Abstract
The opaque relationship between biology and behavior is an intractable problem for psychiatry, and it increasingly challenges longstanding diagnostic categorizations. While various big data sciences have been repeatedly deployed as potential solutions, they have so far complicated more than they have managed to disentangle. Attending to categorical misalignment, this article proposes one reason why this is the case: Datasets have to instantiate clinical categories in order to make biological sense of them, and they do so in different ways. Here, I use mixed methods to examine the role of the reuse of big data in recent genomic research on autism spectrum disorder (ASD). I show how divergent regimes of psychiatric categorization are innately encoded within commonly used datasets from MSSNG and 23andMe, contributing to a rippling disjuncture in the accounts of autism that this body of research has produced. Beyond the specific complications this dynamic introduces for the category of autism, this paper argues for the necessity of critical attention to the role of dataset reuse and recombination across human genomics and beyond.
Big data has often been presented as the final answer to problems in categorization. Rather than attempting to carve nature at its joints via idealized examples, big data projects purport to represent natural diversity in its totality—to simply show things ‘as they are’. In the life sciences, such imaginations have found fertile ground: Fantasies of ‘the total archive’ have long animated ideas about objectivity and universality (Jardine & Drage, 2018; see also Strasser, 2019), and increasing computational capacities have led to widespread reorganizations in the professional ecology of the contemporary biosciences (Leonelli, 2016; Stevens, 2013). While big data biology projects are varied in their specifics, they share a conviction that the messiness of categorization can be made to disappear under the magnitude of sheer, seemingly uncomplicated scale.
Perhaps nowhere has this logic appeared more seductive than in psychiatry—not incidentally a field dogged by its own thorny classification politics in the demarcation and discipline of human difference (Foucault, 1965; Rose, 1998, 2009). In a review in JAMA Psychiatry, Weissman (2020) suggests that big data solutionism has repeatedly appeared in response to recent field-level crises in psychiatry: in psychiatric epidemiology, large cohort studies, and the mass-scale repurposing of Electronic Health Records (EHRs) for research (see also Arribas-Ayllon et al., 2019; Panofsky, 2014). This drive for ever-larger data is now also enshrined within the US National Institute of Mental Health’s (NIMH) Research Domain Criteria (RDoC) program, which proposes to adjudicate the likely intractable classificatory conflicts between symptomatology and neurobiology (Insel et al., 2010; Pickersgill, 2019). Central to NIMH’s efforts is the accumulation of big genomic data, as indicated by the significant outpouring of federal funding into sequencing and data sharing consortia (Senthil et al., 2017).
Submerged within this ethos, however, is something of a chicken and egg problem: If you need (big) data ‘about’ a psychiatric condition to understand it, you need to already have decided what that condition is, or at the very least a firm definition of whom that category includes and an idea about which of their characteristics might be of interest for clinical research. This is not a trivial task: As Nelson (2019) has shown, the relationships between behaviors and categories in psychiatric genetics are hard-won achievements, and are maintained only through significant and ongoing effort. Here, examining the case of autism spectrum disorder (ASD), 1 I show how this effort necessarily extends into the data infrastructures that support genomic research. As I argue, data ecologies with dissensus on categorical boundaries can complicate or obscure just as much as their scale promises to illuminate.
This article excavates how the circulation of datasets in categorical misalignment has contributed to a growing fracture in big data autism genomics, resourcing two distinct accounts of ASD and how it should be studied. By categorical misalignment, I mean that how these datasets demarcate autism—its salient features and the populations that it can be said to describe—are partially disjunct, even though each dataset is assumed to be representative of a unified category. These differences are discursively vivified in the scientific literature each dataset is used to produce; research projects solidify data-based categories by linking them backward to genotype as well as forward to etiologic, diagnostic, or therapeutic implications. In doing so, researchers both biologize and medicalize the innate categorical differences of their data, meaning that what appear to be minor technical differences between datasets can echo into significant redefinitions—or, here, diverging definitions—of what appears to be an otherwise stable diagnostic entity.
I argue that biobanks and other big data repositories are critical intermediaries in the research process, shaping and constraining the kinds of variables that can be analyzed and the populations under study. In the process, these organizations have had surprising influence over the disease entities that biomedical research reproduces, shifting categorical boundaries and realigning the grammar of inquiry with the inbuilt assumptions of their datasets.
To make this claim, I employ mixed methods to analyze a corpus of published big data research on autism genetics, limited to the five-year period between 2017 and 2021 (n = 303 articles). First, I use network mapping to demonstrate a clear bifurcation in the topics and citational literatures that anchor this research. This bifurcation suggests two distinct and increasingly non-overlapping ideas of what autism is and important methodological differences in how it should be studied. Then I turn to a close textual analysis of a smaller set of articles, comparing research produced using two notable datasets: MSSNG (pronounced ‘missing’; n = 38), a biobank linked to parental advocacy organization Autism Speaks, and the direct-to-consumer genetic testing company 23andMe (n = 22). These organizations have significant (and well-historied) differences in their orientations toward autism science, underwriting differences in their data collection strategies and categorical implementations. I link these differences to the conceptual clusters indicated by the network mapping and draw out a finer-grained analysis of their implications for the discursive reproduction of autism(s) through big data genomics.
This project draws from critical data studies (CDS) in its attention to how data shape conceptual categories in biomedical research. CDS scholarship understands data not as a ‘raw’ element of subsequent inquiry, but rather a richly social set of inscriptions that arrive always already ‘cooked’, theory-laden, and replete with meaning (e.g. Bowker, 2008; Gitelman, 2013; Kitchin & Lauriault, 2014). Following broader concern for how research infrastructure (and particularly data infrastructure) shape scientific practice (Borgman, 2016; Bowker & Star, 2000; Edwards, 2010; Edwards et al., 2013; Star & Ruhleder, 1996), CDS research foregrounds questions about how data are produced, how they are maintained and mobilized over time, and in what ways they are interpreted. In doing so, it illuminates how data themselves—rather than just the actors who rely on them—can drive shifts in larger social formations. In particular I take inspiration from Denton et al.’s (2021) genealogy of the ImageNET machine learning dataset, which was similarly positioned as a big data solution to categorization problems. Rather than ‘solving’ these problems, ImageNET reinscribes and naturalizes particular sets of social values while simultaneously invisibilizing the actors who hold them.
CDS accounts (including Denton et al.’s) typically examine the development and deployment of a single, widely used dataset. In most big data sciences, however, notable datasets proliferate, and a research community might rely on a number of common resources. Consider, for example, the datastreams associated with large telescopes or satellite arrays in astronomy, physicists who rely on data from neutrino observatories or particle accelerators, or the small number of influential climate models whose outputs feed into a diverse set of research communities. (It is not incidental that these examples all rely on expensive and/or site-dependent research infrastructures—the more difficult data are to produce, the more researchers are incentivized to rely on shared data. Biobanks are similarly resource-intensive.) If we are to take the supposition that all data are theory-laden seriously, then, it stands to reason that different datasets might bear with them different sorts of theories, values, and categorical imaginations. Comparative studies, like the present case, offer to draw out how the circulation of multiple datasets in a research ecology can smuggle in dissimilar and potentially incompatible representations of their shared object.
While this case study examines the use of biobank data to make claims about a single diagnostic category, similar processes are well underway across the life sciences and big data-driven research broadly. An attention to the histories and applications of large reused datasets, as I argue, offers to reveal occulted processes in the construction of the categories they appear to simply represent.
Multiplying autisms: History of a contested diagnosis
Even among other contested psychiatric diagnoses, it’s worth underlining that autism swims in uniquely muddy waters, and has since long before big data arrived on the scene. ASD’s conceptual evolution and rapidly expanding patient population have sparked significant debate in both the biomedical and social sciences for decades, and its definitional contours have consistently defied stabilization. As Verhoeff (2013, p. 446) has commented: ‘Ideas about autism are not fixed but continually in flux. There is not a single test, definition, article or researcher that marks a definite idea of autism in a specific period.’ Elsewhere, others have identified the autism concept as ontologically ‘heterogeneous’ and ‘indeterminate’ (Hollin, 2017); as Singh (2015a) says, there are ‘multiple autisms’. Indeed, this understanding is broadly uncontroversial even in the autism research community: In one of the most-cited articles of the last 20 years, noted psychiatric geneticists Happé et al. (2006) argue that it is ‘time to give up on a single explanation for autism’. That it is time to give up on a single definition for autism is implicit.
