Abstract
Translational models have played an important role in the rapid development of safe and effective vaccines and therapeutic agents for the ongoing coronavirus disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Animal models recapitulating the clinical and underlying pathological manifestations of COVID-19 have been vital for identification and rational design of safe and effective vaccines and therapies. This manuscript provides an overview of commonly used COVID-19 animal models and the pathologic features of SARS-CoV-2 infection in these models in relation to their clinical presentation in humans. Also discussed are considerations for selecting appropriate animal models for infectious diseases such as COVID-19, the host determinants that can influence species-specific susceptibility to SARS-CoV-2, and the pathogenesis of COVID-19. Finally, the limitations of currently available COVID-19 animal models are highlighted.
The devastating impact of the COVID-19 pandemic has necessitated rapid development of various complementary translational model systems, such as
The acute phase of COVID-19 presents with respiratory symptoms as well as other non-respiratory manifestations most notably renal, cardiovascular, neurological, and gastrointestinal symptoms.30,37,56 Also, a growing body of evidence indicates a chronic syndrome with various long-lasting symptoms in SARS-CoV-2 infected patients (the post-COVID syndrome or long COVID). 46 Suboptimal type I and III interferon responses to the SARS-CoV-2 in individuals with severe COVID-19 suggest a dysregulated host immune response to infection. 40 Animal models that recapitulate various phenotypes of COVID-19, including host responses to SARS-CoV-2, will be vital for elucidating pathogenesis, identifying new therapeutic targets, and developing safe and effective vaccines and therapies.
In consideration of the appropriate animal model for SARS-CoV2, understanding of the natural infection and disease in various species are critical. Only in this way can it be determined whether the disease progression is homologous or not. We do not fully understand the origin of SARS-CoV-2 and the transmission pathway to humans. The most prevalent hypothesis is that SARS-CoV-2 originated in bats (as bat coronavirus BatCoV RaTG13 has 96% genome sequence identity with genome of SARS-CoV-2) and that interspecies transmission to an unknown intermediate host as a reservoir occurred prior to zoonotic transmission to humans in 2019.11,80 A range of intermediate hosts (including pangolin, mink, rabbits, raccoon, dogs, civets, ferret, badgers, snakes, and domesticated cats) are proposed where natural infections have been observed. Direct spillover of SARS-CoV-2 from bats to human (direct animal contact or indirectly through animal product or excreta) is also a possibility.11,80 Additional investigations are needed to fully elucidate the evolution of SARS-CoV-2 and transmission pathways.
Although several animal species are susceptible to natural infection, not all are suitable for modeling. Furthermore, natural infections may not identify all susceptible species. Therefore, several studies have evaluated the susceptibility of various animal species to infection with SARS-CoV-2 in experimental settings. Mouse, hamster, ferret, nonhuman primate (NHP), tree shrew, cat, and mink are susceptible to SARS-CoV-2 infection, whereas chicken, duck, turkey, pig, rabbit, cattle, and guinea pig are not receptive to SARS-CoV-2 infection.36,50,80 One explanation for the susceptibility difference involves the structural differences of the host receptor angiotensin-converting enzyme 2 (ACE2). Understanding various pro- and antiviral host factors (e.g., ACE2, host proteases) that influence susceptibility and drive susceptibility differences among species is key to developing suitable animal models for studying COVID-19. Additionally, factors extrinsic to host-pathogen interactions (e.g., historical control data, cost and availability, husbandry-related factors) are important determinants for selection or rejection of a particular animal model and will be elaborated below.
Although several species are susceptible to experimental infection, most studies of SARS-CoV-2 disease (studying viral pathogenesis and transmission) and efficacy (testing antiviral drugs and vaccines) have used NHP, hamster, mouse, and ferret models;19,22,29,36,50,57,64,68,78,82 each with its own advantages and limitations (Table 1) and a focus of this review. Detailed descriptions of each species/breeds or strains examined (wild type or genetically modified) with regard to specifics of the viral transmission, pathology, and immunology induced by SARS-CoV-2, is discussed elsewhere in this special issue and in several recent reviews.9,21,36,50 Current status of COVID-19 animal models and best practices for their use in COVID-19 research are provided by National Institutes of Health (NIH) (https://opendata.ncats.nih.gov/covid19/animal).
Advantages and limitations of commonly used COVID-19 animal models.

Abbreviations: Tg hACE2, transgenic Human angiotensin-converting enzyme 2; mACE2, mouse angiotensin-converting enzyme 2; BALB/c, Bagg Albino c; MA, Mouse-adapted; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; NHP, nonhuman primate.
Some SARS-CoV-2 variants of concern (Beta & Omicron) can bind mACE2.
Mortality is likely secondary to encephalitis or due to pulmonary disease and encephalitis both.
Single publication.
