Abstract

Evelyn Fox Keller, who died in September 2023, was a physicist, biologist, historian and philosopher of science who made pioneering contributions to both the understanding of gender issues in science and the philosophy of modern biology. That work culminated with two immensely influential books: Refiguring Life in 1995 and Making Sense of Life in 2002. Many of the issues Keller raised in those works are only now becoming more widely recognised and accepted as major themes in the reconsideration of the central narratives of modern biology. These narratives were a key focus for the special issue of this journal in 2020 titled ‘Making Sense of Metaphor: Evelyn Fox Keller and commentators on language and science’. Here I return to the issues Keller raised in her contribution to that volume and explain why they have become increasingly salient. I argue that Keller's philosophical positioning of the narratives and metaphors of biology can now be firmly rooted in the empirical output of the discipline and that they demonstrate the potential value of the ‘outside view’ on a discipline for highlighting both its underlying assumptions and how these can shift in ways that are not always readily apparent to practitioners.
Philosopher of biology Daniel Nicholson has said that ‘Nobody has managed to integrate the history and philosophy of biology as effectively and as elegantly as Evelyn Fox Keller’. That task of integration was, like biology itself, always work in progress. Her 2020 paper in ISR (Keller 2020) offered an update, in which Keller wrote that If, as I claim, recent work in genomics has finally disrupted the narratives of developmental genetics that have prevailed for over a century, geneticists will now need a new narrative to help guide them through the thickets that lie before them.
Hereditary information is encoded in the double helix (i.e. the genome).
DNA sequence codes for proteins which ultimately form an organism's observable characteristics or phenotype.
To many biologists, and certainly to many students of the subject, these two assertions might seem unproblematic, even axiomatic. Keller argued, however, that recent work in molecular biology points to complications. First, it is no longer possible to say that the only parts of the DNA sequence that matter for the phenotype are those that code for proteins. Second, the leap from proteins (i.e. from their sum total or proteome) to phenotype is unsupported not only by (lack of) evidence but also by logic. Third, it is only by eliding these two statements A and B that one can make the claim – still a mainstream one in public messaging about genomics from professional bodies such as the US National Human Genome Research Institute – that the genome is an inherited ‘blueprint’ or ‘instruction book’ for making the person. ‘The difficulty’, writes Keller, ‘… arises when people speak of the hereditary information encoded in the double helix [as equivalent to] the information required not for a set of proteins, but for an organism’.
I have a particular – one might even say vested – interest in Keller's discourse. While her article is in part an introduction to her influential exploration of metaphors in the natural sciences as devices that do cognitive (and not just explanatory or pedagogical) work, in large measure it seems to have been stimulated by an article of mine (Ball 2013) published in Nature as part of the 60th-anniversary retrospective of the publication of James Watson and Francis Crick's announcement of the double-helical structure of DNA – the launching point, one can reasonably say, of the modern era of genomics. My article attempted to point out how much is still unknown about the nature and function of the genome, and thus about the real molecular basis of evolution.
The immediate stimulus for that article was the publication of the results of the international ENCODE project (ENCODE Project Consortium 2012), which set out to ‘to build a comprehensive parts list of functional elements in the human genome, including elements that act at the protein and RNA levels, and regulatory elements that control cells and circumstances in which a gene is active’. ENCODE is still ongoing, but the initial announcement of its findings included a striking claim: that the genome was not, as was commonly implied, composed of a small proportion of protein-coding DNA – the genes – surrounded by vast swathes (about 98% of the total sequence) of meaningless ‘junk’ accumulated by evolution. On the contrary, the results seemed to the ENCODE researchers to indicate that, in some cells, at some time or another, as much as 80% of our DNA is transcribed into RNA, and that all this RNA might have some biochemical function. There is, it appeared, a great deal more to the genome than protein-coding genes. Indeed, the ENCODE consortium felt that a new definition of the gene was demanded – for which they proposed ‘a union of genomic sequences encoding a coherent set of potentially overlapping functional products’ (Gerstein et al. 2007). That rather convoluted definition demands some unpacking and is certainly less transparent than simply ‘a stretch of DNA encoding a protein’.
That ‘non-coding’ (nc)DNA (i.e. genomic DNA which does not encode genes) may be transcribed to RNA was not a new finding. The surprise was in the extent of it – sufficient, at face value, to suggest that making proteins is just a small part of DNA's biological function. This discovery was widely reported in the media, but provoked a heated backlash from some biologists who were convinced that the ENCODE claims were nonsensical, even anti-evolutionary (perhaps the ultimate biological insult).
