Abstract
This article expands on recent studies of machine learning or artificial intelligence (AI) algorithms that crucially depend on benchmark datasets, often called ‘ground truths.’ These ground-truth datasets gather input-data and output-targets, thereby establishing what can be retrieved computationally and evaluated statistically. I explore the case of the Tumor nEoantigen SeLection Alliance (TESLA), a consortium-based ground-truthing project in personalized cancer immunotherapy, where the ‘truth’ of the targets—immunogenic neoantigens—to be retrieved by the would-be AI algorithms depended on a broad technoscientific network whose setting up implied important organizational and material infrastructures. The study shows that instead of grounding an undisputable ‘truth’, the TESLA endeavor ended up establishing a contestable reference, the biology of neoantigens and how to measure their immunogenicity having slightly evolved alongside this four-year project. However, even if this controversy played down the scope of the TESLA ground truth, it did not discredit the whole undertaking. The magnitude of the technoscientific efforts that the TESLA project set into motion and the needs it ultimately succeeded in filling for the scientific and industrial community counterbalanced its metrological uncertainties, effectively instituting its contestable representation of ‘true’ neoantigens within the field of personalized cancer immunotherapy (at least temporarily). More generally, this case study indicates that the enforcement of ground truths, and what it leaves out, is a necessary condition to enable AI technologies in personalized medicine.
Introduction
For about ten years, inquiries in Science & Technology Studies (STS) have documented the constitutive relationships of machine learning or artificial intelligence (AI) algorithms, which are probabilistic models that infer calculation rules from sets of data (e.g. Hoffmann, 2017; Jaton, 2019; Lee, 2021). Among these social inquiries, some have focused on the material infrastructure required for the constitution of new algorithms (Crawford, 2021; Jaton, 2021a), especially in terms of data work and annotation (Gray & Suri, 2019; Tubaro et al., 2020). These works have made visible problematic entities often called ‘ground truths’ (Henriksen & Bechmann, 2020; Jaton, 2017) which are manually constructed benchmark datasets that gather input-data and output-targets, thereby establishing what can be retrieved computationally and evaluated statistically (see Figures 1 and 2). By underlining the centrality of ground-truth datasets for the development AI algorithms and by bringing to the forefront the question of the biases and inequalities inscribed in them (e.g. Crawford, 2016; Jaton, 2021b; Noble, 2018), these works have effectively operated, at their own level, as a counter-fire to the commercial and seductive rhetoric of techno-modernist promoters and AI over-enthusiasts.

Schematic of a ground-truth dataset for face detection (WIDER FACE, by Yang et al., 2016). On the left, one among the 32,203 images of the publicly available dataset for face-detection research. In the middle, the face annotations for this specific image. Since each annotation belongs to the coordinate space of the digital image, it can be expressed by a set of four numerical values, the first two expressing the start position of the label along the x and y axes, the third one expressing the number of pixel wide, the fourth one expressing the number of pixels high.
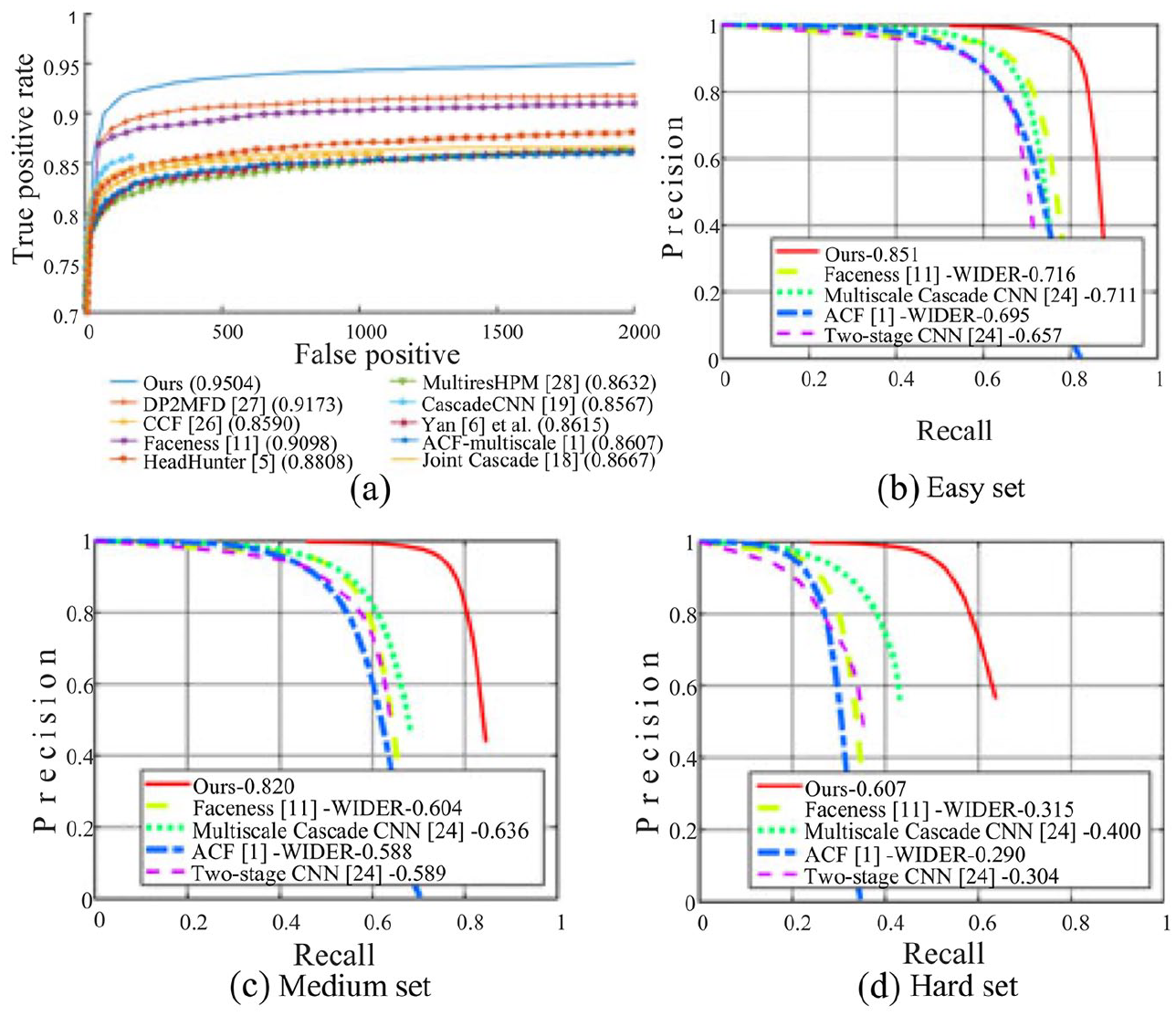
Performance graphs that evaluate competing face-detection algorithms on the WIDER FACE ground truth. All the curbs—called ‘Ours’, ‘DP2MFD’, etc.—refer to competing face-detection algorithms published by different research teams. Graph (a) shows the true/false positive rates of all algorithms; graphs (b) (c) and (d) show the precision (true positives divided by the sum of true positives and false positives) and recall (true positives divided by the sum of true positives and false negatives) of all algorithms on three subsets of the WIDER FACE ground truth, that themselves define the boundary of the coordinate systems (i.e. coordinate [1;1]).