Infectious Disease Animal Models: Considerations and Requirements
Since infectious diseases like COVID-19 involve a myriad of complex host-pathogen interactions, careful consideration must be given during the selection of an animal species for disease modeling in order to generate relevant and translatable scientific data.7,72 Important considerations for selecting appropriate animal models for studying any infectious disease are listed in Box 1. Identifying animals which are susceptible/responsive to the infection is the first important requirement. For this, an animal should have necessary anatomic structures and biochemical pathways (e.g., viral receptors and other proviral factors) which facilitate viral entry, replication, and disease pathogenesis (detail in the section below). The ability to recapitulate critical aspects of human disease manifestations (data extrapolation) is another important attribute of a good animal model. For example, hamsters, which exhibit lower fidelity to humans than monkeys in terms of anatomy, are a better model to study COVID-19 respiratory disease as they closely mimic the severe disease observed in terminally ill patients (discussed later in the manuscript). Since SARS-CoV-2 is a Risk group 3 pathogen and must be handled in biosafety level 3 (BSL-3) facilities, 39 factors such as experience/training of animal care and research staff and cost become even more important, particularly with novel pathogens. Limited institutions around the world have these laboratories and in addition to construction, equipment, and training costs, running and maintenance of a BSL3 laboratory are expensive and complex. Because of higher costs, complex requirements, and time it takes to build a new BSL-3 facility, a lower-cost, BSL-2 murine model has been developed recently to study COVID-19 which utilizes a Risk group 2 pathogen murine hepatitis coronavirus 3. Although not perfect, such surrogate models provide a good insight into the viral biology of SARS-CoV-2. 41
A good literature base and/or historical usage of animal models are also important when modeling novel diseases. For example, many currently used SARS-CoV-2 models have expanded upon animal model data accumulated from studying previous coronavirus diseases including SARS in 2002 to 2003 and Middle East Respiratory Syndrome (MERS) in 2012.23,50,59,70 Finally, it is also preferable if these models have clinical, microscopic, or macroscopic endpoints that can be correlated with virus titers.
Species-Specific Susceptibility to SARS-CoV-2: Role of ACE2 and Other Host Determinants
A first, and probably most important, criterion for selecting experimental animals for modeling a viral disease is animal susceptibility to the virus, allowing virus infection and replication which contributes to disease. Since SARS-CoV-2 entry into host cells is mediated by the interaction between the viral receptor binding domain (RBD) on its spike glycoprotein (S) and its receptor ACE2 and subsequent membrane fusion, ACE2 expression on the cell surface is required for viral entry. In addition, the efficiency with which the virus binds to ACE2 is a key determinant of transmissibility and host susceptibility. 35 Human ACE2 (hACE2) has the highest affinity to bind viral RBD and hence humans are highly susceptible to SARS-CoV-2 infection. 62 Higher affinity binding of SARS-CoV-2 RBD than that of SARS-CoV S RBD to hACE2 partially explains the increased transmissibility of SARS-CoV-2. 35 In fact, increased transmissibility of the Omicron (B.1.1.529), a SARS-CoV-2 variant, may be attributed to higher binding affinity of Omicron RBD to hACE2 than Wuhan-Hu-1 RBD. 17 Therefore, compatibility of ACE2 to RBD on S has been a major determinant in selecting susceptible experimental animals for COVID-19 modeling.34,62 Structural variation in ACE2 of different species likely determines RBD binding and thereby susceptibility to infection. 62 Hamsters, NHPs, and ferrets are considered good models as ACE2 of these species have high RBD binding and hence these animals are permissive to SARS-CoV-2 infection. Although NHP ACE2 has high sequence similarity to hACE2, binding affinity to RBD is lower than hACE2 as revealed by structural analysis. 62 This may explain in part why NHPs don’t develop severe disease despite being permissive to SARS-CoV-2 infection. On the other hand, the reason that pigs, dogs, ducks, rats, and chickens are minimally affected with COVID-19 is because ACE2 in these species is less compatible with RBD binding. Mice, which are the most utilized animal model for studying human disease, are resistant to SARS-CoV2 infection as mouse ACE2 (mACE2) has very low binding affinity with RBD. Therefore, use of mouse models for COVID-19 has been limited to genetically modified mice expressing hACE2 (transgenic mice containing permanent genetic modification to express hACE2 or transduced mice containing adenovirus or adeno-associated virus that encodes for hACE2) or wild-type mice infected with mouse-adapted (MA) SARS-CoV-2 (by sequential passing of virus in mouse lung tissue or by reverse genetics to modify the RBD of the virus) that effectively bind mACE2. 50 In transgenic or transduced hACE2 mice, the hACE2 is ectopically expressed and that may change the tissue or cellular tropism of SARS-CoV-2 and the innate and acquired immune responses. Expression of hACE2 in transgenic (Tg) mouse models is controlled by promoters (e.g., the human cytokeratin [K18] epithelial cell promoter in K18-hACE2 mice) and disease severity in these Tg mouse models correlates with hACE2 expression level. 50 Infection with MA SARS-CoV-2 infection in wild-type mice produced variable disease phenotypes; while SARS-CoV-2 MA10 (ten serial passage of SARS-CoV-2) infection of BALB/c mice produced a lethal mouse model that exhibited weight loss and recapitulated age-related disease severity observed in humans, SARS-CoV-2 MA10 infection of C57BL/6J mice or SARS-CoV-2 MA (virus adapted by reverse genetics) infection of young BALB/c mice produced only very mild disease.44,50 In 2021, several SARS-CoV-variants of concern, including Omicron, acquired ability to infect wild-type mice through increased binding affinity of RBD to mACE2 by naturally acquired mutations. 17 This suggests that wild type mice can be a useful model to study SARS-CoV-2 variants capable of binding to mACE2. However, increased transmissibility hasn’t always led to a severe disease phenotype in wild type mice. While Beta (B.1.351) produced severe clinical COVID-19 disease in wild type mice, 58 recently identified Omicron (B.1.1.529) produced an attenuated disease. 31 Additional studies will be needed to establish wild type mice as a model for SARS-CoV-2 variants.