There were certainly grounds for challenging these claims. In particular, just because a piece of DNA is transcribed into RNA does not mean it plays a significant role in the cell. Perhaps it is simply easier for cells to transcribe a DNA sequence rather freely, regardless of its content, than to have tight controls over where the molecular machinery of transcription should stop and start. Some critics of ENCODE also asserted that evolution will inevitably produce junk-ridden DNA in complex organisms such as humans. Only for species where the populations are huge and the cost of accumulating junk is large should we expect genomes to be streamlined to remove excess baggage – a condition satisfied for bacteria, but not for metazoans. ‘We urge biologists not be afraid of junk DNA’, wrote biologist Dan Graur and his colleagues: The only people that should be afraid are those claiming that natural processes are insufficient to explain life and that evolutionary theory should be supplemented or supplanted by an intelligent designer. ENCODE's take-home message that everything has a function implies purpose, and purpose is the only thing that evolution cannot provide (Graur et al. 2013).
In other words, what seemed to be at stake in these arguments was the very integrity of biology itself – specifically, the need to divorce it from purpose, a longstanding shibboleth. The discourse became not about how to understand the genome in the light of a surprising amount of functional nc transcription (the exact amount can be debated, although some members of ENCODE now feel that 30% is perhaps a more realistic figure), but about the right to retain the ‘junk’ metaphor. It was to some degree a contest of narratives.
This is a common pattern. We see it too with challenges to the biological metaphors of ‘machines’, ‘computing’, ‘selfishness’ and ‘blueprints’. When new findings seem to threaten these metaphors, the impulse is often to look for ways to adapt them to accommodate that new knowledge, rather than to seek new metaphors. This of course then negates the very purpose of metaphor in science as a means to understand or explain the unfamiliar by means of the familiar. Yes, we humans can still be called ‘robots’ – but only in the sense that, if there were robots that had consciousness, feelings, culture, wild and foolish aspirations, then we would be like that kind of robot. Yes, genes can still be considered selfish, if this means that alleles (not genes!) can be considered to compete with one another for resources in a zero-sum game, while genes themselves operate collectively and cooperatively in the development of an organism. We need to recognise that scientists cling as tenaciously to their metaphors as to their theories.
Keller has excavated this territory forensically. She implies that metaphors in science have a life cycle. In their early days they possess a fertile vitality: as Hesse puts it (Hesse 1988), ‘metaphor is interesting only when it is alive – provoking surprise and shock, indicating new thought’. When they lose that vitality, they do not necessarily lose their influence. A ‘dead metaphor’ is not one devoid of power; simply, it has ceased to be recognised as a metaphor at all, while leaving ‘a conceptual framework that continue[s] to shape how scientists actually perceived their objects of study’.
The ‘selfish gene’ metaphor exemplifies this. When Dawkins proposed it in the 1970s, it crystallised the Neodarwinian gene-centred view of evolution in a way that jolted popular thinking out of naive ideas about natural selection and adaptation. Biologists still attest to the value of this perspective as a corrective to the misconceptions about evolution with which many students arrive at their undergraduate classes. The selfish-gene view is devoid of any developmental biology, indeed by design: Dawkins says in The Selfish Gene that ‘Embryonic development is controlled by an interlocking web of relationships so complex that we had best not contemplate it’ (Dawkins 1976). The gene-centric picture therefore has some pedagogical value, although it ignores deeper nuances of evolutionary theory (Walsh 2015). It is, in essence, a narrative form of the models used by evolutionary population biology, in which organisms as integrated agents of action do not appear and genes are given a kind of fictitious agency in their place. In this case, the vitality of the metaphor was itself a mixed blessing, riding roughshod over its own limitations through the sheer narrative power of the image. Today the ‘selfish’ metaphor is indeed dead, in the sense that it is widely perceived (outside of evolutionary biology itself) as simply a description of ‘how things are’ – and is sometimes sustained by peculiar contortions of the original statement (Bird 2020).
In Making Sense of Life, Keller explains what has happened to the notion of a gene to enable the conflation of statements A and B above. Specifically, Much of the theoretical work involved in constructing explanations of development from genetic data is linguistic – more specifically, that it depends on productive use of the cognitive tensions generated by multiple meanings, ambiguity, and more generally, by the introduction of novel metaphors (Keller 2002).
But it was never ‘done’. Having completed this allegedly Apollo-scale enterprise, we are now doing it again, and again, filling vast data banks with genomic information from the diversity of humankind. There are very good reasons why one genome was not enough: only by making comparisons between many genomes can we identify some of the rare genetic mutations associated with certain diseases, for example. Such correlations are already proving of tremendous value for assessing disease risks for individuals based on their genomes. Yet cures have proved harder to access, because they generally demand that we have some sense too of what physiological effects the genes in question govern – if indeed they govern at all. While some genomics-based cures are appearing, it is not with the abundance once hoped. The usual story is that the genome has proved to be rather more complicated than we thought. Keller would, I think, argue that this might rather be a consequence of our having the wrong narrative to start with.