Most of these studies have focused on applied domains of computer science (e.g. computer vision, natural language processing) and few have documented how AI algorithms are mobilized and worked upon in experimental sciences. 1 This is especially, and surprisingly, true for biomedical sciences: While the past few years have seen many important debates about the politics and epistemology of data- and algorithm-driven biomedicine (e.g. Chin-Yee & Upshur, 2019; Green & Svendsen, 2021; Leonelli 2016; Prainsack, 2018; Strasser, 2019), there are still few empirical case studies on the effects of AI algorithms on contemporary biomedical practices (c.f. Carboni et al., 2023; Dahlin, 2023). What is going on today in biomedicine with regards to AI technologies? As more and more utopian and dystopian discourses are being produced about an increasingly personalized medicine that would be powered by AI algorithms (e.g. Roth & Bruni, 2021; Topol, 2019; Zuboff, 2019, chap. 8), it seems important, or at least interesting, to address this question in a down-to-earth manner.
In this article I explore the case of a consortium-based project in personalized cancer immunotherapy—the Tumor nEoantigen SeLection Alliance (TESLA, nothing to do with the electric car company)—that started in 2016 and provisionally ended in 2020 with the release of an online dataset describing neoantigens, which are promising molecules for cancer immunotherapy, and the joint publication of a collective article in the high-impact journal Cell (Wells et al., 2020). Based on a literature review and semi-structured interviews (N = 12) with bioinformaticians who took part in the TESLA project, this article retraces some of the events that led to the publication of the first genuine ground truth for neoantigen prediction in cancer immunotherapy, as well as some of the issues and controversies related to this achievement. In that sense, this article is a practical application of what Kang (2023) recently coined ‘ground-truth tracing’, namely the investigation into the construction processes of ground-truth datasets as well as the habits, desires, and values these processes entail and promote. 2
Using the case of the TESLA project, I first intend to account for ongoing processes related to the development and use of AI technologies in biomedical research. In parallel, I also intend to dig deeper into the topic of ground-truthing processes by exploring an instructive ambiguous case where the ‘grounding’ of the scientific referent—the so-called ‘truth’ to be later approximated algorithmically—depended on measurement protocols that were not yet fully stabilized. Indeed, as we shall see, instead of grounding an undisputable ‘truth’, the TESLA endeavor ended up establishing a contestable reference, the biology of neoantigens and how to measure their immunogenicity having slightly evolved alongside this four-year project. However, despite its controversial results, the magnitude of the technoscientific efforts that the TESLA project set into motion and the need it ultimately succeeded in filling in the scientific and industrial community counterbalanced its metrological uncertainties, thus instituting its representation of ‘true’ neoantigens within the field of personalized cancer immunotherapy (at least temporarily). More generally, this case study indicates that the enforcement of ground truths, and what it leaves out, is a necessary condition to enable AI technologies in personalized medicine, even while they may stifle discordant realities.
Neoantigens for personalized cancer immunotherapy
The recent history of cancer research has been marked by a general reconsideration of the role of the adaptive immune system 3 in cancer development (Löwy, 1996; Pradeu, 2019). Far from being strictly misled by cancer development, the adoptive immune system and its thymic-derived cells (T-cells) have appeared increasingly capable of triggering anti-tumor responses (Graber et al., 2023). And among the many entities (standardized scientific equipment, attested cells, authorized drugs) that have contributed to the resurgence of what is now confidently called ‘cancer immunotherapy’ (Esfahani et al., 2020), there are the so-called neoantigens.
Neoantigens, whose existence has been assumed since 1965, 4 derive from somatic mutations (i.e. genomic variations of a somatic cell) that are now considered one of the main hallmarks of cancer (Hanahan & Weinberg, 2011). In broad strokes, tumor-specific mutations produce novel protein sequences from which derive unprecedented (neo) fragments of proteins (antigens) capable of being displayed on the surface of tumor cells via a molecular arrangement called major histocompatibility complex (MHC). 5 Importantly, neoantigens are deemed to be absent from the ‘normal’ (i.e. non-cancerous) human genome, which make them specific to the tumor and the patient. Hence the immense clinical interest in somehow making endogenous (i.e. deriving from the same body) T-cells capable of recognizing these tumor- and patient-specific molecules since they can, and do (Tran et al., 2017), trigger immune responses confined to the cancerous tissues (since neoantigens are not produced, nor presented, by any cells other than tumor cells). And as of 2014, it was these unique characteristics of neoantigens that fueled the promise of a personalized cancer immunotherapy (Heemskerk et al., 2013, p. 201). 6
In terms of how to make patients’ T cells capable of recognizing neoantigens presented on the surface of cancer cells via the MHC (see Figure 3), two main methods have been developed since the launch of this line of research, that has come to be known as personalized adoptive cell therapy (Rosenberg & Restifo, 2015). The first one involves sampling T-cells from the patient’s blood and then genetically modifying certain properties of their receptors (TCRs) to make them sensitive to a putative neoantigen presented on the tumor surface via the MHC (Yamamoto et al., 2019). This method, called chimeric antigen receptor (CAR) T-cell therapy was developed between the 1990s and 2000s (i.e. before the advent of neoantigens in cancer immunotherapy research) in order to target antigens that are present in abundance on certain types of solid cancer (but also on other types of healthy cells, hence major problems of immune reactions). Although they produce promising results on solid tumors and are the object of important investments in corporate research and development, 7 neoantigen-specific CAR T-cell therapies are, at the time of writing, still experimental and thus restricted to clinical trials (Wang & Cao, 2020).
