Abstract
Staphylococcal infection is associated with significant morbidity and mortality in critically ill patients. Using data from 16,681 patients who had a nasal Staphylococcus aureus polymerase chain reaction (PCR) assay on admission to the intensive care unit (ICU) of Royal Perth Hospital between March 2006 and September 2016, this retrospective cohort study assessed whether nasal S. aureus colonisation on admission to an ICU was predictive of concurrent or subsequent S. aureus infections. Culture-proven S. aureus infections were identified using the hospital microbiology database. Of the 16,681 patients included, 565 (3.4%) had a positive methicillin-resistant S. aureus (MRSA) assay, 146 (0.9%) had a positive methicillin-sensitive S. aureus (MSSA) assay and eight (0.05%) had both positive MRSA and MSSA assays. Of those 565 patients with a positive MRSA PCR assay, 79 (13.8%) had concurrent or subsequent MRSA infections. Of those 146 patients with a positive MSSA PCR assay, only 5 (3.4%) had MSSA infection. The sensitivity and specificity for the MRSA PCR assay in predicting concurrent or subsequent MRSA infection were 72.7% (95% confidence intervals (CI) 63.4%–80.8%) and 97.0% (95% CI 96.8%–97.3%), respectively. The sensitivity and specificity for the MSSA PCR assay in predicting concurrent or subsequent MSSA infection were 3.3% (95% CI 1.1%–7.6%) and 99.1% (95% CI 98.9%–99.2%), respectively. Both nasal MRSA and MSSA PCR assays had a high specificity and negative predictive value in predicting MRSA and MSSA infections, respectively, suggesting that in centres without endemic S. aureus infections, a negative nasal MRSA or MSSA PCR assay may be useful to reduce unnecessary empirical antibiotic therapy against S. aureus.
Introduction
Staphylococcus aureus infections are common 1 and are associated with substantial morbidity and mortality.2–4 In 2015–2016, 1440 cases of hospital-acquired bacteraemia due to either methicillin-resistant S. aureus (MRSA; 19%) or methicillin-sensitive S. aureus (MSSA; 81%), with an overall incidence of 0.73 per 10,000 days of patient care, were reported in Australian public hospitals. 5 S. aureus bacteraemia is associated with >10% hospital mortality,2,3 and MRSA bacteraemia is often empirically treated with antibiotics to which the pathogen is not susceptible in vitro while cultures and susceptibilities are pending, resulting in prolonged bacteraemia and increased incidence of metastatic seeding of infection compared to MSSA bacteraemia. 3 On the other hand, unnecessary anti-MRSA therapy also has significant risks at both patient and community levels, including nephrotoxicity from vancomycin, anaphylaxis to teicoplanin 6 and development of vancomycin-resistant Enterococci. 7 As such, it is imperative to develop rapid and reliable ways to identify patients who are at high risk of MRSA infection—in particular MRSA bacteraemia—so that prompt appropriate empirical antibiotics can be initiated while minimising overuse of broad-spectrum antibiotics. 8
S. aureus can be found in different body sites such as the skin, rectum, vagina, gastrointestinal tract and axilla. Once the bacteria are in contact with the nasal mucosa, they will interact with epithelial cell ligands, such as loricrin and cytokeratin 10 (K10), and colonise the anterior nares. 9 It has been estimated that up to 30% of the human population are asymptomatically and permanently colonised with S. aureus in their anterior nares. 9 In critically ill patients, recent studies have suggested that nasal colonisation with S. aureus is associated with a higher subsequent risk of S. aureus infections, including bacteraemia.10–12 In addition, nasal decolonisation of S. aureus is associated with a reduced risk of S. aureus pneumonia in critically ill patients and surgical site infection prior to surgery.13,14 Nonetheless, whether nasal S. aureus colonisation—confirmed by a polymerase chain reaction (PCR) assay—on admission to an intensive care unit (ICU) is predictive of all forms of subsequent MRSA and MSSA infections is uncertain.
We hypothesised that nasal MRSA and MSSA colonisation in the critically ill on admission to an ICU can be predictive of MRSA and MSSA infections, respectively. In this retrospective cohort study, we aimed to determine the associations between a positive nasal MRSA/MSSA PCR assay on admission to ICU and concurrent or subsequent S. aureus infections during the same hospitalisation.