While autism may already have been multiple, particular biobanks have organized two distinct and relatively clear-cut autisms from within. We might even think of this as a sedimentation of autisms that were previously in solution, now precipitated into identifiable—and increasingly immiscible—layers. This argument is elaborated below. First, however, it is worth spending a moment on how this curious state of affairs has come to be.
The history of autism and autism research has been extensively documented by others at a finer degree of detail than I am able to offer here. Much of this work has focused on the structure of its diagnostic change, tracking how expansions of ASD—through deinstitutionalization, ongoing population shifts, formal revisions to the DSM, and informal changes in diagnostic practices—have repeatedly reconfigured the autism concept (Eyal, 2013; King & Bearman, 2011; Maynard & Turowetz, 2019; Navon & Eyal, 2016).
Importantly, these processes have drawn in a variety of actors with competing modes of expertise, as well as differing claims to knowledge of and experience with autism as a lived condition (Barker & Galardi, 2015; Eyal, 2013). Among the most influential groups both historically and today have been the parents of autistic children, who have driven categorical redefinitions of autism through their contributions to the development of therapeutic strategies (Eyal, 2013; Hart, 2014) and brain science approaches (Fitzgerald, 2014; Rapp, 2016), as well as their own participation as research subjects (Lappé, 2016). Parents have also played a significant role in lobbying for federal support, securing substantial funding for educational, clinical, and research programs through legislation like the US Autism CARES Act (Autism Collaboration, Accountability, Research, Education, and Support Act, passed in 2014 and renewed in 2019; Singh 2015a). While the role of parents in defining the autism concept (rather than autistic people themselves) remains a point of controversy (Rosenblatt, 2018; Stevenson et al., 2011), it is clear that parents have had an outsized effect: Their efforts have not only reshaped ASD in the laboratory and the clinic, but intervened in broader cultural imaginations of autism.
Most importantly to the story I tell here, however, is parents’ work in coordinating large scale genetic resources for autism research—particularly in the US and Canada. As with a number of genetic conditions, 2 parent advocates have routinely sought to accelerate autism genetics by contributing to the development of autism biobanks. Different accounts have highlighted different facets of these efforts. For example, Singh (2018) has explored how contributing biomaterials allows families to access care and support, while Tabor and Lappé (2011) identify how the scale of biomaterials required for this work has driven changes in the institutional and interpersonal relationships between autism families and coordinating clinics. Through these processes, parent advocates have since the early 2000s made autism genetics a particularly well-resourced and attractive site for researchers from a variety of disciplines (Singh 2015a). These affordances helped to solidify the burgeoning field as a ‘trading zone’, bridging numerous guiding interests and imaginations of the autism concept (Navon & Eyal, 2014).
Critically, it is not just that autism has been broadly geneticized through these projects (though this is certainly the case), but that geneticization has in turn driven significant changes to autism as a category. Drawing on Hacking’s (2006) work on ‘looping effects’, Navon and Eyal (2016) have painstakingly documented how notions of autism as a genetic condition shifted how autistic groups are understood and described, leading to cyclical changes in both the boundaries of the category and the underlying genotypes it can be said to include. As they argue, Every time diagnostic criteria are changed—whether to better capture phenotypic variability, to better reflect/validate genetic evidence, or for any other reason—the genetic makeup of the population picked out by the now-changed classification may also be modified. This new population changes the material conditions for examining the genetic etiology of the classification, which in turn can modify expert understandings of the condition and thereby the self-understandings of the people picked out by the classification. When human kinds loop, their genetic makeup can also therefore be rendered a moving target (Navon & Eyal, 2016, p. 1421).
While Navon and Eyal focus on diagnostic change as a core mechanism, we can understand biobank recruiting and phenotyping as tacitly similar practices insofar as they also serve to identify—at least for the purposes of genetic research—who constitutes the autistic population. I specify some implications of this in connection with individual biobanks in a later section. For now, though, it is enough to observe that genetic research can and has had significant impacts on the autism concept, and ones that are surely continuing to unfold.
Conceptual clustering in contemporary autism research
Even in the face of continued disputes over autism’s categorical boundaries, it remains uncontroversial to claim that autism is largely genetic. The DSM-5 is in agreement with mainstream autism research when it suggests that 60-90% of ASD cases are likely to have a genetic component, and roughly 15% are clearly linked to already-known causal genetic variance (American Psychiatric Association [APA], 2013, p. 57; see Rylaarsdam & Guemez-Gamboa, 2019). However, this is where broad consensus ends: How researchers make sense of the genetic architecture of the yet-unexplained 45-75% of cases with likely genetic contributions varies widely in methodology, topical focus, and in their very conceptualization of autism itself.
Beyond changes to the clinical population, new language in the DSM has also driven unanticipated epistemic shifts in academic autism research, and subsequent knowledge claims. The DSM-5 diverges from previous editions in describing autism as a collection of spectra: diagnostic criteria identify several domains across which a patient can experience differently scaled levels of disability, leading to highly individualized diagnoses that can acknowledge areas of strength as well as specific support needs. Although this conceptualization of a multidimensional ASD was intended to lend diagnostic flexibility and specificity, the language of a ‘spectrum’ has been adopted and extended by researchers in ways far exceeding those imagined by the DSM committee. For example, hearkening back to DSM-IV’s multiple diagnostic categories, which identified distinct typologies of autistic difference, many researchers regard the autism spectrum not as smooth continuums across symptomatic domains, but as a fragmented collection of ‘subgroups’ bound together under an umbrella diagnosis. Others treat the spectrum as a continuum from normal to pathological, in which subclinical difference in autistic domains is a common trait, and diagnostic thresholds are somewhat arbitrary impositions. While these orientations lead to significant differences in research practice, they are not always clearly explained by researchers using the same language in manifestly different ways.
Given the scale of the research literature now produced annually, however, it’s difficult to identify clear patterns in this widely heterogeneous body of work. In order to begin to sketch its contours, then, this section employs computational bibliometrics to identify patterns in the topics discussed and the literature cited. First, I examine a keyword co-appearance network: Such networks exceed traditional keyword analysis, suggesting not only what sorts of topics are discussed, but which concerns tend to circulate together—and which are rarely voiced in tandem. Here, I thematize trends in this network to introduce two distinct categorical implementations of ‘autism’, demonstrating a marked disjuncture in how ASD is described and studied in contemporary big data genomics. Then, I turn to a bibliographic coupling analysis, a similarity measure that maps when two studies both cite a common third reference. This allows me to make the case that there are also two increasingly distinct bodies of scholarship cited by autism researchers, with further implications for the conceptual unity of the field.
I deploy these tools in a corpus representing five years of research publications on autism genomics (with mention of both ‘autis*’ and ‘gene*’ and/or ‘genom*’ in the keywords, title, and/or abstract), compiled through Web of Science. Filtering for original research articles published between 2017-2021, this resulted in a collection of 1,472 papers. I screened these papers manually, documenting data provenance for any paper relying on a biobank, data-sharing consortium, or other big data resource. All other papers using other data types were excluded, including cell and animal model studies, small case series, systematic reviews, and duplicate records. This left a final corpus of 303 articles. Comparing the cleaned corpus with the original set of publications, it is clear that large dataset reuse is increasing over time: Only 14.3% of papers published in 2017 used this type of data, but that number nearly doubled to 28.5% in 2021. It seems likely that reuse will continue to increase as sequencing costs continue to decline and large whole genome resources become more available, solidifying the role of biobanks and similar institutions in autism research and human genetics broadly.