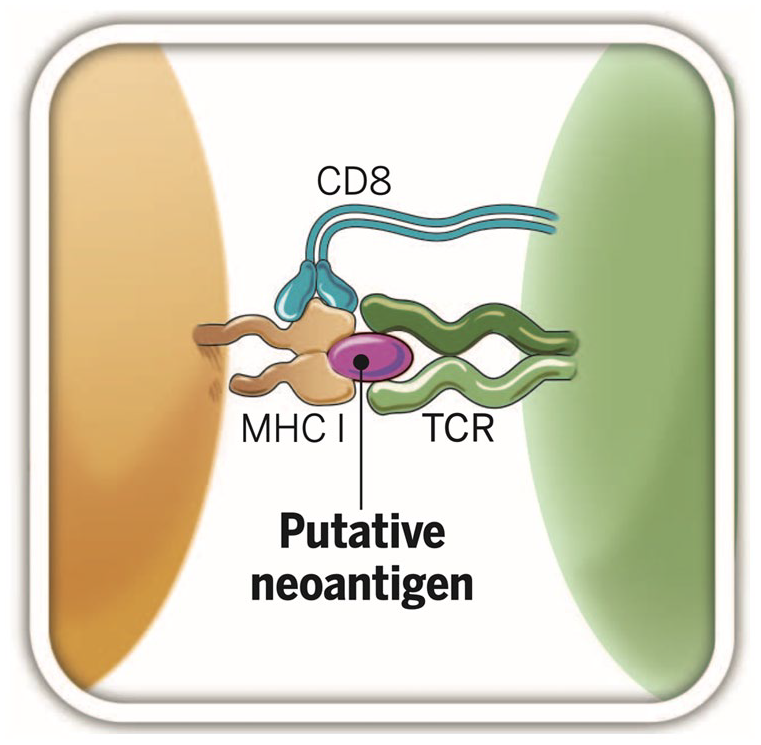
Schematic of a putative neoantigen. The mass on the far left of the figure represents a solid cancer cell. The two growths emanating from it—named MHC I—refer to the major histocompatibility complex, a molecular arrangement whose many functions include displaying peptides derived from turnover proteins on the surface of the cancer cell. The mass on the far right of the figure represents a T-cell. The two outgrowths emanating from it, named TCR (symmetric to MHC I), refer to the T-cell receptor. In the center of the figure lies the putative neoantigen, a peptide that carries great hopes for personalized cancer immunotherapy.
The second type of personalized adoptive cell therapy, which is also in clinical trials at the time of writing, is based on the collection of the patient’s T-cells selected according to their probabilistic affinity for putative neoantigens, their clonal expansion in vitro, and their reinfusion into the patient (Bianchi et al, 2020). This process, sometimes called neoantigen-specific tumor-infiltrating lymphocytes (TILs) therapy, differs from CAR T-cell therapy in that it does not genetically modify TCRs, thus allowing more putative neoantigens to be targeted (Bianchi et al., 2020). As it stands, however, this process requires resetting the patient’s immune system, which involves heavily equipped preparation steps (Cohen et al., 2017).
These two types of neoantigen-specific (and thus potentially personalized) adoptive cell therapies involve identifying neoantigens that can be recognized by T-cells and elicit an effective immune response (i.e. immunogenic neoantigens). This is a condition of possibility of these two experimental treatments 8 : Without the possibility of effectively identifying patients’ immunogenic tumor-specific neoantigens, efforts in terms of neoantigen-centered CAR T-cell or TILs therapies can only be in vain. And achieving this identification is all the more difficult as neoantigens can only be captured indirectly, through probabilistic inferences from comparisons between patients’ normal and tumoral DNA and RNA sequences. It is nearly impossible to detect neoantigens ‘physically’, as the wet and bulky surface of tumor cells are strewn with hundreds of thousands of other very different peptides that have no attraction potential towards T cells. Moreover, since immunogenic neoantigens derive from somatic mutations, they must, in theory, be patient-specific, which prevents the formation of a cumulative taxonomy, as it may be the case for self-antigens. In short, in order to be able to identify immunogenic neoantigens, and to try to realize the promise of personalized cancer immunotherapy, it is necessary to go through algorithmic processes that will compute genomic data to make predictions whose content—that is, existence and biological characteristics of immunogenic neoantigens—can then operate as the targets of personalized CAR T-cell or TILs therapies.
This technoscientific heaviness is certainly one of the current limitations of research on personalized cancer immunotherapy: If each patient produces their own neoantigens, it becomes necessary to profile the cancer transcriptome, sequence the cancer RNA, DNA, and matched normal DNA before proceeding to algorithmic analyses in order to identify abnormal protein-coding regions that could be at the origin of the presence of the immunogenic neoantigens on the surface of the cancer cells (more on this later). And all this only to get the targets for personalized CAR T-cell or TILs experimental treatments, which themselves entail many more convoluted operations (Graber, 2023): quite a heavy apparatus indeed.
But it is also important to note that the technoscientific heaviness of neoantigens is at the same time one of the reasons of their return to the forefront of cancer immunotherapy research. Indeed, as Schumacher and Schreiber (2015, p. 69) argue, without the growing availability (in rich countries) of infrastructures for relatively rapid and affordable high-throughput DNA and RNA sequencing technologies, as well as for big data analysis, the very idea of a cancer immunotherapy based on individual genomic features may not have been developed, let alone supported. In this sense, neoantigens, and the question of their identification, are also the inheritors of the entangled processes related to the deployment of next generation sequencing (NGS) and are imprinted, in their current deployments, of the deep reorganizations of biomedicine that recently took place around sequencing and data analysis technologies.
Reassembled through genomics
Determining the linear order of the components of macromolecules—that is, sequencing—is a process that involves many different actors and devices. This was already the case in the early 1980s when what is now called ‘first generation sequencing’ mobilized, among other things, DNA polymerization, fluorescently-labeled nucleotides, electrophoresis processes (Heather & Chain, 2016, pp. 1–3) but also, sometimes, non-trivial computational models stemming from applied physics and developed by researchers with a hybrid status having access to computing infrastructures of operations research centers (Stevens, 2013, pp. 22–32). But this sequencing-related community of actants (Latour, 2005) has grown dramatically since the 2000s and the advent of so-called ‘post-genomic’ disciplines whose NGS technologies started to involve, among many other things, biopsies, amplification processes, bioinformaticians, digital data formats (e.g. BCL and FASTQ), distributed computing, reference databases, visualization software (e.g. IVG) and countless norms, best practices and gold standards to make, sometimes, genomic big data actionable (Nelson et al., 2013) for diagnosis, prognosis or therapeutics (Hà & Chow-White, 2021). The acronym NGS refers to something far larger than a set of technologies: It refers to a sociotechnical process that comprises concrete locations and instruments, but also people and ways of doing and thinking that mutually constitute each other and, by this fact, participate in the constitution of sequencing-derived biomedical knowledge (Cambrosio, Keating, et al., 2018).