Methods
Patients and setting
Nasal swabs were taken for all patients admitted to the ICU of Royal Perth Hospital within the first hour of admission to identify MRSA colonisation for infection control purposes. Nasal swabs were taken only on admission to the ICU, including readmissions, but not during ICU stay. Contact precaution measures including use of an isolation room would be implemented for patients who were identified as MRSA colonised. Eradication therapy, such as topical mupirocin, was not used. In this study, 16,681 consecutive patients who were admitted to the ICU between 30 March 2006 and 30 September 2016 were included. Patients admitted to the co-located high dependency unit were not screened for MRSA and were excluded from this study unless they were subsequently admitted to the ICU. For patients who had more than one ICU admission during the same hospitalisation, only the nasal MSSA and MRSA PCR results in the index ICU admission were used in this study.
The nasal colonisation for MRSA and MSSA was determined by a PCR assay using either the IDI-MRSA (from March 2006 to June 2013) or the automated BD Max (from July 2013 to the end of study period) system, with the results available within three hours in the study centre.15,16 The first-generation BD Max MRSA assay was not able to identify correctly the mecA empty cassette or the MRSA strains with methicillin resistance gene mecC and some MRSA mec right-extremity junction type strains. 17 In this study, patients with a positive MRSA or MSSA PCR assay, and any concurrent or subsequent culture-confirmed MRSA or MSSA infections during the same hospital stay, were identified using the hospital microbiology database. An infection due to S. aureus was defined as either growth of S. aureus from a sterile site (e.g. blood or joint fluid) or identification of S. aureus from a non-sterile site (e.g. wound or sputum) while there was clinical and/or radiological evidence of infection and no other cause/pathogen could be identified. The microbiology data of the study patients were then merged with the ICU administrative database for this clinical audit (CSQU approval number 13773).
Outcomes
The primary outcome of this study was the ability of a positive nasal MRSA or MSSA PCR assay in predicting concurrent or subsequent culture-proven MRSA or MSSA infection during the same hospital stay. Sensitivity, specificity, positive and negative predictive values, positive and negative likelihood ratios and area under the receiver operating characteristic curve were used to describe the ability of a nasal MRSA and MSSA PCR assay to predict concurrent or subsequent MRSA and MSSA infections, respectively.
Statistical analyses
Because we did not have prior local data on the incidence of a positive nasal MRSA or MSSA PCR assay, the number of patients included was maximised by including all patients who were admitted to the ICU after MRSA/MSSA PCR was initiated for routine use as an infection control measure until initiation of this audit.
Descriptive categorical and continuous data are presented as the number (%) and median (interquartile range (IQR)), respectively. In addition to sensitivity and specificity, the ability of a positive nasal MRSA PCR assay to predict concurrent or subsequent MRSA infection was further assessed by logistic regression with and without adjusting for important covariates in a subgroup of patients with either MRSA/MSSA infection or a positive nasal MRSA/MSSA PCR assay. The covariates used for adjustment were markers of severity of acute and chronic illnesses which might confound the relationship between colonisation and infection. The time from confirming MSSA/MRSA colonisation to infection was not analysed because many of our study patients had concurrent MSSA/MRSA infections instead of development of such infections subsequently. All statistical analyses were conducted using IBM SPSS Statistics for Windows v24.0 (IBM Corp., Armonk, NY, USA) and MedCalc for Windows v12.5 (MedCalc Software Ltd, Ostend, Belgium), and a two-tailed alpha error of <5% was taken as significant. De-identified study data are available on request from the corresponding author.
Results
Of the 16,681 patients eligible and included in the study, 565 (3.4%) had a positive MRSA PCR assay, 146 (0.9%) had a positive MSSA PCR assay, and eight (0.05%) had both positive MRSA and MSSA PCR assays. The rates of MRSA infection (109/16,681; 0.65%) and colonisation (573/16,681; 3.4%) in patients admitted to our ICU were relatively low. Of the 719 patients who had a positive MRSA or MSSA PCR assay, 94 (13%) had coexisting or subsequent MRSA or MSSA infections compared to only 1.0% incidence of MRSA or MSSA infections among those who did not have a positive MRSA or MSSA PCR assay (n = 15,962 patients; Figure 1). Of the 109 and 151 culture-proven MRSA and MSSA infections, 62% and 92% were isolated from sterile sites, including blood culture, joint fluid, pleural fluid or tissue (e.g. heart valve, bone and vascular grafts; Table 1), respectively.
