Abstract
Coronaviruses (CoVs) comprise a large group of positive stranded RNA viruses that infect a diverse host range including birds and mammals. Infection with CoVs typically presents as mild to severe respiratory or enteric disease, but CoVs have the potential to cause significant morbidity or mortality in highly susceptible age groups. CoVs have exhibited a penchant for jumping species barriers throughout history with devastating effects. The emergence of highly pathogenic or infectious CoVs in humans over the past 20 years, including severe acute respiratory syndrome CoV (SARS-CoV), Middle East respiratory syndrome CoV (MERS-CoV), and most recently severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), underscores the significant threat that CoV spillovers pose to humans. Similar to the emergence of SARS-CoV-2, CoVs have been devastating to commercial animal production over the past century, including infectious bronchitis virus in poultry and bovine CoV, as well as the emergence and reemergence of multiple CoVs in swine including transmissible gastroenteritis virus, porcine epidemic diarrhea virus, and porcine deltacoronavirus. These naturally occurring animal CoV infections provide important examples for understanding CoV disease as many animal CoVs have complex pathogenesis similar to SARS-CoV-2 and can shed light on the ongoing SARS-CoV-2 outbreak. We provide an overview and update regarding selected existing animal CoVs and their primary host species, diseases caused by CoVs, how CoVs jump species, whether these CoVs pose an outbreak risk or risk to humans, and how we can mitigate these risks.
Keywords
Coronaviruses (CoVs) are an ancient virus possessing their most recent common ancestor at least 10 000 years and possibly up to 400 million years ago. 143 Despite being an ancient virus, CoVs have also seen many reiterations of their genome in modern recorded history. 61,133 The propensity of CoV genomes to mutate and recombine allows them to overcome natural barriers normally preventing cross-species transmission, and to adapt and proliferate in new species in a spillover event. These CoV spillover and evolution events commonly appear with divergences such as bovine CoV evolving from equine CoV in the early 1700s 134 and subsequently HuCoV OC43 diverging from bovine CoV in the 1890s. 134 Although those events predated modern virology, the recent spillover of bat CoVs into pigs, including porcine epidemic diarrhea virus (PEDV) and swine acute diarrhea syndrome (SADS)-CoV, highlights these types of events. More recently the animal to human transmission of severe acute respiratory syndrome-CoV (SARS-CoV), potentially severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), and Middle East respiratory syndrome-CoV (MERS-CoV) from bats via intermediate host species to humans have been documented. With CoV spillover events likely occurring mostly unmonitored in wildlife, understanding and preventing these spillover events is becoming more important to break the cross-species transmission chain. The animal CoVs demonstrate a remarkable flexibility in hosts, receptors, and tissue targets. Understanding CoV fundamental pathogenesis in multiple species would help develop a repertoire of treatment options. This review summarizes some of the major animal CoVs, their associated diseases, effective prophylaxis, and therapies. Additionally, we discuss how animal CoVs may serve as model systems to help understand emerging highly pathogenic CoVs such as SARS-CoV, MERS-CoV, SARS-CoV-2, and the next pandemic CoV already waiting in the wings to jump into humans or agriculturally important livestock species. 108,110,112,135