In addition to facilitating viral entry, ACE2 plays an important role in the pathophysiology of COVID-19 via deregulating the renin–angiotensin–aldosterone (RAS) axis.5,35 ACE2 is an integral component of the RAS system and its primary role in normal physiology is to promote the formation of angiotensin (Ang) 1-9 from Ang I. In contrast to Ang I, which induces vasoconstriction and promotes fibrosis, oxidative stress, apoptosis, and inflammation, Ang-1-9 has vasodilatory, antifibrotic, anti-apoptotic and anti-inflammatory effects. Coronavirus infection results in downregulation of ACE2, either by endocytosis or by proteolytic cleavage and processing (e.g., transmembrane protease, serine 2 [TMPRSS2]-mediated), diminishing the anti-inflammatory and anti-fibrotic arm of the RAS system and resulting in increased inflammation, fibrosis, and tissue injury.5,35 Accordingly, downregulation of ACE2 is considered one potential mechanism for contributing COVID-19 pathology and fibrosis (details on other mechanisms in section below). Although young children have increased ACE2 expression in respiratory tissue and hence are at increased or at least similar risk of infection to adults, 55 they appear protected from severe lung disease caused by SARS-CoV-2 infection, suggesting a possible protective effect of ACE2. Increased expression of ACE2 was observed in the lungs of young rats, 84 and young NHPs, hamsters, and mice developed less severe pneumonia after SARS-CoV-2 infection, suggesting a possible protective effect of ACE2 in animals as for humans.29,76,87,90
Although important, ACE2 is not the only host determinant of virus entry. Following ACE2 binding, activation of S is required to mediate membrane fusion for viral entry into the host cell. Furin and furin-like proteases (e.g., TMPRSS2 and cathepsins), which participate in proteolytic cleavage or activation of S at S1/2 and S2’ sites, respectively, are important host determinants. TMPRSS2-mediated S activation at S2’ (which is most common) occurs at the plasma membrane, whereas cathepsin-mediated activation at S2’ occurs in the endolysosome. 35 In addition to S, TMPRSS2 also cleaves ACE2 which causes downregulation of ACE2 following infection. 32 Since species-specific differences in the structure of ACE2 affects SARS-CoV-2 susceptibility, it is plausible to assume that furin and TMRSS2, which facilitate ACE2-mediated viral entry, also contribute to susceptibility differences. However, it appears that furin and TMRSS2 are structurally almost similar in most species and hence do not drive susceptibility differences.12,62 In addition to the ACE2 structure, the differential distribution of ACE2 drives differences in tissue and host susceptibility. In humans, SARS-CoV-2 invades and causes injury in organs expressing higher ACE2 such as lungs, heart, gastrointestinal tract, and kidneys.20,52,55 While some studies have shown little or no ACE2 expression in human lungs others have shown adequate protein expression.20,55 Similar to humans, differential tissue distribution of ACE2 was observed among animals which may contribute to variability of host susceptibility to SARS-CoV-2. While some detected higher expression of ACE2 in hamsters and ferrets lungs,74,85,89 which is downregulated in response to SARS-CoV-2 infection, others failed to detect any expression suggesting a role of another cognate host receptor in virus entry.42,71 Although, several other molecules, in addition to ACE2, have been suggested as alternate receptors for SARS-CoV-2, such as neuropilin 1 and neutral amino acid transporter, B0AT1, additional studies are needed to confirm their role in SARS-CoV-2 infection and if they play a role in species-specific disease susceptibility. 35