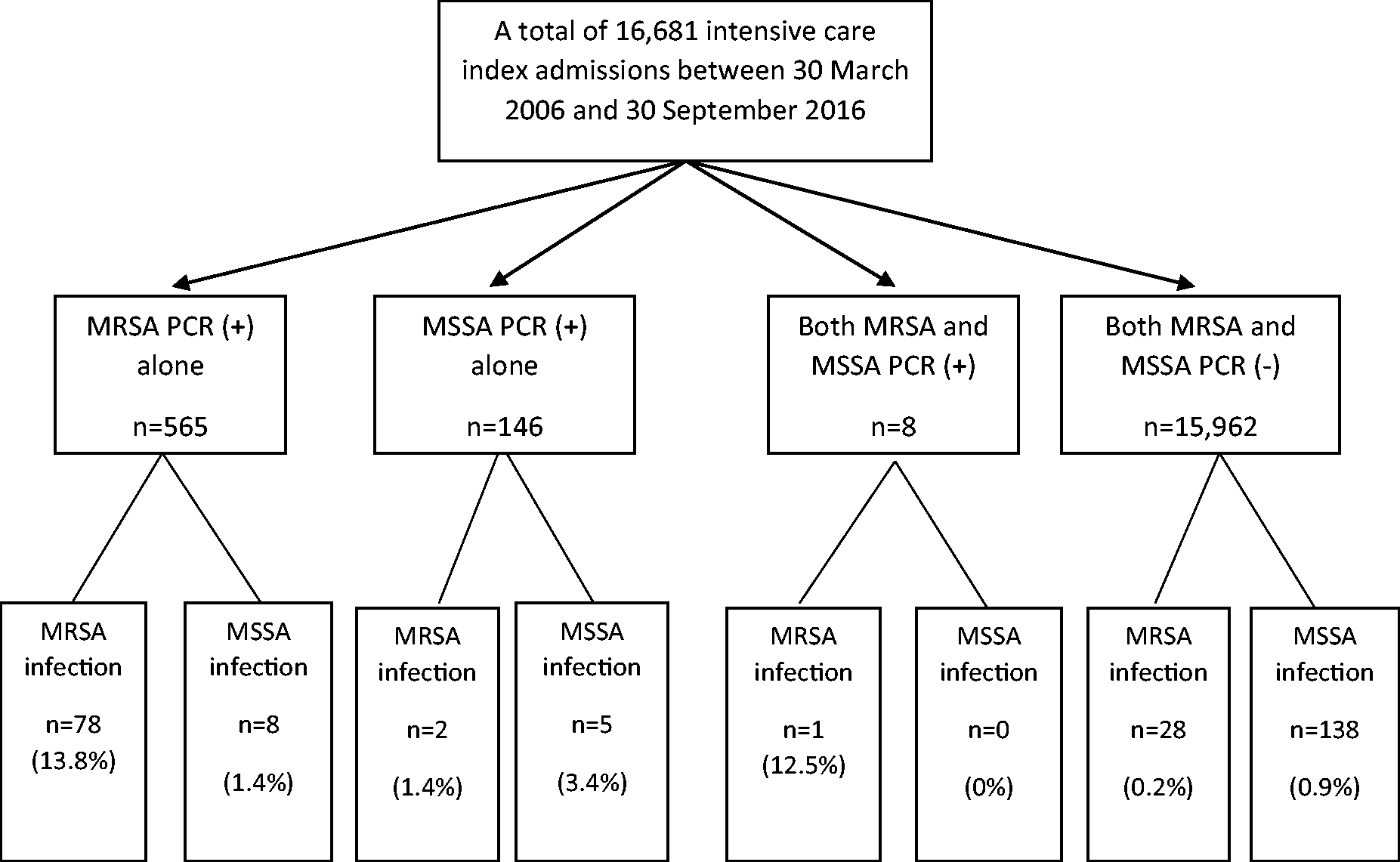
Flow chart showing the patients with and without a positive polymerase chain reaction (PCR) for methicillin-sensitive Staphylococcus aureus (MSSA) or methicillin-resistant Staphylococcus aureus (MRSA) and subsequent risk of MSSA or MRSA culture-proven infection. The proportion of patients and incidence rate of MRSA infections (estimated using median length of hospital stay of 15 days) among those without a positive nasal MRSA PCR assay were 0.18% and 0.44 per 1000 patient years, respectively. Two patients with negative MRSA and MSSA PCR assays had both MRSA and MSSA infections.