Biology of Coronaviruses
Coronaviruses are enveloped viruses approximately 80 to 220 nm in diameter containing a nonsegmented, single-strand, positive-sense RNA genome ranging from 26 to 32 kB in size. 67,142 CoVs belong to the order Nidovirales, suborder Cornidovirineae, and family Coronaviridae. The Coronaviridae family is divided into 2 subfamilies, namely, Letovirinae and Orthocoronavirinae. Letovirinae contains the Alphaletovirus genus, while Orthocoronaviridae is further classified on the basis of phylogenetic analysis and genome structure into 4 genera: Alphacoronavirus (αCoV), Betacoronavirus (βCoV), Deltacoronavirus (δCoV), and Gammacoronavirus (γCoV) including 14, 5, 3, 3 subgenera, respectively (Fig. 1; https://talk.ictvonline.org/taxonomy/ 138 ). Corona in Latin means “crown,” alluding to the spike projections from the virus envelope giving it the shape of a crown under an electron microscope, while Nido means “nest,” due to the ability of the viruses of this order to make a nested set of subgenomic mRNAs. 59
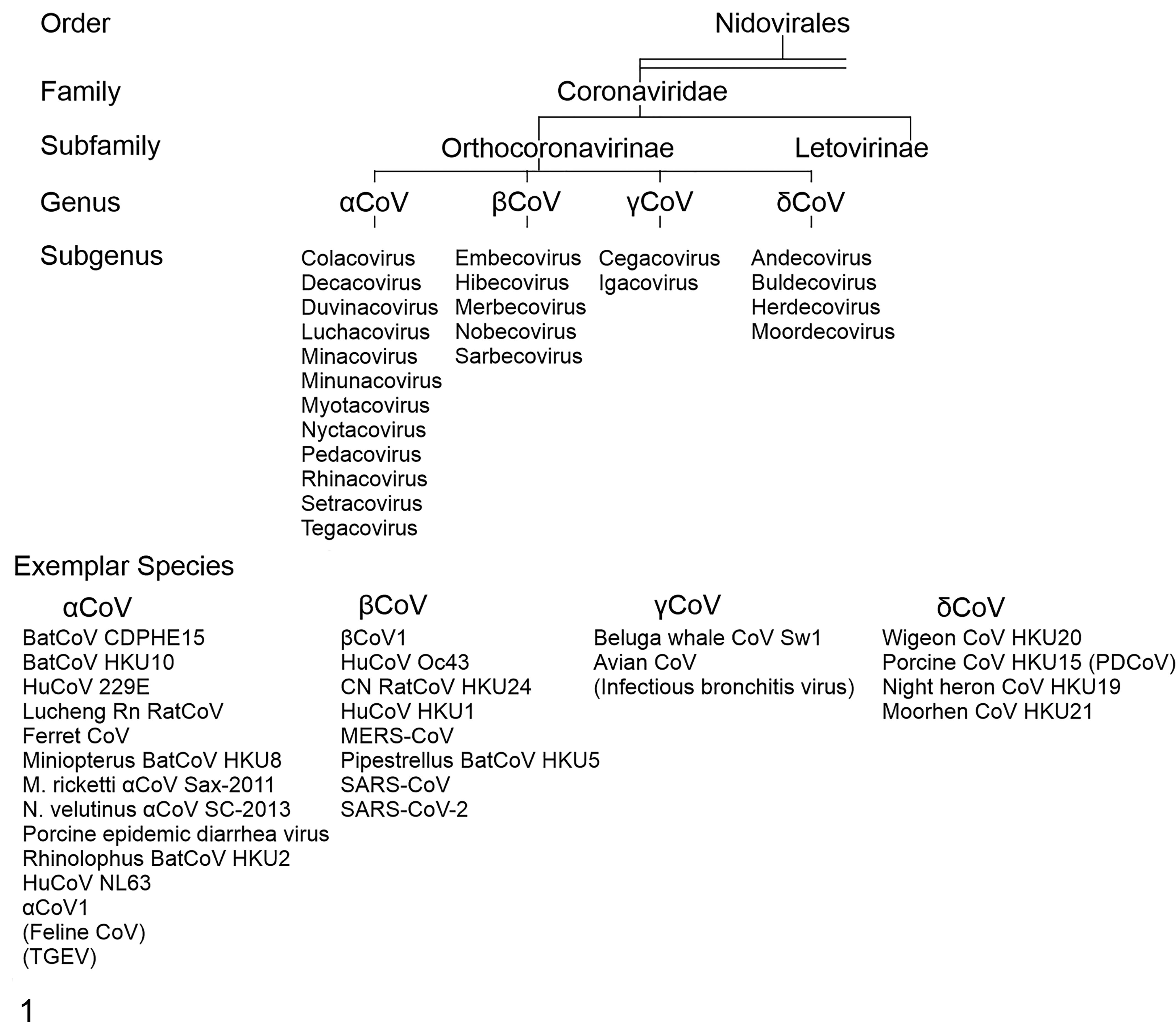
Coronaviruses: taxonomy and exemplary virus strains. The top of the tree shows the broadest categorization with the order Nidovirales, progressing downward through subgenus. The bottom row lists examples of individual viral species divided by genus.
Coronaviruses infect a wide range of wild and domestic animals, including birds. Alphacoronaviruses and βCoVs infect mammals, while γCoVs and δCoVs primarily infect birds with some mammalian spillover. Moreover, several unclassified CoVs exist and many more have yet to be discovered, which means Coronaviridae may be found to infect more species in the future. 31,151 The first human CoV (HCoV) was isolated in 1960 from hospitalized patients who suffered from common cold symptoms. This virus was named B814 from the swab sample in which the virus was isolated and is now grouped with common HCoVs, and more specifically thought to be HCoV 229E. 129 To date, there are 7 different CoVs that infect humans: 2 αCoVs (229E and NL63) and 5 βCoVs (HKU1, OC43, SARS-CoV, MERS-CoV, and SARS-CoV-2; Fig. 2). In 2002–2003, a pandemic caused by SARS-CoV originated in China. 26 In the Middle East, MERS-CoV emerged in 2012, 153 and then the virus spread to 27 countries. SARS-CoV-2, which is closely related to bat SARS-like CoVs, emerged in 2019. Chan et al 14 classified SARS-CoV-2 in the subgenus Sarbecovirus (previously lineage 2b of βCoV), while Zhou et al 158 demonstrated that it is a distinct lineage in the subgenus Sarbecovirus.
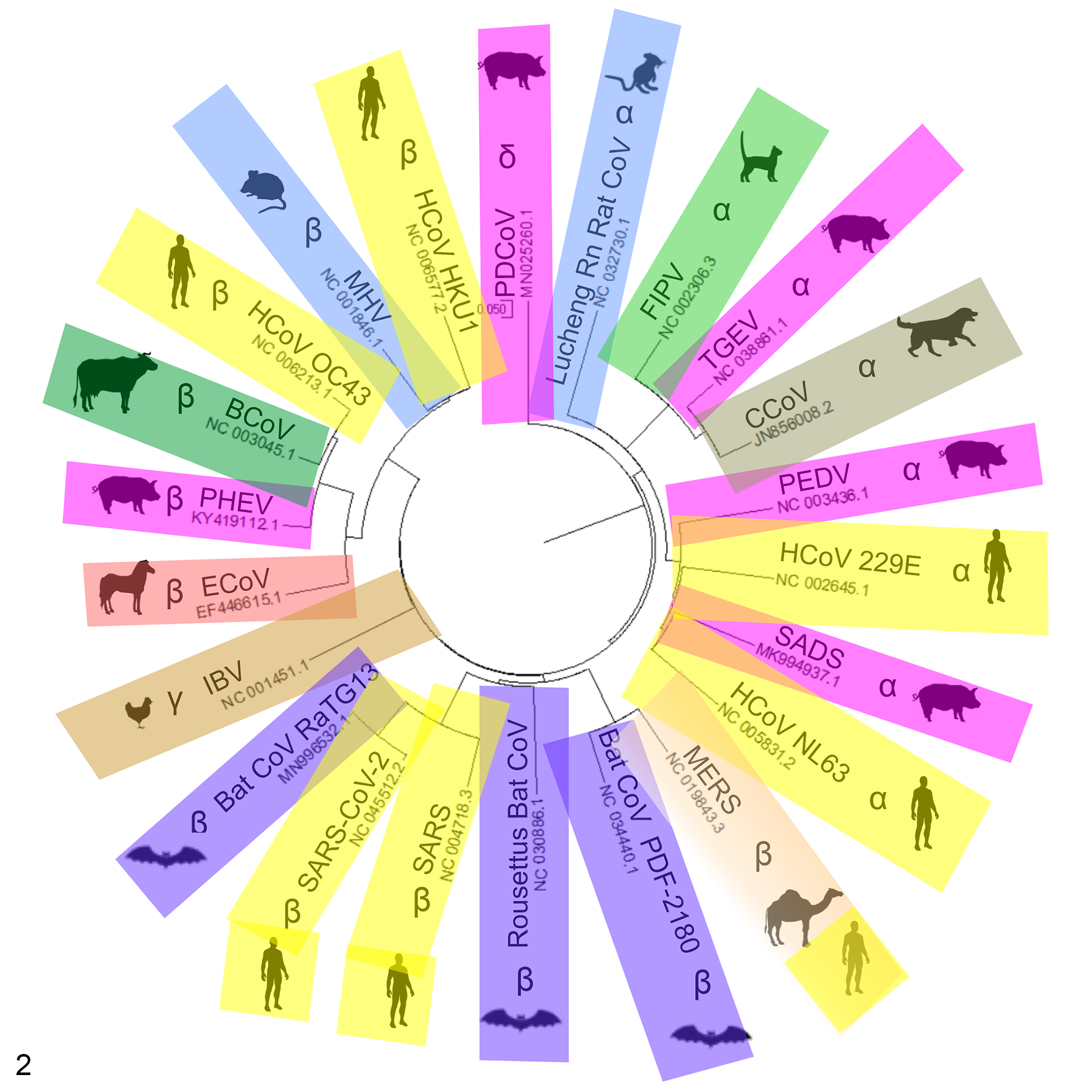
Phylogenetic tree for common animal CoVs, human CoVs, and some bat strains thought to be involved in transmission to humans. Sequence alignments were done using Clustal Omega software, which uses seeded guide trees and HMM profile-profile techniques to generate alignments based on entire genomic sequences for each strain. Alignment files were imported into MegaX64 software and converted to a circular tree. The tree was exported to CorelDraw for additional image modification.