Apart from host factors that facilitate virus entry, other proviral host factors facilitate SARS-CoV-2 infection by supporting either vesicle transport, lipid membrane composition/lipid homeostasis, or viral translation (Box 2). 6 Composition of cholesterol and other lipids in membranes play various roles in SARS-CoV-2 infection as all steps of the viral life cycle are membrane-associated. On the other hand, host restriction factors or antiviral effectors have been identified which restrict SARS-CoV-2 infection by inhibiting viral entry, replication, or impacting viral translation and lipid membrane composition (Box 2). 47 Most of the restriction factors are products of interferon stimulated genes (ISGs) which determine the interferon-related response to SARS-CoV-2 infection. While substantial work has been carried out to identify structural differences in ACE2 and furin/TMRSS2, which govern species specific susceptibility differences, much less is known about other pro- or anti-viral determinants. Understanding how variation among these other host determinants influence host-virus interaction in different species will enhance our ability to study and cure this viral disease in humans. Species-specific inhibition of host restriction factors has been shown in poxvirus and herpes virus infections, 66 but has not been demonstrated yet with SARS-CoV-2. Therefore, it is reasonable to assume that many pro- and anti-SARS-CoV-2 host determinants, in addition to ACE2, have species-specific interaction profiles with viral proteins that can influence the host range and disease severity. Differential interactions of pro- or anti-viral determinants (other than ACE2 and TMRSS2) with SARS-CoV-2 proteins may explain why hamsters, whose ACE2 has almost similar binding affinity with S as NHPs ACE2, develop a more severe respiratory disease than NHPs. Omicron, which has higher transmissibility due to higher affinity with hACE2, appears to produce less severe pulmonary disease in humans and animal models;31,88 it can be assumed that unfavorable interactions with other host determinants in the lungs may in part be responsible for a reduced disease severity.
COVID-19 Pathophysiology and Disease Phenotypes in Animal Models
In humans, direct viral toxicity, dysregulation of the RAS, endothelial cell injury and activation of coagulation pathways, and dysregulation of the immune response have all been proposed as key mechanisms in COVID-19 pathophysiology that lead to multi-organ failure.30,40 ACE2-mediated pathways play an important role in COVID-19 pathogenesis either by virtue of ACE2-mediated entry of SARS-CoV-2 (direct viral-associated tissue damage including endothelium injury and subsequent activation of coagulation pathways) or secondary to downregulation of ACE2-related to viral entry (dysregulated RAS leading to inflammation and tissue injury as previously described). 30 Accordingly, organs/tissues with COVID-19-related damage or effects generally have high levels of ACE2 and furin-like proteases. In addition to viral infection, host factors such as a dysregulated immune system (impaired interferon responses or hyperactivated complement system resulting in an exacerbated inflammatory response), age, and co-morbidities (obesity, cardiovascular disease) also contribute to disease severity in critically ill patients.1,30 Several studies implicate complement activation in the pathogenesis of coronavirus disease and see potential therapeutic benefit of complement inhibition.1,28 Host complement systems can be activated by direct SARS-CoV-2 (through the lectin pathway, the classical pathway and/or the alternative pathway) or by SARS-CoV-2-mediated endothelial injury or thromboinflammation. To date, animal studies addressing the role of complement activation in coronavirus pathogenesis are limited and restricted to SARS-CoV infection. Mice deficient in C-3 exhibited significantly less weight loss and airway lesions compared to wild-type mice in response to SARS-CoV infection. In addition, factor B−/− or C4−/− mice also exhibited less weight loss than wild-type mice. 28 Complement system genes (e.g., lectin pathway component MASP1 and ficolin 1) were significantly upregulated in ferrets following primary infection with SARS-CoV. 16
Common COVID-19 pathological manifestations in humans and those that have been commonly recapitulated in COVID-19 animal models are listed in Table 2.
Commonly reported pathologic manifestation of COVID-19 in human and animal models.