Not all genes code for proteins
Before the ENCODE results were announced, it was already widely suggested that the reason we have considerably fewer genes than expected – no more than the tiny nematode worm Caenorhabditis elegans, and considerably fewer than the onion – was that the complexity of our form results instead from the richness of interactions between genes. For example, the expression of genes is controlled by a host of molecular factors: proteins (especially so-called transcription factors) encoded by other genes, as well as all manner of epigenetic processes (chemical tagging of DNA or the protein component of chromatin, three-dimensional packaging of chromatin, and more) and regulatory DNA elements such as enhancers elsewhere on the double helix. ENCODE's discovery of widespread transcription lent support to the idea that much of this regulation is conducted not by proteins but by ncRNA molecules of all sizes, transcribed from DNA but not translated to protein. RNA is now considered as important for the regulation of genes – and thus the state and activity of cells – as are proteins.
One of the first functional long non-coding (lnc)RNA molecules to be discovered turned out to be the product of an assumed gene called Xist, which is involved in inactivation of one of the two copies of the X chromosome in the cells of females. (‘Long’ here means more than about 200 nucleotides.) Xist lncRNA works by binding to the DNA of the X chromosome that expresses it; the RNA transcripts appear to spread over more or less the entire chromosome and to induce persistent chromosomal changes (Wang et al. 2021).
Not all regulatory ncRNAs are long; some are surprisingly small. The first to be discovered, in 1993, was just 22 nucleotides long (Lee, Feinbaum and Ambros 1993): the first known microRNA. It is now clear that microRNAs play diverse and important roles in regulating genes, ramping their expression up or down. One estimate suggests that around 60% of our genes are regulated by microRNAs. Some small RNAs, such as those encoded in sections of the genome called small open reading frames (smORFs), are even translated into short protein-like molecules called peptides that have regulatory functions. The function, if any, of many smORF peptides remains a puzzle.
In metazoans, regulation of messenger RNA (mRNA) by microRNAs happens promiscuously: a given microRNA might target many (typically more than a hundred) mRNAs, and many different microRNAs might target a specific mRNA. The universe of regulatory ncRNAs is still expanding (Morris and Mattick 2014). There are, for instance, Piwi-interacting RNAs that are involved in silencing errant transposons (‘jumping genes’) in collaboration with so-called Piwi proteins, which play a key role in the differentiation of stem cells and germline cells. There are small nucleolar RNAs that steer and guide chemical modifications of other RNA molecules, such as those in the ribosome or the transfer RNA molecules that ferry amino acids to the ribosome for stitching into proteins. If cells are stressed, this can trigger the conversion of transfer RNA into molecules called tiRNAs that regulate the cells’ coping mechanisms, and which also have roles in the development of cancer.
RNA seems to be a general-purpose molecule that cells – particularly eukaryotic cells like ours in which gene regulation is so important – use to guide, fine-tune and temporarily modify its molecular conversations. With this in mind, biologist John Mattick has said that it is RNA, not DNA, that is ‘the computational engine of the cell’ (Mattick 2010).
That realisation destabilises still further the idea of a gene itself. The very term ‘non-coding’ implies a relegation of importance: the molecules are defined by what they do not do, namely mediate protein production. But there is no reason to regard gene-regulating proteins as any more significant in this regard than gene-regulating RNAs. Indeed, the first known ncRNAs were mistaken for proteins, which is why they were granted standard gene names. Some of the ENCODE team suggested that ‘gene’ be redefined as the DNA encoding any functional molecule (Gerstein et al. 2007), and there is now general acceptance that genes include at least some sequences encoding ncRNAs, even if for many of these the functional relevance remains unclear.
One of the more thoughtful responses – ripostes, even – to my 2013 Nature article came from the distinguished geneticist Adrian Bird (Bird 2013). ‘One could be forgiven for thinking that biology is in turmoil following recent discoveries that seem to undermine conventional wisdom surrounding the role of genomes in evolution’, he wrote in Cell. He went on to warn that ‘the quality bar needs to be kept high – a responsibility that scientists themselves need to shoulder. Without this, biology runs the risk of proclaiming revolutions before their time’. One of the false leads he dismisses is epigenetic inheritance – a hot topic at the time, although not one espoused either by my article or by Keller. Bird observes that “Transgenerational epigenetic inheritance … opposes the notion – unpalatable to some – that many human attributes are genetically ‘hard-wired’” – before showing that the evidence for it is weak, and the popular enthusiasm for such pseudo-Lamarckian inheritance seems to be largely a product of ‘wishful thinking’. He is right (for metazoans, at least; epigenetic inheritance is well established and quite common in plants).
This reconsideration of the genome represents one of the most profound recalibrations of molecular biology over the past several decades – a shift starkly illustrated in a plot of the estimated human gene count since the early days of the Human Genome Project at the start of the 1990s (Figure 1). The estimated number of protein-coding genes has dropped from around 100,000 to a value now thought to be below 20,000 (Amaral et al. 2023). At the same time, since the start of the millennium, the number of nc sequences recognised as genes has increased to the point where it outnumbers coding genes – and estimates anticipate that it might soar in the years ahead. It is simply not credible that the underlying narrative of molecular biology and genetics could remain much the same in the face of such a transformation.