Additionally, other CoVs have caused disease in animals, leading to mortality and/or economic losses (Fig. 2, Table 1). These viruses include infectious bronchitis virus (IBV) in chickens, 155 mink epizootic catarrhal gastroenteritis virus, 136 beluga whale coronavirus SW1 (BWCoV-SW1), 81 bottlenose dolphin CoV (BdCoV HKU22), 149 bat CoVs CDPHE15 and HKU10, porcine epidemic diarrhea virus (PEDV), transmissible gastroenteritis virus (TGEV), and swine acute diarrhea syndrome-CoV (SADS-CoV). 140,157 Bats are now recognized as important reservoir hosts for many CoVs. Surveillance studies have shown an average of 10% SARS-related CoV nucleotide homology present in bats. Bat CoVs have been proposed as the progenitors for MERS-CoV, SARS-CoV, and SADS-CoV. Bats carry CoVs related to human 229E, NL63, and PEDV, in addition to a significant number of αCoVs and βCoVs that have the potential to spill over into humans (Fig. 3), as well as uncharacterized CoVs cycling among bats and native wildlife. 30
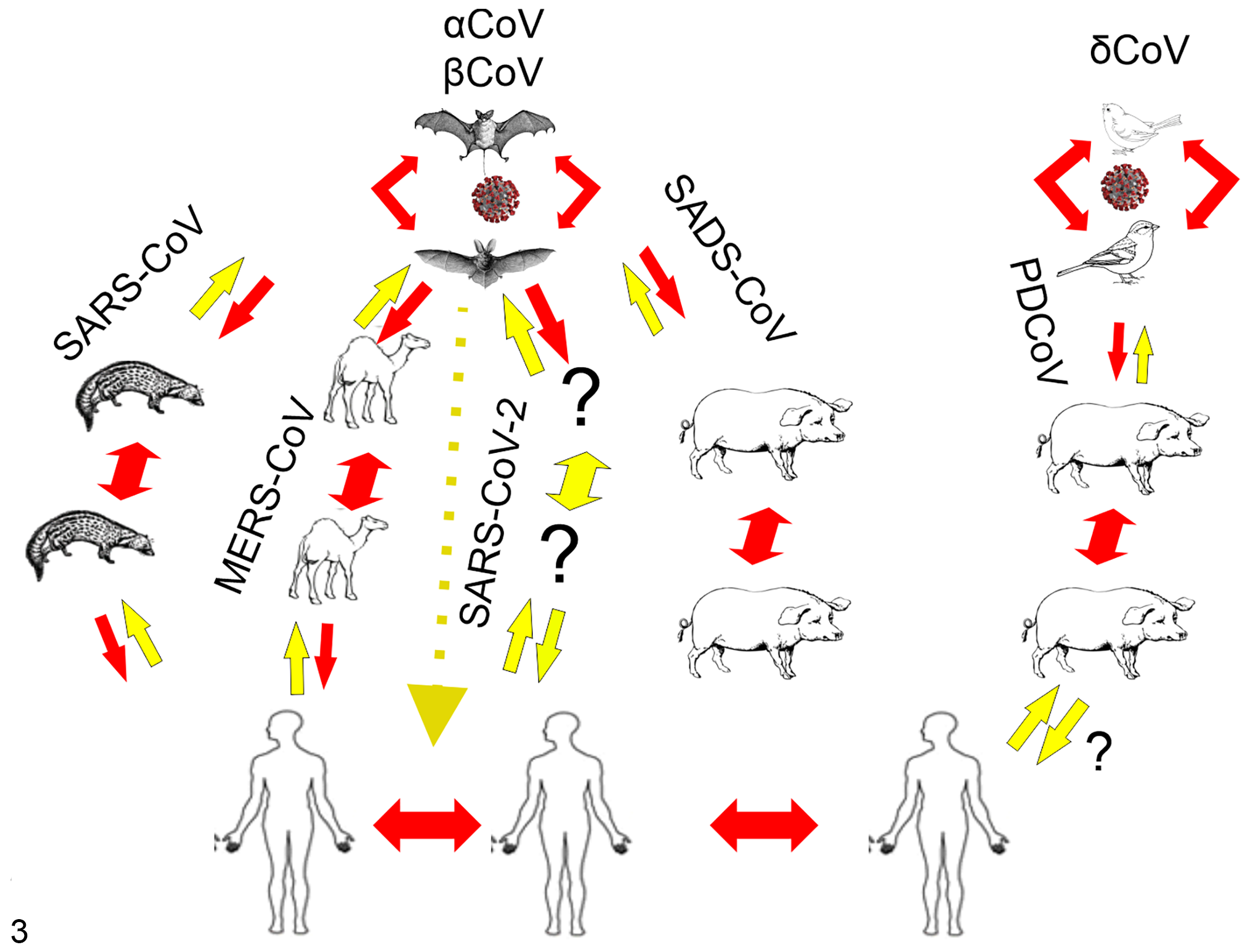
Proposed host switching events for CoV genera with human pathogenic potential. Members of the αCoVs and βCoVs continually circulate in multiple and differing species of bats. For SARS-CoV, MERS-CoV, and SARS-CoV-2, bat strains underwent recombination in an intermediate host such as civet cats for SARS-CoV, dromedary camels for MERS-CoV, and unknown species in SARS-CoV-2 before being transmitted to human hosts. Red lines are either known or highly likely transmission directions. Yellow lines are hypothetical transmission routes based on likelihood of S protein/host receptor compatibility. Like the highly pathogenic human CoVs, swine enteric coronaviral disease (SECD) pathogens also circulate in bats (TGEV, PEDV, SADS) or birds (PDCoV) before crossing into pigs and spreading from animal to animal.