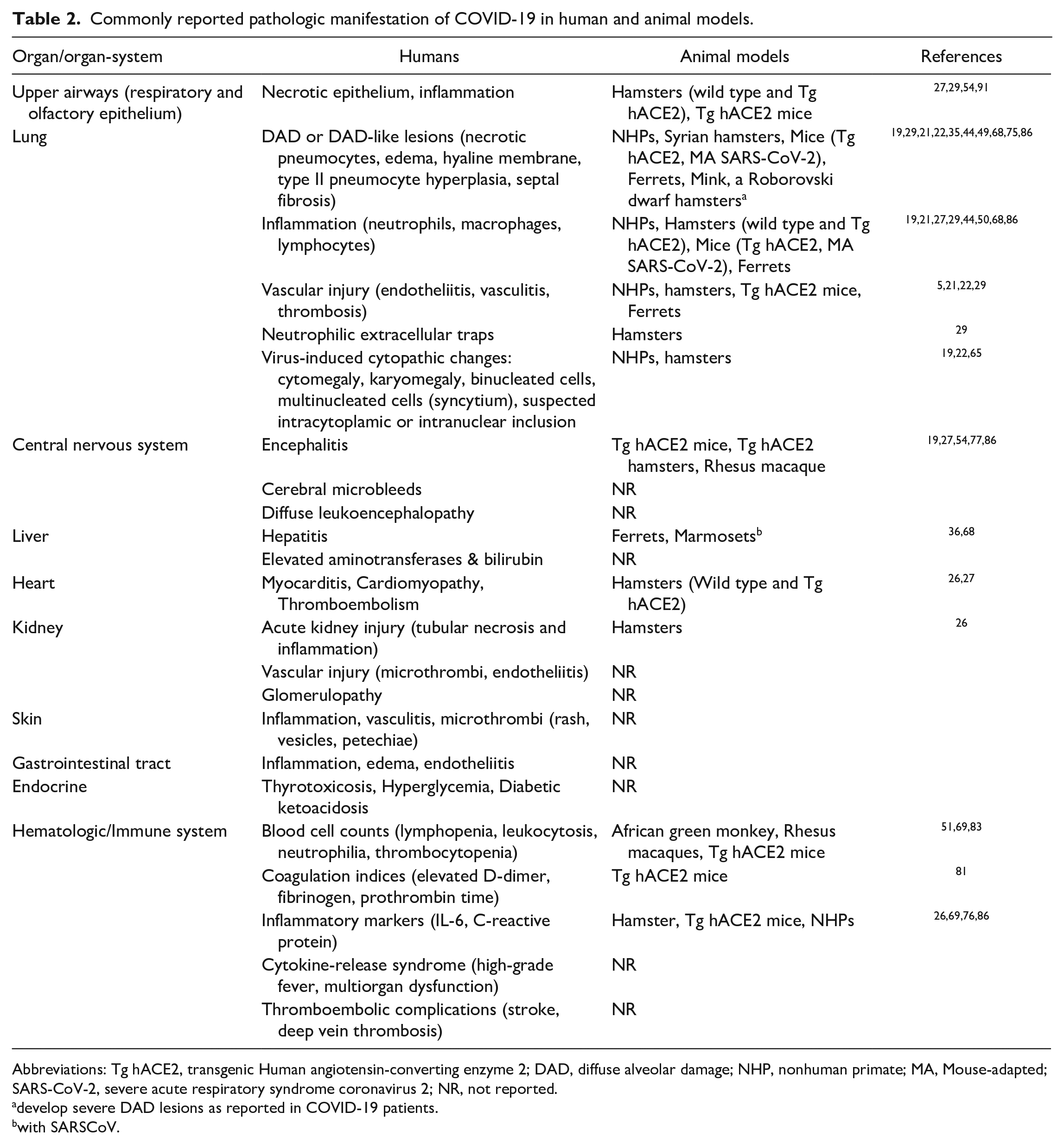
Abbreviations: Tg hACE2, transgenic Human angiotensin-converting enzyme 2; DAD, diffuse alveolar damage; NHP, nonhuman primate; MA, Mouse-adapted; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; NR, not reported.
develop severe DAD lesions as reported in COVID-19 patients.
with SARSCoV.
Pulmonary Manifestations
The pulmonary manifestations of COVID-19 are most prominent clinically and include cough, shortness of breath, sputum production, respiratory failure, and acute respiratory distress syndrome (ARDS). 37 At autopsy, the average lung weight is increased, and gross pathology findings include edematous and congested lungs in deceased patients. The main histopathological manifestations include diffuse alveolar damage (DAD), inflammation, and vascular injury (endotheliitis, vasculitis, and microthrombi) in the lungs.56,61 DAD is the primary histologic manifestation of COVID-19 and consists of pneumocyte injury, edema, fibrin/protein accumulation in the alveoli, and hyaline membrane formation in the early acute phase, and type II pneumocyte hyperplasia and alveolar septal fibrosis in the chronic phase of disease. 73 Hyperplastic pneumocytes with viral cytopathic effects such as cytomegaly, karyomegaly, intracytoplasmic or intranuclear inclusions, and syncytia formation are frequently observed in the lungs of COVID-19 patients. 63 Syncytia formation is thought to be triggered by SARS-CoV-2 S on the infected cell surface, which causes fusion with ACE2-positive neighboring cells. 14 Inflammation is characterized by a massive influx of innate immune cells, primarily neutrophils, macrophages, and lymphocytes in the alveolar space and interstitium. 3 Vascular changes include endothelial damage and subsequent inflammation resulting in endotheliitis, vasculitis, and microthrombus formation. Recently, neutrophilic extracellular trap (NET) formation secondary to SARS-CoV-2 has been shown to contribute to acute disease severity by causing vessel inflammation, endothelial damage, and thrombosis.4,8 Many of these pulmonary lesions have been successfully recapitulated in animal models although disease severity varies, and lethality isn’t commonly observed (Table 1) (Figures 1–6). Generally, hACE2 mice and hamsters develop mild to severe pulmonary disease that simulate severe COVID-19 pulmonary disease observed in patients, whereas NHPs and ferrets exhibit mostly mild to rarely moderate pulmonary disease. In some models such as hACE2 hamsters, pulmonary lesions are localized predominantly in bronchi and bronchioles, whereas in other models such as hACE2 mice, the lesions are predominantly in the alveoli. 27 Although pulmonary vascular changes do occur including endothelial damage/endotheliitis, vasculitis, microthrombi, and NETs, the severity of these findings are generally limited in animal models. Virus cytopathic-like changes observed in animal models can also be associated with regeneration of injured airway epithelium and need further characterization by IHC/ISH and/or ultrastructural evaluation for confirmation.