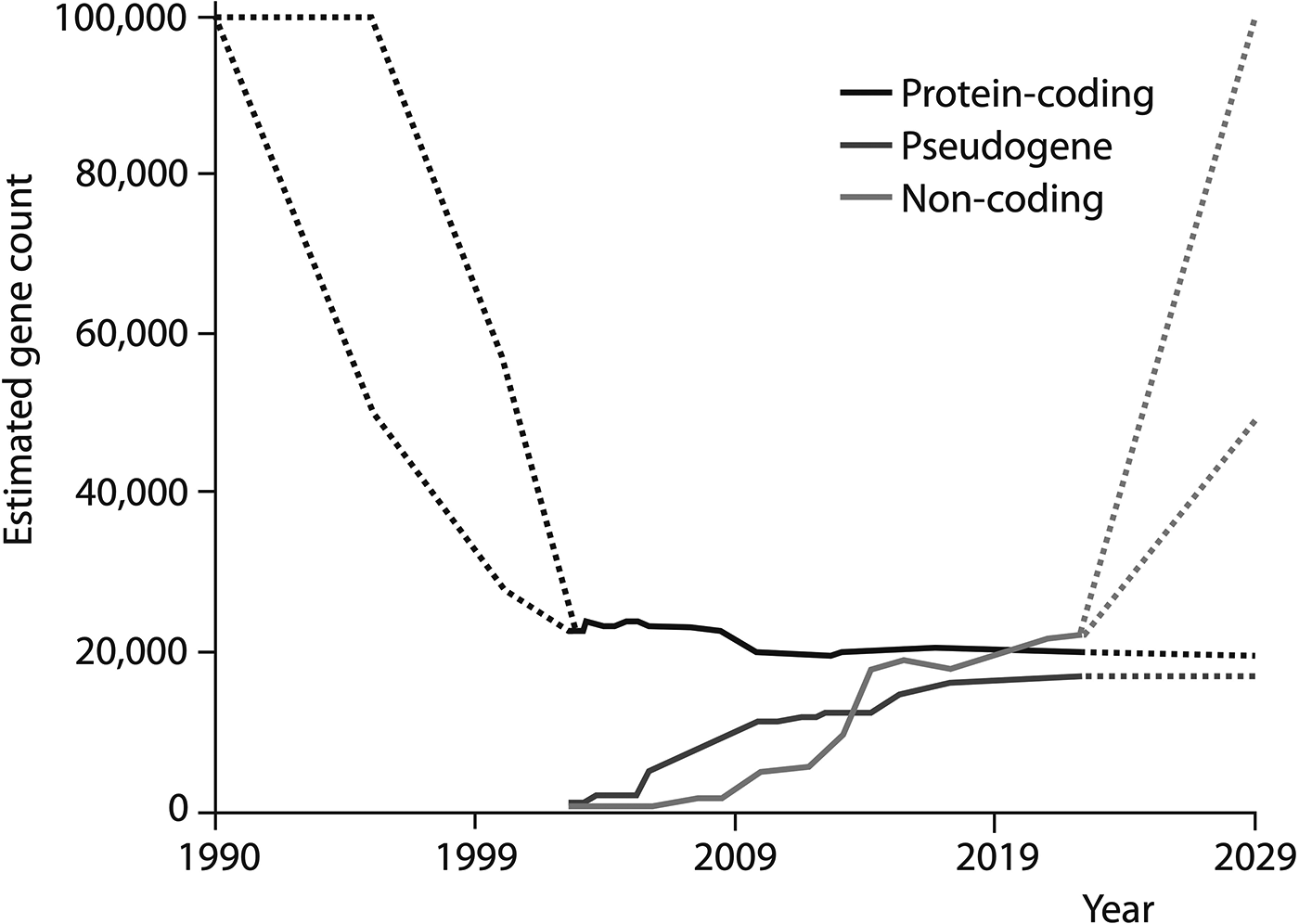
The tally of human genes. Dashed lines indicate the ranges of estimates in the literature. (Pseudogenes are DNA elements once thought to have lost their gene function, but some of which might retain biological functions.) From Amaral et al. (2023).
A question of control
Does this simply mean that there are more (and new types of) molecules involved in the genomic ‘program’ than we thought previously? On the contrary, the extent of eukaryotic RNA regulation further undermines the idea that the genome is somehow a ‘master program’ that the cell passively reads out. In contrast, that readout is itself determined by higher-level inputs, such as chemical, mechanical or electrical signals arriving at the cell surface. The genome is responding epigenetically to those signals.
What, then, is in control – the genome or the environment? Neither; this is a constant feedback process involving dialogue between various aspects of the biological hierarchy, up to and including the cognitive choices that the whole organism makes (for example, diet influences epigenetic gene regulation). As Keller says, Today's genome looks very different from the one with which the science of genetics began. Rather than a set of genes initiating causal chains leading to the formation of traits, it looks far more like an exquisitely sensitive reactive system – a device for regulating the production of specific proteins [and other molecules!] in response to constantly changing signals it receives from its environment (Keller 2020). In the future, attention undoubtedly will be centered on the genome, with greater appreciation of its significance as a highly sensitive organ of the cell that monitors genomic activities and corrects common errors, senses unusual and unexpected events, and responds to them, often by restructuring the genome.