NGS processes, and the social relationships they require and produce, had a significant impact on contemporary biomedicine, particularly in relation to the increasing specialization of its various sub-disciplines and their (assumed) relative distance from clinical practice. Geneticist, oncologist, immunologist, data analyst, bioinformatician, cloud computing expert: The myriad of professions involved in NGS processes had, and have, their own learned societies, academic journals, and even departments that deliver specific PhDs, which progressively became compulsory for researchers wishing to pursue a career and obtain a stable position. And if this specialization dynamic effectively supported major advances in fundamental research, especially in oncology, it has also been criticized, notably for its inability to produce effective differences in routine clinical care (Sung et al., 2003).
From the 2010s onwards, this assumed pitfall of ultra-specialized post-genomic biomedicine prompted major funding agencies (e.g. NIH and NHS) to find out ways to ‘fill in the gap’ between research and the clinic (Butler, 2008). As a result of these institutional reflections, whose precise history remains to be done, a series of measures have been implemented—among them a redefinition of the requirements for obtaining grants and a series of changes in the criteria for evaluating outputs (e.g. counting patents and collaborations with hospitals and firms)—under the umbrella term ‘translational research’ (Woolf, 2008).
There is much debate on whether the hundreds of millions of dollars, pounds, and euros invested since the 2010s in translational research aimed at bringing fundamental research closer to the clinic (‘bench to bedside’) have had positive or negative impacts on Western biomedicine, broadly considered (for a discussion on this debate, see Blümel, 2018). But it seems now established, at least among social students of biomedicine, that instead of leading to a clear translation of fundamental certified knowledge into routine clinical applications, the rearrangements composed under the banner of translational research have contributed to blurring the distinction between research and care, especially in oncology (Besle, 2018; Nelson et al., 2014). In a recent attempt to characterize this new mode of biomedical practice at the interface of basic research and the clinic, Cambrosio, Vignola-Gagné et al. (2018) have proposed the notion of ‘experimental care’: a configuration that brings together clinicians, basic researchers and bioinformaticians—from academia, hospitals, non-profit organizations and, sometimes, the industry—around trials, during which research and care are mutually constituted.
The rearrangements suggested by experimental care configurations, themselves induced by the instauration of translational research, suggest new opportunities, such as renewed connections between public networks and commercial entities, especially those proposing distributed cloud computing services, now mostly based in the Bay Area. 9 Indeed, uncovering correlations between genomic variations (i.e. textual mismatches) and health conditions (i.e. attested phenotypes) during clinical trials implies swiftly navigating trillions of bytes of text data, an operation made possible since the 2010s and the widespread distribution of dedicated algorithms (e.g. Hadoop and Cloudburst) that have in turn enabled the use of professional storage and computing services provided by specialized companies (Stevens, 2016). Biomedicine, in its translational research efforts adapted to NGS processes, has thus progressively constituted a new market, that of biomedical cloud computing infrastructures, which specialized companies and start-ups such as Cloudera and Spiral Genetics, but also industrial giants such as Amazon and Microsoft (via their cloud services), have rushed to enter and shape. This new ecosystem of infrastructure providers has also contributed to the emergence of hybrid non-profit organizations acting as facilitators between industry, regulatory agencies (e.g. U.S. Food and Drug Administration), and translational post-genomic research, one of the latest US-based example being the Parker Institute of Cancer Immunotherapy (PICI) that we will soon closely examine.
Let us recap. NGS processes have contributed to major transformations in biomedicine. The advent of translational research, itself induced by the hyper-specialization required to make sequencing and interpretation technologies operational, has contributed to blurring the distinction between research and clinical care, in turn suggesting experimental care configurations that themselves inherit previous local arrangements and pre-existing networks. This mosaic of relationships implies organizational and governance issues that themselves suggest closures, but also opportunities that may be seized by new actors, especially those close to cloud-based and distributed computing, an obligatory point of passage for any current sequencing operation (Graber et al., 2023). These synergies have, in turn, contributed to the emergence of hybrid entities—halfway between research and industry—operating as coordinators for the implementation of infrastructures aimed at facilitating the relationships between industry, research, and regulatory agencies. And it is in this context of a sort of Silicon Valley-based reappropriation of the Human Genome Project’s imaginaries that appeared PICI, the institution behind the neoantigen-related TESLA project that we are now about to follow.
The TESLA project challenge
The new ecosystem of post-genomic experimental care is looking West, especially at the Bay Area, where companies and capital engaged in the computing industry are clustered. And it is in this specific small part of the world, the epicenter of contemporary digital capitalism, that the TESLA project aiming to address the challenge of neoantigen prediction took shape, in the wake of the creation of PICI, itself emanating from the Parker Foundation built upon the fortune of charismatic Silicon Valley entrepreneur Sean Parker.
Parker may be best known as one of the founders of the peer-to-peer audio file sharing system Napster, which was shut down in 2002 due to lawsuits. But the short-lived success of Napster allowed Parker to raise funds to launch Plaxo, one of the very first social networking services. Although he was soon ousted from the company for reasons that are still unclear today, this experience brought him into early contact with Mark Zuckerberg and Eduardo Savarin, who appointed him president of their nascent company Facebook in 2004. And while Parker’s presidency came to an abrupt halt following a suspected drug possession, it still allowed him to own many Facebook shares, which turned out to be gold mines after Facebook’s IPO on NASDAQ in May 2012. Indeed, by selling his shares when they were at their highest, Parker became one of the youngest Silicon Valley billionaires.
Parker founded the Parker Foundation in 2015 for $600 million, based on the North American model of philanthropy (de Merced, 2015). 10 It is difficult to know the precise reasons, but it seems that stories of family illness along with recurrent meetings with Jeffrey Bluestone—a renowned expert in T-cells, based at the University of California, San Francisco—then prompted Parker to allocate $240 of these $600 millions to the creation of a non-profit research organization focused on cancer immunotherapy. This was PICI, founded in 2016. This non-profit organization was explicitly designed for the management of clinical trials and bioinformatics resources for six US-based partner institutions active in the field of cancer immunotherapy: the Memorial Sloan Kettering Cancer Center (MSKCC), the University of Pennsylvania (UP), the University of Texas M. D. Anderson Cancer Center (MDACC), Stanford Medicine (SM), and the University of California campuses in San Francisco (UCSF) and Los Angeles (UCLA; Pollack, 2016). 11 This positioning as both initiator and coordinator of cross-institutional projects makes PICI function as an ‘operational foundation’ (Anheier & Daly, 2005, p. 162), with fixed-term postdoctoral employees coordinating so-called ‘PICI-funded initiatives’ (see Figure 4). However, PICI can also be considered, at least to some extent, a ‘corporate foundation’ (Anheier & Daly, 2005, p. 162), as many of its board members are also board members of private companies and start-ups.