Selected Coronaviruses of Animals and Humans, Their Disease Signs, Transmission Routes, Control Measures, and Vaccine Status.
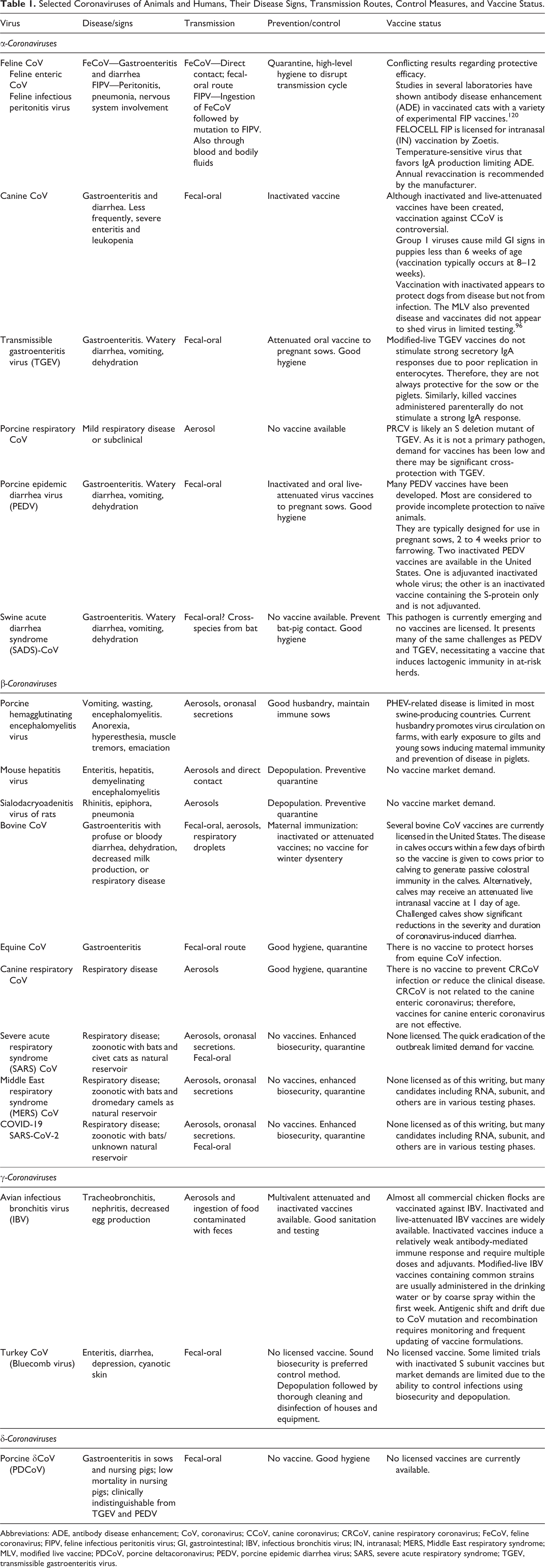
Abbreviations: ADE, antibody disease enhancement; CoV, coronavirus; CCoV, canine coronavirus; CRCoV, canine respiratory coronavirus; FeCoV, feline coronavirus; FIPV, feline infectious peritonitis virus; GI, gastrointestinal; IBV, infectious bronchitis virus; IN, intranasal; MERS, Middle East respiratory syndrome; MLV, modified live vaccine; PDCoV, porcine deltacoronavirus; PEDV, porcine epidemic diarrhea virus; SARS, severe acute respiratory syndrome; TGEV, transmissible gastroenteritis virus.
Animal CoV Spillover Events
Animals are equally at risk of an emerging virus spillover. This has been exemplified multiple times over the past few decades (Fig. 3). Like SARS-CoV-2, which originated in Wuhan, other coronaviruses have emerged within the swine industry in recent or distant spillover events with bats. Transmissible gastroenteritis (TGE) was first identified in United States in swine in 1946 25 and was subsequently identified as a CoV (TGEV) and found in Europe, Asia, Africa, and South America. 15,43,93,99,115,145 While the molecular diagnostic tools of the time lacked the ability to show bat to pig transmission, this was a likely route of introduction to swine. Interestingly, in 1984, porcine respiratory coronavirus (PRCV) evolved from the enteric pathogen TGEV into a respiratory pathogen, PRCV, demonstrating the ability of CoV mutation to confer differing tissue tropism within a host species. 15,115 Porcine epidemic diarrhea virus (PEDV) is an αCoV that is suspected to have originated from a bat reservoir. 45 PEDV first appeared in Europe in the 1970s 92,150 and then spread to Asia, where it has circulated since 1982; however, it was not until 2010 when it suddenly increased in virulence. 77,140 The PEDV epidemic strains that appeared in the United States in 2013 137 likely emerged from related highly virulent strains from pigs in China. 45 PEDV causes diarrhea in naïve pigs with mortality rates ranging up to 100% in suckling piglets, and PED resulted in significant losses to the pork industry as it spread worldwide. 77 There are other ongoing coronavirus spillover events in pigs, including SARS-CoV, an αCoV that appears to have recently jumped directly from bats to pigs in China. 35,88,157 Porcine δCoV, another coronavirus in the δCoV genus (composed of mostly avian coronaviruses), is thought to have originated from a bird to pig transmission event, although the details are unknown. 148 With animal spillover events, there are fewer tools and less funding available to study these diseases when compared with human pathogens. With the threat that these pathogens pose to human health, both by decreasing available human food supplies and possibly by additional interspecies transmission events, stopping these animal virus threats and limiting contact with potential virus reservoirs should be a high priority.