Keyword clusters
To get a sense of the discursive landscape figured by these articles, I used the open-access bibliometrics tool VOSViewer (van Eck & Waltman, 2010) to map the co-occurrence of words and phrases which commonly appear in the titles and abstracts of articles in this corpus. The resulting network is pictured in Figure 1. 3
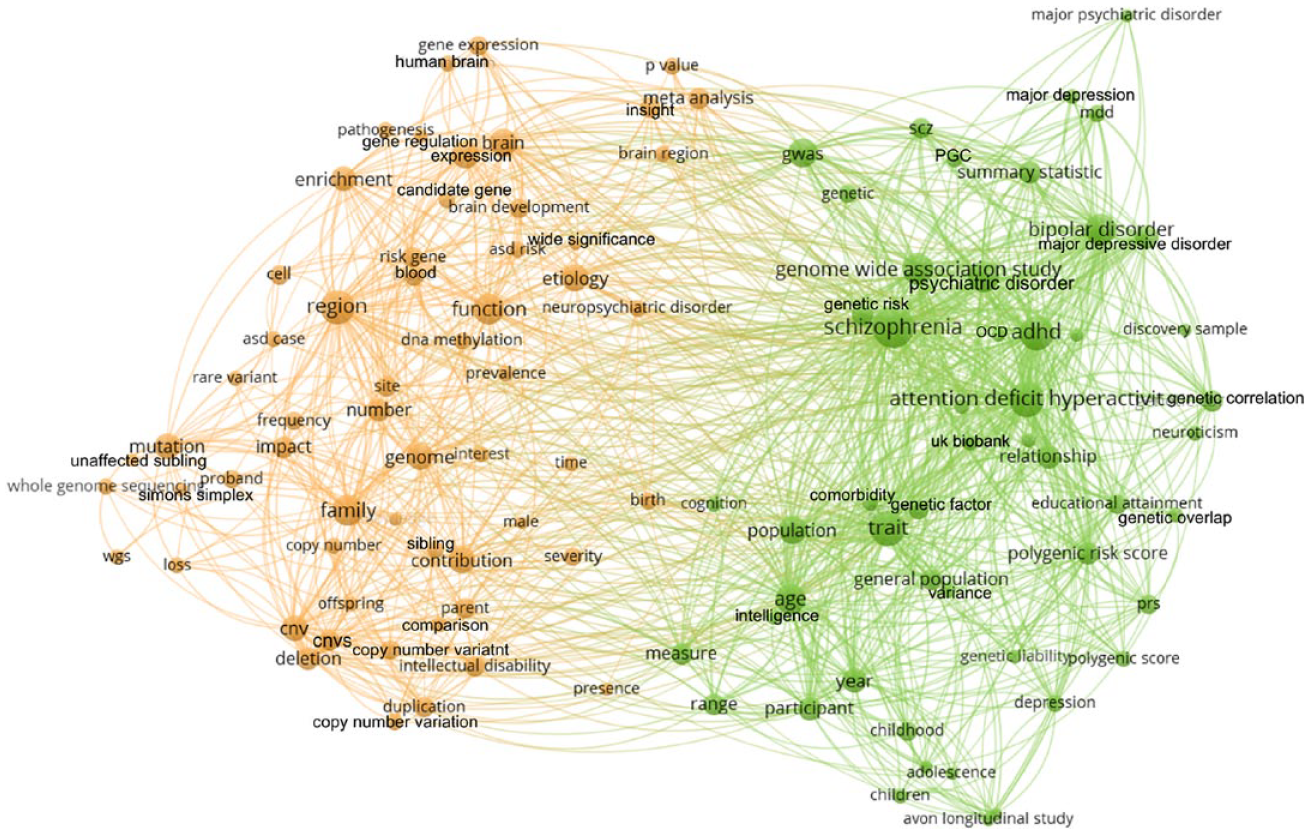
This network shows common terms which appear in the titles and abstracts of corpus publications. Terms that are close to each other frequently appear together in the same publications; distance indicates words which rarely appear in the same publications.
After excluding common words, VOSViewer maps the frequency with which any two terms appear together (positive spring weight) versus independently (negative spring weight) in the titles and abstracts of all papers in the corpus. Those weights are used to construct clusters of commonly co-appearing terms, mapping relationships within and between them. Increased distance between two nodes indicates that those terms typically appear in the absence of one another; nearby nodes are terms which frequently appear together in the same papers.
As the clear bifurcation of this network indicates, two relatively distinct discursive assemblages appear to be at odds in this corpus, reflecting what I will term Categorical Alignments A and B. In this subsection, I explore patterns in this network by linking particular sets of terms to the larger concepts they indicate. What emerge are clear connections between a variety of objects of interest, theories about genetic causation, and problematizations of the autism category that are not necessarily obvious in any individual paper, but yet cohere clearly within the discursive production of the larger field.
Some of the notable features of each categorical alignment will likely not come as a surprise to those interested in the history or sociology of autism—they resemble particular and historied configurations of the autism concept. Nevertheless, I think it is useful to hold these resemblances in abeyance in order to avoid assuming or misreading the actual contents of these categories as they are instantiated within this particular body of research—these will be explored in the final empirical section.
Now let’s consider the keyword network—beginning with Categorical Alignment A, the orange cluster at left. An initially striking feature is a cluster of synonyms in the lower left: ‘copy number’, ‘copy number variant’, ‘copy number variation’, ‘cnv’, and ‘cnvs’. Copy number variants (CNVs) are a form of structural genetic variation in which certain chromosomal regions are duplicated or deleted (‘duplication’ and ‘deletion’ are also tightly coupled here, and ‘region’ is in the center of this cluster). While everyone has CNVs, particular variations in some regions are widely considered causative of or strongly linked to autism (see Vicari et al., 2019). However, given that CNVs and single-nucleotide rare variants (‘rare variants’) are together estimated to account for less than 20% of autism’s total incidence, it’s notable that interest in CNVs specifically seems to characterize such a disproportionately large swath of the network.
We can also identify etiologic concern as primarily situated within Categorical Alignment A. ‘Etiology’ and ‘pathogenesis’ are visible toward the top, and a variety of keywords describing how genes are (or aren’t) expressed are clustered around them: ‘gene expression’, ‘dna methylation’, ‘enrichment’, and ‘function’ all stick out. Note also the prevalence of words like ‘brain’, ‘brain development’, ‘blood’, and ‘cell’, suggesting mechanistic inquiry as to how genetic processes shape larger biological systems—a concern for what autism ‘is’ and how it functions at the level of tissues. Finally, this half of the network includes a collection of words describing populations of interest in these studies, including ‘family’, ‘parent’, ‘offspring’, ‘unaffected sibling’, and ‘proband’ (the first individual in a family to be diagnosed with a particular condition, sparking pedigree or other familial study). This suggests a concern for inheritance, for novel genetic differences that emerge between parents and children, and for a particular pedigree-centric approach to medical genetics.
Taken together, this set of keywords starts to frame a particular set of ideas about what autism is and how it should be studied. Categorical Alignment A can be summarized as follows: Autism Spectrum Disorder is typically the result of CNVs and other rare, highly penetrant genetic differences. These mutations can be studied and understood through their etiologic changes in the brain and other tissues. ASD is usually identified in children.
These terms are siloed together on one side of a relatively bifurcated network. This means, for example, that papers that are concerned with familial inheritance are much more likely to reference CNVs as a proximate cause to the general elision of causes described on the other side of the network—despite the fact that CNVs are not always novel mutations in a particular family, and a variety of other changes that aren’t CNVs have also been linked to autism. In other words, these data make it clear that certain epistemic approaches and topics of interest can travel together in this literature even when they aren’t conceptually reliant on one another. In the next section this will be seen to have critical implications.
Let’s look at Categorical Alignment B, the green cluster on the right side of the network. Here, we can observe a starkly different approach to how autism should be studied—not through concern for its cellular mechanisms, but by the higher-order cognitive and social traits with which it is identified. Toward the bottom are a number of terms suggestive of ‘trait’ genetics projects, including ‘cognition’, ‘educational attainment’, ‘intelligence’, and ‘neuroticism’, as well as descriptive language like ‘measure’ and ‘range’. Interestingly, the only similar trait on the left side of the network is ‘intellectual disability’, suggesting a much more limited engagement with (or measurement of) cognitive difference in those projects. A similar interest informs terms at the top of the cluster, which appear to be concerned with the ‘genetic overlap’ or ‘genetic correlation’ between autism and a variety of other ‘psychiatric disorder[s]’ including ‘schizophrenia’ (commonly abbreviated ‘scz’ in academic genetics), ‘major depressive disorder’ (‘mdd’), ‘bipolar disorder’ (‘bpd’), ‘obsessive compulsive disorder’ (‘ocd’) and ‘attention deficit hyperactiv[e disorder]’ (‘adhd’). While these conditions have well documented genetic similarities with each other and with autism (see Sullivan & Geschwind, 2019), they are diagnostically distinguished by differences in symptoms—or, as we might name here, differences between their associated behavioral and cognitive traits.
A concern with traits also necessitates a different mode of analysis. Rather than cellular etiology, the right side of the network abounds with language describing data-intensive statistical measures, including ‘genome wide association study’ (‘gwas’) and ‘summary statistic’. This follows from an interest in trait genetics: The evidence that complex behavioral traits like educational attainment are meaningfully genetic tends to be quite thin, and these traits are unfailingly correlated with changes in dozens or hundreds of genes. This complexity means that locating causally significant biological regularities (if they even exist) is near-impossible, requiring alternative epistemic strategies that avoid the question of cell or tissue-level etiology. Researchers navigate and even operationalize the large numbers of correlated genes by producing ‘polygenic risk score[s]’ (‘polygenic score’, ‘prs’), which estimate the combined effect of many genes on a given phenotype. These effects are sometimes also described as the ‘genetic liability’ or ‘genetic risk’ of that phenotype, terms included nearby.
