Abstract
Discovered in 2003 at the Royal Veterinary College, London, canine respiratory coronavirus (CRCoV) is a betacoronavirus of dogs and major cause of canine infectious respiratory disease complex. Generally causing mild clinical signs of persistent cough and nasal discharge, the virus is highly infectious and is most prevalent in rehoming shelters worldwide where dogs are often closely housed and infections endemic. As the world grapples with the current COVID-19 pandemic, the scientific community is searching for a greater understanding of a novel virus infecting humans. Similar to other betacoronaviruses, SARS-CoV-2 appears to have crossed the species barrier, most likely from bats, clearly reinforcing the One Health concept. Veterinary pathologists are familiar with coronavirus infections in animals, and now more than ever this knowledge and understanding, based on many years of veterinary research, could provide valuable answers for our medical colleagues. Here I review the early research on CRCoV where seroprevalence, early immune response, and pathogenesis are some of the same key questions being asked by scientists globally during the current SARS-CoV-2 pandemic.
Keywords
In 2004, I began my PhD studies on a novel respiratory coronavirus of dogs that had been discovered the previous year by a colleague and now board-certified veterinary pathologist, Dr Kerstin Erles, at the Royal Veterinary College, London. 5 The group I joined, as an eager and recently qualified veterinarian, had been working with a large rehoming shelter in London to investigate possible causes of respiratory disease in their dogs, despite the use of vaccines. 3 My role was to investigate (a) the prevalence of the virus in the United Kingdom and wider dog population, (b) determine the effects of the virus on the upper respiratory tract, and (c) understand the early innate immune response to the virus in vitro.
Just one year earlier, between November 2002 and June 2003, a novel human respiratory coronavirus (now known as SARS-CoV-1) had emerged in Guangdong province in southern China and spread to neighboring Hong Kong and latterly to other parts of the Asia-Pacific region. 7 Initial reports of a new atypical pneumonia were rather patchy and news somewhat suppressed, but what subsequently emerged was a sudden, albeit rather short-lived, global interest in betacoronaviruses. A further relatively modest flurry of research interest surrounded the emergence of another betacoronavirus in Saudi Arabia in 2012 (now known as MERS-CoV). 19 Although associated with more significant morbidity, its limited geographic reach, perhaps, did not spark the required level of concerted international research effort into these viruses.
The virus I worked with, canine respiratory coronavirus (CRCoV), is genetically closely related to the well-established betacoronavirus of humans HCoV-OC43 (one cause of the common cold) and bovine coronavirus (BCoV). 5 Indeed, it is likely that all 3 viruses have a common ancestor. Many betacoronaviruses, including SARS-CoV-1 and MERS-CoV, are thought to have emerged from bats. Indeed, various species of bats are natural reservoirs for these viruses, which crucially possess both cross-species and zoonotic transmission ability. 2
And so, in spring 2020 with the global COVID-19 pandemic well established, we find ourselves “locked down” or “sheltering in place” but asking the very same questions of another novel betacoronavirus now infecting the respiratory tract of humans, as I did with dogs in 2004. Looking back, almost 16 years since commencing my PhD, I can’t help but think…this all sounds rather familiar…“I better go dust off that thesis!” Here I summarize our early investigations into CRCoV, now known to be a common and widespread cause of canine infectious respiratory disease complex (CIRDC), which has many parallels with the current research efforts underway globally with SARS-CoV-2. 14
Prevalence of the Virus
From our initial work investigating the possible etiological agents of CIRDC in a large UK rehoming center, we demonstrated a strong association between exposure to CRCoV and the development of respiratory disease in dogs entering the shelter. We knew, based on reverse transcription polymerase chain reaction (RT-PCR) testing, that the virus was present and circulating within this one large shelter, but we did not know if it was present more widely within the UK canine population, or globally.