Determinants of Species and Tissue Tropisms
Most CoVs infect cells of respiratory or enteric origin, primarily due to the presence of host proteins that bind to the viral spike (S) protein. CoVs attach and enter cells via interactions of the S protein with host cell-surface receptors. 6 The S1 region of S contains 2 distinct domains: the N-terminal domain (NTD) and the C-terminal domain (CTD); both can function as receptor-binding domains (RBDs). 67 These S RBDs display a complex receptor recognition pattern. Highly similar CoV S1-CTDs within the same genus can recognize different receptors, whereas different CoV S1-CTDs from different genera can recognize the same receptor. 66 Additionally, CoV S1-NTDs can recognize either protein or sugar receptors. There is no common receptor usage pattern among CoV genera. For instance, HCoV-NL63 belongs to the αCoV-genus while SARS-CoV and SARS-CoV-2 belong to the βCoV-genus, yet their S1-CTDs recognize the same receptor, human angiotensin-converting enzyme 2 (hACE2). 41,72,139 Conversely, HCoV-NL63, PEDV, and TGEV belong to the αCoV-genus, but their S1-CTDs recognize different receptors: the TGEV S1-CTD utilizes aminopeptidase N (APN), and although PEDV also binds APN, the functionality of APN as a receptor remains debated, with APN knockout pigs remaining susceptible to infection. 21,34,71,78,144 Similarly, SARS-CoV, SARS-CoV-2, and MERS-CoV all belong to the βCoV-genus, although the MERS-CoV S1-CTD recognizes a different receptor (dipeptidyl peptidase 4 [DPP4]) than SARS-CoV and SARS-CoV-2. 102 Additional members of the βCoV-genus, MHV and BCoV, have S1-NTDs, which recognize carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1) or 5-N-acetyl-9-O-acetylneuraminic acid (sugar), respectively. 27,91,117,146 It has also been shown that host glycans are utilized by SARS and SARS-CoV-2. 28,39 The S1-NTDs of the αCoV TGEV and γCoV IBV also recognize sialoglycoproteins such as sialic acid in addition to the protein receptor. 57,100,116,118,147 Overall, coronaviruses have evolved a complex and seemingly redundant receptor recognition usage pattern including use of one or both S1 domains as RBDs, recognition of different protein receptors by highly similar coronavirus S1-CTDs within the same genus, and CoV S1-NTDs that can recognize either protein or sugar receptors. These complex interactions between S protein and host receptors are considered a major determinant of host restriction, 76 while the ability of CoVs to utilize multiple and diverse receptors may contribute to cross-species transmissions.
CoV receptors have their own host physiological functions and contribute to viral-associated disease. ACE2 is a zinc-dependent carboxypeptidase functioning in blood pressure regulation 24,54,89 and also protects against severe acute lung failure. SARS-CoV-induced downregulation of ACE2 contributes to lung injury. 46,58 APN is a zinc-dependent aminopeptidase that plays multifunctional roles in pain regulation, blood pressure regulation, and tumor cell angiogenesis. 16,83 DPP4 is a serine exoprotease that functions in immune regulation, signal transduction, and apoptosis. 53,103,104 CEACAM1 is a cell adhesion molecule and functions in cell-cell adhesion. 4,40,126 Sugar-based carbohydrates adorn many proteins and lipids on cell surfaces and function in biological processes such as immunity and cell to cell communication. 33 Factors driving CoV evolution to select these specific receptors have yet to be identified.
Although receptor specificity plays a key role in modulating host susceptibility and pathogenesis, it is not the sole determining factor. This is exemplified by the case of porcine δCoV. In vitro, porcine δCoV spike binds to a highly conserved region of the APN protein of cells from differing host species including pigs, chickens, and humans. 70 While porcine δCoV-infected neonatal pigs have significant morbidity and mortality, no porcine δCoV infections have been confirmed in humans. Experimental infection of 3-week-old chickens, while producing mild disease, did not result in mortality. 9 This can be further exemplified by the ability of MERS-CoV to infect pigs but without the resultant disease severity seen in human infections. 132 Similar scenarios are evident in bat hosts for multiple CoVs and often in the intermediate animal hosts. These examples suggest that although a CoV can enter host cells, this does not mean that the virus will induce significant morbidity and mortality. This is likely due to contributions from the host innate immune response and host-specific differences required for efficient virus infection. 68
Clinical Disease and Pathogenesis
Highly Pathogenic Human βCoVs
During SARS-CoV and SARS-CoV-2 infection, patients present with flu-like symptoms including fever, chills, cough, and malaise. 23 For SARS-CoV, 30% of SARS patients showed clinical improvement after 1 week while 70% of patients progressed to shortness of breath with recurrent or persistent fever. 37 Up to 30% of patients required intensive care treatment including mechanical ventilation. Primary tissue targets are the respiratory tract and, in many patients, also the gastrointestinal tract. 65,154 In the lungs, diffuse alveolar damage, including formation of membranes composed of protein and dead cells around the alveolus present, making gas exchange difficult and resulting in varying degrees of morbidity. 37 Macrophage infiltration, multinucleated giant cells, atypical reactive pneumocytes, and vascular injury are also observed. Viral RNA can be detected in pneumocytes, lymphocytes, and macrophages. SARS-CoV replicates in the gastrointestinal tract, although no obvious pathological changes or only nonspecific changes are typically observed. The mucosal lymphoid tissue is often depleted with viral RNA found in mucosal epithelial cells. 22
COVID-19 has an incubation period of 1 to 14 days, with typical onset ranging from 3 to 7 days after infection. 62 The COVID-19 clinical disease associated with SARS-CoV-2 infection commonly presents as fever (88% to 98%), cough (68% to 76%), and myalgia or fatigue (36% to 44%). Less common symptoms are sputum production (28%), headache (8% to 12%), hemoptysis (5%), and diarrhea (3% to 5%). Many patients (22% to 55%) developed dyspnea. 44,69 Clinical laboratory tests showed the most common abnormalities were lymphocytopenia (64%), elevated C-reactive protein (44%), elevated lactic dehydrogenase (28%), and leukocytopenia (29%). 69 Thrombotic occurrences may arise during acute infection or during convalescence. Complications in patients with COVID-19 have included pulmonary embolism, femoral vein thrombosis, phlegmasia caerulea dolens, cerebral venous sinus thrombosis, aortic thrombosis, aorto-iliac thrombosis, humeral artery thrombosis, acro-ischemic presentations, and strokes including in young patients. 5,63,85,87,94,95,128,130 Males are more likely to present with serious illness (60%). 105 Comorbidities play a significant role in severe disease presentations, with the most common being hypertension (17% to 57%), obesity (42%), and diabetes (8% to 34%). 38,105 In terms of case fatality rates, SARS-CoV has killed 11% of infected individuals, 13 MERS-CoV has killed 34.4% of infected patients, 2 and SARS-CoV-2 has killed approximately 4% of total infected individuals, with case mortality rates by country ranging from 0.3% to 11.8% that do not typically account for untested asymptomatic infections. Significantly more deaths occur in older people (3.6% of those aged >60, 8% age >70, 15% age >80) and those with underlying health issues. 125