Here we see not only evidence of a shared epistemic style, but a theory of the autism genome. As we might summarize, in Categorical Alignment B, Autism Spectrum Disorder is linked to various differences in cognition, behavior, and life course. It is the product of complex, multifactorial genetic contributions in which any individual gene is minimally penetrant, requiring statistically intensive methods of study. Both the phenotypic traits and underlying genetics of ASD often overlap with those associated with other major psychiatric disorders.
As we can see, Categorical Alignments A and B contrast sharply, both in how they understand the genetic architecture of autism as well as how they situate it in relation to other conceptual entities.
Citational clusters
So: Researchers are talking about different sorts of autistic traits, in relation to different groups, and they are using different analytic methods to do so. But there is one more bifurcation worth exploring in this corpus—a division in their reference materials. Again, using VOSViewer, Figure 2 maps bibliographic coupling patterns for all cited references used in five or more corpus publications. 4
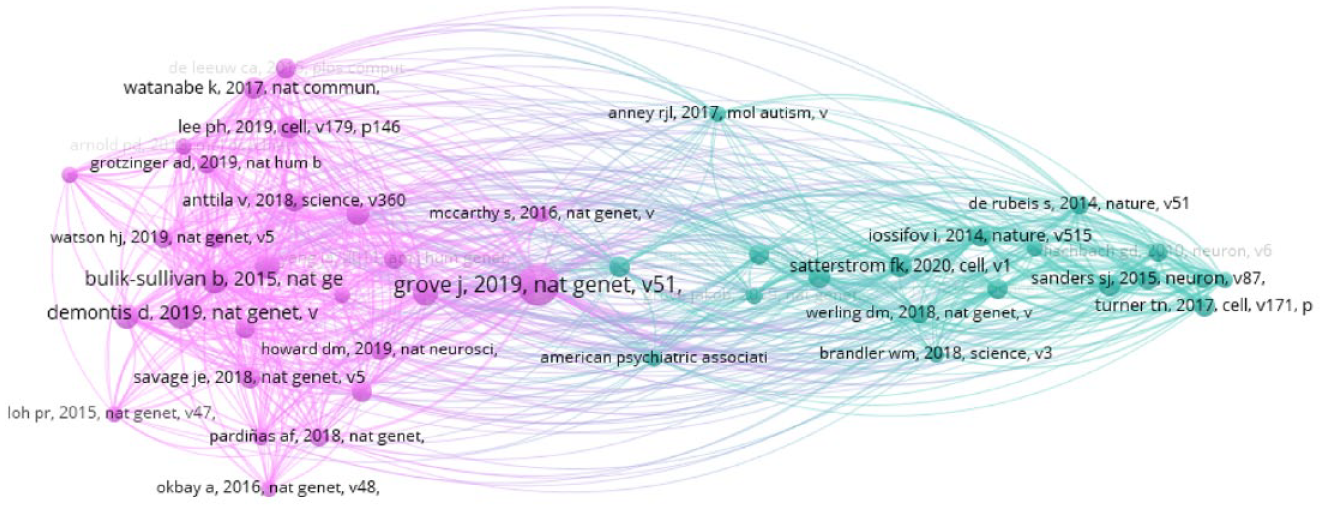
This network shows which references are cited by five or more corpus publications. References that appear close to each other tend to be cited by the same articles; references which are farther apart are rarely cited together.
Labels refer to the first author, year and journal of each reference. Here, nearby nodes are those that are frequently cited by the same papers. We can observe again that there is a marked division in the network, suggesting that papers in this corpus are not citing the same body of foundational texts in autism genetics, but are instead drawing from two somewhat distinct sets of literature.
The bibliometric methods employed mean that it is not necessarily the case that this division maps neatly onto the last one—but there is good reason to believe so. Importantly, it is not just that these clusters represent different citations, but that they represent different types of citations. Table 1 thematically groups the references in each cluster.
Thematic characterization of all references included in the network in Figure 2.

The teal cluster is primarily concerned with twin and family studies, and it includes a prominent reference to the Simons Simplex Collection—a biobank which indexes ‘simplex’ (single autistic child) families. Conversely, the pink cluster is anchored by GWAS studies, and includes a number of technical papers which validate statistical methods for things like complex- and multi-trait analyses using GWAS data. It also figures references to the UK Biobank—perhaps the most notable multipurpose biobank in the world—as well as to iPSYCH, a large Danish case-control cohort which contains five psychiatric disorders (including autism). It is clear how the types of resources and statistical approaches indicated by the pink cluster would facilitate the kinds of cross-disorder trait analyses common to Categorical Alignment B.
Taken together, these networks are suggestive of a body of literature that is growing increasingly disjoint. While autism may have long been multiple, as the previous section described, researchers have historically worked to produce a ‘coordinate unity’ (Potochnik, 2020) which could encompass multiple understandings of both the disorder and its causes. Indeed, most researchers would nominally agree that ASD includes cases that resemble both Categorical Alignments A and B as well as mixtures between them (e.g., Happé et al., 2006; Weiner et. al, 2017). This said, and as Mol (2003) has demonstrated, the coordination of such multiplicity is not trivial work: Autism cannot simply be ‘both’ of these things without the ongoing and effortful coordination of the research communities who produce them. If researchers are using ‘ASD’ to describe markedly different ontologies of autism, evinced through divergent methods, and in conversation with different literatures, the ties holding them together would appear increasingly threadbare.
To reiterate, this is not to suggest that these particular categorical instantiations of autism have become entirely irreconcilable, nor that they are wholly novel: They are neither. It is the assemblages and divisions of concepts that make these Categorical Alignments interesting, not the novelty of the concepts themselves. Nevertheless, it seems that this research community is moving away from—rather than toward—a consensus framework that figures both within a shared discursive landscape. Moreover, it appears to be the case that data (and data-intensive methods) have something to do with it. What remains to be shown is how those differences link to particular datasets, and how these categorical gestalts are animated in specific research programs.
Database(d) differences: MSSNG and 23andMe
A brief review of the larger corpus, however, suggests how challenging this is to do. There are: hundreds of large datasets represented in these articles, including population registers like the Avon Longitudinal Study of Parents and Children; national biobanks such as Generation Scotland; autism-specific resources like the Autism Genome Project; biobanks specific to conditions other than autism, such as the Atherosclerosis Risk in Communities cohort; data-sharing consortia like the Psychiatric Genomics Consortium; and a variety of other personal and institutional collections described only by reference to their cities of origin (Adelaide, Brussels, San Diego). Complicating matters further, most studies combine data from multiple sources, often relying on between three and ten biobanks to assemble both autistic and control populations of sufficient size.
Rather than attempting to disentangle this larger set, I track the contributions of two organizations: the parent advocacy-linked biobank MSSNG, and the direct-to-consumer genetic testing company 23andMe. Each is a notable data resource in contemporary autism genetics, and—as I return to in the conclusion—both are partially representative of broader categories of similar actors. First, I describe these two biobanks in more detail, and highlight how historical dissimilarities in their scientific goals and recruiting strategies have led to fundamental differences in both the autistic populations they index and the types of information they collect. These differences model the above keyword analysis, demonstrating that the topical disjuncture evinced in the larger corpus falls along the lines of data origin and specifying some of its less-obvious properties. Then, in the final empirical section, I provide a close textual analysis of the subset of articles produced using 23andMe or MSSNG data. This illuminates how organic differences between groups of people are assembled into biobank data sets, ultimately resourcing a fundamental division of categorical definition.
MSSNG
Originally introduced in 2014 as the soon-to-be-renamed AUT10K project, MSSNG is a collaborative effort between the parental advocacy organization Autism Speaks, Verily (formerly Google Life Sciences), and the Toronto Hospital for Sick Children (SickKids). MSSNG’s data represent the whole genome sequences of over 11,000 research participants as of 2022 (MSSNG, n.d.), making it the largest resource of its kind for autism genomics. Of particular interest here, its self-stated aims strongly resemble Categorical Alignment A. As its research team described in a recent publication: ‘Our study provides a guidebook for exploring genotype-phenotype correlations in the 15-20% of ASD families who carry ASD-associated rare variants, as well as an entry point to the larger and more diverse studies that will be required to dissect the etiology in the >80% of the ASD population that remains idiopathic’ (Trost et al., 2022, emphasis mine). I previously suggested that this particular combination of interests doesn’t hang together because of scientific necessity: An interest in families, in rare variants, or in etiology could be pursued in the absence of the others. MSSNG’s institutional history makes clear, however, there are other reasons for these concepts to travel together, and for the broader durability of this particular categorical alignment.