Given the very close genetic relationship to BCoV, we developed an ELISA (enzyme-linked immunosorbent assay) based on a commercially available BCoV antigen, which we then used to test sera from UK dogs through a collaboration with a large UK diagnostic laboratory. Using leftover sera that would otherwise have been discarded we were able to show that 36% of UK client-owned dogs had antibodies to a betacoronavirus. 12
Until this point we had not been successful in culturing the virus in vitro, but a breakthrough came with the use of human rectal tumor cells (HRT-18G, derived from a colorectal adenocarcinoma) in which the virus grew to a high titer and without requirement for adaptation (Figs. 1, 2). 4 This latter finding was particularly interesting in light of the current pandemic in which SARS-CoV-2 has been demonstrated to infect secretory epithelia of the human gastrointestinal tract, possibly suggesting a dual tissue tropism of the supposed “respiratory” betacoronaviruses. 18
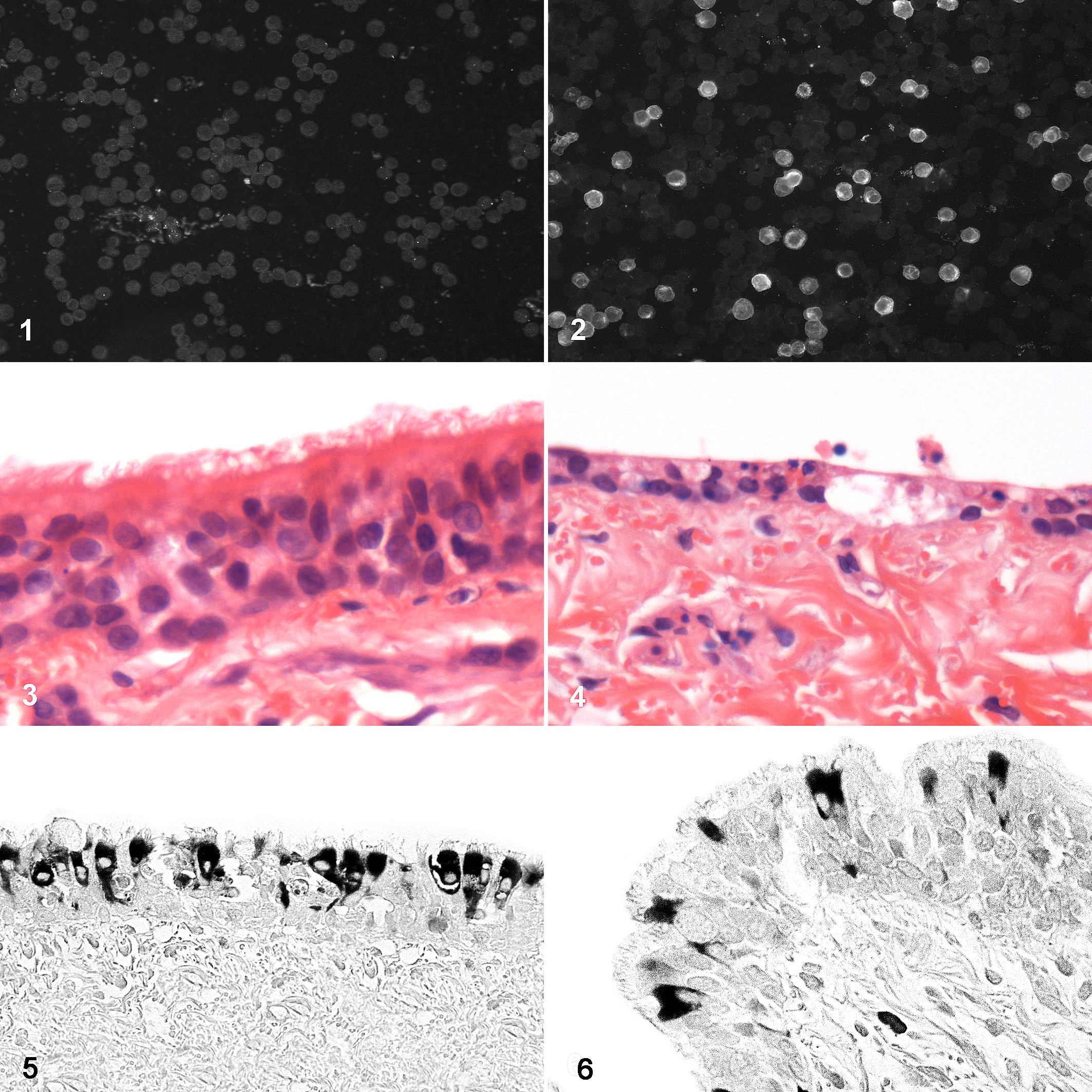
Human rectal tumor (HRT-18G) cells. Uninfected (
With viral culture optimized, we were able to refine the ELISA technique using an antigen prepared as a whole-cell lysate of CRCoV-infected cells and confirmed the previously estimated seroprevalence in the United Kingdom. Through a collaboration with University of Bari in Italy and Cornell University in the United States, we then showed the presence of antibodies to the virus in dogs from Italy (23.3%) and the United States (54.7%). 12,15 Subsequently, other groups have demonstrated a global distribution based on presence of antibodies to the virus.