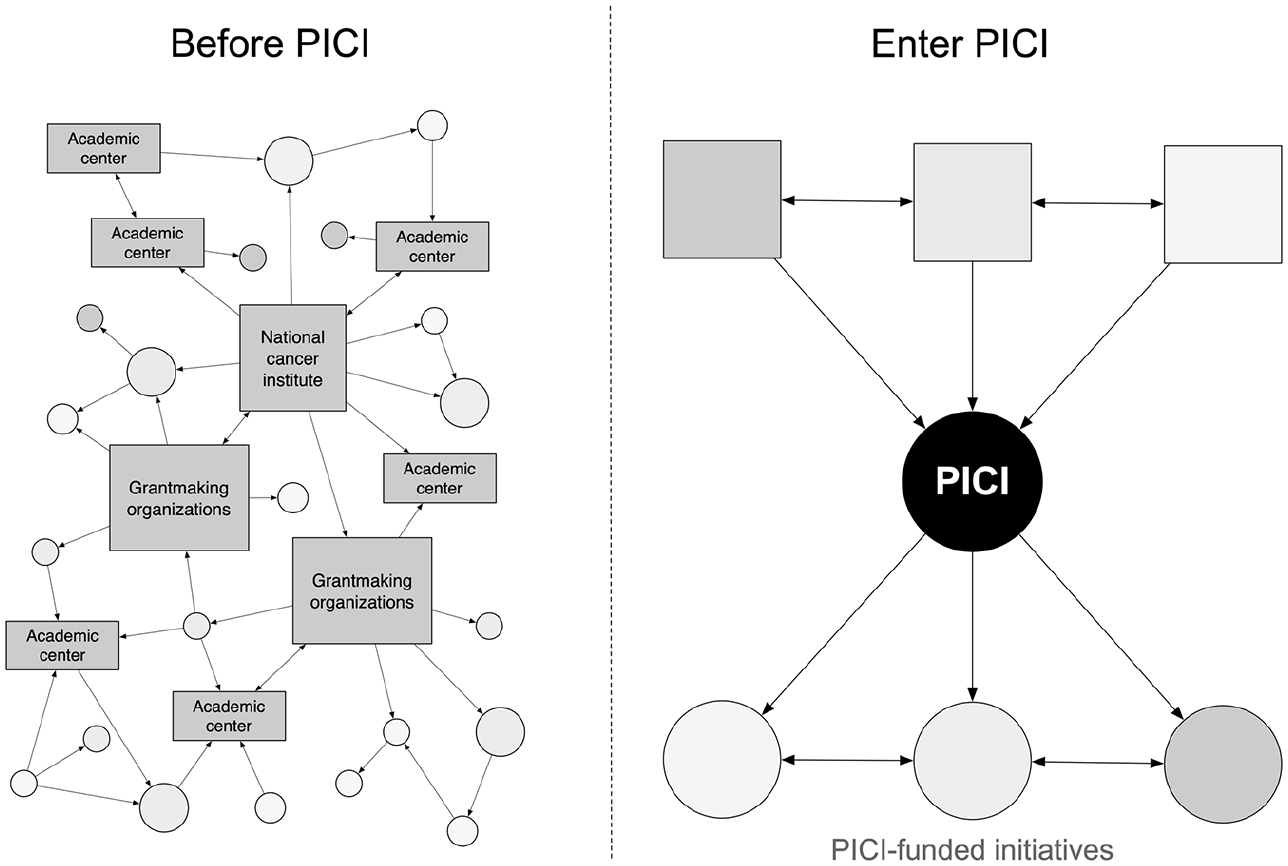
Schematics derived from PICI’s website representing the desired positioning of PICI within US-based cancer immunotherapy research. On the left, the situation as considered by PICI promoters before its creation: Numerous stakeholders (rectangles) have disordered interactions (arrows) that lead to many scattered projects (circles). On the right, the situation as hoped for by PICI promoters following its creation: Interactions are structured around PICI, which takes care of the coordination between the different stakeholders, notably via PICI-funded initiatives.
PICI’s explicit area of expertise (i.e. cancer immunotherapy), its numerous relationships with the computing industry (e.g. its location in the Bay Area, its being chaired by Silicon Valley billionaire Sean Parker) as well as, more generally, the prominent place of data analytics in post-genomic biomedicine (e.g. the necessity of navigating through trillions of bytes of text data using distributed infrastructures) soon prompted PICI to take on the issue of neoantigens—the then-most promising molecules for cancer immunotherapy—and frame it in bioinformatics terms. Specifically, due in part to the recruitment of a team of biostatistician postdocs at the launch of PICI in 2016, an opportunity for a PICI-funded initiative quickly emerged around the development of an infrastructure to support the development of neoantigen prediction algorithms. As one of my informants put it: At that time [2015-2016] many [research] teams were developing custom algorithms to predict good peptides [i.e., immunogenic neoantigens], some with fancy machine learning tools. But the problem was that you could only self-assess your model because there was no benchmark. (Informant 3, biostatistician)
In other words, the rush towards neoantigen-related experimental care went together with the development of various computerized methods of calculation—which today we would call AI algorithms—aiming to predict the existence of immunogenic neoantigens capable of triggering immune responses by means of made-greedy endogenous T cells. At the time of PICI’s launch in 2016, these algorithms—which rely on more or less arbitrary assumptions about the genomic derivation of neoantigens as well as their ecology within, and around, cancer cells—could not be compared with each other because the scientific community did not have any reference repository. In computational biology, and in applied computer science in general (Jaton, 2017), this is the well-known problem of self-assessing the superiority of one model over another by mobilizing custom references and metrics. In these problematic situations, the same informant continues: [T]he general rule is that your algorithm is better than the others because you know better how it works and so you can evaluate it accordingly. … [W]ithout a benchmark, you can’t get out of this problem, and you don’t know which kind of algorithm performs the best. (Informant 3, biostatistician)
At the launch of PICI, a first potential PICI-driven initiative could then try to answer the following questions: How can one shape a list of immunogenic neoantigens that could serve as a basis for training and evaluating prediction algorithms? How can one build a reference dataset that could serve as a benchmark for the development and evaluation of neoantigen prediction algorithms? These were genuine scientific questions because cancer immunotherapy research indeed lacked an authoritative ground truth to distinguish between good and less good algorithms for predicting neoantigens (algorithms that could in turn be mobilized in experimental care configurations). But they were also genuine strategic questions because it may also enable PICI to effectively make its way into an existing network of prestigious cancer-related research institutions (MSKCC, UP, etc.). And as many classical STS on metrology have shown, establishing oneself as a reference scale often equates to positioning oneself as an obligatory passage point (Callon, 1999; O’Connell, 1993; Mallard, 1998). From PICI’s perspective in 2016, then, without a benchmarked ground truth, it was impossible to get cancer immunotherapy breakthroughs (powered by prediction algorithms) out of the local networks that generated them. But with a benchmarked ground truth, preferably built through a PICI-funded initiative (and thus officially associated with PICI), the results of cancer immunotherapy research—at least for its bioinformatics side—could show their significance and robustness in other networks, for example industrial or regulatory.