As an early press release touted, MSSNG was meant to become ‘the world’s largest collection of autism genomes’ in order to ‘transform the autism research landscape’ (Autism Speaks, 2014). However, MSSNG was not built as a novel resource from the ground up: Rather than recruiting thousands of participants, it instead reassembled data (and resequenced biomaterials) from a number of prior autism genomic databases. Those collections often represented similar imbrications of philanthropic funding and academic coordination as does MSSNG, sometimes even through the same institutions. For example, the largest contributing collection, the ASD: Genomes to Outcomes Study (representing 5,903 MSSNG participants as of 2022), shares MSSNG research director Stephen Scherer as its PI, and SickKids as a home institution (Prasad et al., 2012). Other notable contributing organizations include public-private partnerships like the Autism Simplex Collection (529 participants) as well as advocacy-funded projects at academic research centers like REACH (1,662 participants), iTARGET (463 participants), and the Autism Phenome Project (231 participants).
Perhaps most notable, however, is MSSNG’s inclusion of participants from the Autism Genetic Resource Exchange (AGRE, pronounced ‘agree’, 2,303 participants). AGRE was the first large-scale autism biobank in the US, founded in 1997 by the now-defunct advocacy organization Cure Autism Now! (CAN!). Its debut represented a turning point within both the research and advocacy movements coalescing around ASD at the dawn of the postgenomic era: as Singh (2015a, pp. 56–77) accounts, AGRE solidified the role of parental advocates as organizers who could coordinate resources, researchers, and federal funding. In the process, they established autism genetics as a particularly attractive and well-resourced subfield for up-and-coming molecular biologists, creating the conditions for the explosion of big data autism genomics through the 2000s and into the present day. In this sense, AGRE is the project that made MSSNG possible—not only by recruiting a plurality of its participants, but by fertilizing the landscape of autism research such that both researchers and funding would be plentiful by the time it arrived. Put another way, MSSNG is the logical end result of decades of parent advocacy, and can be seen to stand in for (and, in the case of these particular institutions, literally continue) the work of a host of previous groups with partially overlapping interests.
All of MSSNG’s contributing organizations are focused on infants and young children, and many tout affiliations ranging from pediatric hospitals to educational early intervention programs. The majority describe themselves as ‘family-centered’ in their public communications or on their websites. Unsurprisingly, then, MSSNG’s reassembled data are also familial, primarily representing grouped DNA samples from parents and their autistic child (‘simplex’ families) or children (‘multiplex’ families). These children often experience significant disability, as is typical of families that become involved in disease advocacy—more subtle phenotypic differences don’t necessarily create the same impetus for participation in time-intensive clinical research. Such families are often motivated by imaginations that research will result in meaningful differences in their children’s lives (Lappé, 2014; Silverman, 2011). For genomic research, then, that often means orienting toward the discovery of genetic markers which are likely to be linked to cellular or tissue-based differences that can become the basis of novel therapeutics—again joining etiology to this conceptual cluster not scientifically, but socially.
This aim is shared and specified in MSSNG’s own mission, which describes its ultimate goal as to contribute to ‘personalized and more accurate treatments’ via ‘the identification of many subtypes of autism’ (MSSNG, n.d.). ‘Subtyping’ is a commonly-used epistemic strategy toward the management and treatment of complex conditions, as with the identification of biomarkers to distinguish previously synonymous diseases (e.g. the recently mobilized distinction between HER2 ± breast cancers). In contradistinction to approaches resembling Categorical Alignment B, which seeks to expand its frame of interest outward across multiple conditions with genetic or phenotypic overlap, MSSNG proposes to do the opposite—to disintegrate the category of autism from the inside out, amplifying elusive differences in order to identify potential avenues of therapeutic ingress. This is, in short, a ‘splitters’ and ‘lumpers’ distinction. While ‘subtype’ does not appear in the keyword network, then, we can understand this epistemic opposition as driving much of the distance between the two categorical alignments.
23andMe
In contrast to MSSNG’s clearly defined conceptual orientation, 23andMe seems somewhat unmoored from any particular stake in ASD communities or research—indeed, it seems barely invested in basic research at all. However, while the company is better known for its direct-to-consumer genetic testing service, it has courted a robust secondary market circulating its customers’ genetic data for biomedical research. Much of this effort derives from its economic model, in which the company is able to use its data toward preliminary analyses establishing potentially lucrative drug targets or pharmaceutical pathways. In recent years this approach has begun to pay out, as 23andMe has sold a significant stake in the company to pharmaceutical giant to GlaxoSmithKline (Molteni, 2018), and rights to its first drug candidate to the Spanish biotech company Almirall (Wetsman, 2020). Beyond the spectacle of these high-dollar exchanges, 23andMe additionally provides data to academic and other nonprofit researchers. It also employs its own research team, which frequently collaborates and publishes with academic investigators on a diverse set of conditions, including autism and other neuropsychiatric disorders.
Importantly, 23andMe’s genomic data are single nucleotide polymorphism (SNP) sequences—not, like MSSNG, whole genomes. While humans share the overwhelming majority of their DNA, SNPs are the roughly 1% of base pairs that represent common points of variance between individuals. While some SNPs are strongly associated with disease, they are definitionally common: They do not include the ‘rare variants’ that can be found in the remaining expanse of the genome and constitute the main interest of Categorical Alignment A. They also cannot be readily used to identify CNVs. To genotype an individual on a SNP chip costs hundreds of dollars less than whole genome sequencing, making it a generally more suitable tool for the cost constraints of a consumer product like 23andMe. However, SNP genotyping does significantly limit the kinds of genomic inquiry that can be pursued with the resulting data, demanding different epistemic strategies that orient toward other affordances.
Instead, 23andMe produces data with two other notable qualities: scale, and multidimensionality. First, scale. Of its more than 12 million accounts, nearly 80% have open consent for their data to be used for research purposes (23andMe Research Team, n.d.), vastly outnumbering MSSNG’s 11,000 participants. Second, those users contribute a huge variety of data about their health and personal traits, far exceeding the sort of clinical data collection common in autism biobank curation. 5 This function allows the company to produce polygenic scores for their participants across a variety of both health and novelty traits, such as having a higher likelihood of developing glaucoma, or disliking cilantro. More interestingly, however, the expansive collection of health and trait data provides more potential variables to researchers looking for genotype-phenotype correlations, thus improving their likelihood of finding a statistically significant association. This flexibility is sometimes described as ‘researcher degrees of freedom’, and while it can be productive—particularly when dealing with extremely subtle statistical signals, as is often the case in psychiatric genetics—it is often linked to concerns about ‘data dredging’: exhaustively searching all variables including those likely to be in spurious correlation. Nevertheless, the abundance of traits (and the absence of more comprehensive genomic data) represented in 23andMe datasets are what makes statistically intensive trait genetics approaches like those represented in Categorical Alignment B not only possible, but practical epistemic strategies tied to the particular affordances of this data.
Population differences
To this point I have argued that the research imaginations of the actors who shape MSSNG and 23andMe’s data diverge on a number of points, which systematically orient them toward different knowledge production practices broadly resembling Categorical Alignments A and B. However, there is one more critical difference between these data sets—or rather, between the populations that they index. As has been well documented, 23andMe users are not, on the whole, population-representative: they are significantly more educated and of a higher socioeconomic status than the average US citizen (Tung et al., 2011). Because research cohorts are selected from within 23andMe’s large user base, it is not always clear if and how they resemble its overall makeup. However, within the studies I analyze below, several describe their autistic cohorts as having higher-than-average IQ and educational attainment, and the majority rely on survey instruments like the Systematizing Quotient-Revised (SQ-R) that are indicated for people of average or higher intelligence (Wheelwright et al., 2006). In MSSNG’s cohort, however, IQ ranges from well above to well below the population average, with roughly 20% exhibiting clinically defined intellectual disability (Yuen et al., 2015). Many of the studies produced using MSSNG data rely solely on participants drawn from that 20%, exploring genetic markers correlated with what they describe as ‘severe’ or ‘low-functioning’ autism.