Limitations of serological methods are clear and highly relevant to the current research underway with COVID-19. Seroprevalence gives an indication of previous viral infection, not active infection, and importantly we are still unclear how long antibodies to CRCoV remain detectable following infection. This is a particular problem for animal shelters—the very places where, largely due to the efficient spread between closely housed dogs, the virus is most prevalent. Since the turnover of dogs in these shelters is generally high, repeated exposure to an endemic virus also likely acts to repeatedly “boost” the immune response, and the ability to retest an individual dog is limited as many have left the shelter within a 3- to 4-week period. Specificity of the CRCoV ELISA was assessed and antibodies to the canine (enteric) coronavirus, an alphacoronavirus and the only known other coronavirus of dogs, were not cross-reactive. 15
Antibodies to CRCoV were detected in 99% of dogs within 3 weeks of entry to the large UK shelter, versus only 30% on entry, indicating a highly contagious virus. 3 Spread of the infection does not require direct contact between animals, and inhalation of air-borne respiratory secretions is the most likely route of transmission. Supporting the infectivity of the virus, CRCoV infection typically occurs in the first week after arrival to a re-homing shelter, while infections with canine parainfluenza virus and canine herpesvirus occur later, in weeks 2 and 3, respectively. 3 Indeed, CRCoV has in most countries emerged as the predominant cause of CIRDC, and no vaccine is as yet commercially available.
By way of an aside, a recent study from Italy demonstrated 100% identity between one of the main antigenic epitopes (and >80% for 2 other epitopes) of the CRCoV spike protein and SARS-CoV-2, highlighting how studying the immune response to this natural infection in the dog could offer useful insights on COVID-19. 16
Clinical Disease and Pathogenesis
Our group showed that dogs possessing neutralizing antibodies to CRCoV on the day of entry to the shelter were at much less risk of developing CIRDC. 5 Collective findings from outbreaks and clinical cases indicated that CRCoV is associated with generally mild respiratory disease, with typical clinical signs including a dry cough and nasal discharge, but what we needed was a greater understanding of the infection at the tissue and cellular level.
Using canine tracheal explant cultures, we created an air-interface (“organotypic”) model to recapitulate as closely as possible the environment of the canine upper respiratory tract. These 1 × 1 cm explants were used to assess the functional effects of the virus on mucociliary clearance and identify which cells are infected using immunohistochemistry. This technique was both technically challenging and time-consuming but provided data to show that there was a clear and consistent association between CRCoV infection and inflammation in the nares and trachea, accompanied by significant injury such as shortening and clumping or loss of tracheal cilia (Figs. 3, 4). 10 CRCoV significantly reduced mucociliary clearance and that both ciliated and goblet cells were infected by the virus (Fig. 5). 13 Later studies using experimentally infected dogs revealed identical immunohistochemical labelling of bronchiolar epithelium in the lower airways (Fig. 6). Perivascular and peribronchiolar lymphoid aggregates were observed in the lungs, but changes were less significant and more variable than those within the upper airways. Histopathological changes were apparent at the earliest euthanasia time point at 3 days post-infection and were still clearly evident at 14 days post-infection, although some recovery was noted. 10 Together these findings support the hypothesis that CRCoV causes damage to the mucociliary clearance mechanisms of the upper airways, which may predispose dogs to secondary infections.
In the first experimental CRCoV infection study to be reported, the pathogenesis of the disease resulting from infection with 5 CRCoV isolates that originated from the United Kingdom and the United States was investigated. 10 Following intranasal challenge, all 5 isolates gave rise to clinical signs of respiratory disease (nasal discharge, sneezing, and coughing), consistent with clinical signs of naturally occurring disease. 4 Viral shedding from the oropharynx was readily detected up to 10 days post-infection by RT-PCR and 6 days post-infection by virus isolation. 10 CRCoV was also detected and re-isolated from a wide range of respiratory tissues and respiratory-associated lymphoid tissues including trachea, palatine tonsil, nasal cavity, nasal tonsil, bronchial lymph node, cranial and caudal lung lobes, and lung lavage fluids. Consistent with findings from kenneled dogs, the trachea and nasal tonsil were the most common sites of infection and were also reported to have the highest viral loads. The re-isolation of CRCoV from the tissues of experimentally infected dogs with clinical disease fulfilled the final criteria for Koch’s postulates and definitively proved a causal relationship between CRCoV and respiratory disease in dogs.