For MERS-CoV, greater than 60% of initial cases experienced severe disease which demanded intensive care treatments such as mechanical ventilation, in contrast to SARS-CoV-2 where 3.2% of total patients required ventilation 80 and 75% to 99% of patients with comorbidities required ventilation. 8,36 Hematological aberrations observed during MERS-CoV infection included neutrophilia (8%), lymphopenia (34%), thrombocytopenia (36%), and lymphocytosis (11%). 123,131 MERS-CoV readily infects human respiratory epithelial cells, macrophages, T-cells, and dendritic cells, and influences the production and induction of pro-inflammatory cytokines and chemokines. 156
Feline CoV and Feline Infectious Peritonitis Virus
Feline CoV is typically a mild enteric or chronic asymptomatic infection of domestic cats. The first evidence that this virus was a CoV was reported by Ward in 1970 based on electron microscopic studies. 141 Antigenically, feline CoVs (FCoVs) can be classified in 2 serotypes: I and II based on differential antibody neutralization associated with variability in the amino acid sequence of the S protein. 56 Serotype I FCoV is the predominant circulating strain. FCoV serotype II viruses resulted from a double recombination between serotype I FCoV and canine CoV. FCoVs infect mature intestinal epithelial cells including enterocytes, resulting in disruption of the brush border and villi. 10,106 Autogenous FCoV strains infecting enterocytes develop random mutations to become feline infectious peritonitis virus (FIPV). While there is no definitive evidence of what makes FIPV highly virulent compared to FCoV, there are several hypotheses, with the most commonly accepted relating to a specific change in FIPV tropism from epithelial cells and enterocytes to monocytes and macrophages. 90 The tropism switch from enteric epithelium for FCoV to monocytes and macrophages for FIPV allows systemic rather than localized distribution of the virus and is considered the major reason for FIPV virulence. The FIPV form of the virus causes an incurable and fatal disease in cats. 48 FIPV is characterized by severe systemic pyogranulomatous inflammatory damage that may target serosal membranes and occur in the lungs, liver, lymphoid tissues, and brain. It has been postulated that the host immune system is crucial in pathogenesis, as T-cell depletion from the periphery and lymphatic tissues as well as changes in cytokine expression are observed in end-stage FIPV. Hypergammaglobulinemia-associated with FIPV infection is indicative of virus-induced immune dysregulation with macrophages thought to be important for the development of vasculitis. 10,55 There is a vaccine currently available for FCoV, a modified live intranasal product labeled for use in cats >16 weeks of age, which is given as a series of 2 doses 3 to 4 weeks apart. Results of efficacy studies with this vaccine are variable, some showing no efficacy, and others showing limited efficacy under certain circumstances. 56,120 Preliminary studies suggesting cats are susceptible to SARS-CoV-2 have led to great interest in establishing cats as an infection model. Concerns that feline species may contribute to viral spread also currently exist. 121
Canine CoV
The first case of canine CoV was reported in 1971 in Germany. Since that time several canine CoV outbreaks have occurred worldwide. 19 Canine CoV utilizes the host protein aminopeptidase (APN) to infect enterocytes lining the villi of the small intestine. Intestinal pathogenesis is likely related to virus-induced apoptosis of small intestinal mucosal epithelial cells, as evident in vitro. 107 Pantropic strains of canine CoV have been described as a putative cause of systemic disease in dogs characterized by pyrexia, anorexia, depression, vomiting, diarrhea, leukopenia, and neurologic signs including ataxia and seizures. Killed vaccines have been available since the 1980s that limit disease but not infection. The typically mild infection and timing of vaccination versus when natural infection occurs have made the use of available vaccines debatable.
More recently, a second canine CoV that is classified as a βCoV, making it more closely related to HCoV-OC43 and BCoV, was identified and named canine respiratory CoV (CRCoV). It was first isolated in the United Kingdom in 2003 from lung samples of dogs 29,97 and has since been found in Europe 20 and in Japan. 152 Serological studies in 2006 showed antibodies to CRCoV in dogs in Canada and the United States. 98 CRCoV is affiliated with canine infectious respiratory disease complex, being spread in areas where dogs are housed in close contact with one another or via infected surfaces. In severe cases, clinical signs include lethargy, inappetence, vomiting, hemorrhagic diarrhea, and neurologic signs (ataxia, seizures). Infected animals may have hemorrhagic enteritis, abundant serosanguineous fluid in the abdominal cavity, and severe lesions in parenchymatous organs. The lungs present with patchy areas of consolidation. Livers can be yellow-brown and congested with hemorrhages on their surfaces, and spleens can be enlarged with subcapsular hemorrhages. Gross changes in other organs include multifocal hemorrhagic renal cortical infarcts and petechial hemorrhages in lymph nodes. CRCoV antigen is detectable in organs with gross lesions including lungs, kidneys, liver, spleen, gut, and lymph nodes. In contrast to canine CoV (the αCoV causing mainly enteric disease), no vaccine currently exists for CRCoV.
Porcine Enteric CoVs
Pigs appear to be targeted frequently by coronavirus spillover events. Pigs serve as a natural host for at least 6 different CoVs representing the αCoV, βCoV, and δCoV genera. Porcine αCoVs include TGEV, PEDV, SADS-CoV, and the TGEV variant porcine respiratory coronavirus (PRCV). 112,115,140 Porcine hemagglutinating encephalomyelitis virus is a porcine βCoV, and porcine δCoV is aptly named for its susceptible primary host and genus. 115 TGEV, PEDV, SADS-CoV, and porcine δCoV cause swine enteric coronavirus diseases (SECD). They are age-dependent, with moderate to high morbidity and high mortality in neonatal pigs. 112,115,140 Seronegative TGEV- or PEDV-infected piglets show considerably higher morbidity and mortality compared with infections of piglets with SARS-CoV or porcine δCoV that emerged recently; the latter viruses appear to be relatively less adapted to the new host, pigs. 140 All produce severe gastrointestinal diseases in neonatal pigs. These diseases are virtually indistinguishable from one another based on clinical manifestations and pathologic findings alone. 112,115,140 Necrosis of infected intestinal epithelial cells occurs, causing villous atrophy that results in malabsorptive diarrhea. The severe diarrhea and vomiting leads to dehydration and death of piglets.