In pointing out these differences, I do not mean to naturalize ‘high-functioning/low-functioning’ distinctions, or to forward intelligence (particularly as proxied by instruments as fundamentally flawed as IQ tests) as the most important marker of autistic diversity. As disability studies scholars and advocates have articulated, the rhetoric of functional severity relies on a deficit model, obscures how ‘functioning’ is itself a product of social environments and redirects attention from widely heterogeneous individual needs toward binaristic understandings of ability (Anderson-Chavarria, 2022; Baker, 2006). However, it is clear that these datasets index phenotypically—and, thus, almost certainly also genotypically—different populations. As Navon and Eyal (2016) have shown, expanding the population under analysis in autism genetics has already driven ‘looping’ effects, fundamentally altering the contours of the category. What, then, might happen when that population is not simply expanded, but splintered?
From data to disorder
A first glance at the published research produced using 23andMe (n = 22 articles) and MSSNG (n = 38 articles) 6 data does not suggest this split—indeed, the two sets of articles appear rather similar on the surface. These papers were published across a similar range of journals, from high impact generalist venues like Science and Nature to specialist publications like Molecular Psychiatry and Molecular Autism, and both groups of articles have average citation counts north of 30. The set of author-elected keywords (the keywords requested by the publisher, not the keywords examined above) attached to the articles is also broadly shared. Taken together, these publication metrics are (albeit roughly) indicative of a group of generally well-regarded papers contributing to a relatively well-defined research community.
However, as the rest of this section shows, these studies ultimately describe very different ideas about what autism is and rely on divergent approaches as to how it should be studied. Here, I attend to the disciplinary authorship, topics of concern, analytic approaches, and rhetorical strategies deployed in these articles to trace how differences in their data have echoed into much larger disjunctures in their accounts of autism. In doing so, I specify further differences between the categorical alignments embedded in their data and point to several resulting incompatibilities between these bodies of research literature.
First, though, it bears considering how these sets of articles square against the keyword network presented in Figure 1, which mapped co-appearance between common topics across the entire body of big data autism genetics published during this period. Figure 3 shows the frequency with which a selection of the respectively most common keywords appeared in 23andMe (blue) and MSSNG (red) articles.
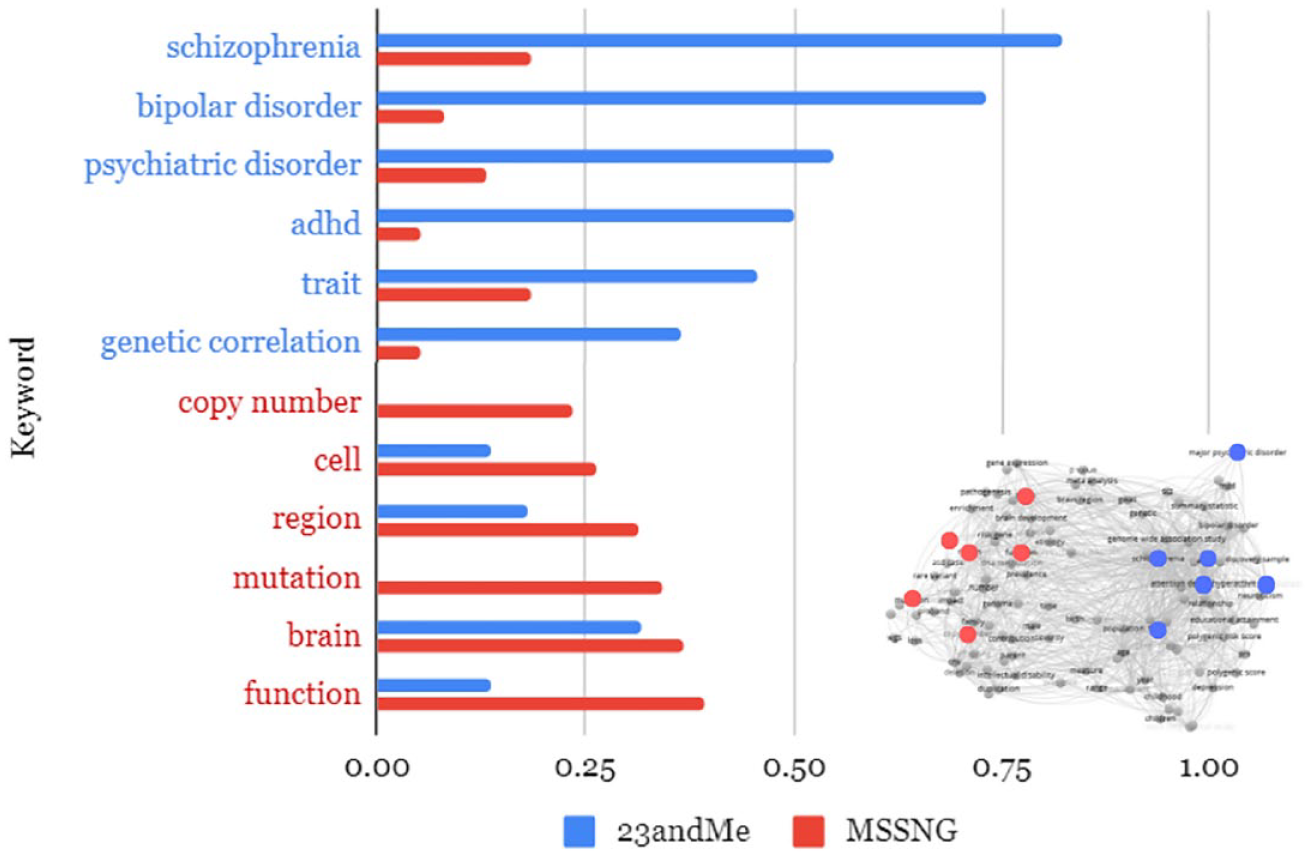
As the inversion of frequency suggests, these bodies of literature have some overlap but are largely concerned with different conceptual sets. Further, and as the recolored overlay of Figure 1 in the right corner of Figure 3 shows, those sets clearly map onto the two clusters representing Categorical Alignments A and B. This should come as no surprise—as the previous section showed, 23andMe data are poorly suited for work investigating CNVs, rare variants, or cellular etiology, but have scalar affordances that enable cross-disorder investigation of various traits; it is the opposite for MSSNG. Datasets shape and constrain the kinds of topics that animate subsequent research.
There is one more notable difference between these sets of articles worth examining before digging into their contents—their authorship. Table 2 presents the disciplinary affiliations of the last author (conventionally the Principle Investigator [PI] of the project) of each paper using publicly available biographical information.
Disciplinary affiliations of the last authors of papers in the 23andMe and MSSNG corpuses based on publicly available biographical data.

It’s notable that a majority of the last authors using MSSNG data are molecular biologists—mostly appointed in genetics or genomics departments. 23andMe last authors, in contrast, tend to have joint appointments or to work solely in a psy-science. While this is a small sample of the larger group of PIs working in autism genetics during this period, it points to something interesting: there’s a relationship between home discipline and choice of dataset—and with it, its particular affordances and constraints.
The fragmentation of behavioral genetics is well-documented: as Panofsky (2014) describes, behavioral genetics is a disciplinarily antagonistic field in which mutually distrusting groups of scientists compete for authority within controversial problem spaces. Along these lines it makes sense that geneticists would develop projects that seek to identify causal rare variants and trace occulted cellular pathways—the imagined use-case for which MSSNG was designed. Similarly, a concern with symptomatology and the relationship between disorders aligns with the psy-science's longstanding authority over the evolution of ASD’s diagnostic categorization. Work in that tradition requires phenotypically detailed data such as 23andMe’s surveys. All of this is to say, another way of telling this story with different protagonists could follow preexisting disagreements between disciplinary researchers, who select the data that seem sufficient for their diverging interests and goals.
But, and as I will argue in the remainder of this section, there is more to gain from focusing on the datasets. The disciplinary fragmentation at play certainly helps to explain some of the differences between these two sets of papers, as well as the citational split within the larger corpus. However, it is not simply that the papers authored by psy-scientists tend to build on similar discourses, but that the papers using shared data do so even when authored across disciplines. Rhetorical, epistemic, and categorical patterns in these papers are distinctly shaped by the imaginations and affordances of their data.