Immunogenic neoantigens are very small, rare, NGS-dependent, patient- and tumor-specific, and therefore extremely complicated and expensive to detect and assess reliably. Still from PICI’s perspective, how could it shape this precious list of attested immunogenic neoantigens that could serve as a basis for the training and evaluation of algorithms for the research community? How could it generate attested ‘true’ neoantigens, robust enough to be accepted and used by bioinformaticians working in the (restricted) field of computational cancer immunotherapy? This is where the notion of ‘challenge’ comes in.
In applied computer science, a challenge is a competition most often organized by an independent organization (e.g. learned societies) that makes several teams compete on the same dataset. The computational results of each team are then evaluated against the ‘true’ answers, known only to the competition organizers. The team that comes closest to the ‘true’ answers—that is, the ground truth—wins the challenge, which leads to a ranking of the competing teams. The best performing computational methods, according to the metrics used by the challenge organizers, are then published online and made available to the rest of the scientific community. Bioinformatics, as a discipline, has been built around and through this type of competitions, which have long been integral part of conferences and symposia: In bioinformatics, there is a great history of competition between different research groups. For example, there was already the CASP [Critical Assessment of protein Structure Prediction] challenge, where the DeepMind [Google] people recently outclassed everyone. There was also the DREAM [Dialogue on Reverse-Engineering Assessment and Methods] challenge, financed by IBM and which was a bottom-up project where people from the engineering side [of research] tried to avoid the ‘my method is better than yours’ kind of self-assessment. And then DREAM evolved into Synapse, which was the next big challenge and then became also a platform. And TESLA came after all that, and built on challenges that were already there. (Informant 2, bioinformatician)
In 2016, the launch of one of the first PICI-funded initiatives, the Tumor nEoantigen SeLection Alliance (TESLA) project challenge, followed from an alignment of factors: the return of neoantigens to the forefront of cancer immunotherapy research; a problematic and widespread self-assessment habit in the field of neoantigen algorithmic prediction; a new non-profit organization eager to make its way within a network of already established, and renowned, partner institutions; and the history of bioinformatics shaped by challenges where research teams compete on the same dataset whose correct answers are known only by the challenge organizers. But the TESLA challenge was different from most of the other challenges in bioinformatics because it was primarily about producing some of the data materials needed to constitute a new ground truth. In other words, unlike most bioinformatics challenges that are ends in themselves, the TESLA project challenge was mainly a means to an end, in this case the creation of a ground truth to help predict neoantigens in a reliable way. This is what the same informant explains when he says: But here, the new thing is that they used the challenge to build the benchmark. You first had teams submitting results, then [the TESLA organizers] used these results to do the validation [i.e., the ground truth]. (Informant 2, bioinformatician)
Yet to enroll participants and form a consortium capable of collectively producing—via the ‘challenge’ form—a PICI-stamped ground truth for algorithmic neoantigen prediction, the TESLA organizers had to come up with a protocol that was both time-efficient and interesting for the participants, but also serious enough to result in a robust dataset that could be used by the broader scientific community. In an attempt to accomplish this delicate exercise, the TESLA organizers decided to proceed in five steps. First, they would make clinical grade genomic data from cancer patients available to participants on a secure server. Second, participants would download the data and mobilize their own custom algorithms to predict putative neoantigens (called ‘neoantigen candidates’). No later than six weeks after downloading the data, each team would be required to provide an Excel sheet describing the characteristics of their predicted neoantigen candidates. Third, based on the content of these Excel sheets, the challenge organizers would synthesize the candidates and make them pass a series of tests in order to validate or not their status of ‘true’ immunogenic neoantigens, that is, neoantigens capable of triggering effective immune responses. Fourth, based on the relative number of validated candidates, the organizers would announce the ranking of the different competing teams. Fifth, a collective article reviewing the challenge would be published and, in the wake of this, the ground truth will be made available online.
However, as audacious and innovative as it was, this protocol was nothing without an initial effective dataset: In order to enroll participants in the challenge, the TESLA organizers would have to start by having reliable and relevant data for neoantigen prediction in cancer immunotherapy. Fortunately—and this helped suggest the idea of a consortium-based project challenge in the first place—among PICI’s partner institutions were several university hospitals engaged in clinical research involving cancer patients who had given formal consent for the reuse of their genomic data. And it was from this pool of tissue samples—in this case, standard-prepared by MSKCC and UCLA lab technicians and then sequenced and translated into a workable format (FASTQ) by the bioinformatics teams at the same two institutions—that the TESLA challenge organizers were able to tap into. These institutional supports allowed the TESLA organizers to quickly assemble a dataset consisting of the normal and cancerous exon DNA and RNA sequences of nine patients with metastatic melanoma or lung cancer. 12 And by putting forward both the scientific interest of a challenge-based ground truth to support neoantigen prediction and also interesting clinical grade genomic data made available for free on a secure server (Wells et al., 2020, p. e3), the TESLA organizers progressively managed to enroll 28 computational biology teams. From there, once the consortium was formally established in December 2016, the TESLA project challenge could officially begin.
Scarcity of contested positives
In Fall 2017, the TESLA organizers made available on the Synapse platform the anonymized working data of the challenge, namely the FASTQ files of normal and tumor DNA and RNA sequences, along with clinical grade HLA typing information 13 of nine cancer patients. The 28 teams that had agreed to take part in the challenge were given six weeks to make their predictions of neoantigens using their own custom algorithms. 14 And the ranked list of these so-called neoantigen candidates, which took the form of Excel sheets describing the properties of the peptides, were then sent back to the challenge organizers.
Once all the ranked lists received, the problem that quickly arose for the organizers was the important difference in the number of neoantigen candidates proposed by the different teams: While some teams were limited—by virtue of the parameters of their algorithm—to the identification of 50 candidates, others went up to 100,000 candidates. Besides constituting an advantage for the teams that proposed more candidates, this variance also posed a problem for the validation step of the challenge. Indeed, given the many costly operations necessary to determine if a candidate is immunogenic—and therefore a true positive—it was impossible for the organizers to test all the proposed candidates.
To somehow reduce the number of candidates, the organizers decided to retain, as far as possible, the five best candidates from each team as well as all those that were ranked highest in the top 50 of all teams. Interestingly, this procedure did not please all the participants, some of whom complained that they had proposed candidates that were potentially immunogenic but not sufficiently present in the competing lists to be selected as tested candidates. Nevertheless, based on these two criteria—top five ranked and most recurrently ranked in the top 50—the organizers came up with a final to-be-validated list that included 608 neoantigen candidates.