Natural infection induces protective immunity, but there is no known cross-protection among the 3 viruses. 140 Much research is being conducted to combat these economically important food animal diseases. The appearance and widespread prevalence of the TGEV variant PRCV, which usually causes mild respiratory disease, has induced protective herd immunity, and lessened the clinical impact of TGEV. 115 Live attenuated vaccines for PEDV based on the classical G1 strains have been in use in Asia since the early 2000s, typically reducing neonatal pig mortality but not morbidity. 52,64,124 These vaccines were minimally protective against the emerging highly virulent G2 strains. In 2014, two PEDV vaccines were introduced in the United States: a replication-deficient Venezuelan equine encephalitis virus expressing the PEDV S protein and an inactivated PEDV vaccine. 18,60 Europe also has access to inactivated PEDV vaccines but does not use them widely. 32 While the use of these vaccines is often beneficial in previously exposed herds, they often fail in naïve herds. 18 One reason for the variable vaccine efficacy may be the use of vaccines given intramuscularly rather than live virus vaccines given orally to induce a strong local enteric immunity. 60,112,115,119 Serum PEDV-specific IgA and IgG antibodies in naturally infected sows remain high for 6 months and then decline, whereas fecal PEDV-specific secretory IgA antibodies receded 1 to 2 months after infection. 86 Based on that, long-lived plasma cells in the bone marrow of infected sows might not persist for their lifetime, indicating the necessity of repeated immunizations for the persistence of humoral immunity. 115,119 Similarly, whether patients recovered from SARS-CoV-2 or immunized once a commercial vaccine is available will also require regular booster vaccinations for persistence of humoral immunity needs to be studied. 113 Similar vaccine strategies are being devised for porcine δCoV, and a similar strategy could function with all swine enteric CoV disease, with the cognate viruses.
Porcine Respiratory CoV
The remaining αCoV, PRCV, is actually derived from TGEV whereby a large S deletion (207-227-aa) changed the major tissue tropism from an enteric to respiratory pathogen. 119 Like SARS-CoV or SARS-CoV-2, PRCV infects the epithelial cells lining the upper and lower respiratory tract, such as type 2 pneumocytes. Although PRCV disease is generally mild, it can cause atypical pneumonia in infected pigs that is exacerbated by stress or co-infections. 49,51
Porcine Hemagglutinating Encephalomyelitis Virus (PHEV)
To date, PHEV is the only porcine βCoV and the only known neurotropic coronavirus affecting pigs. It produces vomiting and wasting disease (VWD) and/or encephalomyelitis. 84,115 PHEV was first reported in Ontario, Canada, circa 1957. Since then, several outbreaks of PHEV infection have been reported. 74,84,101 Although PHEV is prevalent in most swine herds worldwide and often does not cause clinical signs, morbidity and mortality were age-dependent and typically reported only in piglets under 4 weeks of age. Affected piglets exhibit VWD and/or encephalomyelitis, with the overall mortality rates for affected piglets under 3 weeks of age ranging from 30% to 100%. 84 Recently, one study reported that PHEV probably caused influenza-like illness in market-age pigs at state fairs in Michigan, Ohio, and Ilinois. 79 PHEV replicates primarily in the upper and lower respiratory tract with some replication in the small intestine. Virus travels to the central nervous system via peripheral nerves in 1 of 3 pathways: vagal nerves to the vagal sensory nuclei in the brainstem; nasal mucosa and tonsils to trigeminal ganglia and trigeminal sensory nuclei; or submucosal and myenteric plexuses to the central nervous system (CNS), targeting neurons in ganglia and vagal nuclei of the brainstem as replication sites, resulting in severe neuronal damage in the medulla oblongata progressing to the spinal cord, and occasionally cerebrum and cerebellum. 75,84 Although pigs are the only species susceptible to natural PHEV infection, the virus is also neurotropic in experimental infections of mice and Wistar rats. No vaccine or challenge studies have been described in swine.
Bovine CoV
Bovine CoV infections can cause 3 clinical syndromes in cattle: calf diarrhea, winter dysentery (hemorrhagic diarrhea) in adult cattle, and respiratory infections in cattle of varying ages. 111,112,114 One of the more serious manifestations is bovine respiratory disease complex (shipping fever) in feedlot cattle. 111 In 1973, bovine CoV was first reported as a cause of diarrhea in calves in the United States. 7,42 It has since been recognized worldwide in association with the aforementioned cattle diseases. 17 Diarrhea in calves occurs most commonly at about 1 to 3 weeks of age, when maternal antibody titers in the cow’s milk begin to wane. Pathogenesis of enteric coronavirus infection of calves is similar to that described for porcine enteric CoVs. The viral infection destroys mature absorptive cells lining the small intestinal villi and colonic mucosa. This damage leads to malabsorption, with rapid loss of water and electrolytes. The resultant hypoglycemia, acidosis, and hypovolemia can progress to circulatory failure and death, typically in young animals.
Winter dysentery is characterized by severe diarrhea, which often contains blood, accompanied by depression, anorexia, decreased milk production, and frequent respiratory signs. 114 The pathogenesis of winter dysentery resembles that of calf diarrhea, but with marked petechial hemorrhages in the colonic mucosa and extensive necrosis of cells within the crypts in the large intestinal mucosa. Morbidity rates range from 20% to 100% in affected herds with mortality rates around 1% to 2%. 1,111
Bovine CoV also causes mild respiratory signs such as coughing, nasal discharge, sneezing, fever, rhinitis, or pneumonia in 2- to 6-month-old calves. 17,111,112 Primary viral replication sites include the nasal and tracheal epithelium, and occasionally the lung. Bovine CoV is closely related on antigenic and genetic levels to HCoV OC43 that causes the common cold in humans. 42,134 In severe cases, clinical signs include fever and severe respiratory distress. Gross lung lesions due to associated bacterial infection consist of subacute exudative and necrotizing lobar pneumonia involving 50% to 80% of the lung volume. Histologic lung lesions are characterized as fibrinous, necrotizing lobar pneumonia, but with moderate to severe bronchitis and bronchiolitis. 111 OC43 has been proposed to have originated from historical zoonotic transmission of bovine CoV. 134 Bovine CoV has also been shown to subclinically infect dogs and turkey poults, leading to fecal virus shedding, diarrhea, seroconversion, and transmission to contact controls. 47,111 Importantly, one of the first reports documenting the interspecies transmission of CoVs with spillover from wildlife reservoirs was the demonstration that CoVs from captive ruminants and a child could experimentally infect calves, suggesting a transmission cycle that could involve wildlife to domestic animals to humans or any permutation of these infection events. 127 This information is directly relevant to SARS-CoV and MERS-CoV with spillover from wildlife (bats) and intermediate animals (eg, camels) to humans.