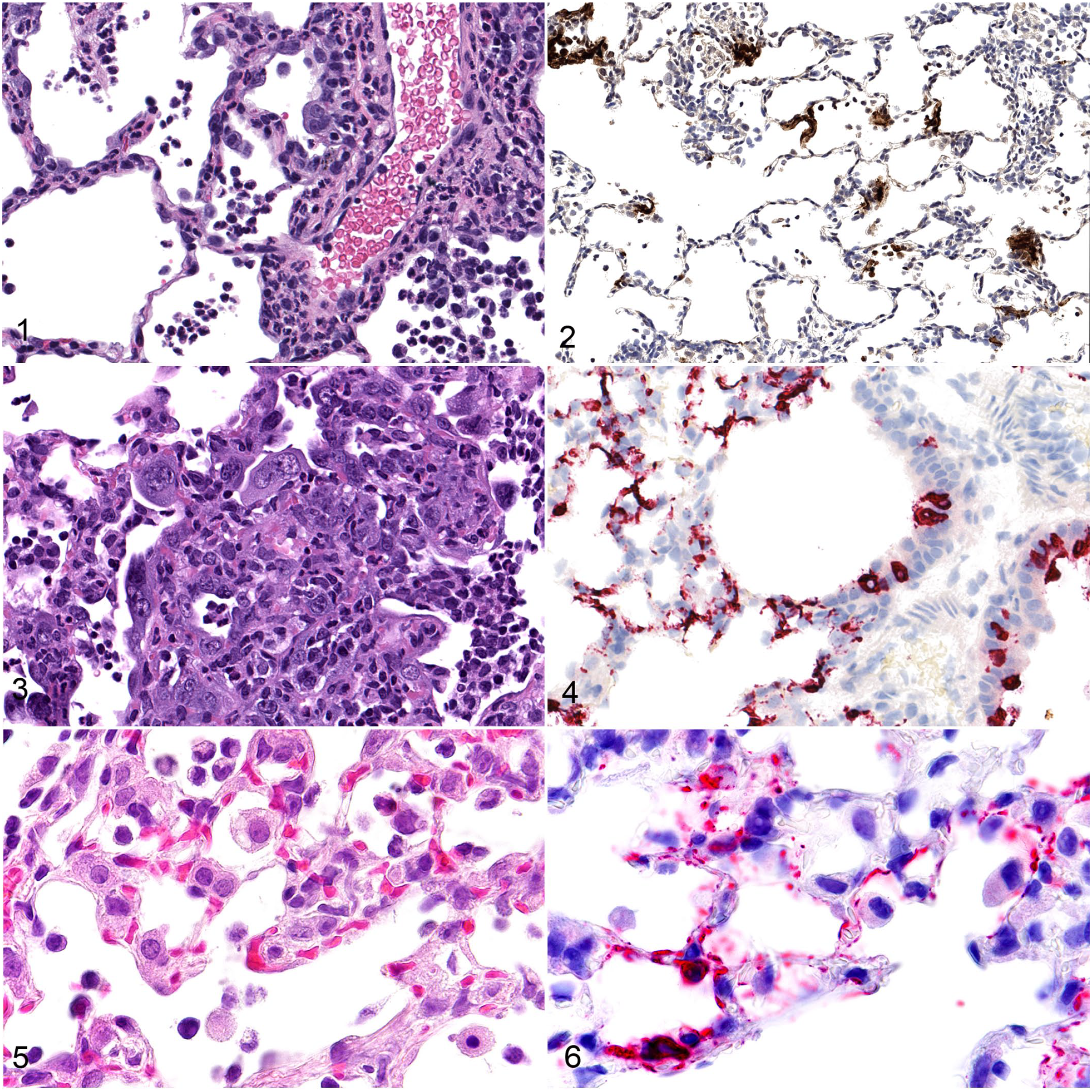
Figures 1–2. SARS-CoV-2 infection, lung, rhesus macaque, 7 days post-exposure.