To get validated neoantigens, the organizers had to set up convoluted and costly validation experiments. In a nutshell, building on several authoritative papers on antigen immunogenicity assessment (e.g. Kvistborg et al., 2012; Sidney et al., 2013), the TESLA organizers decided to use two validation experiments—HLA binding and flow cytometry—which consist, in very broad strokes, of testing the candidates’ ability to associate with purified MHCs and T-cells. Precisely describing the numerous steps and operations involved in these two validation experiments is beyond the scope of this article. 15 For our purposes, it is sufficient to mention that these experiments involved, for each of the 608 candidates, the creation of about ten samples co-incubated, under different modalities, with purified MHC and linked with specially primed T-cells. These numerous operations required a great deal of laboratory benchwork, but also ultra-specialized pieces of equipment (e.g. flow cytometers, microscintillation counters) to which even PICI partner institutions did not have access. To carry out their validation experiments on all the selected candidates, the TESLA organizers had therefore to include in the consortium two other research centers specialized in immunological assessment: the ImmunoMonitoring Lab at Washington University and the Netherland Cancer Institute in Amsterdam (Wells et al., 2020, p. e5).
Both HLA typing and flow cytometry validation experiments thus involved a lot of specialized equipment and practical operations done by trained personnel. It is little surprise, then, that they took time and money—about 6 months and between $5 and $10 million, according to my informants. Nevertheless, once this long, expensive, and tedious work was done, the TESLA organizers were able to affirm that they had thirty-seven immunogenic neoantigens. In other words, after double-testing the 608 candidates, it turned out that thirty-seven of them were ‘true’ neoantigens.
Thirty-seven true positives—or output-targets, according to the terminology sometimes used in applied computer science—may not seem a lot in view of the trillions of bytes of genomic text data that constituted the input-data of the ground truth. However, these thirty-seven immunogenic neoantigens have the important merit of existing and therefore making it possible to build the first benchmarked ground-truth dataset for neoantigen prediction, made available online in November 2020 after the publication of a collective article in the high-impact journal Cell presenting the TESLA project challenge and its implications for personalized cancer immunotherapy research (Wells et al., 2020). 16
But are the neoantigens described in the TESLA ground-truth dataset really immunogenic? After all, they do not directly derive from patient testing; biopsies of patients were brought to laboratories that then synthesized neoantigen candidates and conducted in vitro experiments to attest to their binding potential with reconstituted T cells and MHCs. Is this indirect methodology robust enough to attest to the veracity of the immunogenic neoantigens of the TESLA project? It certainly was in 2018, when standard validation tests were performed on the 608 candidates provided by the challenge participants. But what if, in parallel to the TESLA project challenge, basic research on neoantigens led to different conclusions with regards to immunogenicity testing? Would the veracity of the TESLA ground truth’s ‘true’ neoantigens be questioned? In sum, as the foundations of the TESLA ground-truth dataset rest on the fragile state of the art of basic immunology research, what would happen if the biology of neoantigens happened to change?
Interestingly, this question arose directly after the release of the TESLA ground truth. During this four-year symposium-based project—which, again, required great organizational (e.g. setting up a challenge, collecting candidate lists) and material resources (e.g. production of genomic data, validation experiments)—the biology of neoantigens as well as how to attest their immunogenicity has partially evolved, as indicated by the following excerpt explicitly referring to the TESLA project: However, neoepitope immunogenicity in this [TESLA] and other studies was validated by experimentally measuring reactivity of existing T cells in patient blood or tumor. Given that tumor-specific T cells in advanced tumors are dysfunctional, it is likely that functional readouts of these assays are limited by low sensitivity. Additionally, it is possible that therapeutic priming against neoantigens overlooked by these assays could unleash productive T-cell responses from naive T cells or reservoirs of clonally expanded precursors in lymphoid tissues. (Westcott et al., 2021, p. 1081)
According to the authors of this recent publication on neoantigen biology (but also others, such as Borden et al., 2022; Jaeger et al., 2022), there are robust indications that the standard in vitro methods used to attest to the immunogenicity of neoantigens are at odds with the complex in vivo interactions between neoantigens and potentially dysfunctional T-cells. In short, the results of testbed validation experiments no longer correspond entirely, in 2021, to what happens on the surface of solid cancers; there tends to be a mismatch between what happens routinely in heavily equipped laboratories and what happens in the bodies of cancer patients. And far from only concerning researchers on the fringe of bioinformatics cooperation networks, these results are now also recognized by computational biologists, as one of the contributors to the TESLA project acknowledges: I think people realized it for the last two years now that the whole story is more challenging that what we thought at first. I think that now [in early 2021], the biology of neoantigens is much more complicated. And I think that people understand that the whole story is more challenging that what we thought. They understand that even if we assess something as immunogenic, it’s not necessarily a clinically effective target. So there are new features here and we will need more specific validation to train predictors to deconvolute those [neoantigens] that are immunogenic and those that are clinically effective. (Informant 11, bioinformatician)
But does this increased complexity of in vivo neoantigen biology, which calls into question the usual in vitro validation experiments, also call into question the relevance of the TESLA ground-truth dataset? Not completely, as it still is—at the time of writing—the most comprehensive source of information on the problem of algorithmic neoantigen prediction. As the same protagonist of the challenge continues: So overall now with this [TESLA] data, it’s very useful for us, even if it has limits. And we’re using it for benchmarking our pipelines and so on and so forth. You have to recognize the quality, the challenge of collecting all these predictions, validating them, testing the peptides, etc. So it’s still an important benchmark. (Informant 11, bioinformatician).
The room to maneuver for the algorithmic prediction of neoantigens is quite small. Indeed, to include new elements regarding the biology of neoantigens and the way to attest to their immunogenicity would imply starting over the whole time-consuming and expensive validation process. And in any case, as it stands, there is no standardized assay capable of taking into account the new issues raised by basic immunology research. In sum, for the TESLA project, and also for the many public and private research groups that mobilize this ground truth to train and evaluate their predictive algorithms, the medium must be the message. The TESLA project has done everything it could to produce an acceptable ground truth: The initial working data are clinical grade, the selection of candidates is justified by relevant arguments (notably financial ones), and the validation tests are based on well-established state-of-the-art procedures. It is not doable, as it stands, to integrate recent discoveries on the biology of neoantigens: The TESLA ground truth, because of the massive infrastructure that allowed it to come into existence, operates as the foundation of what immunogenic neoantigens actually are for research in computational cancer immunotherapy.