Characteristics of patients who had either a positive PCR assay for MSSA/MRSA or MSSA/MRSA culture-proven infection (N = 883).
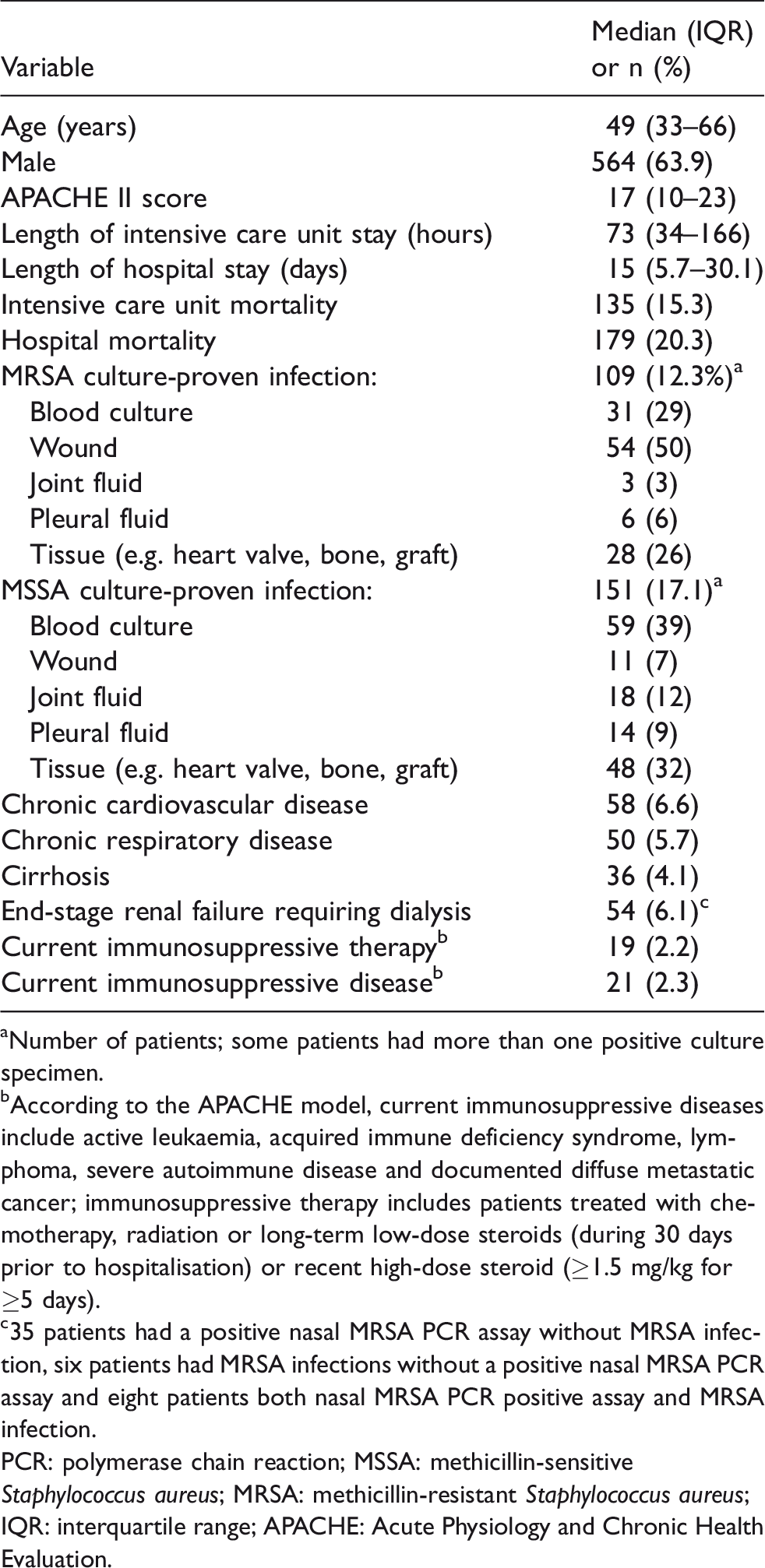
aNumber of patients; some patients had more than one positive culture specimen.