Avian γCoVs
The main representative of the γCoVs is avian CoV, a taxonomic name that encompasses the highly contagious infectious bronchitis viruses (IBVs) in chickens and similar viruses infecting other domestic birds such as turkeys, guinea fowl, or quail. In chickens, IBV initially replicates in respiratory tissues (including the nasal turbinates, trachea, lungs, and air sacs) causing deciliation and disease, with resultant clinical signs similar to an upper respiratory tract infection. IBV can also infect kidney (causing nephritis), oviduct, and the alimentary tract including esophagus, proventriculus, duodenum, jejunum, bursa of Fabricius, rectum, and cloaca, where it is usually asymptomatic. The virus can persist, with hens beginning to shed virus again at onset of egg laying (4 to 5 months of age). This phenomenon is believed to be a consequence of the stress of coming into lay potentially inducing immune suppression. 12 Additionally, IBVs have been detected in healthy wild birds along with significant nucleotide homologies among virus strains, suggesting that they may act as a transfer vector between domestic and wild birds. 82 Vaccines against chicken IBV have perhaps been the most widely used vaccines for coronaviral diseases, the others being against bovine, canine, feline, and porcine coronaviruses. 12 Live attenuated IBV strains were introduced as vaccines in the 1950s. 122 This was followed by inactivated vaccines for boosting protection in egg-laying birds. Even though 100% of chickens may be protected against clinical signs by inactivated vaccines, as many as 10% of vaccinated chicks do not achieve a protective immune response and can shed virus. Protection is also short lived. The decline in effectiveness becomes apparent 9 weeks after vaccination, with vaccines based on highly attenuated strains. Single applications of inactivated virus generally led to protection of <50% of chickens. Two applications have led to variable protection from below 50% to 100% protection depending on the study. 11 A major problem is that the immunity induced by inoculation with one IBV serotype protects poorly against infection by other serotypes. 3
Applications of Studies With Animal CoVs to Highly Pathogenic Human CoVs
Aspects of disease caused by highly pathogenic human CoV can be recapitulated in one or another of the animal CoV diseases described above. The animal CoVs covered within this review originated via relatively recent host switching events, allowing for the potential study of virus and host factors contributing to such spillovers. The morphing of TGEV into PRCV allows for the study of determinants of tissue tropisms relating to S protein structure and function and other accessory genes. Although PRCV-related clinical disease is mild, tissue or cellular tropism of the virus to the epithelial cells lining the upper and lower respiratory tract occurs and the resulting pneumonia 49,51 resembles that of SARS-CoV or SARS-CoV-2. PRCV infection has common histological characteristics to SARS-CoV infection including interstitial pneumonia with type 2 pneumocyte hypertrophy and hyperplasia, with leukocyte infiltration in the late phase of pneumonia. Although extrapulmonary lesions of PRCV infection are rare, the virus was detected in the small intestinal lamina propria of young pigs but without intestinal villous atrophy, and it was present at low titers in the liver and enlarged lymph nodes. 49 Many animal CoV infections begin as respiratory or enteric infections before progressing to involve other organ systems, allowing researchers to assess determinants of viral spread within the host. PHEV progresses from a respiratory disease to a systemic infection, and FeCoV undergoes mutation allowing it to infect macrophages and progress to a deadly systemic infection. Bovine CoV infection presents as enteric and respiratory disease depending on the age of the animal, stress level, viral and bacterial co-infections, and other factors, 111,112 thereby suggesting it may represent an useful model to understand SARS-CoV-2 infection, which also seems to have an age-related component to disease severity and tissue tropisms. Similar to some pathogenic or clinical features of SARS-CoV-2 infection and although PEDV is enteropathogenic, infected neonatal pigs often shed viral RNA in the feces even during the incubation stage of infection, acutely followed by massive destruction of infected epithelial cells, increased systemic innate and pro-inflammatory cytokine responses, and viremia. 50 The dual tissue tropism of PEDV for the epithelial cells lining the upper respiratory (nasal cavity) 73 and gastrointestinal tracts also resembles those of bovine CoVs, SARS-CoV, and SARS-CoV-2.
Conclusions
Knowledge acquired from animal CoV diseases provides a window into the steps needed to combat highly pathogenic human CoVs, while also highlighting their utility as surrogate models which do not require restrictive BSL-3 experiments. From our experiences with animal CoVs, many aspects related to emerging human βCoVs can be defined. CoVs infect multiple cell types within a host while utilizing divergent receptors to jump between hosts. They may be asymptomatic or highly pathogenic depending on adaptation to the host via receptor and other mutations. Vaccines are often ineffective, requiring multiple doses and live attenuated vaccines to prime immune responses in naïve animals followed by killed or subunit vaccines to boost the immune responses. 109,112,113,115 CoVs have proven their ability to adapt and survive for millions of years. Scientists will need to use every available tool in their arsenals to combat and defeat this serious pathogen. Given the ubiquitous nature of CoVs, it is essential to continue learning about these viruses to fight COVID-19 and potential future pandemics caused by CoV spillovers into humans due to continuous intensification of food animal production and numerous anthropogenic influences.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was partially supported by National Institute of Food and Agriculture, US Department of Agriculture (USDA; Award Number 2015-67015-23067, 2015) and USDA Agricultural Research Service (Award Number 58-3625-4-087, 2014). Salaries and research support were provided by state and federal funds provided to the Ohio Agricultural Research and Development Center (OARDC), The Ohio State University.