Looking first at the MSSNG papers, a notable feature is in fact how little they discuss what they understand autism to be. Articles routinely start with a single sentence describing the core DSM diagnostic domains of ASD before immediately turning to genetic analysis. Consider the first lines of these papers: Autism is a neurodevelopmental condition currently defined by atypical social communication and interaction, intense interests, and repetitive behaviour. (Douard et al., 2021) Autism spectrum disorder (ASD) is a complex neurodevelopmental disorder that often involves impaired cognition, communication difficulties and restrictive, repetitive behaviors. (Jangjoo et al., 2021)
Others elide diagnostic criteria entirely, skipping clinical definition and beginning directly with epidemiological or mechanistic descriptions of autism: Autism spectrum disorder (ASD) is a phenotypically heterogeneous disorder affecting about 1 in 59 children in the United States. (Wilfert et al., 2021) The genetic basis of autism spectrum disorder (ASD) is known to consist of contributions from de novo mutations in variant-intolerant genes. (Brandler et al., 2018)
While behavioral genetics research does not tend to spill substantial ink on questions of symptomatology, it is typical to include at least some discussion of the behaviors under investigation. The descriptions of ASD in this corpus are strikingly brief, and are generally not elaborated further in the bodies of these articles. To these researchers, then, it seems that what autism is as a socio-behavioral category is entirely self-evident—so obvious, in some cases, as to not even merit a clinical description. Moreover, as the unchallenged repetition (or assumption) of a DSM-adjacent definition of ASD suggests, the expanse of the diagnostic category appears synonymous with the population under investigation in these studies. It is worth repeating here that MSSNG’s cohort is not necessarily broadly representative of the total population diagnosable under DSM-5, as the previous section showed.
Despite these narrative beats, however, this body of literature is not entirely bought into the DSM’s conceptual schema. Several papers deal with disease ‘subtyping’ by name, and many more implicitly rely on subtyping as their primary epistemic approach. The DSM does not allow for conditions with the ‘same’ symptoms to be broken out separately: Subtyping challenges the DSM’s categorical unity by identifying genotypes that can be linked to a clear, if subtle, pattern of phenotypic difference within the broader diagnostic category. Importantly, however, the phenotypic ‘subtypes’ in these articles are near-totally limited to the autistic domains associated with DSM criteria: IQ, Language, Social Responsiveness, Social Communication, and Repetitive Behaviors. These traits are formalized and measured by the standardized diagnostic instruments with which MSSNG collects data. However, by assessing the comparative ‘severity’ of individual domains when associated with particular genetic variants, these articles attempt to make the case that genetically distinct subtypes exist. These subtypes are commonly linked to CNVs and other rare variants in these papers, as the frequency of ‘copy number’ and ‘mutation’ in the corpus also indicates.
After making these genotype-phenotype links through statistical analysis, the majority of the papers in this corpus then follow their gene(s) of interest into mechanistic investigation. These projects take a variety of forms, ranging from animal model experimentation to in vitro cell culture studies to neural mapping and brain tissue analysis. In all cases the goal is to demonstrate that the gene(s) are not merely associated with the phenotype but are definitively causal—or at least, causal of an isolable biological change that can then be speculatively linked to autistic symptomatology. In doing so, these works serve to construct autism as an increasingly biologized disorder. This also serves to orient their larger contribution toward the development of future clinical interventions: ‘therapeutics’ and ‘diagnostics’ are topics referenced in the majority of discussion sections in this corpus, but uncommonly in the 23andMe papers. However, and while the particular biological pathways examined here may one day be clinically actionable, it’s important to note that research in this corpus is extremely distant from drug development projects: the invocation of therapeutics is as much a rhetorical strategy as it is a plan for future research, discursively constructing an autism that can be genetically decomposed into readily interpretable and manipulable biological pathways.
The 23andMe corpus paints a starkly different picture of autism—or, rather, pictures. Consider these opening lines, representative of the how this literature tends to frame a variety of concerns far exceeding DSM diagnostic criteria: People who experience childhood abuse are at increased risk of mental illness. Twin studies suggest that inherited genetic risk for mental illness may account for some of these associations. (Ratanatharathorn et al., 2021) Use of tobacco is (still) prevalent in the Western world: about 20% of the population (15+ years) in Europe and the United States is a regular smoker. This percentage is remarkably higher in people with psychiatric disorders. (Vink et al., 2021) Empathy is the ability to recognize and respond to the emotional states of other individuals. It is an important psychological process that facilitates navigating social interactions and maintaining relationships, which are important for well-being. (Warrier et al., 2018)
The word ‘autism’ doesn’t appear here, despite being a central topic in all three papers. Instead, unnamed ‘psychological process[es]’, ‘mental illness[es]’, and ‘psychiatric disorders’ mediate traits of more immediate interest. In turn these accounts push back against the assumed boundaries of autism as a category: Beyond classic cardinal symptoms, readers are invited to consider autistic populations who smoke, or have experienced childhood abuse. This narrative style and topical framing is common throughout the 23andMe articles, examining how autism is genetically associated with traits as diverse as the frequency of mouth ulcers, aneurysms, and left handedness.
Moreover, several of these articles attempt to render internal attitudes and emotions as measurable—and (at least partially) genetically-determined—autistic traits. While the DSM criteria are firmly rooted in psychiatric measures, those are strictly external (and generally interpersonal) displays common to clinical evaluation. For example, ‘repetitive behavior’ is easily quantified, and ‘social responsiveness’ is distilled within diagnostic interview guides as discrete actions like eye contact, facial expressions, or verbal reactions. In contrast, something like ‘empathy’—or ‘neuroticism’, another common topic in these articles—is a much more abstract psychological (rather than clinically psychiatric) concept, and the survey-based measures deployed through 23andMe rely on self-reflective subjects capable of participating in extended documentation of their emotions and personal histories. Here, we can observe that these instruments don’t simply assess one phenotype, but necessitate others: it would be straightforwardly impossible for many of the children represented in the MSSNG data to complete such an exercise (nor are these instruments designed or validated for children younger than teenagers). Thus, while overtly expanding the sorts of traits that we might associate with autism, we can observe that these articles also implicitly foreclose a variety of others.
Due in part to the complexity of the behaviors under investigation, all but one of these studies evince dozens of correlated genes, and often describe their findings as indicative of ‘polygenic risk’. Given the sometimes tenuous relationship between autism and these traits, many papers also talk about ‘pleiotropy’, the idea that one gene can have multiple and apparently unrelated phenotypic effects. This means that—and unlike the MSSNG papers, which often include biological experimentation—projects in this corpus by and large cannot directly examine biological pathways. The tens of genes implicated in something like a polygenic risk score can be involved in many more cellular processes, with no clear indication which might play an important role. Instead, the majority of these studies have to rely solely on computational/statistical approaches to substantiate their findings, and turn to biological intermediaries between genes and behavior only through theoretical speculation.
In the absence of biologized ‘proof’, then, many of these authors turn to other approaches to underline the importance of their work. Because each gene in a polygenic score represents an extraordinarily subtle statistical signal, this type of research requires significantly larger research cohorts than does the identification of a highly-penetrant rare variant. Indeed, the 23andMe papers average a staggering 913,515 participants per study, while the MSSNG publications average only 10,611. This is an important point of contrast: While both 23andMe and MSSNG describe themselves using the language of ‘big data’—and while both are big, compared to the autism data of a decade ago—this demonstrates that ‘bigness’ cannot be treated as an epistemic monolith. MSSNG’s data are ‘big’ in part because they are whole genomes, but include minimal phenotypic information per participant. 23andMe have ‘small’ SNP data but huge numbers of participants and access to other types of data about them. These scalar differences are tied to different affordances and favor different meaning-making strategies. Indeed, as we see in the 23andMe papers, the repeated invocation of scale serves as much of a rhetorical purpose as it does a scientific one: Nearly all of these articles reference the size of their participant cohort in the abstract, and several underline its magnitude as an epistemic virtue setting their study apart from previous work within their methods sections. These features are uncommon in MSSNG papers.
Further, the trait-first approach allowed many of the 23andMe projects other means by which to maximize their study size. Traditional genetic analyses of a diagnostic entity like ASD look for genetic patterns in research participants with that diagnosis. Here, however, several projects seek to determine the underlying genetics of ‘autistic traits’ in participants without a diagnosis of autism, including those with other psychiatric diagnoses representing partially overlapping symptoms, and those without psychiatric history who may yet identify with a certain trait (e.g., a high ‘neuroticism’ score on a personality survey). Genetic correlates in those groups then become candidate genes for ‘autism risk’, providing additional avenues for triangulating very weakly-penetrant genes. Through these strategies, autism itself begins to appear as an accumulation of dissociable traits, and correlation is justified as a meaningful epistemic approach to autism genetics even in the absence of testable biological mechanisms.