bAccording to the APACHE model, current immunosuppressive diseases include active leukaemia, acquired immune deficiency syndrome, lymphoma, severe autoimmune disease and documented diffuse metastatic cancer; immunosuppressive therapy includes patients treated with chemotherapy, radiation or long-term low-dose steroids (during 30 days prior to hospitalisation) or recent high-dose steroid (≥1.5 mg/kg for ≥5 days).
c35 patients had a positive nasal MRSA PCR assay without MRSA infection, six patients had MRSA infections without a positive nasal MRSA PCR assay and eight patients both nasal MRSA PCR positive assay and MRSA infection.
PCR: polymerase chain reaction; MSSA: methicillin-sensitive Staphylococcus aureus; MRSA: methicillin-resistant Staphylococcus aureus; IQR: interquartile range; APACHE: Acute Physiology and Chronic Health Evaluation.
Of the 883 patients who had either a positive nasal MRSA/MSSA PCR assay or MRSA/MSSA infection, infection/sepsis was the commonest clinical admission diagnosis (n = 160), followed by trauma (n = 75), postoperative management after major surgery (n = 63), acute cardiovascular diseases (n = 51), acute neurological diseases (n = 50) and acute respiratory diseases (n = 44). The characteristics of these 883 patients are described in Table 1.
Of those 565 patients with a positive MRSA PCR assay, 79 (13.8%) had concurrent MRSA infections on ICU admission (n = 72) or developed MRSA infections subsequently (n = 7; median = 2 days, IQR 1–11); 78/79 patients had a positive MRSA PCR assay alone, but one patient had both positive MRSA and MSSA PCR assays. Of those 146 patients with a positive MSSA PCR assay, five (3.4%) had concurrent MSSA infection (n = 1) or developed subsequent MSSA infection (n = 4; median = 3 days, IQR 1–6), and two (1.4%) developed MRSA infection (Figure 1). The proportion of patients and incidence rate of MRSA infections (estimated using median length of hospital stay of 15 days) among those without a positive nasal MRSA PCR assay were 0.18% and 0.44 per 1000 patient years, respectively.
Both nasal MRSA and MSSA PCR assays had a high specificity and negative predictive value but low sensitivity and positive predictive value in predicting concurrent or subsequent MRSA and MSSA infections, respectively. The sensitivity and specificity for a positive MRSA PCR assay for MRSA infection were 72.7% (95% confidence interval (CI) 63.4%–80.8%) and 97.0% (95% CI 96.8%–97.3%), respectively. The sensitivity and specificity for a positive MSSA PCR assay for MSSA infection were 3.3% (95% CI 1.1%–7.6%) and 99.1% (95% CI 98.9%–99.2%), respectively (Table 2).
Predictive parameters of MSSA and MRSA PCR assays for concurrent or subsequent MSSA and MRSA culture-proven infection, respectively.