In addition to the lungs, SARS-CoV-2 frequently affects upper airways including the nasal cavity. Anosmia observed in COVID-19 patients can be related to SARS-CoV-2-mediated damage to olfactory epithelium in the nasal cavity, which is rich in ACE2 and TRMPSS2.10,25 Damage (inflammation and necrosis) to olfactory epithelium as well as respiratory epithelium of the nasal cavity is observed in hACE2-expressing mouse and hamster models.13,27
Extrapulmonary Manifestations
On the other hand, extrapulmonary pathologic manifestations are limited and have not been fully recapitulated in various animal models (Table 2). For example, the prevalence of neurological symptoms (e.g., headache, dizziness, anosmia) is as high as 21% in COVID-19 cases,25,60 and up to 80% among hospitalized COVID patients. 18 COVID-19 associated brain lesions such as encephalitis, cerebral microbleeds, and diffuse leukoencephalopathy may explain reported neurological symptoms. 61 Among animal models, encephalitis, as well as associated neurological signs (e.g., ataxic gait, tremors, or anosmia) have been reported in K18-hACE2 mice following SARS-CoV-2 infections; 91 however, neurological infection by SARS-CoV-2 is less severe in mice than for SARS-CoV. 50 SARS-CoV-2-related histologic lesions (e.g., inflammation and necrosis) in the central nervous system is reported in the rhesus macaque and Tg hACE2 hamster models,19,27 however, no neurological signs were observed. Recently, non-mammalian models, such as zebrafish, were used to investigate neurologic features of COVID-19. 24 To date, neurological signs and associated lesions are not reported in other animal models, including ferrets.
Among other organs, pathologic findings are frequently observed in the kidney, heart, liver, gastrointestinal tract, skin, and endocrine system of patients with COVID-19 on biopsy and autopsy; this range of organ involvement explains the correlating clinical manifestations (renal, cardiovascular, gastrointestinal, dermatologic, and endocrinologic).30,43,45,56 Currently, reports on animal models exhibiting the above-mentioned pathological manifestations are rare (Table 2). Recently, SARS-CoV-2-related lesions in the heart (myocarditis, ventricular hypertrophy, and microthrombi) and kidney (tubular inflammation) was observed in hamster models (wild-type or Tg hACE2)26,27 and in the liver of ferrets (hepatitis). 68 Extrapulmonary effects in wild type hamsters or ferrets are mostly transcriptomic or lipidomic changes with less severe microscopic alterations than in humans and live virus is not detected in these organs.26,68 There are a few reports of extrapulmonary manifestations following coronavirus (dilated cardiomyopathy in rabbit) or SARS-CoV (hepatitis in marmoset) infections,2,50 but similar organ effects are not repeated with SARS-CoV-2 infection. To date, pathologic findings or clinical manifestations observed in the skin, endocrine system, and gastrointestinal tract of humans have not been reported in animals.
Finally, several hematologic and immune system-related manifestations are observed in COVID-19 patients including clinical pathological findings (abnormal cell counts, elevated coagulation indices), inflammatory markers, thromboembolic complications (e.g., stroke), and cytokine-release syndrome. 30 Only some of these manifestations have been recapitulated in animal models (Table 2). Multiple anatomic and clinical pathology endpoints were not included in the study design of many published animal studies to date. This may be one reason that some of the tissue or hematological manifestations occurring in human COVID-19 patients have not been reported in animal models. Additional studies with more extensive and/or specific study designs are critical for identification of representative animal models to recapitulate extrapulmonary manifestations of human disease.