This seems an exciting and surprising shift in the narrative. What seems peculiar is that molecular biology has been so muted about it. It is not as if this new picture is in any doubt; indeed it has been one of the triumphs of post-genomic biology to elucidate some of the ways in which it operates. Yet one senses – at any rate, I sense – a determination to present this as ‘business as usual’, embedded within the same old narratives. This diffidence might be seen as endearingly self-effacing, or at worst as lacking in self-awareness, were it not that efforts to suggest otherwise tend to be met with resistance bordering on hostility. As biologist and ENCODE researcher Thomas Gingeras has said [personal communication], Instead of highlighting the novel and unique aspects of lncRNAs, much time and effort has been spent on trying to promote the notion that lncRNAs are only marginally functional and that a lot of the work published in this field is plagued with incomplete and poor science. It is puzzling why there is such attention to persuade colleagues to move from a sense of interest and curiosity in the ncRNA field to a more dubious and critical one.
The new narrative is fuzzy
Keller suggests that the notion of a reactive, responsive genome might supply the beginnings of a new narrative. I believe that, while this is true, we can now go further, precisely because biological research has done so. The new story emerges, I believe, from the different molecular grammar that has come to light.
The idea of a genetic program – Erwin Schrödinger's ‘code-script’ (Schrödinger 1944) – mobilises a computational analogy, whereby information is passed around the cell along well-defined channels of molecular communication. Such is the paradigm of the cell signalling pathway. In this view, the robustness of the cell to life's stochasticity – cell-to-cell fluctuations in gene expression levels, say – comes from redundancies in the signalling networks. If one route fails, there is generally another.
It now seems clear that much of the molecular logic (to use a metaphor that itself requires scrutiny) of the eukaryotic, and especially the metazoan, cell is not of this sort at all. Many of the molecular interactions at the heart of gene regulation and signal transduction are now known to be remarkably fuzzy. Rather than the precise lock-and-key recognition advertised as the secret to the apparent specificity of information processing in the cell, these processes often involve promiscuous interactions with low specificity: key molecules will ‘speak to’ a wide range of others, to varying degrees. How these loose unions conspire to produce well-defined outcomes at the cell and physiological level is still not clear, but it seems to involve a combinatorial logic (Klumpe et al. 2022) somewhat akin to that which produces a vast range of highly specific odours from the collective action of just 400 or so olfactory receptor proteins. Crucial to the stability of these outcomes is also Conrad Waddington's concept of canalisation (Waddington 1942): the channelling of possible states of biological systems into just a limited number, irrespective of the fine details. Such processes are now understood in terms of the theory of dynamical systems, whereby high-dimensional systems can be described using just a few key parameters whose dynamical behaviour produces attractor states (Sáez, Briscoe and Rand 2022).
Central to this kind of molecular logic is the relatively new discovery that many proteins are not, as has been traditionally supposed, highly and intricately structured (‘ordered’) but contain some degree (and sometimes a predominant degree) of intrinsic disorder (Uversky 2014; Wright and Dyson 2015). This means that their polypeptide chains do not adopt the well-defined, characteristic structural motifs identified by much of the protein crystallography to date, but remain ill-defined in solution and roam rather freely over a conformational landscape (Figure 2).
