043). The research enrolled participants who all provided written informed consent.
Consent for publication
Not applicable.
Author contributions/CRediT
The research concept and design were contributed by Yongqing Lai and Ling Ji. Experiments were conducted by Zhenjian Ge and Pengwu Zhang, who also authored the manuscript. Yong Xia, Shengjie Lin, Yutong Wu, Xutai Li, Huimei Zhou, Wuping Wang, and Siwei Chen collected the samples. Chong Lu, Chen Sun, Zhenyu Wen, Wenkang Chen, and Yingqi Li performed the statistical analysis. All the authors have thoroughly reviewed and given their endorsement to the final version of the manuscript.
Zhenjian Ge and Pengwu Zhang contributed equally to this work.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Shenzhen Clinical Research Center for Urology and Nephrology, Shenzhen High-level Hospital Construction Fund, Clinical Research Project of Peking University Shenzhen Hospital, (grant number LCYSSQ20220823091403008, LCYJ2017001, LCYJ2020002, LCYJ2020015, LCYJ2020020).
Conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Statement on the accessibility of data
The corresponding author can provide the data supporting this study upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