At this point, it is important to note that disregarding the results of basic research on neoantigen immunogenicity is a practical imperative: Computational biologists need large datasets to train their algorithms, evaluate their performances, and value their results, especially to the public and private institutions that fund them. But this imperative, which is a condition for the development of AI algorithms aiming at the personalization of cancer immunotherapy, has a cost: The algorithms that emerge from infrastructurally massive benchmarks such as the TESLA ground truth can only reproduce, and thus promote, the specific version of the immunogenic neoantigens inscribed within these datasets.
The TESLA project was thus an act of science, but also an act of faith and power. The heaviness of its infrastructure, and the applications that its ground truth enables, tend to stifle the other reality of immunogenic neoantigens that is emerging in fundamental immunology. In short, because of the stabilized references algorithmic entities require in order to come into existence and circulate, applied bioinformatics performs a specific reality: In computational cancer immunotherapy—in academia, in the industry, and in the numerous start-ups that invest in this promising field of AI-enabled cancer immunotherapy—the immunogenic neoantigens are those represented in the TESLA ground-truth dataset.
Discussion and conclusion
As a conclusion, and at the end of this foray into personalized cancer immunotherapy in the making, I would like to discuss two provisional claims.
The first one has to do with the centrality of ground-truth datasets in algorithmic design. As it has already been said elsewhere (e.g. Mackenzie, 2017), algorithms—or rather, those who build them—must approximate the functions that supposedly organize the relationships between input-data and output-targets of ground-truth datasets. In this sense, for both AI and non-AI algorithms, ground-truth datasets operate as initial matrices and unsurpassable horizons: As the algorithmic form requires a concrete initial reference, every algorithm emanates from at least one specific ground-truth dataset that cannot be transcended (Jaton, 2021b).
Yet just like the algorithms they engender and confine, ground-truth datasets do not pre-exist: they derive from collective construction processes. These ground-truthing processes engage people, efforts, and resources. Yet, in principle, the products of these processes (i.e. ground-truth datasets) remain limited, arbitrary, and socio-culturally oriented. Consequently, algorithms—as devices that approximate relationships among ground-truth datasets—are also limited, arbitrary and socio-culturally oriented.
But this basic assertion leads to questions that remain largely unanswered today. For example, do disciplinary contexts affect the deployment of ground-truthing processes? And if so, how are these differences expressed? And how do they affect algorithmic design, as well as the effects these algorithms, once constituted, may produce? The case of the TESLA project indicates that the construction of a biomedical ground-truth dataset cannot go beyond the metrological tools available. As we have seen, in order to become reliable and publishable (and therefore mobilizable by others), the TESLA ground truth had to be based on clinical grade sequencing and immunogenicity validation procedures integrated into state-of-the-art pipelines. A biomedical ground-truth dataset like the one emanating from the TESLA project is thus as strong, but also as fragile, as the measurement devices that allow it to be constructed. In short, if we get the algorithms of our ground truths (Jaton, 2017), we get the ground truths of our organizations and metrological equipment. If the measurement rules happen to change, or are the object of controversies, it is the whole ground-truth dataset that oscillates.
However, as we have seen, despite a series of doubts about the protocols for measuring the immunogenicity of neoantigen candidates, the TESLA ground-truth dataset continues to be used and praised. This is because it is currently the only large ground truth available: without the TESLA ground truth, there is no way to build comparable neoantigen prediction algorithms. But the continuous use of the TESLA ground truth is also linked to the quality of its constituent relationships, the project’s coordinators having endeavored to stick to the then-most recent standards throughout the whole ground-truthing process. In this sense, the case of the TESLA challenge seems to indicate the existence of a more or less intuitive threshold from which a ground-truth dataset, even if potentially problematic, is robust enough for the practitioners who use it. In the case of the TESLA project, many precautions have been taken and important means have been mobilized. It is rather unfortunate that new fundamental findings in immunology soon jeopardized part of the ground truth’s content, but the dataset remains sufficiently equipped to support valuable, or at least promising, propositions. In sum, even though the TESLA ground-truth dataset comes up against the current limitations of state-of-the-art immunogenicity assessment protocols, it still manages to reach the quality standards of these protocols, thus justifying its continued use for academic and industrial purposes. 17
My second claim deals with what Lee (2022) recently coined ‘ontological overflow’, namely the politics of enacting objects. As we saw in the present case study, one of the arguments in favor of the TESLA ground-truth dataset is that it is for the moment the most robust. The doability (Fujimura, 1987) it generates—immunogenic neoantigens being able to be predicted in experimental care configurations by means of commensurable algorithms—is considered both precious and unfinished. The actual use of the TESLA ground-truth dataset is thus justified by ground-truthing processes to come, which will be closer to recent advances in basic immunology research. But for the case of neoantigen prediction, how long will it take to build a new ground-truth dataset that will incorporate the latest findings on immunogenicity assessment? And in the meantime, where will have circulated the algorithms trained and evaluated against the TESLA ground truth as well as their predicted neoantigens? And perhaps even more problematically, could the existence of the limited-yet-functional TESLA ground truth not also encourage to play down discordant results in order to keep the relevance of this ground truth? After all, it is the only existing dataset that can benchmark neoantigen prediction algorithms, which are themselves key elements of clinical trials that may lead to approvals of new neoantigen-specific therapies from regulatory agencies. Without the TESLA ground truth, personalized cancer immunotherapy loses a key element, and hence the importance to actively maintain its relevance and ‘construct a particular blindness’ (Lee, 2022).
The purpose here is not to criticize AI-enabled personalized medicine as such. AI technologies do seem to be able to help patients with terrible diseases, which is wonderfully promising. But it is also important to keep in mind the potentially detrimental effects of the ground truths subtending biomedical AI algorithms, notably their readiness to stifle discordant realities by virtue of the applications they permit, as well as the more or less strategic justifications their usability may suggest. Enforcing limited and socio-culturally oriented ground-truth datasets is a necessary condition for an AI-enabled personalized medicine. In this sense, it is certainly important to remain attentive to the inertia generated by these often-massive biomedical ground truths as well as to attempts to over-advocate their use.
Footnotes
Acknowledgements
I am immensely grateful to the bioinformaticians who accepted to answer my questions and more generally helped me with my research. I also want to express my deepest gratitude to Nolwenn Bühler, Luca Chiapperino, Nils Graber, Francesco Panese, Loïc Riom, Philippe Sormani, Mylène Tanferri Machado, Dominique Vinck, Isabelle Zinn, the anonymous reviewers, and the editors of this journal for their support, help, and insightful suggestions.
Funding
The author disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Schweizerischer Nationalfonds zur Förderung der Wissenschaftlichen Forschung, Sinergia 180350, ‘Development of Personalized Health in Switzerland: Social Sciences Perspectives’