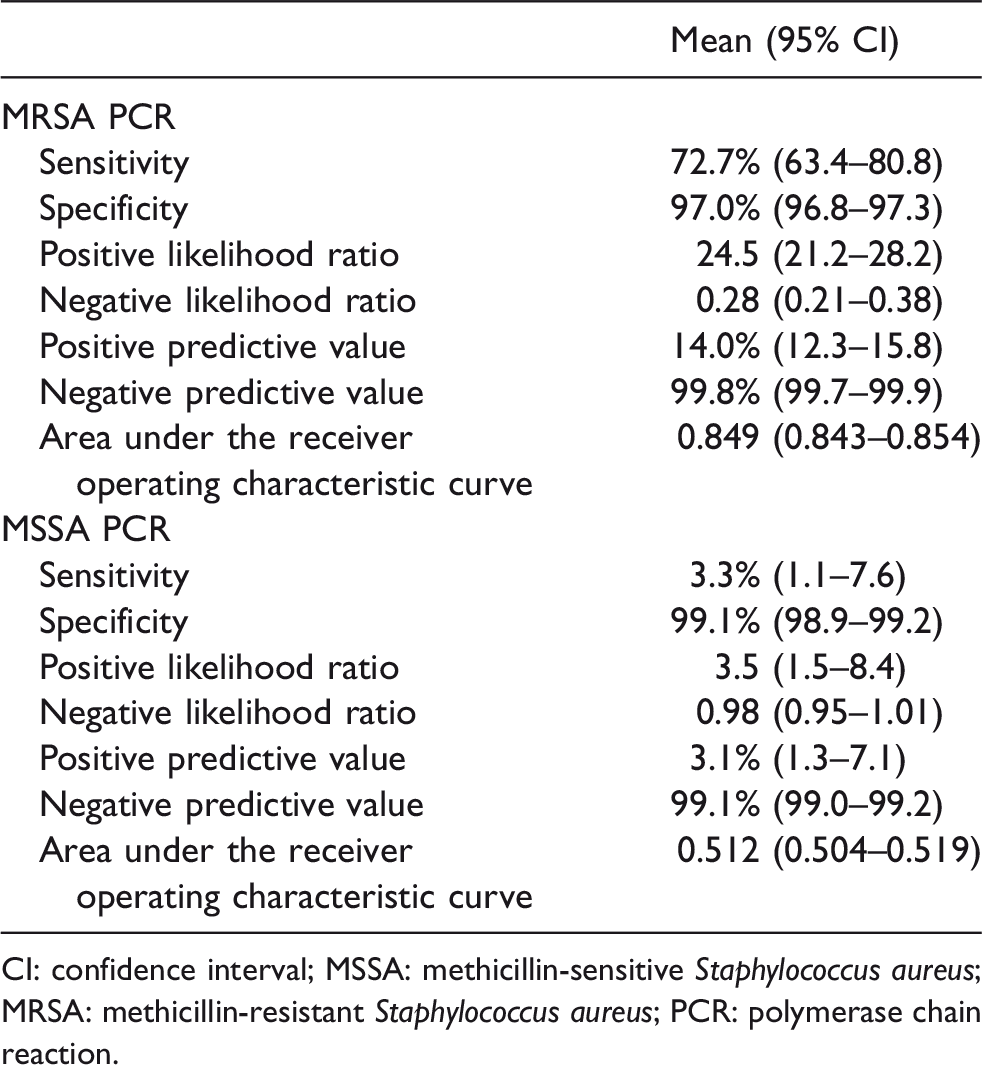
CI: confidence interval; MSSA: methicillin-sensitive Staphylococcus aureus; MRSA: methicillin-resistant Staphylococcus aureus; PCR: polymerase chain reaction.
In the univariable logistic regression analysis, there was a suggestion that a positive nasal MRSA PCR was associated with an increased risk of having MRSA infection, but this was not statistically significant (odds ratio (OR) = 1.55, 95% CI 0.99–2.43, P =0.056). This result remained unchanged (OR = 1.64, 95% CI 0.96–2.79, P = 0.068) after adjusting for age, Acute Physiology and Chronic Health Evaluation II score and coexisting diseases (Table 3). End-stage renal failure requiring dialysis was the only major comorbidity associated with an increased risk of MRSA infection (OR = 2.08, 95% CI 1.01–4.26, P = 0.047).
Logistic regression showing the relationship between a positive nasal MRSA PCR assay and risk of having concurrent or subsequent MRSA infection after adjusting for other covariates (N = 883).