An implication of a trait-driven epistemology is that the majority of these articles aren’t singularly ‘about’ autism—as nearly all of the MSSNG papers are—but are instead about clusters of neurodevelopmental and psychiatric disorders that include autism along with a number of other conditions (as also indicated by the high keyword frequency of ‘schizophrenia’ and ‘bipolar disorder’ in Figure 3). Because these diagnostic entities include some symptomatic overlap, regarding those symptoms as genetically dissociable explains the additional (and well-documented) genetic overlap between these conditions. This is to say, if individual psychiatric traits are genetically determined, two diagnostic entities that share a trait must also share those genes. Some articles go a step further and refer to diagnostic categories themselves as ‘psychiatric traits’, invoking a flat ontology in which diagnostic categorization has no more authority than any other approach to the categorization of human behavioral difference. This is in stark contrast to the MSSNG papers, which analyze cross-disorder symptomatology only when specific genetic variants are linked to multiple diagnoses—a gene-first, rather than trait-first, approach.
As this section has shown, both bodies of literature challenge ideas of ASD as a unifiable category, but they do so from opposite directions. Articles using MSSNG data attempt to decompose autism from the inside out, identifying genetic subtypes that can be biologically analyzed toward the development of eventual clinical interventions. In contrast, 23andMe papers leverage ‘traits’ to disintegrate the category from the outside in, using autism’s phenotypic similarity with other diagnostic entities and populations to make sense of its polygenic complexity. In both cases researchers draw on the strengths of their respective datasets, deploying methodological strategies that would not be practical (or even possible) with the other data. This demonstrates that datasets do not shape subsequent research simply by constraining the available variables: rather, their inbuilt affordances echo the epistemic values of their creators and users in a variety of subtle but impactful ways.
Critically, and as the previous section also argued, these datasets additionally represent groups of research participants with markedly different phenotypes—almost certainly indicating underlying genetic differences between them as well. As a result, even if direct replications of these studies using the other dataset were possible (which they are typically not), it is unclear if results produced using MSSNG data would hold in the 23andMe population or vice versa. Indeed, underlying population differences are likely to be a significant factor in repeated failures to reproduce candidate gene findings across different cohorts—and while accounts of failed replication attempts often argue that even larger populations will resolve this issue (e.g., Torrico et al., 2017), what this study suggests is that scale alone has not and likely cannot solve that problem. Moreover, that these data resource incomparable levels of explanation and epistemic approaches serves to further obscure this fact.
Conclusion
The differences we see in the above analysis also appear to characterize the larger body of research literature on autism genomics, mapped in the earlier keyword analysis (see Figure 1). Despite the fact that those publications draw from dozens of other datasets, this is not necessarily surprising: In many ways MSSNG is a paradigmatic example of autism advocacy biobanks, which have played a major role in the field since the 1990s (Navon & Eyal, 2014; Singh, 2015a). Resources like the Simons Simplex Collection share research imaginations, recruiting strategies, and data collection practices with MSSNG, and it stands to reason that those data would produce discursively similar accounts of ASD. Indeed, in publications using multiple datasets, MSSNG’s data were combined with those from Simons and/or the Autism Genome Project (another family biobank) to the near exclusion of all other organizations. In contrast, 23andMe better resembles a more recent wave of databasing projects like the UK Biobank or the Psychiatric Genomics Consortium (PGC). These organizations work to collect massive amounts of highly dimensional data in order to facilitate data reuse across diagnostic conditions and topical concerns. Unsurprisingly, then, the biobanks whose data were most commonly combined with 23andMe were PGC and iPSYCH.
Although there are certainly important differences between individual datasets in both groups, this broad dichotomy appears to be a proximate cause for the disjuncture across the field. Datasets must have common instruments or commensurable variables in order to be combined or compared. It is not just that the categorical imaginations of autism in these datasets differ, but that their pragmatic approaches to producing inscriptions are not, on the whole, compatible: they measure largely non-overlapping sets of traits. These sets of traits, in turn, shape what can be said about autism, and the methods with which it can be evinced. When these organizations additionally index different populations, it becomes impossible to make claims using one set of categorical concepts regarding the other group. For example, there are no publications about the genetics of neuroticism or empathy in autistic children with significant disability because large scale data about those traits in that group do not exist. There are few publications about autism cases linked to CNVs or rare variants in independent adults for the same reason. It is not simply that research accounts of autism produced using divergent data don’t overlap, it’s that they largely can’t.
I suggested above that research produced using these data might be productively described with the metaphor of sedimentation: While elements of these different imaginations have long coexisted within the landscape of autism research, dataset disjunctures have precipitated them into distinct and increasingly immiscible layers. But these layers do not exhaust all the possible categorical configurations of autism—far from it. Here, it is interesting to note that recent years have seen a spate of new autism biobanking projects that are oriented toward different uses and situated within different national contexts. Both MSSNG and 23andMe are North American institutions: While their data may be used globally, their epistemic genealogies can be firmly situated within US histories of genetics and autism research. As Evans (2013, 2017) has explored, however, ideas about autism in the UK have evolved along different cultural lines and in response to different political pressures. Take, for example, British psychologist Simon Baron-Cohen’s controversial proposal 7 for Spectrum 10K—a UK-based autism biobank which includes MSSNG and its scientific director Daniel Geschwind as ‘partners’. It is interesting to note that the project includes a wide, 23andMe-like array of trait questionnaires and is designed for research into ‘co-occurring disorders’, apparently responsive to a very different set of research trajectories and institutional actors than MSSNG. If the project comes to fruition, it will be interesting to observe how these data circulate. It is easy to imagine a future in which they bolster 23andMe-adjacent research literature while resituating traits of particular interest in the UK.
Moreover, differences between autism science in the US and UK narrow when compared with the broader global landscape: The huge number of culturally-sensitive concepts involved in autism research makes it quite difficult to even begin to work across contexts, much less collect large-scale data for comparative analysis (de Leeuw et al., 2020). Currently, projects like the NeuroDev Study—an autism and ADHD biobank recruiting in South Africa and Kenya, affiliated with the Broad Institute in the US—use instruments standardized in Global North research in order to produce commensurable data (de Menil et al., 2019). Nevertheless, it’s unclear how the sorts of contextual specificities, population differences, and local knowledges that de Leeuw et al. (2020) identify will shape the production and interpretation of these and similar data. It is certainly possible that datasets like this could precipitate very different categorical configurations than the ones I’ve described here.
This is not to give datasets the final word on which research trends take hold or which categorical alignments become definitive. Instead, this case study shows the messy coproduction of data, field, and object over time. In the institutional histories of MSSNG and 23andMe we can observe formative encounters with a variety of other actors: parents and their children, lay genetics enthusiasts and pharmaceutical investors, and the networks of researchers that form with and around them. Here, decisions about desirable research futures and the data they require far exceed the laboratory and the clinic, and the datasets they produce shape the kinds of big data inquiry that become possible in their wake. These datasets then enter a research ecology already rich with available concepts, epistemic approaches, and disciplinary intuitions. The affordances of particular data encourage, but don’t determine, what assemblages can form out of this mass.
In focusing on databases, I have pointed to a set of critical but often neglected actors in psychiatric genomic research. An attention to the origins of data reveals dynamics that are often obscured in research reporting, particularly regarding how choices about participant recruitment and data collection are managed. When multiple databases enact conflicting practices, then, the larger research ecologies that rely on them can echo and elaborate these points of divergence—even while appearing to represent ‘the same’ populations or phenomena. For autism genomics, this has led to a marked split not only in the conceptual category of ASD, but in the epistemology by which it is known. Given the rapid rise of large open access databases and other forms of dataset reuse across the sciences, then, similar forms of categorical misalignment are likely to represent important sources of controversy and reproducibility problems in the years to come.
Footnotes
Acknowledgements
Thank you first to Daniel Navon, who has been supportive of this project’s development from its earliest iterations and who has read far too many drafts along the way. Stuart Geiger, Nima Boscarino, and Jeremy Kemball kindly provided technical support and troubleshooting on the computation elements. Finally, thank you to the anonymous reviewers, as well as to audiences at 4S and ISHPSSB, for questions and comments which have quite improved the final manuscript.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Kathryne Metcalf’s work is supported by a National Science Foundation Doctoral Dissertation Research Improvement Grant (2341622).
Data Statement
Publication data included herein are derived from Clarivate Web of Science. (Copyright Clarivate 2023. All rights reserved.) Data are available upon request to the corresponding author.