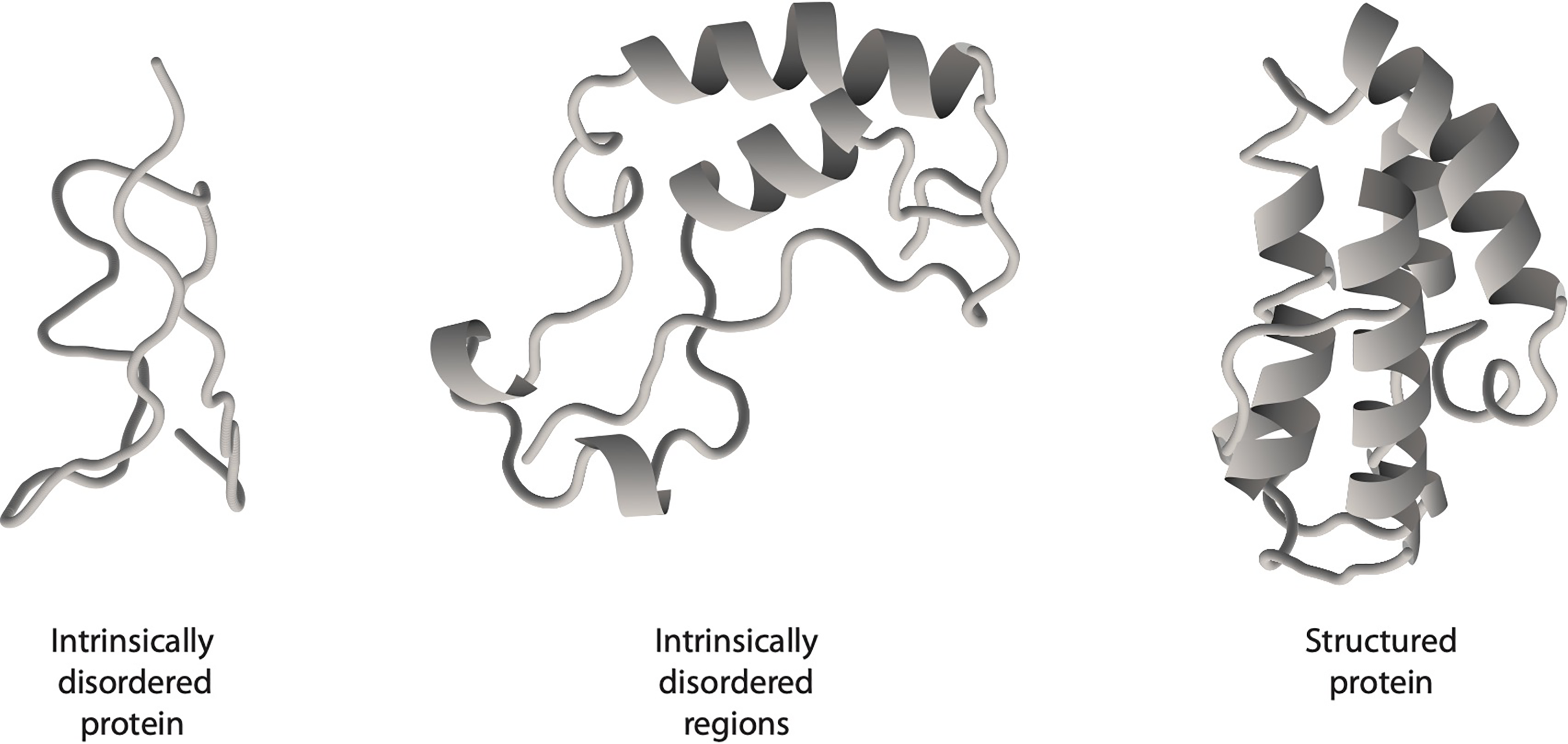
Disorder and order in protein molecules. The loose, open segments of the polypeptide chain have no fixed shape but instead explore a diverse ‘landscape’ of conformations.
In contrast to the typical lock-and-key specificity with which ordered proteins bind their targets, intrinsically disordered proteins (IDPs) can wrap themselves around or stick to a variety of partners, often including other proteins and DNA sequences. To a greater or lesser degree, they are promiscuous in their interactions. (Some IDPs do show a fair amount of binding specificity, perhaps because they adopt more ordered structures when they enter such unions.) Their tendency to aggregate rather unselectively with other proteins means that IDPs may act as ‘hub’ molecules that create connections between distinct networks of molecular interactions: many of the most deeply embedded and well-connected proteins in these networks have appreciable amounts of disorder. It is widely thought that it is by forging such links – and not by acquiring new genes – that complex organisms evolve (Gerhart and Kirschner 2007). Disordered proteins also seem to engage in low-specificity interactions with regulatory RNA molecules in processes such as splicing (in which fragments of an RNA transcript of a gene are rearranged before being translated into a protein) and gene regulation.
One reason the importance of IDPs was not sooner apparent is that there are so many ordered proteins to study, and well-developed techniques to do so. Disordered proteins tend not to crystallise well, whereas protein crystallography relies on the availability of good crystals. So even though it has been estimated that 37% to 50% of human proteins have disordered regions, their importance and prevalence is a relatively recent discovery – and indeed, is still not widely known even among some biologists (Wright and Dyson 2015).
This centrality of IDPs to biomolecular interaction networks makes them potentially important targets for therapeutic interventions. But the disorder also makes it challenging to design drugs to bind to them and alter their functionality (Lim et al. 2023). The ease with which IDPs associate is also thought to lead to pathological conditions such as the formation of the cytotoxic protein aggregates responsible for Alzheimer's and other neurodegenerative conditions – a property that also makes it important to better understand how IDPs behave and interact.
Crucially, this fuzziness of molecular interactions is not a consequence of nature's limits on how precisely molecular interactions can be engineered. Nature can do much better, and often does: gene regulation in bacteria, for example, tends to involve transcription factors that are rather sequence-specific and have larger DNA binding sites, while IDPs are rarer in prokaryotes: perhaps just 4% of the proteome, compared to an estimated 30% to 50% for metazoans. Evolution has, so to speak, chosen to do things this way in more complex organisms. It seems reasonable to suppose that this is because that is the only way such organisms are viable in the first place: reliance on weak interactions of low specificity may be the key to making organisms that are both adaptable quickly to changing circumstances (a requirement more commonly encountered for complex higher organisms with a long generational timescale) and able to evolve without mutations simply being deleterious.