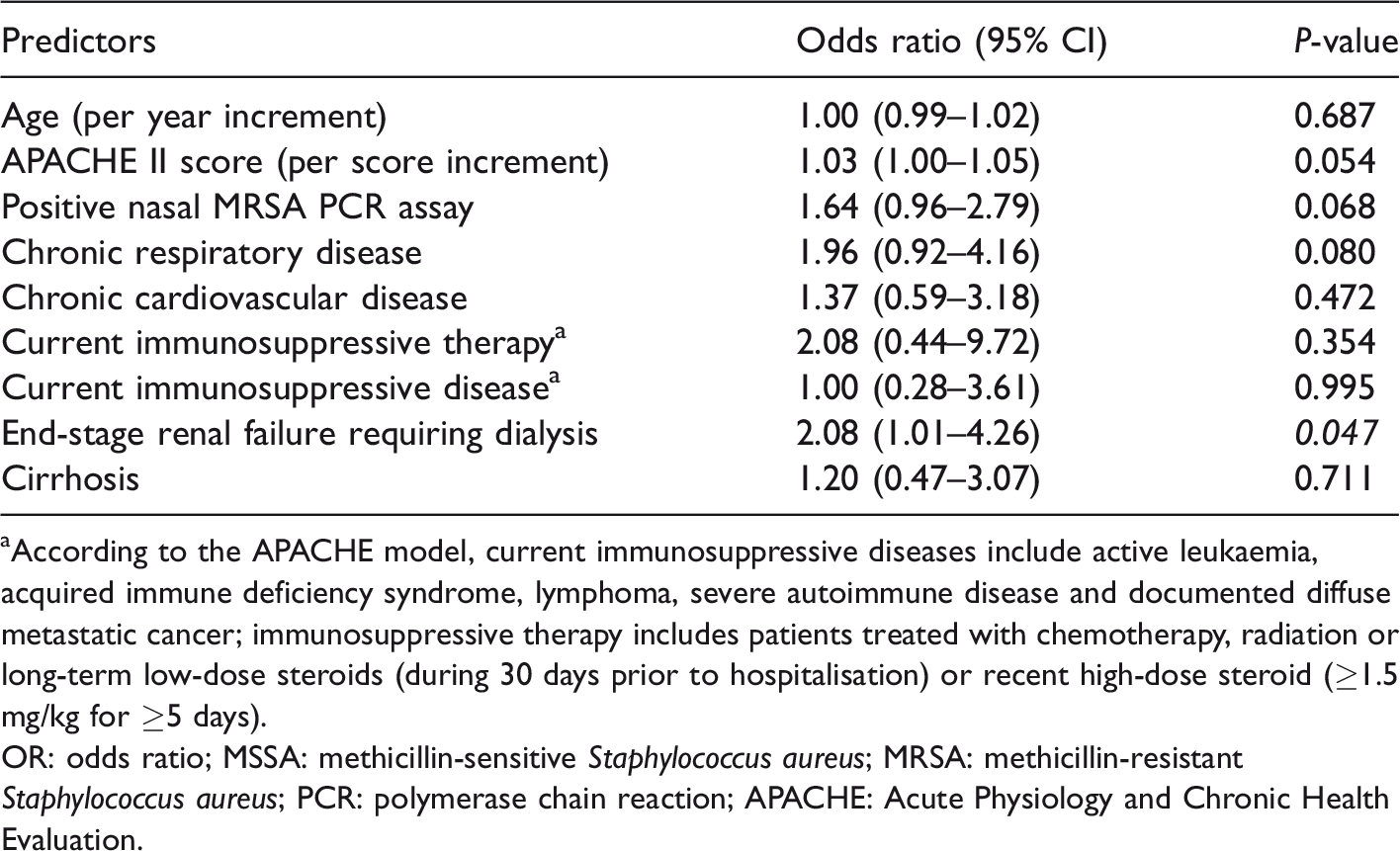
aAccording to the APACHE model, current immunosuppressive diseases include active leukaemia, acquired immune deficiency syndrome, lymphoma, severe autoimmune disease and documented diffuse metastatic cancer; immunosuppressive therapy includes patients treated with chemotherapy, radiation or long-term low-dose steroids (during 30 days prior to hospitalisation) or recent high-dose steroid (≥1.5 mg/kg for ≥5 days).
OR: odds ratio; MSSA: methicillin-sensitive Staphylococcus aureus; MRSA: methicillin-resistant Staphylococcus aureus; PCR: polymerase chain reaction; APACHE: Acute Physiology and Chronic Health Evaluation.