During an investigation of 10 kenneled dogs with respiratory disease, CRCoV was predominantly detected in respiratory tissues. However, in 2 dogs, CRCoV was also detected in the spleen, mesenteric lymph node, and colon. 9 The detection of CRCoV in gastrointestinal samples suggests that, like some bovine coronavirus strains, CRCoV may have the potential to exhibit a dual tropism. 11 Further investigation of this and the potential clinical consequences is required, although again there is a growing body of evidence to suggest SARS-CoV-2 can also infect the gastrointestinal tract of humans and clinical signs of enteric disease have been reported in a significant number of patients. 1,18
Early Antiviral Immune Response
Coronaviruses use multifaceted approaches to counteract host antiviral responses, providing an opportunity for viral replication, dissemination within the host and, ultimately, the opportunity to infect new hosts. If they can achieve the latter undetected by the host immune system (ie, asymptomatically) then they would be considered perfectly host-adapted viruses. Other betacoronaviruses, including SARS-CoV-1, have been shown to suppress the activation of antiviral effector genes, and hence reduce the production of pro-inflammatory cytokines, by both passive and active inhibition of transcription factor IRF-3. 6,17
Using the air-interface tracheal explant culture system, we examined the messenger RNA (mRNA) levels of 3 key pro-inflammatory cytokine genes: interleukin (IL)-6, interleukin-8, and tumor necrosis factor-α following CRCoV inoculation. mRNA levels of all 3 genes were significantly suppressed, versus medium-only controls, at 24 to 48 hours post-inoculation. 13 During this time, viral load (assessed by RT-qPCR) within the tissue remained stable. At 96 hours, there was an increase in viral load and a corresponding increase in the mRNA levels of all 3 cytokines. The implication is that the virus is able, by a mechanism not yet fully understood, to suppress the host’s early pro-inflammatory immune response, perhaps “buying time” for the infection to fully develop.
This potential ability of betacoronaviruses to suppress early innate immune responses for at least 4 days post-infection may offer clues to the widely held view that asymptomatic or presymptomatic individuals are spreading SARS-CoV-2 as “covert transmitters.” 8 Additionally, although not yet assessed in vivo or if there are strain-associated differences, the significant rise in IL-6 mRNA levels observed in canine tracheal explants at 96 hours (corresponding to the highest levels of virus in the tissue) may offer some insights into the most severe expressions of disease (including an acute respiratory distress syndrome–like disease) occurring in some patients. 13 IL-6 in particular has been suggested as critical for the cytokine release syndrome (CRS, or “cytokine storm”) occurring in humans with the most severe COVID-19 disease. 20
A Final Word
Our knowledge and understanding of SARS-CoV-2 and the resulting disease COVID-19 is growing by the day, and while there is a lot still to discover, it would be worth taking the time not only to look at historical records of the emergence of betacoronaviruses in the human population but also engage with the veterinary community who have been working with coronaviruses in animals for many years. The concept of One Health, which had fallen a little out the limelight in recent years, is now back full center stage. The veterinary pathologist has a crucial role in bridging the gap between human and animal disease using knowledge of infectious diseases in animals to inform, collaborate, and push forward the One Health concept.
The exponentially expanding body of peer-reviewed (and non-reviewed) literature on SARS-CoV-2 and COVID-19 suggests roles for various wild and domestic animals including dogs in the current pandemic, variously as possible sources of the virus, intermediate vectors, or simply unwitting fomites. Indeed, SARS-CoV-1 is widely thought to have infected humans from bats via masked palm civets, while dromedary camels are considered the hosts for MERS-CoV. Time will tell what species, if any, played a role in transmission of SARS-CoV-2 to humans. While some reports have definite merit, others are very much computer-based modeling exercises that, while offering intriguing speculation, present no concrete proof.
2020 will surely become the year when we as a global society finally move from a position of reactivity to a one of proactive horizon-scanning for future threats, reasonably safe in the knowledge that it is not IF the next coronavirus will cross the species boundary, but WHEN—and we need to be better prepared for that time.
Footnotes
Acknowledgement
The technical assistance of Dr Alejandro Suárez-Bonnet with the figures is greatly appreciated.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