So in addition to Keller's ‘sensitive genome’, I would suggest that the new narrative of molecular biology that is now emerging should include the following considerations (Ball 2023):
- Increasing organismic complexity necessitates qualitatively different principles of operation at the molecular level. - Those principles involve fuzzy, promiscuous interactions that achieve robust outcomes via combinatorial rules and dynamical canalisation. - The consequence of this ‘sloppiness’ in molecular dialogue is causal spreading: a decreased reliance on genetic causation in favour of that occurring at higher levels of integration. Information and control simply cannot, under these conditions, be reliable ‘fed upwards’ from the genome; it would be like trying to write on jelly. This is not a flaw but a feature: with causal spreading, the fine details may not matter and stochasticity at the molecular level does not beget fragility at the cell or organismal levels.
The importance of metaphor and narrative
It was with a wry smile that I read Keller's ‘two questions of particular relevance to the history and philosophy of science’ – one of which is ‘What makes it possible now for Ball to write – in the pages of Nature no less – that ‘the conventional narrative … is as misleading as the popular narrative of gene function itself’?’ She continued: ‘Has Ball in fact escaped the confines of the prevailing discourse? And if he has, what has made it possible for him to do so?’ (Keller 2020)
I am not clear quite how Keller would have answered those questions directly – except to say perhaps (and rightly) that, to the extent that I had indeed ‘escaped the confines of the prevailing discourse’, my argument needed the kind of refinement that she was able to bring to it: to point out that the problem lies in the elision of ‘genome’ with ‘protein-coding genes’, as well as the blithe black-boxing that takes us from proteome to phenotype. What is more striking is Keller's remark: … is the realization that it/they [the popular narrative(s)] are misleading a new realization? Not really. Then how is it that such ‘misleading’ narratives are so routinely perpetuated in the teaching of Molecular Biology, indeed in so much of the technical, the lay, and even the philosophical literature?
This is surely a little peculiar. How can it be that Keller – a penetrating thinker but hardly an iconoclast in modern biology – was ready to consider as unremarkable and well known what to other biologists seemed almost heretical?
But perhaps we should not be too surprised. It is insufficiently acknowledged that what seems unremarkable and even prosaic from one perspective in science might be unfamiliar and even shocking from another. Population geneticists may sometimes not even recognise concepts or points of view that are widely held among developmental biologists, for example. Even scientific subdisciplines develop norms and narratives that become an invisible part of their cognitive landscape while being alien to neighbouring subdisciplines. But that such a situation is common does not of course mean that it is unproblematic.
Keller's own work was important for understanding why such circumstances arise. It is not through mere intellectual drift of communities that rarely converse. As she notes (Keller 2020), the slippages between elements of the conventional narrative(s) of molecular biology ‘are not casual but systematic, not an occasional or accidental feature of molecular discourse but a systemic one. As such, they seem to serve a function’.
A part of that function, she suggests, is about status: ‘they enhance the image of what has been achieved (good of course for both funding and egos)’. Particle and high-energy physicists know that the common conflation of their subdiscipline with all of physics in the public perception is not a true reflection of what the field does or is, but they hardly have an incentive to correct the misperception, and so they rarely do.
But Keller says that such slippages are also perhaps an essential way of making progress in science: they ‘prevent (or at least minimise) distraction by problems that appear to be impossible to address with the tools at hand; they work to focus attention on problems that can be so engaged, and that in itself can be useful.’ Science has of course always been not so much an investigation of the natural world as an investigation of those aspects we have the (experimental and conceptual) tools to investigate (that is why we have come so late to IDPs, for example). But nowhere has this been more apparent than in modern biology. The questions raised by genomics-era projects such as ENCODE are profound and deeply puzzling. Yet often the response has been to collect even more data – not necessarily because it is required to test hypotheses, but because we possess or can develop tools for doing so. The enormous challenge of understanding what the Human Genome Project has told us is, one can argue, being postponed until we have decoded lots more genomes too – and added to it innumerable other “-omes”, from the proteome to the metabolome and the microbiome. The same tendency is evident in brain science, where vast data-collecting and neuron-mapping resources are not accompanied by a corresponding expansion in theoretical ideas about the brain. We make progress on the fronts we can reach.
It is rather ironic, then, that Bird (2013) claims that ‘in biology, ideas are relatively cheap’. I would suggest that rather, in biology (especially molecular biology and neuroscience) ideas are rather rare. It is data that are, if not cheap, then abundant to the point of excess, and it is possible to make a career collecting data that one is never obliged to interpret. It is rather in physics that ideas tend to come cheaply and almost casually, especially when data are scarce – witness the flood of theoretical papers always submitted to the physics preprint servers to ‘explain’ some new, tentative anomaly in high-energy physics that turns out to evaporate on closer inspection.