Discussion
A delay in initiating appropriate antimicrobial therapy for infection, in particular for those with bacteraemia, is associated with an increased risk of organ failure, metastatic infection and mortality.3,18–21 In this study, our results showed that both nasal MRSA and MSSA PCR assays have a high specificity and negative predictive value but a low sensitivity, especially the MSSA PCR assay, to predict concurrent or subsequent clinical staphylococcal infections. These results are consistent with a recent systematic review on the utility of nasal MRSA PCR assays in predicting MRSA infections (pooled specificity was >90%, but positive predictive value was only 50%), 22 suggesting that only a negative MRSA PCR assay may have clinical utility when the probability of having subsequent MRSA infection would be low and empirical vancomycin therapy for any suspected nosocomial sepsis may not be indicated, especially in clinical areas where MRSA is not endemic, as in our study centre. Conversely, a positive MRSA PCR assay is not particularly helpful with a low sensitivity and low positive predictive value. Initiating empirical vancomycin based on a positive MRSA PCR assay result would thus lead to overuse of vancomycin, resulting in increased healthcare costs, risks of nephrotoxicity and development of organisms resistant to vancomycin.7,23
Unlike the association between MRSA infection and nasal MRSA PCR assay, the association between a positive nasal MSSA swab and systemic MSSA infection has so far received little attention. 24 In this study, we showed that the prevalence of a positive nasal MSSA PCR assay in our patients was much lower than expected compared to the general population. 9 This result may be related to the fact that many study patients might have received broad-spectrum antibiotics prior to their ICU admissions, or the nasal MSSA PCR assay was just not as sensitive. In a study of 79 episodes of S. aureus (MRSA 12% and MSSA 87%) bacteraemia, 24 the incidence of a positive nasal swab culture for MSSA or MRSA was <50%, suggesting that the sensitivity of a nasal swab culture for MSSA is low. If this study’s results as well as ours are confirmed by other studies, use of a MSSA PCR assay to identify patients who are at increased risk of MSSA infection including MSSA bacteraemia will not be cost effective. Nonetheless, due to its high negative predictive value, a negative MSSA PCR assay may still be useful to exclude concurrent MSSA infection, similar to the utility of a negative MRSA PCR assay.
This study showed that patients with end-stage renal failure requiring dialysis admitted to our ICU were at increased risk of having MRSA infections independent of their nasal MRSA colonisation status. This result is consistent with other reports showing end-stage renal failure is a risk factor of MRSA colonisation, infection and infection-related mortality.25–30 Within the nested cohort of our patients who were either colonised or infected with MSSA/MRSA, the MRSA infection rate among those with end-stage renal failure was high (14/54; 26%) compared to those without end-stage renal failure (73/668; 12.3%). If our results are confirmed by other Australian ICUs, perhaps empirical vancomycin therapy should be considered in critically ill patients who have end-stage renal failure when serious systemic infection or bacteraemia is suspected, especially if they are known to be colonised with MRSA, or in centres where MRSA infection is endemic.
Finally, we need to acknowledge the limitations of this study. First, staphylococcal infections are not endemic in our centre, and this would increase the negative predictive values of both nasal MSSA and MRSA PCR assays in predicting staphylococcal infection. As such, our results may not be generalisable to centres that have endemic staphylococcal infections. Second, we only analysed patients with culture-proven staphylococcal infection. It is possible that some staphylococcal infections might have occurred among those who had been colonised with MRSA or MSSA, but the microbiological cultures were not taken or they were treated empirically with antibiotics, reducing the yield of subsequent microbiological cultures. In addition, we also did not have data on MRSA contact prior to index ICU admission. Prospective research including contact history will be useful to assess whether this can strengthen the ability of nasal MRSA PCR assay in predicting MRSA infections. Third, the sensitivity of using nasal swabs to detect bacterial colonisation may be subject to variations in the techniques in obtaining the nasal swabs, as well as how many nasal swabs are taken. A recent study suggested that significantly more (P = 0.016) patients with two negative nasal swabs returned with a recurrent MRSA infection or colonisation than those who had three consecutive negative nasal swabs (27.8% versus 17.0%, respectively). 31 Finally, the sensitivity of a PCR assay is known to be dependent on the strains of staphylococcal bacteria, which can vary substantially between different populations. 32 This important limitation has recently been overcome with the development of two new generations of more sensitive MRSA PCR assays. 17
Conclusions
Both nasal MRSA and MSSA PCR screening assays obtained on admission to the ICU had a high specificity in predicting concurrent or subsequent S. aureus infections, respectively. However, the sensitivity of both MRSA and MSSA PCR assays were low, especially for the MSSA PCR assay. These results suggest that only a negative MSSA and MRSA PCR assay may have some degree of clinical utility when the probability of having staphylococcal infection would be low and empirical anti-staphylococcal antibiotic therapy would not be indicated for patients with nosocomial sepsis, especially in ICUs where S. aureus infections are not an endemic problem.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship and/or publication of this article.