Limitations of Current COVID-19 Animal Models
Ideal COVID-19 models should reproduce the full spectrum of COVID-19 phenotypes (clinical manifestations and pathology). To date, however, no single animal model reproduces the full spectrum of human COVID-19 (Tables 1 & 2). A limitation of modeling severe disease is that the number of animals with severe disease in individual groups do not have statistical power. There are other limitations (Box 3), most notably the limited lethality and lack of extrapulmonary manifestations among current models. Animal models that demonstrate severe to lethal disease provide proof-of-concept data and hence are ideal for testing vaccine and therapeutic efficacy. 50 Although transgenic mouse models (K18-hACE2 mice) exhibit a lethal phenotype and result in mild to severe pulmonary disease and mild encephalitis, it has not been determined whether the animals primarily died of lung injury and/or brain lesions. In addition, other extrapulmonary manifestations are usually mild and therefore, do not completely recapitulate severe clinical symptoms observed in humans.54,86 Differences in the severity of lesions in lung vs other tissues are likely due to non-physiological tissue distribution of hACE2 in mice (higher expression in lungs but limited expression in other tissues) and the protective effect exhibited by intact mACE2 and angiotensin 1 to 9. 5 Hamsters, particularly Roborovski dwarf hamsters, mimic clinical and pathological manifestations of severe human COVID-19, including devastating DAD and coagulopathy, but mortality is not observed and studies using Roborovksi dwarf hamsters are limited. 75 Although large animal models such as NHPs are physiologically, immunologically, and genetically more closer to humans and hence are critical to understanding how the immune system responds to SARS-CoV-2 infection, they mostly develop mild disease. In addition, NHPs are not easily infected via nasal exposure (which is the natural route of infection in humans) but instead need intratracheal and/or intrabronchial routes of exposures. 38 Changes which often precipitate acute death, such as cytokine storm, hypoxemic respiratory failure, and multiple organ failure, have not been reported in animal models. Similarly, conditions that linger after COVID-19 illness, such as long COVID and multisystem inflammatory syndrome in humans, are rarely studied in animal models. 82 Recently, a humanized mice model was reported (transiently expressing hACE2 in lungs) and recapitulated some features of chronic COVID-19 including persistent viral RNA and failure to recover body weight. 69
Current animal models also lack the relevant risk factors that humans do for developing severe COVID-19, such as old age, and comorbidities (e.g., asthma, diabetes, cardiovascular diseases, and obesity). Compared to younger animals, aged NHPs, hamsters, and BALB/c mice have been reported to develop more severe pneumonia and higher viral titers after SARS-CoV-2 challenge.29,76,87,90 However, studies addressing the association between advanced age and severe disease have been limited so far. Alterations in the eicosanoid pathway are implicated and targeted for possible therapies using mouse models. 67 In addition to age, obesity-associated severe COVID-19 phenotype was shown in mouse models using a mouse-adapted SARS-CoV-2 strain,44,90 suggesting mice may be valuable to study various COVID-19-associated comorbidities. Superinfection with community-acquired or nosocomial organisms and ventilator-associated lung injury are other risk factors known to complicate the course of severe COVID-19; 15 it is not possible to fully recapitulate these conditions in animal models.
Finally, efforts to develop COVID-19 animal models are hampered by enhanced (BSL-3) requirements (as explained in the previous section), increasingly restrictive but important ethical considerations, limited immunological and molecular research tools (particularly in species, such as hamster & ferrets), high demand for experimental animals during the pandemic which exceeds the capacity of commercial breeders, and interspecies anatomical and immune system differences. 48
Conclusion
Although there is no single animal model to study the full spectrum of clinical COVID-19 phenotypes in humans, several do allow study of SARS-CoV-2 induced respiratory disease. Since COVID-19 pneumonia in humans has a wide range of presentations, ranging from no or mild symptoms to severe illness, we need more than one model mimicking varying degrees of clinical disease manifestations. With new, poorly understood diseases, like COVID-19, we should be open and flexible to pick the best model to answer specific questions. For example, severe respiratory disease models (hACE2 mice and hamsters) are being used to test efficacy of vaccines and antiviral drugs, whereas mild to moderate disease models (e.g., NHPs and ferrets) can be used for mechanistic studies to understand the disease pathobiology. Observations of encephalitis in mice and rhesus macaques suggest that these models can be used to study COVID-19 related neurological manifestations, although more work is required to establish them as good models. Similarly, more studies are needed to establish hamsters as a model for several extrapulmonary disease manifestations (e.g., cardiovascular, renal) observed in COVID-19 patients.
Since COVID-19 is primarily seen as an acute disease affecting the lungs, most of the efforts have been devoted to developing animal models simulating various pulmonary manifestations in patients. With more and more evidence that COVID-related damage occurs in many organs other than the lung, there is an urgent need to develop animal models exhibiting extrapulmonary manifestations of human disease to extend our understanding of disease pathogenesis and evaluate new treatments; these models would also be helpful to increase our understanding of chronic COVID-19 such as multisystem inflammatory syndrome and recently described long COVID (post-acute sequelae of SARS-CoV-2 infection), both of which involve damage to multiple organs. There are many ongoing efforts, including those led by NIH, 53 on identifying well-characterized small and large animal models for various COVID-19 manifestations (acute and chronic); many of these include grant opportunities in the academic setting. Lastly, existing models may require validation and refinement based on feedback from human data/clinical trials and in light of newly identified SARS-CoV-2 variants of concern to elevate their translational utility.
Footnotes
Acknowledgements
The authors thank Jerrold Ward for the K18-hACE2 transgenic mouse lung images. We also thank the 3 anonymous reviewers for their constructive suggestions.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.