It makes good sense for science not to remain snagged on problems it lacks the resources to solve – but the problem is that those problems can then not be noticed at all. The language of molecular biology has become one of exquisite molecular recognition between its components, although we now see that some of the most important interactions in the cell have rather low specificity – and this by design (to risk a fraught term), in the sense that evolution requires it for developing metazoan complexity. Experts in the field of gene regulation previously had no language, nor even any conceptual landscape, for thinking about such processes. It is to this issue that Keller speaks so eloquently (Keller 2020): Yet this style or habit of chronic slippage from one set of meanings to the other has prevailed for over 50 years; it has become so deeply ensconsed as to have been effectively invisible to most readers of the biological literature. This feature I suggest qualifies it as a Foucauldian discourse – by which I means a discourse that operates by historically specific rules of exclusion, a discourse that is constituted by what can be said and thought, by what remains unsaid and unthought, and by who can speak, when, and with what authority.
Some of the responses to my 2013 article had the same flavour: why tell a complicated but more accurate story if you can tell a simple but misleading one? One philosopher of science responded to the article (Salleh 2013) by saying that ‘While simplistic communication about genetics can be used to hype the importance of research, and it can encourage the impression that genes determine everything … [nevertheless] the answer is [not] to communicate more complexity’. Meanwhile, an academic specialist in science communication asked (Salleh 2013) of the piece ‘Is there a problem that we need to know about here?’ and went on to say that ‘there are dangers in telling the simple story, but he hasn’t spelt out the advantages of embracing complexity in public communication’. Embracing complexity has its complications, to be sure – but ignoring it means staying with an outmoded narrative that, in this age of genomic editing, gene screening and mass genomic sequencing, is not just misleading but potentially dangerous.
While Keller was kind enough to ask ‘what has made it possible for [Ball to] escape the confines of the prevailing discourse?’, I am more interested in what made it possible for Keller herself to recognise, well ahead of the curve, that a new narrative is required. In a review of Making Sense of Life, biologist Matthew Cobb wryly noted (Cobb 2004) that Some people will be irritated by the very existence of this book … The problem will lie not so much in the subject as the author, Evelyn Fox Keller, who many biologists will consider to be doubly damned: she was trained as a theoretical physicist before becoming a philosopher of science.
The tension here is that the history of ‘physicists intervening in biology’ is a chequered one. The molecular biology revolution was in some ways orchestrated by former physicists such as Max Delbrück, Sidney Brenner, Francis Crick and Seymour Benzer, and on the influential fringe Erwin Schrödinger and George Gamow. On the other hand, physicists are notorious for coming to biology with grand ideas too vague and generic to offer any purchase for the lab biologist and often poorly informed about the tortuous and confounding facts of actual biological research.
Certainly, physics can claim no privileged status for understanding a science as contingent, path-dependent and exception-strewn as biology. What it does offer, however, is a kind of ground-state view of the generic behaviours of matter. For example, scepticism (to which I can testify) towards the suggestion that condensed-matter physics can have any relevance to cell behaviour has been motivated in part by a sense – expressed explicitly by evolutionary biologist Ernst Mayr (2004) – that living matter has a unique status beyond the reach of the physical sciences. Yet it is surely only through the evolution of active processes to subvert or suppress them that living systems could avoid the collective modes of behaviour known to govern inanimate matter. While that might sometimes be possible and even necessary, there is abundant evidence now that more often evolution exploits such resources, which can, for example, provide organisation ‘for free’. That is what we see, for instance, in phase separation processes in cells, now a major locus of biological research (Shin and Brangwynne 2017).
In Making Sense of Life Keller already pinpoints the source of tension that arises in speaking across disciplinary boundaries. It is not so much that outsiders might attempt to speak to disciplinary questions without the knowledge that might restrain them from simplistic and hubristic answers (although that surely happens). Rather, Keller says, the problems come from different perceptions of what counts as an explanation of phenomena. For a molecular biologist, for example, an explanation of development might entail identifying the molecules involved and the interactions between them. To a developmental biologist, the question is more about how cells and tissues give rise to biological form. A biophysicist might see those processes as being directed as much by mechanical factors (such as stretching, bending and flow) as by molecular ones. Meanwhile, the wider question of how, if at all, a target morphology is encoded in the nascent organism, or how it remains robust to stochastic variability at the level of cells and molecules, has until recently barely been considered at all. Keller's advice that, in consequence, ‘we attend closely to the meanings of the words we use and the ways in which we use them, and take seriously the linguistic and narrative dimensions of explanation’, is relevant across all the sciences but probably nowhere more so than in the life sciences. Yet many researchers would, I suspect, simply be puzzled by that position, asking: what relevance does it have for the experiments I am conducting?
Doing science is a delicate dance between conservatism and innovation. If it indulges every new idea that come along, it will waste energy and resources down countless rabbit-holes. But if it clings stubbornly to old ideas, beliefs and metaphors, it will never progress. How to tell the difference between what is worth pursuing and what is not? How to know when it really is time for a new narrative?
It would have been wonderful to know what Keller had to say about that. She concluded her 2020 article by saying that the dramatic shift underway in biology's narratives meant ‘I only wish I was just starting rather than ending my career.’ We might wish that too.
Footnotes
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author received no financial support for the research, authorship, and/or publication of this article.
